Abstract
Three different wheat varieties were the subject of an experiment to see how salt affected seed germination and seedling development. BARI Gom20 (Gourab) was employed as a salt sensitive variety, whereas BARI Gom28 and BARI Gom25 were utilized on salt tolerant types. Three replications, three salinity levels (0, 150, and 200 mM of NaCl), and a complete randomized design (CRD) were used to set up the experiment. All the wheat varieties had a noticeable growth loss due to salt stress. Wheat growth was inhibited more in salt-sensitive cultivars compared to salt-tolerant cultivars, BARI Gom28 showed overall stronger salt tolerance. When wheat was tested for salt tolerance, there were striking variations in the antioxidant enzymes (catalase, peroxidase, and ascorbate peroxidase) that were present. The effect of rising salt content, catalase (CAT), ascorbate peroxidase (APX), and peroxidase (POD) activity varied between salt-sensitive and salt-tolerant cultivars. Intriguingly, TaAOX1a and TaAOX1c gene expression levels increased in BARI Gom28 as the degree of salt concentration increased. Additionally, the salt-tolerant BARI Gom28 had smaller accumulations of hydrogen peroxide, malondialdehyde, and higher activity of the antioxidant enzymes than the salt-sensitive Gourab, indicating that the latter had comparatively less oxidative damage. However, the contribution of antioxidant enzymes and AOX gene family to salinity stress response in wheat is further requisite to unlock the molecular functions. Exploring the molecular mechanism for lowering salt stress in wheat cultivars will require more research.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In our modern world, the most concerning issue is climate change which actually refers to the change in temperature and weather conditions on a long-term basis. Though it is happening due to natural causes, human activities are also responsible for climate change. The whole world is facing various problems among them salinity rising is one of the most challenging issues. Increasing the salinity of soil by evaporating the water from soil putting back the salt. Soil and water containing highly concentrated soluble salt like NaF, Na2SO4, CaCO3, MgCO3, etc. hampers irrigated and rainfed agriculture worldwide (Hopmans et al. 2021). Around 954 million hectares (ha) of land are affected by salinity worldwide and about 25–30% of irrigated lands remain unproductive because of salinity (Shahid et al. 2018). In the case of Bangladesh, salinity covered area is about 950,780 ha which is about 20% of its total area (SRDI 2010). Though wheat originated in the Southern part of Asia, it’s become very popular in the whole world according to its food value as people are changing their food habits due to health issues. Actually, people use wheat as main staple food in Asia and around one third of world’s total population (about 35%) (Shirazi et al. 2015). The second-most significant crop grown in Bangladesh after rice is wheat (Hossain et al. 2015). There is an anticipation that around 11% of winter cropped (4% in total) area is occupied by wheat cultivation which contributes significantly by producing 7% of the world’s total grain supply in Bangladesh (Shirazi et al. 2015). Many attempts have been taken in Bangladesh to develop high yield varieties to keep pace with the increasing demand for wheat (Hossain and Teixeira da Silva 2013). However, as the saline level rose and the yield of existing types decreased as a result of the adverse weather brought on by global warming, such efforts were substantially hampered. Thus, it has become a burning issue to explore the maximum production pathway of wheat in salinity invaded areas. As a sessile, plants have to face various stresses of environment, viz., various temperatures, drought, toxicity of metals, nutrient deficiency, salinity, hypoxia, pathogen attack, etc. by which the lifestyle or lifespan are hampered a lot. Salinity causes various problems on agricultural side. Such as lower growth, and lower yield in the crop cultivars. The production of wheat is also hampered due to the abiotic stress (Salinity) all over the world. The various salinity ranges have an impact on around 53% of the coastal regions in Bangladesh.
Wheat is a salt-sensitive crop, and salinity has a significant impact on it throughout its life cycle. It is extremely susceptible to salt, especially during the vegetative and early reproductive phases (Darwish et al. 2009; Kumar et al. 2013). Alternative oxidase (AOX) is basically a terminal oxidase acting as a key enzyme in the mitochondrial alternative pathway (Vanlerberghe and McIntosh 1992). AOX activity can be increased by oxidative stress so it can act as an antioxidant enzyme to alleviate ROS accumulation due to impaired or restricted respiration activity in mitochondria (Sugie et al. 2006). As a result of the production of ROS, wheat cells die narcotically (Sugie et al. 2007; Mizuno et al. 2010). So, the AOX genes of wheat are crucial. Under the stress of drought, AOX protein levels of wheat increased (Carlos et al. 2004).
To meet the current demand for food in the world, farmers need to cultivate salt tolerant crops (Rengasamy 2006). The tolerance is generally followed by 2 distinct mechanisms. One is exclusion of sodium and another is tissue tolerance (Munns et al. 2008). Salt stress causes plant cells to accumulate ROS (e.g. O2−, H2O2 and OH−) (Guo et al. 2009; Xu et al. 2009). For both intracellular and intercellular signaling, they are crucial (Foyer and Noctor 1999) though their high concentration can hinder the normal metabolism of plants (Hernández 2001). Such as excess ROS production can lead to ultimate death of plant cell (Banu et al. 2009, 2010). Consequently, it’s critical to guard against ROS harming cellular components. This has led to the development of numerous detoxification mechanisms in plants, including the production of different enzymes and antioxidant compounds (Yamane et al. 2009; Hoque et al. 2007, 2007b). Catalase (CAT: EC 1.11.1.6), peroxidase (POD: EC 1.11.1.7), and ascorbate peroxidase (APX: EC 1.11.1.11) are the key ROS scavenging antioxidant enzymes (Alscher et al. 2002; Arora et al. 2002). Due to higher salinity, while POD breaks down H2O2 to detoxify ROS, CAT transforms H2O2 into water and molecular oxygen. Several studies suggest that endogenous oxidative stress tolerance can be increased by regulating the activity of antioxidant enzymes (Saiema et al. 2012; Saeedipour 2013). Various mechanisms are followed by plants to survive these salinity conditions as osmotic potential (π) of soil is lowered by salinity due to osmotic stress. For this reason, significant overlap occurs in plants. The strategies followed by plants to tolerate salinity stress are osmotic mitigation stress and ionic stress. The current study’s objective was to ascertain how salt stress affected ROS accumulation as well as antioxidant enzyme activities and AOX gene of wheat cultivars at seedling stage in response to salt stress for BARI Gom28, BARI Gom25, and Gourab.
Materials and methods
Seed sowing and seedling growth
BARI Gom28 and BARI Gom25 were used as salt tolerant whereas BARI Gom20 was used as salt sensitive. Petri dishes or pot were used to sow the seeds and to avoid any interruption in proper growth, Hogland’s nutrient solution was treated in this experiment. The nutrient solution of Hogland was made as previously described by previous author (Hoagland and Arnon 1950).
Design of experiment
It was a completely randomized design (CRD) layout having three replications. There are two factors viz. various concentrations of NaCl and three wheat cultivars. A petri dish or pot contained twenty-five seeds sown at an equal distance from each other. During the seed sowing, 20 ml of each 150 mM, and 200 mM NaCl were applied, and distilled water was in control to the seed so that it could be moist in each Petri dish or pot. After seed germination, 3 ml of Hoagland’s nutrient solution was sprayed on the seedlings of wheat cultivars at two day intervals for two weeks so that sufficient essential nutrients could be supplied for ensuring proper growth.
Data collection
21 days after sowing (DAS) seeds, the length and weight (g) of shoot and root were recorded. Randomly uprooted five seedlings from each treatment were washed. The shoot and root length were measured with scale and fresh weight of whole seedling was measured.
Antioxidant enzyme assay
Leaf sample was homogenized at 4oC in 50mM potassium phosphate buffer (pH 8.0), containing 5mM cysteine and 1% (w/v) polyvinylpyrrolidone. This homogenized mixture was filtered, and centrifuged for 20 min at 15,000 rpm and the antioxidant enzyme was tested in the supernatant.
Catalase activity measurement
Catalase (EC: 1.11.1.6) activity was measured with a few adjustments as previously described (Aebi 1984). A cuvette was filled with exactly 50 mM, 0.7 ml potassium phosphate buffer with a pH of 8.0, and 0.1 ml each of EDTA and H2O2 were added. At 4 °C, they were well mixed. When 0.1 ml enzyme extract was added, the reaction started. Meanwhile, absorbance changes were recorded at 240 nm wavelength for 2 min at a regular interval of 30 s.
On the basis of the minutely drop in absorbance, the catalase activity was determined when the extinction coefficient of H2O2 was 40 M− 1cm− 1. The following formula was used to determine the enzyme activity -.
CAT (mMoleg− 1FW)\(=\frac{\left(\text{A}\text{b}\text{s}\text{o}\text{r}\text{b}\text{a}\text{n}\text{c}\text{e} \text{d}\text{i}\text{f}\text{f}\text{e}\text{r}\text{e}\text{n}\text{c}\text{e} /\text{m}\text{i}\text{n}\right)\times \text{D}\text{i}\text{l}\text{u}\text{t}\text{i}\text{o}\text{n} \text{f}\text{a}\text{c}\text{t}\text{o}\text{r} \left(\text{D}\text{F}\right) }{40}\)
Peroxidase (EC: 1.11.1.6) activity was assessed as described by Nakano and Asada (1981). In brief, about 50 mM 0.6 ml potassium phosphate buffer having pH of 8.0 was taken in cuvette, and subsequently both EDTA, H2O2, and guaiacol were added 0.1 ml each and mixed perfectly. When 0.1 ml enzyme extract was added, the reaction started. Meanwhile, absorbance changes were recorded at 470 nm wavelength for 2 min at a regular interval of 30 s. The peroxidase activity was calculated from the absorbance as it increased per minute when the extinction coefficient of guaiacol was 26.6 mM− 1cm− 1. The equation below was used to compute the enzyme’s activity-.
POD (µMoleg− 1FW) = \(\frac{(\text{A}\text{b}\text{s}\text{o}\text{r}\text{b}\text{a}\text{n}\text{c}\text{e} \text{d}\text{i}\text{f}\text{f}\text{e}\text{r}\text{e}\text{n}\text{c}\text{e}/\text{min}\times \text{D}\text{F} }{26.6}\)
Ascorbate peroxidase (EC: 1.11.1.6) activity was used to evaluate by Nakano and Asada (1981) with slight modifications. A cuvette containing 0.1 ml each of EDTA, H2O2, and ascorbate was filled with around 50 mM, 0.6 ml of potassium phosphate buffer at a pH of 8.0. When 0.1 ml enzyme extract was added, the reaction started. Meanwhile, absorbance changes were recorded at 290 nm wavelength for 2 min at a regular interval of 30 s. The peroxidase activity was calculated from the absorbance as it decreased per minute when the ascorbate extinction coefficient was 2.8 mM− 1cm− 1. The activity of enzyme was calculated by the equation given below-.
APX (U− 1gFW) =\(\frac{(Absorbancedifference /\text{min})\times DF}{2.8}\)
Determination of hydrogen peroxide (H2O2) content
To determine the H2O2 content, fresh leaves were collected at seedling stage and preserved at 20 °C until analysis as described previously (Rasel et al. 2019). Homogenization was performed on 0.1 g of fresh leaf material with 1 ml of 0.1% trichloroacetic acid (TCA), keeping the temperature at 4°C, to estimate the H2O2 concentration. The supernatant was maintained for one hour in the dark after mixing with 1 M potassium iodide and 10 mM phosphate buffer (the ratio was- 0.5 ml: 0.5 ml: 1 ml) following centrifugation for 15 min at 10,000 rpm. The solution was used to record the absorbance at 390 nm wavelength. 4 °C temperature was maintained in all steps except absorbance measurement.
Determination of malondialdehyde (MDA) content
0.1 g of leaf tissue from a young, enlarged leaf of comparable age was pulverized with liquid nitrogen to determine the MDA level. In order to homogenize the leaf tissue, the powder was then added to a tube containing 1 ml of 0.1% (w/v) TCA. The mixture was centrifuged for 10 min at 10,000 rpm to homogenize it and the supernatant was then transferred to a new tube. It was blended with 4 ml of 20% TCA that contained 0.5% TBA. This concoction was heated for 15 min at 95 °C before being swiftly cooled on ice. Because TBA and MDA can interact to form a red compound in an acidic buffer, concentration of MDA was determined by measuring the density of the red complex with a spectrophotometer at 532 nm wavelength. High temperatures can speed up the process, whereas low temperatures can prevent it. Supernatant was removed after the mixture had been centrifuged for five minutes at 10,000 rpm. The resulting solution was used to measure absorbance at 532 nm wavelength. 4 °C temperature was maintained in all steps without absorbance measurement.
Expression analysis of TaAOX1a and TaAOX1c (Triticum aestivum, Ta) and bioinformatics analysis
Utilizing the RNeasy Plant micro kit (Qiagen) for gene expression research, total RNA was extracted from wheat shoots. ReverTra Ace® qPCR RT kit (Toyobo) and qRT-PCR were used to synthesize cDNA. They were conducted with SYBR Green Real-time PCR Master Mix-Plus (Toyobo) with the help of Thermal Cycler Dice (TP800; Takara Bio) instrument as mentioned earlier (Sayed et al. 2016). According to the gene bank which was sequenced previously (Marcussen et al. 2014; Takumi et al. 2002) gene-specific primers were designed (Supplementary Table 1). The designed primers were blasted in the whole genome sequence of wheat (https://plants.ensembl.org) to find the distribution pattern of High Scoring Segment Pairs (HSPs) across a genome as well as the highest number of matches, suggesting they effectively target the desired gene in the respective chromosome.
Statistical evaluation
Two-way analysis of variance (ANOVA) using GraphPad Prism7 (https://www.graphpad.com) was used to analyze data. Here mean data were presented with standard deviation (SD) and in the graph, values with different letters indicate that there was statistically significant variation among the treatments and cultivars at P < 0.05 which follows Tukey’s numerous comparisons test.
Result
Impact of salinity stress on growth responses in various wheat cultivars
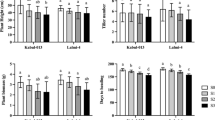
In the presence of 150 mM NaCl solution, a significant reduction in shoot length was observed in the salt-sensitive wheat cultivar Gourab (P < 0.05). Conversely, there were no significant changes (P > 0.05) in shoot length observed for the BARI Gom25 and BARI Gom28 cultivars at same concentration. At the 200 mM NaCl concentration, BARI Gom28 exhibited significantly greater shoot length compared to all other varieties surprisingly (Fig. 1a). All wheat cultivars demonstrated a decrease in root length (P < 0.05) when exposed to 200 mM NaCl relative to the BARI Gom28 cultivar (Fig. 1b). The weight of Gourab seedlings exhibited a significant decrease (P < 0.05) while the fresh weight of BARI Gom25 and BARI Gom28 remained unaffected when treated with 150 and 200 mM NaCl solutions (Fig. 1c and supplementary Fig. 1).
The influence of salinity stress on the activity of antioxidant enzymes in wheat
The APX activity exhibited a significant increase in BARI Gom28, while it remained unchanged in Gourab and BARI Gom25. At 200 mM NaCl stress, BARI Gom28 displayed the highest APX activity, whereas Gourab and BARI Gom25 showed a contrasting response (Fig. 2a). CAT activity demonstrated variation with increasing salt concentrations. At 200 mM NaCl stress, Gourab, BARI Gom25, and BARI Gom28 exhibited significant differences (P < 0.05) in CAT activity. The control condition displayed the lowest CAT activity (Fig. 2b). POD activity showed no statistically significant changes across all wheat cultivars under different salinity levels (Fig. 2c). However, at 150 mM NaCl stress, both BARI Gom25 and Gourab exhibited decreased POD activity, whereas the highest POD activity was observed in BARI Gom28. H2O2 content remained consistent in BARI Gom28, but increased in the other two cultivars. The control condition showed the highest content of H2O2 in BARI Gom28, while it was the lowest in Gourab. Under 200 mM NaCl stress, Gourab exhibited the highest H2O2 content, followed by BARI Gom25 and BARI Gom28 (Fig. 3a). MDA content remained similar across all wheat cultivars. Notably, BARI Gom25 displayed the highest content of MDA compared to other varieties under 200 mM NaCl stress, although there was no substantial increase in BARI Gom25 compared to other cultivars at 150 mM NaCl stress (Fig. 3b).
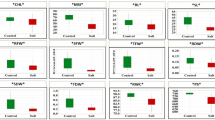
Expression analysis of TaAOX1a and TaAOX1c revealed similar patterns in three wheat cultivars and showed significant differences in BARI gom25 and BARI gom28 than Gourab. However, at 150 mM NaCl stress, slightly higher expression was observed in BARI Gom25 and BARI Gom28 compared to Gourab. Surprisingly, under 200 mM NaCl stress, both TaAOX1a and TaAOX1c transcripts exhibited higher expression in BARI Gom28 compared to the other two varieties (Fig. 4). Furthermore, we blasted our designed primers of AOX gene in the data base of whole genome sequencing of spring wheat (https://plants.ensembl.org) and subsequently got various hits for all primers (Alaux et al. 2018) For instance, we got 21 hits in the case of AOX1aF, 5 hits for AOX1aR, 210 hits for AOX1cF, 2034 hits for AOX1cR, 37 hits for ADPaseF and 15 hits in case of ADPaseR (Supplementary Figs. 1–6). But we got the maximum match when blasting with AOX1aF, AOX1aR, AOX1cF, and AOX1cR (Supplementary file 1–6). Therefore, we can assume that these four primers are our desired AOX gene.
Effect of salinity on gene expression of Ta A0Xla and Ta A0X1c. ADPase transcripts were used as a normalization control. Experiments were performed in triplicate for each sample. Data are expressed as the mean ± standard deviation. Values with different letters in the graph indicated statistically significantly different (n = 3; P<0.05)
Discussion
In the coastal belt of Bangladesh, salinity causes a reduction in wheat yield as the growth is reduced under salt stress. When cultivated under salt stress, reduced plant growth is a common occurrence and often manifests as stunted shoots. Roots are the first to come into contact with the saline medium and may also be the initial site of harm since they are in direct contact with the surrounding saline. All of the wheat varieties’ root development was inhibited by the application of salt. The results from this study indicated a significant decline in root length with the increase of salt concentration (Fig. 1b). Other researchers have observed similar outcomes in wheat as well as other crops (Momayezi et al. 2010). It was presumably because salt may have an impact on both the size of final cells and the pace of cell formation thereby producing shorter roots (Azaizeh et al. 1992). These findings, however, conflict with those of Cramer and Nowak (1992), who claimed that roots were less susceptible to salt stress. According to certain research, the most typical reaction to salinity is the suppression of shoot and root development, which is also one of the most significant agricultural markers of salt stress resistance (Koca et al. 2007). The findings of this investigation showed that increased salt levels caused a considerable decrease in shoot length (Fig. 1a). A significant decline in the length of root with the increase of salinity concentration was indicated in this study (Fig. 1b). Similarity of reduction in fresh weight with the increase of salt stress was noticed (Fig. 1c) as it was reported by (Grieve and Fujiyama 1987) and (Shannon and Grieve 1998). Growth inhibition may be caused by decreasing new cell production as per the report of (Shabala 2000; Hakim et al. 2010; Talat et al. 2013). As the report of Sweet et al. (1990), there might be a dry weight loss. According to the current study, salt-sensitive (Gourab) genotypes had a considerable decrease in the activity of antioxidant enzymes compared to salt-tolerant genotypes (BARI Gom28 and BARI Gom25) in response to salinity stress. Without BARI Gom28, the activity of POD and CAT was considerably either unchanged or decreased in all types of tested genotypes in compare to control (Fig. 2a-c). In wheat genotypes that can withstand salt stress, APX activity showed an upward trend, whereas salt sensitive genotypes showed a downward trend (Gourab) (Fig. 2a) which did not agree with the report of previous study (Turan and Tripathy 2013; Dogan et al. 2011). Though consistent results were reported by some researchers (Vaidyanathan et al. 2003; Meloni et al. 2003; Demiral and Türkan 2005). The three major enzymes CAT, POD and APX are used to detoxify harmful reactive oxygen species under abiotic and oxidative stress (Mittova et al. 2004). Salt-tolerant wheat genotypes showed significantly higher CAT, POD and APX activities under increasing salt stress than the salt-sensitive genotype which indicated that tolerance to salt stress is conferred to these genotypes via the greater antioxidant enzyme activity. With regard to salt stress and normal circumstances, BARI Gom28 with H2O2 and MDA showed a substantial difference, demonstrating the capacity of salt-tolerant cultivars to reduce oxidative damage brought on by salt stress (Fig. 3a&b) and this result combined well with the earlier finding that BARI Gom25 and BARI Gom28 are tolerant of salt stress (Siddiqui et al. 2017).
Analysis of gene expression in TaAOX1a and TaAOX1c exhibited a consistent pattern across three wheat cultivars, with notable variations observed in BARI gom25 and BARI gom28 compared to BARI gom20 with increasing salt stress (Fig. 4). Our study revealed that AOX gene family of bread wheat showed high expression during abiotic stress like salt stress. AOX gene family consists of a number of subclass located in different chromosomes. Recently, genome-wide identified 20 AOX genes and subsequently classified AOX1a, AOX1c, AOX1d, and AOX1e in hexaploid wheat which were expressed under abiotic stress (Brew-Appiah et al. 2018). Another study found 17 AOX genes which were the similar sub-class and showed significant levels of expression under salt stress in spring wheat (Zhang et al. 2023). Interestingly, all of these AOX genes are located in the different chromosome that is also suggested our primer blast results or bioinformatics analysis. Our bioinformatics analysis revealed that AOX1a is located on 2 A, 2B, 2D, 3B, 5 A, 6B, 7B and AOX1c is located on 6 A, 6B 6D and other chromosomes (Supplementary Figs. 1–6 and file 1–6) that results were found similar as described previously (Brew-Appiah et al. 2018; Zhang et al. 2023).
In contrast to monocots, which only contain members of the AOX1 subfamily, plant AOXs are encoded by a small nuclear multigene family that is split into two subfamilies: AOX1 and AOX2. Various plant species have a range of AOX genes. The number of copies of each subfamily has varied amongst various eudicot plants. In contrast to soybean, which contains one AOX1-type and two AOX2-type genes, Arabidopsis has four AOX1-type and one AOX2-type gene. AOX2-type genes have therefore increased in legumes like soybean and cowpea, whereas AOX1-type genes have increased in Arabidopsis (McCabe et al. 1998). Carrot is the only known instance when each of the two extended AOX subfamilies has two members. Normally, only one of the two subfamilies has more than one member (Costa et al. 2009a). A tandem gene arrangement has been reported for Arabidopsis AOX1b and AOX1a (Saisho et al. 1997) as well as for soybean AOX2b and AOX2a (Thirkettle-Watts et al. 2003) and rice AOX1b and AOX1a (Ito et al.1997), probably because of gene duplication. But subsequent research has shown that the two carrot AOX2 genes were connected to two linkage groups. There is minimal evidence for variation in allelic sequences of the AOX gene among species or in individual plants, despite the availability of genomic and transcript level information for many plant species’ AOX gene sequences. There are limited reports of single nucleotide polymorphisms (SNPs) in the AOX genes of rice and tomato, which may be connected to differential gene expression and stress tolerance (Holtzapffel et al. 2003) The two subfamilies of AOX genes found in eudicots are AOX1 and AOX2, with AOX1 often being thought of as constitutively or developmentally regulated (Considine et al. 2002). In Arabidopsis, different tissue, developmental and stress regulation of the AOX genes has been found (Clifton et al. 2006) and in soybean (Considine et al. 2002). In all higher plants, AOX is a component of the mitochondrial respiration pathway that uses electrons to catalyze the four-electron reduction of oxygen to water by diverting them from the energy-saving cytochrome route. Its potential can be used in molecular plant breeding as a functional marker (Arnholdt-Schmitt et al. 2006). As a result, the AOX may serve as both a key regulator of plant growth and development and an integrator of stress signals for the deployment of defenses under stress. Salt stress poses a significant threat to crop productivity worldwide, including wheat, one of the most important cereal crops. In response to salt stress, plants activate various molecular mechanisms to mitigate its adverse effects and maintain cellular homeostasis. Among these mechanisms, the expression of AOX genes has gained considerable attention due to their role in mitigating oxidative stress and sustaining mitochondrial function under adverse conditions (Vanlerberghe and McIntosh 1992). Several studies have reported the upregulation of AOX gene expression in wheat under salt stress conditions. This upregulation suggests a potential role for AOX in conferring salt tolerance by alleviating oxidative damage and maintaining cellular energy metabolism. The activation of AOX may represent an adaptive response aimed at sustaining mitochondrial function and ATP production under salt-induced oxidative stress (Brew-Appiah et al. 2018; Zhang et al. 2023). The precise mechanisms by which AOX contributes to salt tolerance in wheat remain a subject of ongoing research. It is hypothesized that AOX may alleviate salt-induced oxidative stress by diverting electrons from the respiratory chain, thus reducing the production of reactive oxygen species (ROS) and preventing oxidative damage to cellular components (Vanlerberghe 2013). Additionally, AOX may facilitate the maintenance of mitochondrial membrane potential and ATP synthesis under salt stress conditions, thereby ensuring cellular energy homeostasis. Furthermore, the regulation of AOX gene expression in response to salt stress appears to be complex and multifaceted, involving various signaling pathways and transcriptional regulators. Understanding the regulatory networks governing AOX expression under salt stress will provide valuable insights into the molecular mechanisms underlying salt tolerance in wheat. In addition to its role in mitigating oxidative stress, AOX may also interact with other stress-responsive pathways to confer comprehensive stress tolerance in wheat. Cross-talk between AOX-mediated mitochondrial signaling and other stress-responsive pathways, such as those involved in osmotic stress and ion homeostasis, merits further investigation to elucidate the integrated response of wheat to salt stress. Overall, the elucidation of the role of AOX gene expression in wheat under salt stress represents a crucial step toward the development of salt-tolerant wheat varieties through targeted genetic manipulation or breeding strategies. By harnessing the potential of AOX-mediated stress tolerance mechanisms, researchers can contribute to enhancing the resilience of wheat crops to salt stress and ensuring global food security in the face of changing environmental conditions.
Conclusion
It is clear from the current analysis that no one parameter could be recommended as the primary predictor for salt stress resistance in wheat. These traits individually or in combination can help wheat to tolerate salt. Salt-tolerant wheat variety (BARI Gom28) may have higher defense against oxidative damage by enhancing the activity of antioxidant enzymes (CAT and APX) under salt stress. Although BARI Gom25 also demonstrated a reasonable level of salt tolerance, it exhibited less physiological and biochemical traits than BARI Gom28. The salt-tolerant genotype BARI Gom28 was able to endure a greater salinity level (200 mM) than the tolerant and sensitive genotypes. In addition to integrating stress signals for defense deployment under stress, AOX may serve as a major regulator of plant growth and development. This gene has thus been suggested as a possible option for functional marker-assisted breeding approaches for stress tolerance. In conclusion, the upregulation of AOX gene expression and antioxidant activity in wheat under salt stress represents a crucial adaptive response aimed at counteracting oxidative damage and maintaining cellular function. Elucidating the intricate interplay between AOX-mediated mitochondrial signaling and antioxidant defense mechanisms will pave the way for the development of salt-tolerant wheat varieties, thereby ensuring food security in regions affected by soil salinity.
References
Aebi H (1984) Catalase in vitro- methods. Enzymol 105:121–126
Alaux M, Rogers J, Letellier T, Flores R, Alfama F (2018) Linking the International Wheat Genome Sequencing Consortium bread wheat reference genome sequence to wheat genetic and phenomic data. Genome Biol. 19(1)
Alscher RG, Erturk N, Heath LS (2002) Role of superoxide dismutase (SODs) in controlling oxidative stress in plants. J Exp Bot 53:133–141
Arnholdt-Schmitt B, Costa JH, Fernandes De Melo D (2006) In International Congress on Plant Mitochondrial Biology, Abstract Book, Vol. 34, Issue 6
Arora A, Sairam RK, Srivastava GC (2002) Oxidative stress and antioxidative systems in plants. Curr Sci 82:1227–1238
Azaizeh H, Gunse B, Steudle E (1992) Effects of NaCI and CaCI2 on Water Transport across Root cells of Maize (Zea mays L.) Seedlings. Plant Physiol, p 99
Banu MNA, Hoque MA, Watanabe-Sugimoto M, Matsuoka K, Nakamura Y et al (2009) Proline and glycinebetaine induce antioxidant defense gene expression and suppress cell death in cultured tobacco cells under salt stress. J Plant Physiol 166:146–156
Banu MNA, Hoque MA, Watanable, Sugimoto M, Islam MA et al (2010) Proline and glycinebetaine ameliorated NaCl stress via scavenging of hydrogen peroxide and methylglyoxal but not superoxide or nitric oxide in tobacco cultured cells. Biosci Biotechnol Biochem 74(10):2043–2049
Brew-Appiah RAT, York ZB, Krishnan V, Roalson EH, Sanguinet KA (2018) Genome-wide identification and analysis of the alternative oxidase gene family in diploid and hexaploid wheat. PLoS ONE 13(8):e0201439
Carlos B, Guillermo F, Go´mez D, Ethel, Martı´nez, Juan Jose´ G (2004) Mitochondria are the main target for oxidative damage in leaves of wheat (Triticum aestivum L. J Exp Bot 40:1663–1669
Clifton R, Millar AH, Whelan J (2006) Alternative oxidasesin Arabidopsis: a comparative analysis of differential expression in the gene family provides new insights into function of non-phosphorylating bypasses. Biochim Biophys Acta 17(57):730–741
Considine MJ, Holtzapffel RC, Day DA, Whelan J, Millar AH (2002) Molecular distinction between alternative oxidase from monocots and dicots. J Plant Physio. (129): 949–953
Costa JH, Cardoso HG, Campos MD, Zavattieri A, Frederico AM (2009) Daucus carota L. – an old model for cell reprogramming gains new importance through a novel expansion pattern of alternative oxidase (AOX) genes. Plant Physiol Biochem (47): 753–759
Cramer GR, Cramer RSN, Nowak GR, Cramer GR (1992) Supplemental manganese improves the relative growth, net assimilation and photosynthetic rates of salt-stressed barley. 84(4):600–605
Darwish E, Testerink C, Khalil M, El-Shihy O, Munnik T (2009) Phospholipid signalling responses in salt-stressed wheat leaves. Plant Cell Physiol 50:986–997
Demiral T, Türkan I (2005) Comparative lipid peroxidation, antioxidant defense systems and proline content in roots of two rice cultivars differing in salt tolerance. Environ Exp Bot 53(3):247–257
Dogan M (2011) Antioxidative and proline potentials as a protective mechanism in soybean plants under salinity stress. Afr J Biotechnol 10(32):5972–5978
Foyer CH, Noctor G (1999) Leaves in the dark see the light. Science 284:599–601
Grieve CM, Fujiyama H (1987) The response of two rice cultivars to external Na/Ca ratio
Guo B, Liang Y, Zhu Y (2009) Salicylic acid regulates antioxidant defense system, cell death, cadmium uptake and partitioning to acquire cadmium tolerance in rice. J Plant Physiol 166:20–31
Hakim MA, Juraimi AS, Begum M, Hanafi MM, Ismail MR et al (2010) Effect of salt stress on germination and early seedling growth of rice (Oryza sativa L.). Afr. J Biotechnol 9(13):1911–1918
Hernández JA (2001) INTRODUCTION
Hoagland DR, Arnon DI (1950) The water culture method for growing plants without soil, Circ. 347. California Agricultural Experiment Station, Berkeley, CA
Holtzapffel RC, Castelli J, Finnegan PM, Millar AH, Whelan J et al (2003) A tomato alternative oxidase protein with altered regulatory properties. 1606 (2003) 153–162
Hopmans JW, Qureshi AS, Kisekka I, Munns R, Grattan SR et al (2021) Advances in Agronomy critical knowledge gaps and research priorities in global soil salinity, vol 169. Academic Press Inc, pp 1–191
Hoque MA, Okuma E, Banu MNA, Nakamura Y, Shimoishi Y et al (2007) Exogenous proline mitigates the detrimental effects of salt stress more than exogenous betaine by increasing antioxidant enzyme activities. 164:553–561
Hossain A, Teixeira da Silva JA (2013) Wheat production in Bangladesh: its future in the light of global warming. AoB Plants 5:pls042
Hossain MI, Hossain MI, Rahman MA, Haque MA, Hasan S et al (2015) Yield potentials of recently released wheat varieties and advanced lines under different soil fertility. Agron Res 6:91–99
Ito Y, Saisho D, Nakazono M, Trutsumi N, Hirai A (1997) Transcript levels of tandem-arranged alternative oxidase genes in rice are increased by low temperature. Gene 203:121–129
Koca H, Bor M, Özdemir F, Türkan I (2007) The effect of salt stress on lipid peroxidation, antioxidative enzymes and proline content of sesame cultivars. Environ Exp Bot (60): 344–351
Kumar K, Kumar M, Kim SR, Ryu H, Cho YG (2013) Insights into genomics of salt stress response in rice. Rice 6:1–15
Marcussen T, Sandve SR, Heier L, Pfeifer M, Kugler KG et al (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome ancient hybridizations among the ancestral genomes of bread wheat Genome interplay in the grain transcriptome of hexaploid bread wheat Structural and functional pa. Science (New York, N.Y.) 345(6194), 1250092
McCabe TC, Finnegan PM, Harvey Millar A, Day DA, Whelan J (1998) Differential expression of alternative oxidase genes in soybean cotyledons during post germinative development. Plant Physiol. (118): 675–682
Meloni DA, Oliva MA, Martinez CA, Cambraia J (2003) Photosynthesis and activity of superoxide dismutase, peroxidase and glutathione reductase in cotton under salt stress. Environ Exp Bot 49:69–79
Mittova V, Guy M, Tal M, Volokita M (2004) Salinity up-regulates the antioxidative system in root mitochondria and peroxisomes of the wild salt-tolerant tomato species (Lycopersicon Pennellii). J Exp Bot 55:1105–1113
Mizuno N, Hosogi N, Park P, Takumi S (2010) Hypersensitive cell death-like reaction is associated with hybrid necrosis in interspecific crosses between tetraploid wheat and Aegilops tauschii Coss. PLoS ONE 5(6):113–126
Momayezi MR, Zaharah AR, Hanafi MM, Ismail MR (2010) Effect of chloride and sulfate salinity on nutrient uptake in Iranian rice (Oryza sativa L.). 19th World Congress of Soil Science, Soil solutions for a changing world, Brisbane, Australia. 36–39
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Physiol 59:651–681
Nakano Y, Asada K (1981) Hydrogen peroxide is scavenged by ascorbate-specific peroxidase in spinach chloroplasts. Plant Cell Physiol 22:867–880
Rasel M, Tahjib-Ul-Arif M, Hossain MA, Sayed MA, Hassan L (2019) Discerning of Rice Landraces (Oryza sativa L.) for Morpho-physiological, antioxidant enzyme activity, and molecular markers’ responses to Induced Salt stress at the Seedling Stage. J. Plant Growth Regul
Rengasamy P (2006) World salinization with emphasis on Australia. J Exp Bot 57:1017–1023
Saeedipour S (2013) Relationship of grain yield, ABA and proline accumulation in tolerant and sensitive wheat cultivars as affected by water stress. Proceedings of the National Academy of Sciences. 83(3): 311–315
Saiema R, Altaf A (2012) Differential response of chickpea genotypes under salt stress. J Funct Environ Bot (2): 59–64
Saisho D, Nambara E, Naito S, Tsutsumi N, Hirai A et al (1997) Characterization of the gene family for alternative oxidase from Arabidopsis thaliana. Plant Mol Biol (35): 585–559
Sayed MA, Umekawa Y, Ito K (2016) Metabolic interplay between cytosolic phosphoenolpyruvate carboxylase and mitochondrial alternative oxidase in thermogenic skunk cabbage, Symplocarpus Renifolius. Plant Signal Behav 11:1–7
Shabala S (2000) Ionic and osmotic components of salt stress specifically modulate net ion fluxes from bean leaf mesophyll. Plant cell Environ 23(8):825–837
Shahid SA, Zaman M, Heng L (2018) Soil salinity: historical perspectives and a World overview of the Problem. Guideline for Salinity Assessment, Mitigation and Adaptation using Nuclear and related techniques. Springer International Publishing, pp 43–53
Shannon MC, Grieve CM (1998) Tolerance of vegetable crops to salinity
Shirazi MU, Khan MA, Mujtaba SM, Shereen A, Hood RC, Mayr L, Khan MA, Mahboob W (2015) Evaluation of salt tolerance in wheat genotypes on growth and carbon isotopes discrimination technique. Pak J Bot 47:829–833
Siddiqui MN, Mostofa MG, Akter MM, Srivastava AK, Sayed MA et al (2017) Impact of salt-induced toxicity on growth and yield-potential of local wheat cultivars: oxidative stress and ion toxicity are among the major determinants of salt-tolerant capacity. Chemosphere 187:385–394
SRDI (2010) Saline soils of Bangladesh. SRMAF Project, Ministry of Agriculture, Bangladesh, pp 1–60
Sugie A, Naydenov N, Mizuno N, Nakamura C, Takumi S (2006) Overexpression of wheat alternative oxidase gene Waox1a alters respiration capacity and response to reactive oxygen species under low temperature in transgenic Arabidopsis. In Genes Genet Syst (81)
Sugie A, Murai K, Takumi S (2007) Alteration of respiration capacity and transcript accumulation level of alternative oxidase genes in necrosis lines of common wheat. Genes Genet Syst 82:231–239
Sweet WJ, Morrison JC, Labavitch JM, Matthews MA (1990) Altered synthesis and composition of cell wall of grape (Vitis vinifera L.) leaves during expansion and growth inhibiting water deficits. Plant Cell Physiol 31:407–414
Takumi S, Tomioka M, Eto K, Naydenov N, Nakamura C (2002) Characterization of two non-homoeologous nuclear genes encoding mitochondrial alternative oxidase in common wheat. Genes Genet Syst 77:81–88
Talat A, Nawaz K, Hussian K, Bhatti KH, Siddiqi EH et al (2013) Foliar application of proline for salt tolerance of two wheat (Triticum aestivum) cultivars. World Appl Sci J 22(4):547–554
Thirkettle-Watts D, McCabe TC, Clifton R, Moore C, Finnegan PM et al (2003) Analysis of the alternative oxidase promoters from soybean. Plant Physiol 133(3):1158–1169
Turan S, Tripathy BC (2013) Salt and genotype impact on antioxidative enzymes and lipid peroxidation in two rice cultivars during de-etiolation. Protoplasma 250(1):209–222
Vaidyanathan H, Sivakumar P, Chakrabarty R, Thomas G (2003) Scavenging of reactive oxygen species in NaCl-stressed rice differential response in salt tolerant and sensitive varieties. Plant Sci 165:1411–1441
Vanlerberghe GC (2013) Alternative oxidase: a mitochondrial respiratory pathway to maintain metabolic and signaling homeostasis during abiotic and biotic stress in plants. Int J Mol Sci 14(4):6805–6847
Vanlerberghe GC, McIntosh L (1992) Lower growth temperature increases alternative pathway capacity and alternative oxidase protein in tobacco. Plant Physiol 100:115–119
Xu J, Yin HX, Li X (2009) Protective effects of proline against cadmium toxicity in micro propagated hyper accumulator (Solanum nigrum L). Plant Cell Rep 28:325–333
Yamane K, Mitsuya S, Kawasaki M, Taniguchi M, Miyake H (2009) Antioxidant capacity and damages caused by salinity stress in apical and basal regions of rice leaf. Plant Prod Sci (12): 319–326
Zhang S, Yan C, Lu T, Fan Y, Ren Y et al (2023) New insights into molecular features of the genome-wide AOX family and their responses to various stresses in common wheat (Triticum aestivum L). Gene 20:888
Acknowledgements
We are grateful to Institute of Research and Training (IRT), HSTU, Dinajpur for financial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest in this study.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Omar, A.A.A., Zaman, M.M.U., Abedin, M.T. et al. Unraveling the dynamics of salt stress response in wheat cultivars: insights into growth inhibition, antioxidant enzyme activity, and AOX gene expression. J. Crop Sci. Biotechnol. 27, 439–448 (2024). https://doi.org/10.1007/s12892-024-00242-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12892-024-00242-7