Abstract
Alpha-gliadins are the predominant immunogenic fractions of gluten proteins with strong T cell stimulatory epitopes that affect celiac disease (CD) patients. To obtain essential information on the CD epitopes of Ethiopian durum wheat, molecular characterizations were conducted using α-gliadin gene clones of three elite Ethiopian durum wheat cultivars. Deduced amino acid sequences of the three selected clones share all the known primary features of α-gliadins and core sequences of CD epitopes. Several insertions and deletions (InDels), and substitutions in the core and adjacent sequences of the epitopes are dissimilar among the deduced α-gliadins. Due mainly to these sequence variations, the core sequences of all the toxic epitopes are in one to four mismatches to their canonical sequences, showing distinct patterns among the deduced α-gliadins, except one canonical epitope from cv. Assasa. These sequence variations also result in a drastic difference in peptide cleavage patterns by proteases contained in human gastric and duodenal juices. In addition, characteristic differences in disulfide connectivity, and hydropathy and polarity of amino acid residues correlate well with noticeable variations in conformational ensembles of the peptides, especially at the region harboring multiple CD epitopes at the N-terminal region. α-Gliadin from cv. Assasa is unique in that it has mostly variant toxic epitopes and all the epitope motifs are cleaved by multiple enzymes at up to five positions. Ethiopian germplasms expressing α-gliadins with divergent sequences at the epitope motifs together with higher susceptibility to proteases in human digestive systems could be potential resources for the development of hypoallergenic cultivars.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gliadins refer to a mixture of proteins of similar amino acids with both hydrophilic and hydrophobic properties. Depending on the variety type, gliadins comprise 30–60% of wheat gluten proteins. These proteins have a special elasticity effect on gluten structure and mainly impart viscosity to dough (Delcour and Hoseney 2010). Among the four classes of gliadin proteins, based on their mobility in the A-PAGE gels, the α-gliadins are the predominant immunogenic fractions of gluten proteins with strong T cell stimulatory epitopes that affect celiac disease (CD) patients (Arentz-Hansen et al. 2002; Asri et al. 2021; Ciccocioppo et al. 2005; Molberg et al. 2005; Sollid et al. 2012; Vader et al. 2003). CD is an immune-mediated and gluten-related chronic inflammatory disorder that causes gastrointestinal signs and symptoms in sensitive individuals. Individuals that carry the susceptible human leukocyte antigen (HLA) gene DQ2 and/or DQ8 suffer from CD. Increased consumption of wheat and wheat-based products is considered a major cause of CD.
Gliadins and glutenins contain the DQ2- and DQ8-binding epitopes (Battais et al. 2005; Srinivasan et al. 2015). Four typical epitopes, such as DQ2.5-glia-α1a (the nine amino acid core sequence, PFPQPQLPY), DQ2.5-glia-α1b (PYPQPQLPY), DQ2.5-glia-α2 (PQPQLPYPQ), DQ2.5-glia-α3 (FRPQQPYPQ), are known to be digestion-resistant and the main toxic peptides for CD patients (Bromilow et al. 2017; Shan et al. 2002; Sollid et al. 2012). Once toxic peptides are absorbed in the digestive system, one or two glutamine residues in the core sequence of the epitopes are good substrates for the tissue transglutaminase (tTG) in small intestine (Nakamura et al. 2013; Palosuo et al. 2003; Di Sabatino et al. 2012). Deamidation of glutamine to glutamic acid by the enzyme increases the binding affinity of the peptides containing these epitopes for HLA-DQ2 and -DQ8 (Qiao et al. 2011). Binding of these CD epitopes to DQ2 and/or DQ8 potentiates the subsequent inflammatory T cell reaction (Schuppan et al. 2009).
Alpha-gliadin comprises 15–30% of the wheat grain protein, with an average molecular weight of 33 kDa (Huo et al. 2018). Considering that wheat is the most consumed crop in the world, much attention is given to the development and promotion of wheat cultivars that lack immunogenic gluten peptides but retaining good processing quality (Rashtak and Murray 2012; Shewry and Tatham 2016).
Studies have shown that, in durum wheat, α-gliadins are encoded by approximately 100 genes and pseudogenes per haploid genome at the Gli-2 loci on the short arm of chromosomes 6A and 6B, and Gli-1 and Gli-3 loci on the short arm of chromosomes 1A and 1B (Ozuna et al. 2015; Payne 1987; Shewry et al. 2003). The α-gliadin gene is usually arranged in the following order: 20-amino acid signal peptide followed by a repetitive N-terminal domain and then followed by two longer non-repetitive C-terminal domains that are separated by polyglutamine repeats (Anderson and Greene 1997; Shewry et al. 2003). In most instances, α-gliadin contains five to six conserved cysteine residues in the non-repetitive domains, which form interchain disulfide bonds (Shewry et al. 2003; Wang et al. 2017). The presence of additional cysteine residues in the α-gliadins allows the formation of strong intramolecular interchain disulfide bonds, and this leads to have positive effects on pasta quality (Shewry and Tatham 1997).
Various research findings have shown that the levels of T cell stimulatory epitopes vary among different cultivars and biotypes (Vader et al. 2003; Van Herpen et al. 2006; Ozuna et al. 2015). Generally, fewer immunogenic peptides were released from monoploid and tetraploid cultivars than from hexaploidy cultivars after human gastrointestinal digestion ex vivo (Asledottir et al. 2020) though there are some contrasting reports (Prandi et al. 2017). This gives a chance to select cultivars with reduced α-gliadin toxicity levels, and hence include them in breeding programs as parental lines to develop novel wheat cultivars with reduced CD-causing epitopes.
In this study, we have employed gene-specific PCR and in silico sequence analysis to investigate the genetic organization and molecular characteristics of CD-toxic epitopes in the conserved coding sequences of α-gliadin genes from the elite Ethiopian durum wheat genotypes.
Materials and methods
Experimental materials
In the previous study, the alleles of gliadin loci were investigated for the thirteen durum wheat cultivars with diverse genetic backgrounds that represent the modern Ethiopian durum wheat gene pool. The three cultivars, Assas, Denbi, and Mukiye, which are classified into the three different clade groups based on the genetic distance calculated for the alleles of α-gliadin loci (Hailegiorgis et al. 2017), were selected and used for the molecular analysis of α-gliadins of the selected cultivars.
Cloning and sequencing of α-gliadin genes
Gene-specific primers targeted for the conserved sequences of the α-gliadin genes were designed to amplify the target DNA (Chen et al. 2008; Dubois et al. 2016; Van Herpen et al. 2006). The primers used to amplify the whole frame of the α-gliadin genes from genomic DNA were: GL1F 5′-ATGAAGACCTTTCTCATCC-3′ and GL1R 5′-GTTAGTACCGAAGATGCC-3′. PCR amplification was performed with the cycling consisted of an initial denaturation at 95 °C for 2 min, 33 cycles of denaturation at 95 °C for 20 s, annealing at 55 °C for 10 s, extension at 72 °C for 1 min, and a final extension at 72 °C for 4 min. The amplified PCR products were cloned in pGEM-T-Easy vector (Promega, Madison, USA) following the manufacturer’s instructions. Subsequently, recombinant positive clones containing an insert of the expected size were identified for nucleotide sequence analysis. DNA sequencing was conducted by Sanger sequencing through a commercial service (SolGent Co., Ltd., Daejeon, ROK). Three clones, one from each cultivar, harboring the full-length open reading frame (ORF) were selected for the analysis of CD-causing epitopes. The DNA and amino acid sequences reported in this work were deposited in NCBI GenBank database (MT510704–MT510706; https://www.ncbi.nlm.nih.gov/nuccore/).
Identification of CD epitopes and structural analyses
The deduced amino acid sequences were also systematically investigated for the major T cell stimulatory epitopes using the alignment tools for protein sequences, SIM (https://web.expasy.org/sim/) and ClustalW2 (https://www.ebi.ac.uk/Tools/msa/clustalw2/). Disulfide connectivity was predicted using DiANNA algorithm (Ferrè and Clote 2005) at the web interface (http://clavius.bc.edu/~clotelab/DiANNA/). The three-state secondary structure was predicted based on position-specific scoring matrices (Jones 1999) using the program PSIPRED at http://bioinf.cs.ucl.ac.uk/psipred/. The hydrophobicity profile was generated based on the amino acid scale by Kyte and Doolittle using the ProtScale at Expasy (https://web.expasy.org/protscale/). Potential cleavage sites on the deduced polypeptides were predicted for proteases contained in human gastric and duodenal juices using the tool, PeptideCutter, on Expasy (https://web.expasy.org/peptide_cutter/). Disordered protein regions were identified using IUPred2A (Mészáros et al. 2018) web server at https://iupred2a.elte.hu/. Linear net charge per residue (NCPR) plots were generated using CIDER tool (http://pappulab.wustl.edu/CIDER/) (Holehouse et al. 2014). In the plots, Blob index indicates the number of residues beyond which the balance of chain–chain, chain–solvent, and solvent–solvent energies is of order κT (Das and Pappu, 2013), in which κ and T denote Boltzmann’s constant and the absolute temperature, respectively.
Results and discussion
Primary structure of deduced α-gliadin polypeptides
The deduced amino acid sequences of the selected PCR clones share all the typical primary structural features of α-gliadins, such as an N-terminal signal peptide (SP) of 20 residues, followed by N-terminal repetitive domain (NRD) of 93–96 residues, and a C-terminal region of 165–171 residues. The C-terminal region is interrupted by the two separate poly-glutamine regions, Poly-Q1 (PQ1) of 17–18 residues and Poly-Q2 (PQ2) of 8–10 residues, respectively, generating additional two sub-regions, C-terminal non-repetitive domain 1 (CND1) and 2 (CND2). CND1 of 69–71 residues is in between PQ1 and PQ2, and CND2 of 69–74 residue at the C-terminal end (Fig. 1).
Multiple alignments of the amino acid sequences deduced form the α-gliadin gene clones of the three Ethiopian durum wheat cultivars. The known undigested gliadin peptide p31–43 and the toxic 33-mer are indicated by the broken underlines. Boxed sequences in the rounded rectangles represent epitopes and their variants including DQ2.2-glia-α1 QGSVQPQQL; DQ2.5-glia-α1a, PFPQPQLPY; DQ2.5-glia-α1b, PYPQPQLPY; DQ2.5-glia-α2, PQPQLPPQ; DQ2.5-glia-α3, FRPQQPYPQ; DQ8-α-Glia, QGSFQPSQQ, respectively. A region containing the most significant CD epitopes in overlap is indicated by a broken-rectangle. Indicated at p237 and p231 are the disruption of epitope DQ2.2-glia-α1 by a single amino acid change from Q to R and conserved L, respectively, which are found typically in T. monococcum (A genome). The sequence CT is conserved in α-gliadins encoded by A genome. The sequence PIS is conserved in most gliadins and frame shift or premature stop codon only appear downstream of the PIS motif. The conserved cysteine residues are indicated by shade. An asterisk (*) indicates the position which has a single, fully conserved residue. A colon (:) indicates conservation between groups of strongly similar properties. A period (.) indicates conservation between groups of weakly similar properties
Basically, multiple comparisons of the deduced α-gliadin sequences from the three cultivars show about 88–92% similarity. Genetic relationship among the cultivars estimated with α-gliadin sequences is similar to that by the allele composition of gliadin loci (Hailegiorgis et al. 2017). Frequent single-nucleotide polymorphisms (SNPs), insertions and deletions (InDels) in each α-gliadin sequence resulted in variations in amino acid number from 278 (Assasa) to 287 (Denbi), molecular weight from 32.2 kD (Assasa) to 32.9 kD (Denbi), and pI from 6.99 (Assasa) to 9.10 (Denbi). The sequence CT at p284–285 is conserved in α-gliadins encoded by A genome (Wang et al. 2017). The conserved residues L at p231 and R at p237 are typical signatures in T. monococcum (A genome) among diploid Triticum genomes (Van Herpen et al. 2006). The sequence PIS at p114–116 is conserved in most gliadins and frame shifts only appear downstream of the PIS motif (Ozuna et al. 2015; Wang et al. 2017). The cysteine residues conserved are five in the α-gliadin from cv. Denbi and six in those from cvs. Assasa and Mukiye.
Variations in amino acid composition
Amino acid composition of α-gliadins, a major wheat protein, is important in many aspects including nutritional values, processing properties, and health effects. Content of essential amino acids is about 30%, whereas that of nonessential amino acids is about 70%, in the α-gliadins from the three cultivars. Notably, the content of nutritionally important tryptophan and methionine is lower than 0.4% and 1.5%, respectively. These levels fall into the range of average values of the α-gliadins from the thirteen durum wheat accessions (Table 1).
Cysteine residue is considered most significant concerning processing quality and conformational features of gluten proteins. There are five cysteine residues in the α-gliadin from cv. Denbi, but six cysteine residues in that from cvs. Assasa and Mukiye. In most instances, α-gliadins contain five to six conserved cysteine residues in the C-terminal non-repetitive domain, and they form intramolecular disulfide bonds, which is the chemical basis for physical properties of baked products (Shewry et al. 2003; Wang et al. 2017).
Conformation of proteins is affected significantly by the sequence of amino acids residues and local interactions among the stretches of polypeptide chains. Amino acids can be classified into distinct groups based on the tendency of the side group to interact with water, such as negatively charged (D, E) or positively charged (K, R, H), aromatic (F, Y, W), polar (S, T, C, N, P, Q), and non-polar (G, A, V, L, M, I) groups (Radivojac et al. 2007). The α-gliadins from the three cultivars show a similar characteristic composition for each group of amino acids. Polar residues are most highly abundant (about 59–60%) followed by non-polar (25–26%), aromatic (7–9%), positively charged (~ 5%), and negatively charged (1 ~ 3%) ones. The α-gliadin from cv. Assasa is different from the others in that it has significantly higher negatively charged and lower aromatic amino acid residues. In the negatively charged group, glutamic acid and aspartic acid residues are similarly higher (Fig. 2).
Regarding health effects, α-gliadins have several unique features that contribute to their immunogenic properties. One of the features is related to their extremely high proline and glutamine contents. Gastric and pancreatic enzymes lack post-proline cleaving activities and this results in resistance to proteolytic degradation within the gastrointestinal tract (Shan et al. 2002). Content of glutamine and proline ranges from about 30–33% and from 13 to 16%, respectively, in the α-gliadins of the three tested cultivars, encompassing average values of α-gliadins from general durum wheat accessions. Interestingly, the α-gliadin from cv. Assasa is unique in its glutamine and proline contents, in that it shows higher and lower contents of these amino acids, respectively, than the α-gliadins from the other two cultivars (Table 1).
In general, the composition of amino acid residues is within the range of previous reports (Arentz-Hansen et al. 2002; Shan et al. 2002; Shewry et al. 2003). The high glutamine content makes gliadins a good substrate for tTG (Nakamura et al. 2013; Palosuo et al. 2003). Under physiological conditions, tTG can convert glutamine into the negatively charged glutamic acid, leading to enhanced immunogenicity of the resulting modified peptides, which can preferentially bind to HLA-DQ2 or HLA-DQ8 (Kim et al. 2004; Sollid et al. 2012). Deamidation is an important event in the generation of gluten-specific T cell response and concomitant celiac disease development (Di Sabatino et al. 2012).
CD-toxic epitopes divergent from the canonical sequences
CD-toxic and immunogenic epitopes are present in canonical or variant forms in the NRD and CND2 (Fig. 1 and Table 2). Except DQ2.5-glia-α3 which is in two copies, all the other epitopes are present in one copy. Interestingly, the epitopes in the canonical sequence are rare. In the α-gliadin from cv. Assasa, one of the two DQ2.5-glia-α3 epitopes and one immunogenic DQ2.2-glia-α1 epitope are present in the canonical sequence. In the α-gliadins from cvs. Denbi and Mukiye, however, only DQ2.2-glia-α1 is present in the canonical sequence. Risk of CD occurrence is decreased in DQ2.2 than in DQ2.5 (Koning 2012; Dørum et al. 2014) among the HLA variants. The sequences of the toxic epitopes, such as DQ2.5-glia-α1, DQ2.5-glia-α2, DQ2.5-glia-α3, have one to three mismatches to their canonical sequences, showing distinct patterns among the α-gliadin from the three cultivars. Interestingly, the target glutamines for deamidation are completely conserved in all DQ2.5-glia epitopes in the α-gliadins from the three cultivars. Conversely, the target glutamine in DQ8-α-glia is substituted to arginine in the three α-gliadins. This substituted arginine is well conserved in the α-gliadins encoded by A genome of Triticum species (Van Herpen et al. 2006). The p31–34 and the 33-mer are present in partial fractions in the α-gliadins from the three cultivars due to these sequence variations in the epitope sequences, and SNPs and InDels. These variations are more evident especially in the region encompassing the multiple overlapping epitopes (Figs. 1 and 3). In the three α-gliadins, the canonical repeat unit P(F/Y)PQPQL in the region of 33-mer is variable either to PYPQPQP or PY(P/S)Q(P/A)QP and the variant unit is repeated twice. So, these three α-gliadins from the tested cultivars are most similar to the subtype 1.2–2 according to the number of repeat unit (2 after the dot) and the number of variant CD epitope present in the region (2 after the hyphen) under the type 1 group, by the rule proposed by Ozuna et al. (2015).
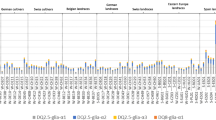
Potential cleavage sites in the selected region encompassing the multiple overlapping epitopes of the α-gliadins from the three Ethiopian durum wheat cultivars. Cleavage sites for proteases contained in human gastric and duodenal juices were predicted by PeptideCutter on the ExPASy Server (https://web.expasy.org/peptide_cutter/). Only those enzymes that cleave any site encompassing the selected region from p51 to p116 are presented
Abundance of the canonical and variant epitopes is different by the types of DQ2.5-glia epitopes. Only one epitope is in the canonical sequence among the total six DQ2.5-glia-α3 present in the three α-gliadins. No epitopes are in the canonical sequence among each of the three DQ2.5-glia-α1a, DQ2.5-glia-α1b, and DQ2.5-glia-α2 epitopes. The percentage of canonical epitope with respect to variants is higher in DQ2.5-glia-α1 and DQ2.5-glia-α3 than that in DQ2.5-glia-α2 in wheat α-gliadins (Ruiz-Carnicer et al. 2019). Regarding the DQ2.5-glia-α3, Ozuna et al. (2015) identified the variants of DQ2.5-glia-α3 epitope and named FR-, FP-, and FS-type according to the first two amino acids of their sequences. The most frequent FP-type-variant F1P2P3Q4Q5P6Y7P8Q9 (with R to P substitution at p2) that are present in most Triticum species (Ruiz-Carnicer et al. 2019) is also present in the α-gliadins from cvs. Denbi and Mukiye. The other forms of DQ2.5-glia-α3 variants that are present in the three α-gliadins are not the frequent variants reported in Triticum species (Ozuna et al. 2015; Ruiz-Carnicer et al. 2019; Mitea et al. 2010), and they also are the variants of FP-type. As the canonical DQ2.5-glia-α3 in the α-gliadin from cv. Assasa belongs to FR-type, FS-type is absent in the α-gliadins from the tested cultivars.
About the DQ2.5-glia-α1 and DQ2.5-glia-α2, the most frequent forms of variants reported in several Triticum species (Ruiz-Carnicer et al. 2019; Mitea et al. 2010) are absent in the α-gliadins from the tested cultivars. The substitution of the deamidation target residue Q to R at p5 in DQ8-α-glia is unique as all the deamination target Q residues are strictly conserved in all the other variant epitopes (Table 2). Deamidation of the target residue Q by tTG increases affinity of the peptides that bind to HLA-DQ2.5 or -DQ8, potentiating amplification of gluten-specific T cell response (Koning 2012).
In most cases, a single substitution of amino acid residue in the canonical sequence eliminate T cell proliferation and the binding capacity of the epitopes to DQ2 (Ellis et al. 2003; Ruiz-Carnicer et al. 2019). However, in several single or dual substitutions, T cell proliferation and the binding capacity of the epitopes to DQ2 are little affected. For example, in single substitutions, such as at P3 to S in DQ2.5-glia-α1, at P8 to S in DQ2.5-glia-α2, and in dual substitutions, such as at R2 to L together with at Q5 to L and at R2 to P together with at Q5 to L in DQ2.5-glia-α3, T cell proliferation and the binding capacity of the variant epitopes are either similar to or higher than those of their canonical epitopes (Ruiz-Carnicer et al. 2019). Among the variants observed in this study, only the variant with a substitution at R2 to P in DQ2.5-glia-α3 could retain significant levels of T cell proliferation and the binding capacity to DQ2 (Ruiz-Carnicer et al. 2019). In other words, it is suggested that most of the substitutions in the epitope sequences in the α-gliadins from the tested cultivars may hinder the variant epitopes from stimulating T cells, thereby contributing to significantly lower immunogenic response of the α-gliadins.
Variations in protease cleavage sites in the region harboring multiple CD-toxic epitopes
Differential SNPs and InDels in the α-gliadins from the test cultivars which produce variant epitopes with up to four amino acid substitutions in the major canonical CD-epitopes (Table 2) also result in significant variations in protease cleavage sites as seen in the region from p51 to p116, that harbor multiple immunotoxic CD epitopes (Fig. 3). Prediction for cleavage-site positions reveals a similar difference as seen in the variant CD epitopes. Patterns of the cleavage-site positions are generally identical in the α-gliadins from Denbi and Mukiye. Basically, there are a single cleavage site each for chymotrypsins and elastases, and triple sites for pepsins. No cleavage site is anticipated for all the enzymes included in prediction in the two copies of DQ2.5-glia-α3 epitopes. Only one cleavage position is predicted for the overlapping variant DQ2.5-glia-α1a and DQ2.5-glia-α1b. On the contrary, in the α-gliadin from cv. Assasa, multiple additional sites are predicted for cleavage by various types of endo-peptidases while one site for elastase, which is in the other two cultivars, is predicted to be lost. As a result, all the epitope sequences are cleaved by multiple enzymes at up to five positions.
This difference in protease cleavage sites in the three α-gliadins suggests contrasting outcomes for the expected peptides from the region after digestion by proteases in gastric and duodenal juices. Peptides which may harbor the p31–43 and DQ2.5-glia-α3 epitope intact are clearly expected from the α-gliadins from cvs. Denbi and Mukiye. Peptides containing a significantly long part of the variant 33-mer are also expected in the α-gliadins from the two cultivars. As outcomes from partial cleavages, therefore, peptides harboring the entire variant 33-mer cannot be ruled out to be produced. On the contrary, it is hardly expected from the α-gliadin of cv. Assasa to get any peptide that harbor a significant portion of either the p31-43 or the 33-mer.
α-Gliadins are generally inaccessible to human proteases of the gastrointestinal tract due largely to the high proline and glutamine content. It is well documented that proteolytically stable proline-rich peptides play a critical role in immunogenic response to α-gliadins in CD patients. Most notable ones among them are the p31–43 and the 33-mer harboring one or more major immunotoxic CD epitopes (Shan et al. 2002). Recently, the pivotal roles of the p31–43 and the 33-mer peptides in triggering CD are related to the oligomerization, via their selective abilities to self-organize in solution, and followed by structural transformation of the oligomerized peptides (Amundarain et al. 2019; Castro et al. 2019). One of the alternative therapies for CD patients is dietary supplementation of enzyme preparations having capacity to digest the toxic peptides in the stomach. Endopeptidases exhibiting potent activities to degrade the toxic gluten peptides in the gastric and upper intestinal tracts have been discovered in various organisms and suggested to be used to develop as oral supplement to support gluten-free diet and protect from unintentional gluten exposures (Cavaletti et al. 2019). Therefore, genotypes expressing gliadins with multiple cleavage sites for various proteases inside or surrounding the toxic epitopes could render practical advantages of reducing immunogenic risks from wheat consumption.
Significant conformational variations in the N-terminal region
Among the cysteine residues in the three α-gliadins, three to four are in CND1 and the remaining two in CND2. Intramolecular disulfide connectivity is predicted to be 1–2, 3–5 for the α-gliadin from cv. Denbi, 1–4, 2–6, 3–5 for the one from cv. Mukiye, and 1–5, 2–6, 3–4 for the one from cv. Assasa, respectively. Different connectivities among the α-gliadins are reflections of the differences in the number and position of cysteine residues and in the residues surrounding each of the cysteine residues (Figs. 1 and 4). Primarily, disulfide bonds are necessary to correct the folding and stability of the protein structure. The contrasting conformational difference between the N- and C-terminal regions of γ-gliadin is significantly attributed to the presence of disulfide bonds only in the C-terminal region (Sahli et al. 2019).
Structural features predicted for the α-gliadins from the three Ethiopian durum wheat cultivars. S–S, 2nd structure, and IUPred indicate disulfide connectivity, three-state secondary structure, and disordered protein regions, respectively. Amino acid sequence of the α-gliadin from each cultivar is in each rectangle on the top. Positions of CD-epitopes and cysteine residues are indicated on the upper part of the rectangle. The canonical CD-epitope is squared. For cysteine connectivity, each cysteine residue is denoted by a numerical number from one to six by the position order from the N-terminal end. In the three-state secondary structure cartoons, a slashed box and a broken line indicate α-helix and coil, respectively. The hydropathy is plotted on a scale by Kyte and Doolittle from -4.5 to 4.5. IUPred2A is plotted on a scale from 0 to 1, where disordered region (gray area) scores greater than 0.5
α-Gliadins are monomeric in their native state; thus, cysteine residues only form intramolecular disulfide bonds. However, during cooking and baking, these cysteine residues might form variable inter- and intramolecular disulfide bonds, thus contributing to the functionality of wheat flour (Delcour and Hoseney, 2010).
Predictions on secondary structures suggest the three α-gliadins are mainly composed of coils and α-helixes. The proportions of coil and α-helix are about 63% and 37% in the α-gliadin from cv. Denbi, and about 56% and 44% in the α-gliadin from cvs. Mukiye and Assasa, respectively, revealing a considerable difference among the three α-gliadins. The region-specific difference is also distinct in a conformational ensemble in the N- and C-terminal regions. The N-terminal is predominantly composed of coils, conversely, the C-terminal is mainly of α-helixes with stretches of coil interspersed between the helixes (Fig. 4).
Similarly, hydropathy profiles also reveal contrasting features in the two regions of the three α-gliadins: the N-terminal chiefly being hydrophilic, while the C-terminal being hydrophobic and hydrophilic alternating in stretches along the region (Fig. 4).
Generally, the N-terminal region is predicted to be predominantly disordered, while most of the C-terminal region to be ordered by IUPred2 scores, suggesting the three α-gliadins to be partially disordered proteins. Interestingly, a significant difference is indicated in the disordered N-terminal region among the three α-gliadins, specifically at the stretches of residues p45–51 and p76–85. Average IUPred2 score for the residues at p45-51 is 0.76, 0.51, and 0.49 for the α-gliadins from cvs. Denbi, Mukiye and Assasa, respectively. The average score at p76-85 is 0.84, 0.82, and 0.47 for the α-gliadins from cvs. Denbi, Mukiye and Assasa, respectively. Consequently, the N-terminal of the α-gliadins from cv. Denbi is predicted to be a highly disordered domain. However, the disordered N-terminal domain is predicted to be interrupted by a short ordered-stretch at p47–50 in the α-gliadin from cv. Mukiye, and by the two ordered patches at p45–51 and p76–85 in the α-gliadin from cv. Assasa. The variation at p76–85 is particularly intriguing as the section is a core part of the region harboring the overlapping toxic epitopes DQ2.5-glia-α1a and DQ2.5-glia-α1b and also of the 33-mer.
The NCPR plots of the three α-gliadins also reveal a characteristic difference in the polarity at the N- and C-terminal domains (Fig. 5). Essentially, four to five patches of positive and negative net charge clusters are distributed in the C-terminal region in the three α-gliadins. Nonetheless, the NCPR plots of the N-terminal region show a significant difference in the three α-gliadins. In the α-gliadins from cvs. Denbi and Mukiye, only three patches of positive net charge clusters are distributed near the end of the region. Conversely, in the α-gliadin from cv. Assasa, four each of positive and negative clusters are scattered at the region with the three negative ones at the region encompassing p45–51and p76–85. In fact, proportions of order-promoting amino acids (Campen et al. 2008) at the two sections are strikingly different among the three α-gliadins. At the p76–85, the proportion is 10%, 20%, and 40%, respectively, in the α-gliadins from cvs. Denbi, Mukiye and Assasa, respectively (Figs. 2 and 5).
The ordered state of the C-terminal region of γ-gliadins is significantly attributed to attractive electrostatic interactions among the unevenly distributed oppositely charged amino acids in the region (Sahli et al. 2019; Uversky 2011). An uneven charge distribution along the fraction of a protein may promote directional interactions (Nott et al. 2015), which may cause a conformational transition in the region. Therefore, the predicted intermittent sectional transitions to ordered state in the N-terminal region in the α-gliadin from cv. Assasa could be a result of accumulated mutations that caused a few deletions and several substitutions of amino acids. For example, compared to the α-gliadin from cv. Denbi, there are three deletions and two P to L and one S to L substitutions, resulting in increase in the proportion of order-promoting amino acids, especially at p76–85, where toxic CD epitopes overlap. These substitutions may alter self-organizing ability of the peptides harboring toxic epitopes, thus hampering the oligomerization and structural transformation required to trigger CD response (Amundarain et al. 2019; Castro et al. 2019).
Conclusion
The three α-gliadins from Ethiopian durum wheat share all the structural common features previously reported in wheat α-gliadins. However, they show significant variations in amino acid sequences along the peptides, especially at the region including the p31–43 and the 33-mer, which are well known for their CD-toxic immune responses. Most significant variations are two to three deletions of amino acids and several amino acid substitutions from the disorder-promoting to order-promoting ones, such as P to L and S to L. These changes cause CD-toxic epitopes, such as DQ2.5-glia-α1, DQ2.5-glia-α2, DQ2.5-glia-α3, DQ8-α-glia, to vary significantly from their canonical ones except a DQ2.5-glia-α3. Varietal difference in these changes lead to a significant variation in accessibility to digestive proteases and conformational features, especially at the N-terminal region. The α-gliadin from cv. Assasa is most interesting in several aspects; it has mostly variant CD epitopes that are highly accessible to multiple digestive proteases and the disordered N-terminal region interrupted by the two interspersed-ordered sections that may alter oligomerization of the peptides. It is interesting to investigate whether these features could render cv. Assasa comparative advantages for low allergenicity. Lists of important indices for the selection of hypoallergenic α-gliadin genetic sources may include sequence variability at the epitope motifs, susceptibility to proteases in human digestive systems, and peptides’ ability of self-organization.
References
Amundarain MJ, Herrera MG, Zamarreño F, Viso JF, Costabel MD, Dodero VI (2019) Molecular mechanisms of 33-mer gliadin peptide oligomerisation. Phys Chem Chem Phys 21:22539–22552. https://doi.org/10.1039/c9cp02338k
Anderson OD, Greene FC (1997) The α-gliadin gene family. II. DNA and protein sequence variation, subfamily structure, and origins of pseudogenes. Theor Appl Genet 95:59–65. https://doi.org/10.1007/s001220050532
Arentz-Hansen H, McAdam SN, Molberg O, Fleckenstein B, Lundin KEA et al (2002) Celiac lesion T-cells recognize epitopes that cluster in regions of gliadins rich in proline residues. Gastroenterol 123:803–809. https://doi.org/10.1053/gast.2002.35381
Asledottir T, Rehman R, Mamone G, Picariello G, Devold TG, Vegarud GE, Røseth A, Lea TE, Halstensen TS, Ferranti P, Uhlen AK (2020) Ancestral wheat types release fewer celiac disease related T cell epitopes than common wheat upon ex vivo human gastrointestinal digestion. Foods 9:1173. https://doi.org/10.3390/foods9091173
Asri N, Rostami-Nejad M, Anderson RP, Rostami K (2021) The gluten gene: unlocking the understanding of gluten sensitivity and intolerance. Appl Clin Genet 11(14):37–50. https://doi.org/10.2147/TACG.S276596
Battais F, Mothes T, Moneret-Vautrin DA, Pineau F, Kanny G, Popineau Y et al (2005) Identification of IgE-binding epitopes on gliadins for patients with food allergy to wheat. Allergy 60:815–821. https://doi.org/10.1111/j.1398-9995.2005.00795.x
Bromilow S, Gethings L, Langridge J, Shewry P, Buckley M, Roley M, Mills ENC (2017) Comprehensive proteomic profiling of wheat gluten using a combination of data-independent and data-dependent acquisition. Front Plant Sci 7:2020. https://doi.org/10.3389/fpls.2016.02020
Campen A, Williams RM, Brown CJ, Meng J, Uversky VN, Dunker AK (2008) TOP-IDP-scale: a new amino acid scale measuring propensity for intrinsic disorder. Protein Pept Lett 15(9):956–963. https://doi.org/10.2174/092986608785849164
Castro MFG, Miculán E, Herrera MG, Ruera C, Perez F, Prieto ED, Barrera E, Pantano S, Carasi P, Chirdo FG (2019) p31–43 gliadin peptide forms oligomers and induces NLRP3 inflammasome/caspase 1-dependent mucosal damage in small intestine. Front Immunol 10:31. https://doi.org/10.3389/fimmu.2019.00031
Cavaletti L, Taravella A, Carrano L et al (2019) E40, a novel microbial protease efficiently detoxifying gluten proteins, for the dietary management of gluten intolerance. Sci Rep 9:13147. https://doi.org/10.1038/s41598-019-48299-7
Chen FG, Xu CH, Chen MZ, Wang YH, Xia GM (2008) A new alpha-gliadin gene family for wheat breeding: somatic introgression line II-12 derived from Triticum aestivum and Agropyron elongatum. Mol Breed 22:675–685. https://doi.org/10.1007/s11032-008-9208-0
Ciccocioppo R, Di Sabatino A, Corazza GR (2005) The immune recognition of gluten in coeliac disease. Clin Exp Immunol 140:408–416. https://doi.org/10.1111/j.1365-2249.2005.02783.x
Das RK, Pappu RV (2013) Conformations of intrinsically disordered proteins are influenced by linear sequence distributions of oppositely charged residues. Proc Natl Acad Sci USA 110(33):13392–13397. https://doi.org/10.1073/pnas.1304749110
Delcour JA, Hoseney RC (2010) Proteins of cereal. In: Delcour JA, Hoseney RC (eds) Principles of cereal science and technology. American Association of Cereal Chemists, St. Paul, pp 23–80
Di Sabatino A, Vanoli A, Giuffrida P, Luinetti O, Solcia E, Corazza GR (2012) The function of tissue transglutaminase in celiac disease. Autoimmun Rev 11(10):746–753. https://doi.org/10.1016/j.autrev.2012.01.007
Dørum S, Bodd M, Fallang L-E, Bergseng E, Christophersen A, Johannesen MK, Qiao SW, Stamnaes J, de Souza GA, Sollid LM (2014) HLA-DQ Molecules as affinity matrix for identification of gluten T cell epitopes. J Immunol 193:4497–4506. https://doi.org/10.4049/jimmunol.1301466
Dubois B, Bertin P, Mingeot D (2016) Molecular diversity of α-gliadin expressed genes in genetically contrasted spelt (Triticum aestivum ssp. spelta) accessions and comparison with bread wheat (T. aestivum ssp. aestivum) and related diploid Triticum and Aegilops species. Mol Breed 36:152. https://doi.org/10.1007/s11032-016-0569-5
Ellis HJ, Pollock EL, Engel W et al (2003) Investigation of the putative immunodominant T cell epitopes in coeliac disease. Gut 52:212–217. https://doi.org/10.1136/gut.52.2.212
Ferrè F, Clote P (2005) DiANNA: a web server for disulfide connectivity prediction. Nucleic Acids Res 33(s2):W230–W232
Hailegiorgis D, Lee CA, Yun SJ (2017) Allelic variation at the gliadin coding loci of improved Ethiopian durum wheat cultivars. J Crop Sci Biotech 20:287–293. https://doi.org/10.1007/s12892-017-0106-0
Holehouse AS, Ahad J, Das RK, Pappu RV (2014) CIDER: classification of intrinsically disordered ensemble regions. Biophys J 108(2):228a. https://doi.org/10.1016/j.bpj.2014.11.1260
Huo N, Zhu T, Altenbach S, Dong L et al (2018) Dynamic evolution of α-gliadin prolamin gene family in homeologous genomes of hexaploid wheat. Sci Rep 8(1):5181. https://doi.org/10.1038/s41598-018-23570-5
Jones DT (1999) Protein secondary structure prediction based on position-specific scoring matrices. J Mol Biol 292(2):195–202. https://doi.org/10.1006/jmbi.1999.3091
Kim CY, Quarsten H, Bergseng E, Khosla C, Sollid LM (2004) Structural basis for HLA-DQ2- mediated presentation of gluten epitopes in celiac disease. Proc Natl Acad Sci USA 101(12):4175–4179. https://doi.org/10.1073/pnas.0306885101
Koning F (2012) Celiac disease: quantity matters. Semin Immunopathol 34(4):541–549. https://doi.org/10.1007/s00281-012-0321-0
Mészáros B, Erdős G, Dosztányi G (2018) IUPred2A: context-dependent prediction of protein disorder as a function of redox state and protein binding. Nucleic Acids Res 46(W1):W329–W337. https://doi.org/10.1093/nar/gky384
Mitea C, Salentijn EMJ, van Veelen P, Goryunova SV, van der Meer IM et al (2010) A universal approach to eliminate antigenic properties of alpha-gliadin peptides in celiac disease. PLoS ONE 5(12):e15637. https://doi.org/10.1371/journal.pone.0015637
Molberg Ø, Uhlen AK, Jensen T, Flaete NS, Fleckenstein B, ArentzHansen H, Raki M, Lundin KEA, Sollid LM (2005) Mapping of gluten T-cell epitopes in the bread wheat ancestors: implications for celiac disease. Gastroenterology 128:393–401. https://doi.org/10.1053/j.gastro.2004.11.003
Nakamura R, Sakai S, Adachi R, Hachisuka A, Urisu A et al (2013) Tissue transglutaminase generates deamidated epitopes on gluten, increasing reactivity with hydrolyzed wheat protein-sensitized IgE. J Allergy Clin Immunol 132:1436–1438. https://doi.org/10.1016/j.jaci.2013.07.017
Nott TJ, Petsalaki E, Farber P, Jervis D, Fussner E, Plochowietz A, Craggs TD, Bazett-Jones DP, Pawson T, Forman-Kay JD, Baldwin AJ (2015) Phase transition of a disordered nuage protein generates environmentally responsive membraneless organelles. Mol Cell 57:936–947. https://doi.org/10.1016/j.molcel.2015.01.013
Ozuna CV, Iehisa JCM, Giménez MJ, Alvarez JB, Sousa C, Barro F (2015) Diversification of the celiac disease α-gliadin complex in wheat: a 33-mer peptide with six overlapping epitopes, evolved following polyploidization. Plant J 82(5):794–805. https://doi.org/10.1111/tpj.12851
Palosuo K, Varjonen E, Nurkkala J, Kalkkinen N, Harvima R, Reunala T et al (2003) Transglutaminase-mediated cross-linking of a peptic fraction of omega-5 gliadin enhances IgE reactivity in wheat-dependent, exercise-induced anaphylaxis. J Allergy Clin Immunol 111:1386–1392. https://doi.org/10.1067/mai.2003.1498
Payne P (1987) Genetics of wheat storage proteins and the effect of allelic variation on bread-making quality. Ann Rev Plant Physiol 38:141–153. https://doi.org/10.1146/annurev.pp.38.060187.001041
Prandi B, Tedeschi T, Folloni S, Galaverna G, Sforza S (2017) Peptides from gluten digestion: a comparison between old and modern wheat cultivars. Food Res Int 1:92–102. https://doi.org/10.1016/j.foodres.2016.11.034
Qiao SW, Ráki M, Gunnarsen KS, Løset GÅ, Lundin KE, Sandlie I, Sollid LM (2011) Posttranslational modification of gluten shapes TCR usage in celiac disease. J Immunol 187:3064–3071. https://doi.org/10.4049/jimmunol.1101526
Radivojac P, Iakoucheva LM, Oldfield CJ, Obradovic Z, Uversky VN, Dunker AK (2007) Intrinsic disorder and functional proteomics. Biophys J 92(5):1439–1456. https://doi.org/10.1529/biophysj.106.094045
Rashtak S, Murray JA (2012) Review article: coeliac disease, new approaches to therapy. Aliment Pharmacol Ther 35:768–781. https://doi.org/10.1111/j.1365-2036.2012.05013.x
Ruiz-Carnicer Á, Comino I, Segura V et al (2019) Celiac immunogenic potential of α-gliadin epitope variants from Triticum and Aegilops species. Nutrients 11(2):220. https://doi.org/10.3390/nu11020220
Sahli L, Renard D, Solé-Jamault V, Giuliani A, Boire A (2019) Role of protein conformation and weak interactions on γ-gliadin liquid-liquid phase separation. Sci Rep 9:13391. https://doi.org/10.1038/s41598-019-49745-2
Schuppan D, Junker Y, Barisani D (2009) Celiac disease: from pathogenesis to novel therapies. Gastroenterology 137:1912–1933. https://doi.org/10.1053/j.gastro.2009.09.008
Shan L, Molberg O, Parrot I, Hausch F, Filiz F, Gray GM, Sollid LM, Khosla C (2002) Structural basis for gluten intolerance in celiac sprue. Science 297:2275–2279. https://doi.org/10.1126/science.1074129
Shewry PR, Tatham AS (1997) Disulphide bonds in wheat gluten proteins. J Cereal Sci 25:207–227
Shewry PR, Tatham AS (2016) Improving wheat to remove coeliac epitopes but retain functionality. J Cereal Sci 67:12–21. https://doi.org/10.1016/j.jcs.2015.06.005
Shewry PR, Halford NG, Lafiandra D (2003) Genetics of wheat gluten proteins. Adv Genet 49:111–184. https://doi.org/10.1016/s0065-2660(03)01003-4
Sollid LM, Qiao SW, Anderson RP, Gianfrani C, Koning F (2012) Nomenclature and listing of celiac disease relevant gluten T-cell epitopes restricted by HLA-DQ molecules. Immunogenetics 64(6):455–460. https://doi.org/10.1007/s00251-012-0599-z
Srinivasan B, Focke-Tejkl M, Weber M, Pahr S, Baar A, Atreya R et al (2015) Usefulness of recombinant γ-gliadin 1 for identifying patients with celiac disease and monitoring adherence to a gluten-free diet. J Allergy Clin Immunol 136:1607–1618. https://doi.org/10.1016/j.jaci.2015.04.040
Uversky VN (2011) Intrinsically disordered proteins from A to Z. Int J Biochem Cell Biol 43:1090–1093. https://doi.org/10.1016/j.biocel.2011.04.001
Vader LW, Stepniak DT, Bunnik EM et al (2003) Characterization of cereal toxicity for celiac disease patients based on protein homology in grains. Gastroenterology 125:1105–1113. https://doi.org/10.1016/s0016-5085(03)01204-6
Van Herpen TWJM, Goryunova SV, Van der Schoot J, Mitreva M, Salentijn E, Vorst O et al (2006) Alpha-gliadin genes from the A, B, and D genomes of wheat contain different sets of celiac disease epitopes. BMC Genome 7:1. https://doi.org/10.1186/1471-2164-7-1
Wang DW, Li D, Wang J, Zhao Y, Wang Z, Yue G, Liu X, Qin H, Zhang K, Dong L, Wang D (2017) Genome-wide analysis of complex wheat gliadins, the dominant carriers of celiac disease epitopes. Sci Rep 7:44609. https://doi.org/10.1038/srep44609
Acknowledgements
This work was supported by a grant from Regional Subgenebank Support Program of Rural Development Administration, Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Hailegiorgis, D., Seid, E., Lee, C.A. et al. Variations in immunodominant epitope and molecular conformation of alpha-gliadins in elite Ethiopian durum wheat cultivars. J. Crop Sci. Biotechnol. 25, 325–336 (2022). https://doi.org/10.1007/s12892-021-00134-0
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12892-021-00134-0