Abstract
Biodiversity and wild populations are globally threatened by a wide range of actors. The genus Iguana, widely distributed throughout the Americas, is under threat by invasive species, hybridization, the global pet trade, and habitat destruction. This holds especially true for the insular lineages, with the Critically Endangered I. delicatissima having experienced a > 75% range decrease, primarily through hybridization with non-native iguanas. We collated published microsatellite data and genotyped samples from new localities to construct a distribution-wide Iguana dataset built from 190 individuals for 17 microsatellite loci. This enabled us to identify patterns of genetic differentiation within and among populations, and identify key loci and private alleles for use in conservation management. Our analyses reveal clear separation between I. delicatissima and the I. iguana complex, highlighting the power of eight key microsatellite loci for the study of hybridization dynamics. Genetic differentiation within I. delicatissima identifies four clusters that aid decision making for conservation management action. Within the I. iguana complex, we increase mainland localities by 11-fold and recover 3.5 × more alleles across all loci than previously known. Overall, we identify 112 (48% private) and 76 (25% private) alleles for mainland and island lineages, respectively. We highlight loci sets to identify (1) non-native or hybrid iguanas in insular populations and their genetic origin, and (2) genetic origin of insular iguanas in the global pet trade. Overall, we provide a reference for Iguana microsatellite loci in order to allow standardization and comparison among studies, aiding broader assessment of research and conservation hypotheses.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Genetic tools are broadly utilized to gain understanding of wild populations and their conservation needs; e.g., examining metapopulation dynamics (Rowe et al. 2003), tracking illegal collection and trade (Coetzer et al. 2017), and characterizing genetic variation and inbreeding (Luís et al. 2007; Engelhardt et al. 2017). Although development of novel techniques continues, 2nd generation methodologies (microsatellites) are still broadly applied (Vashistha et al. 2020) as these are overall cheap to obtain data from, are preferred for small sample sizes, and can be processed and analyzed rapidly. Especially within conservation science, where funding is limited and results tend to be time sensitive, microsatellites are an important go-to genetic tool.
Among Iguaninae, Iguana is the widest ranging genus, with mainland populations occurring from central Mexico through to central Brazil, and widespread presence on Caribbean islands (Belezian, Colombian, Honduran, Mexican, Nicaraguan, and Tobago and Trinidad islands, as well as in the Southern and Lesser Antilles; Iguana Taxonomy Working Group 2016; Hedges et al. 2019). Taxonomically, this genus consists of two well defined lineages, I. delicatissima and the I. iguana complex. Within the latter, hypotheses of species boundaries are unresolved, though are currently under investigation, see Table 1 for nomenclature and geographic ranges (Stephen et al. 2013; Breuil et al. 2019, 2020, submitted, van den Burg et al. in prep.).
As is true for many Iguanids (IUCN 2020), insular lineages within Iguana are highly endangered. The main threat to these species is the continuous spread and recurrent incursion of non-native iguanas, both through natural events and, since at least the 19th Century (Breuil 2003, 2013), anthropogenic assistance, which is expected to continue in the absence of stricter biosecurity measures (Falcón et al. 2012; Bock et al. 2018; van den Burg et al. 2018a, 2020a, 2020b, 2021b; Breuil et al. 2019, 2020; Knapp et al. 2020). These invasive, non-native iguanas have profoundly impacted native insular iguana populations throughout the Lesser Antilles, as the former have larger clutch and body sizes and show higher aggressiveness than native iguanas (van den Burg et al. 2018a; van Wagensveld and van den Burg 2018; Breuil et al. 2019). These incursions from different nonnative sources have resulted in region-wide hybridization, best documented for the Critically Endangered I. delicatissima (Vuillaume et al. 2015; van den Burg et al. 2018a, 2018b; Pounder et al. 2020), and to a lesser extent for the distinct lineages within the I. iguana complex (Breuil et al. 2019). Given this lengthy, continuous period of hybridization and subsequent within-region movement of non-native iguanas (Censky et al. 1998), morphologically distinguishing between native and (hybridized) non-native iguanas is often not straightforward (Breuil 2013; Vuillaume et al. 2015). This status quo clearly demonstrates the urgent need for robust genetic tools to distinguish native from non-native (or hybrid) animals. Additionally, the increased activity in illegal trade of iguanas from Lesser Antillean islands (Noseworthy 2017; van den Burg and Weissgold 2020) further highlights the need for island-level identification.
Within Iguana, microsatellite research has focused on a wide range of topics; invasion biology, population dynamics, as well as conservation (Valette et al. 2013; Vuillaume et al. 2015; Martin Judson et al. 2018; van den Burg et al. 2018b; De Jesús Villanueva et al. 2021). However, different genetic laboratories have not standardized their data sets and employ different loci sets (Table 2), limiting the vital ability to compare results, efficiently build upon prior work, and improve our understanding of these vulnerable populations. This knowledge gap of variation from across the genus’ range limits our ability to meaningfully assess population structure within lineages, and hinders the essential identification of Iguana hybrids and conservation of endangered lineages. From the perspective of hybridization issues, the research community currently lacks an understanding of allelic variation among known Iguana species and clades because data from native samples are almost entirely limited to island populations. The one exception is a 2015 study by Vuillaume et al. that included a single sample from each of two mainland localities with known origin. Hence, a genus-wide dataset is urgently needed in order to provide a foundation upon which all stakeholders can build, and use to guide conservation management decisions.
Materials and methods
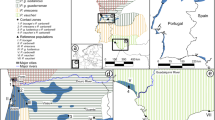
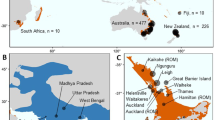
The microsatellite loci set that has been most implemented among Iguana studies is that of Valette et al. (2013). We compiled data generated from all or a subset of these 25 loci (Valette et al. 2013; Vuillaume et al. 2015; Breuil et al. 2019, 2020, submitted). Furthermore, we collected data from new mainland (23) and insular (5) localities for a total of 53 native iguanas, to represent a wide-spread Iguana dataset (Fig. 1, Table 1). We also included six morphologically identified hybrids between I. delicatissima and non-native iguanas; two samples from St. Barths, and Basse Terre and Grande Terre (Guadeloupe) (Vuillaume et al. 2015).
Map showing sample localities and current taxonomic status with Iguana: Inset shows the Lesser Antilles. Further geographic details for each lineage can be found in Table 1. ‘aff.’ indicates the uncertain taxonomic status, which is undergoing assessment (van den Burg et al. in prep)
Laboratory procedures were performed at Labofarm-GenIndexe (France) where PCR settings and primers outlined by Valette et al. (2013) were used to amplify 17 loci (named as IgdL1-25 or L1-25 in previous studies; Valette et al. 2013; Vuillaume et al. 2015). Subsequently, PCR products were ran with Thermo ROX500 and analyzed using an ABI 3130 XL; alleles were next scored using Genemapper, while samples IGU039–43 were used to standardize allele sizes with aforementioned studies (see van den Burg et al. 2021a).
Data analyses
After sample processing, we retained specimens with ≥ 80% scored loci for data analyses (for sample sizes used in analyses see Table 3 and figure captions). Considering our sample-set characteristics, only one population, from Ilet Chancel (29 samples), was suitable to test the presence of null alleles. Following Dabrowski et al (2014), we implemented both Genepop and micro-checker software to assess the presence of null alleles (Raymond and Rousset 1995; van Oosterhout et al. 2004). The remaining sample-set included groups not amenable to this question as they were comprised of; (1) wide geographic sampling within lineages (producing a Wahlund Effect), or (2) low sample size of breeding populations (non-representative of existing variation), or (3) impacted by recent bottlenecks (strong effects of genetic drift).
Data were split into two sets, as previous studies have used different loci (Table 2). Our 12-loci dataset includes all native Iguana locations and the six Iguana-delicatissima hybrids, and our 17-loci dataset includes all native locations from the I. iguana complex. For these datasets we then calculated relative allelic richness using the GenPopReport package within the R environment (R Core Team 2019).
Allele variation across all samples was assessed and visualized in color-coded tables to identify genetic variation within and between Iguana lineages (see Results). Within these tables we also include allelic data from the non-native and hybrid iguanas from Vuillaume et al. (2015) and Breuil et al. (2019).
Genetic differentiation and clustering were assessed using STRUCTURE as implemented within GenoDive 3 (Pritchard et al. 2000; Meirmans 2020). For the 12-loci dataset, we applied this methodology to assess its utility to successfully assign native pure and hybrid specimens, as well as to assess the genetic structure among I. delicatissima populations. For the 17-loci dataset, we assessed genetic structuring within the I. iguana complex, for which we standardized sample sizes among lineages by reducing the sample size of I. melanoderma and I. insularis to those of the other lineages by randomly sampling while maintaining sample size-island proportions. STRUCTURE runs used identical parameters including an admixture model, correlated allele frequencies, and without linkage. Runs composed of 10 repeats for K ranges 1–10 using 100 K burnin and 500 K MCMC repeats. Subsequently, we assessed and visualized delta-K estimates for different K values using Structure Harvester (Evanno et al. 2005; Earl and vonHoldt 2012), and employed distruct to create final structure barplots (Rosenberg 2004). Lastly, within R through the loadingplot() command in the adegenet package, we assessed which loci and alleles most contribute to differentiation of I. delicatissima and non-delicatissima iguanas through a discriminant principal component analysis (DAPC; Jombart 2008).
Results
For the 12-loci dataset, mean allelic richness among I. delicatissima populations ranges between 1.03–1.16, and between 1.05–1.43 within the I. iguana complex. For the 17-loci dataset, mean allelic richness shows higher values with a similar pattern among the I. iguana lineages, 1.06–1.98 (Table 3). Within the complex, I. melanoderma and I. insularis show lower mean allelic richness compared to taxonomic units that mostly have a mainland range, though Iguana aff. iguana Clade I is here represented only by island populations.
Null-allele detection tests show inconsistent results with micro-checker suggesting presence for five loci (L3, L5, L12, L18, L24), whereas Genepop only for L20. We retain all loci in further discussion given that no single locus was flagged by both methods. We recommend these analyses be performed on definable populations that have been adequately sampled and meet the a priori assumption of Hardy–Weinberg equilibrium.
Hybridization
Figure 2 provides an overview of results for hybridization purposes between native I. delicatissima and native non-delicatissima, as well as six hybrids. Structure results are visualized for K = 2, which clusters I. delicatissima and non-delicatissima separately, except for the six known hybrids (Fig. 2a). Surprisingly, assignment of Dominica and St. Eustatius samples to I. delicatissima was < 75% (Fig. 2a), likely due to their low sample sizes and alleles with proportionally higher representation in non-delicatissima samples; all samples in these two populations were fixed for unique I. delicatissima alleles. Visual inspection of patterns of allele variation and size range overlap between I. delicatissima and non-delicatissima reveals four loci with a complete lack of allele variation overlap, three of which include data from all I. delicatissima locations: L8, L18, and L25 (Fig. 2b). An additional two loci (L5 and L24) also indicate a high degree of usefulness for hybridization studies, but show a single allele overlap. Lastly, DAPC analysis identified a set of six loci with the highest contribution to differentiating I. delicatissima and non-delicatissima: L5, L8, L14, L18, L20 and L25 (Fig. 2c).
Genetic differentiation results to study hybridization in Iguana delicatissima, using 12 loci dataset. a Structure plot for pure I. delicatissima (67), pure non-delicatissima (117), and hybrid (6) samples, see Table 3; for K = 2. Asterisks indicate individuals from St. Eustatius and Dominica. b Overview of allele variation between I. delicatissima and non-delicatissima individuals, excluding hybrids. C DAPC loading contribution of loci and alleles for clustering between I. delicatissima – non-delicatissima, excluding hybrids
Iguana delicatissima
Within I. delicatissima, delta-K identified four clusters with highest support (Fig. 3b): (1) Ilet Chancel and St. Barths, (2) Dominica and St. Eustatius, (3) Petite Terre and La Désirade, and, (4) Guadeloupe. The first two groups contain islands not in close geographic proximity. For delta-K of 2 (but see Janes et al. 2017), assignment followed that of Ilet Chancel and St. Barths vs. all other islands. Allelic variation is highest for larger islands or those with high population sizes; Dominica, Guadeloupe, Petite Terre and Ilet Chancel (Table 3, Fig. 3c).
Genetic variation among Iguana delicatissima island populations. a Structure and b delta K plot for population differentiation among pure I. delicatissima individuals (67; Table 3), using 12 loci dataset; K = 4. c Allele variation among seven island populations for 20 loci; islands assigned to identical Structure cluster (a) have equal-tone colors
Iguana iguana complex
Genetic differentiation and allele variation within the I. iguana complex are presented in Fig. 4. Optimal delta-K support indicated four clusters, with sub-structural support for seven clusters (Fig. 4a-c). Allele variation patterns between mainland (generally non-native) and insular lineages highlight loci of interest for identifying the genetic origin of non-native iguanas within those insular lineages; L3, L5, L6, L8, L9, L14 L16, L17, L19, L20, L23–25 (Fig. 4d). Loci that appear to be useful for characterizing insular groups, differs per lineage; I. melanoderma, L6, L14, L20, L25; I. insularis, L6, L13, L14, L17, L20; I. aff. iguana Clade I, L2, L5, L16, L17, L20, L23–25.
Genetic variation within Iguana iguana complex for 17 microsatellite loci. a + b Structure and c delta K plots for population differentiation among seven taxonomic units; optimal K = 4, suboptimal K = 7. d Allele variation among seven taxonomic units. Sample sizes as in Table 3, except for I. insularis insularis (10), and I. melanoderma (10)
Discussion
Here we present a nuclear microsatellite database for Iguana that includes all current taxonomic groups with broad geographic representation across its range, demonstrating clear population genetic structure within I. delicatissima and the I. iguana complex. Furthermore, our results identify key sets of loci and private alleles useful in the study of conservation-hybridization issues and for narrowing down the geographic origin of commercially available pet iguanas and illegally transported individuals. This database fills an important gap given that nuclear loci sequence in prior work show low variability within the I. iguana complex (Stephen et al. 2013), prohibiting the fine scale variation at biparentally inherited loci required to address such issues. Overall, our work demonstrates the successful amplification of these markers and their utility within the Iguana genus. Importantly, the benefits of this database will increase dramatically with further input from the Iguana research community, which we promote through free availability on figshare (van den Burg et al. 2021a).
Assignment of loci and private alleles to address research questions is dependent on sampling scheme, both its geographical width and density, as well as sample size per population. Given the geographic range of Iguana, the coverage of the presented database is wide, though mainland sampling density is still low, with few samples per location. Notably, the database greatly increases coverage and lineage inclusion compared to Valette et al. (2013), which assigned 19 loci as useful for I. delicatissima hybridization studies based on one I. delicatissima (Ilet Chancel, Martinique), and one introgressed (I. delicatissima – I. iguana) population (Grande-Terre, Guadeloupe). Here, including data from almost all I. delicatissima populations (only lacking Anguilla), we identify 5–8 loci of specific use for identification of hybrids between I. delicatissima and the I. iguana complex (Fig. 2b, c). Although these loci are fewer in number, their identification is based on a significant data increase from throughout the genus’ range; allowing conservationists to target fewer, highly informative loci and lower research costs. Furthermore, the need to standardize allele sizes across different studies and laboratories is exemplified in our finding that the three samples from St. Eustatius rescored in this study had different allele size for all five loci compared to results in van den Burg et al. (2018b), shifting higher or lower by 5–22 bps, depending on the locus. Similarly, sample reanalyzes in the same laboratory can indicate allele size deviations, as data of several loci presented by Vuillaume et al. (2015) have since been updated (2–16 bps; Breuil et al. 2019). Lastly, with ongoing conservation priorities aimed at insular lineages, the characterization of their allele diversity will carry on, though we address the importance of characterizing new mainland localities. This continuous data collection highlights the need for a “live” dynamic database that provides a crucial, up-to-date, Iguana conservation tool.
The identification and continued usefulness of microsatellites in research on wild populations lies in their rapid mutation rate (Schlötterer 2000), demonstrating their use in studies of recently separated populations (Kuehn et al. 2007), but highlighting their poor informative value for long evolutionary timelines (Carsten et al. 2013). Indeed, this high mutation rate prevents the determination of shared alleles as synapomorphic or homoplasious, when comparing highly divergent lineages. This is evident in our database for shared alleles (e.g., L2) between I. iguana lineages and two I. delicatissima populations (St. Eustatius and Dominica) (Fig. 2a). Genetic samples from these I. delicatissima populations were obtained prior to first documented hybridization on both islands (van den Burg et al. 2018b, 2020a), though historic hybridization might have occurred. Another explanation for shared L2 alleles is independent mutation as might be expected after a long period of evolutionary divergence.
The Critically Endangered, Lesser Antillean iguana has seen a > 75% decline from its pre-Colombian range due to extirpations subsequent to human settlement, and more recently, hybridization with non-native iguanas of the I. iguana complex (van den Burg et al. 2018a). Given the extremely low mtDNA variation throughout the range of I. delicatissima (Martin et al. 2015), microsatellites are better suited to guide research and conservation (Vuillaume et al. 2015; Martin Judson et al. 2018; van den Burg et al. 2018b). Small populations will lose diversity quickly and randomly diverge due to founder effects and/or drift over short time periods (Wright 1931), which has been demonstrated in Iguanids as well (Colosimo et al. 2014; Welch et al. 2017). Thus, to maintain long term genetic variation within I. delicatissima it is essential that all island populations are genotyped. Our database builds on Vuillaume et al. (2015) by inclusion of two additional island populations, St. Eustatius and Dominica. Genetic clustering among currently sampled populations is not correlated with geographic proximity, given two identified clusters: St. Eustatius–Dominica, and St. Barths–Ilet Chancel (Fig. 3a). Their genetic similarity could originate from within-region dispersal facilitated by hurricanes, Amerindians or post-Colombian colonizers (Censky et al. 1998; Bryan et al. 2007; Bochaton et al. 2016). Although this database represents the most inclusive to date, we call for sampling and genotyping of the Anguilla population and those on isolated islets around St. Barths, as well as additional genotyping of samples from Dominica and La Désirade. These genetic data can furthermore help identify the remaining pure animals on Guadeloupe and Martinique, which are in need of urgent translocation to secure their genetic diversity. Lastly, these data and identified key loci will aid other conservation management decisions as identified by conservation action plans (e.g., translocations (Scott et al. 2020), captive breeding, and reintroductions (Knapp et al. 2014; Angin 2017; ANT/ATE/STENAPA 2018; Breuil 2021)).
We provide an initial assessment of allele variation among recently described lineages within the I. iguana complex. Optimal support was found for four genetic clusters, which grouped currently known lineages as (1) I. rhinolopha, (2) I. aff. iguana Clade IIA, (3) I. insularis, (4) I. iguana and I. melanoderma, and with an admixed assignment for I. aff. iguana Clade I (Fig. 4a, c). A suboptimal 7-cluster division distinguished I. iguana and I. melanoderma, as well as uniquely assigned I. aff. iguana Clade I (Fig. 4b, c). Allelic diversity indicated higher variation in mainland lineages and I. aff. iguana Clade I compared to solely insular lineages, with especially low diversity in I. i. sanctaluciae (Table 3). For vulnerable island populations (I. melanoderma, I. insularis, I. aff. iguana Clade I), we find three loci sets (4–8 loci) for origin assessment (see Results text for specifics). For the current dataset, we note the skewed sampling effort towards recently described insular lineages (Breuil et al. 20192020, submitted), and highlight the need for improved sampling of the mainland lineages. Although threats within the I. iguana complex are less studied compared to those in I. delicatissima, it is well-known that mainland populations can be locally threatened due to overharvesting, either for food or to supplement the pet trade (Stephen et al. 2011). More recently, hybridization between non-native iguanas and native island populations has been described, as well as illegal trade of I. insularis and I. melanoderma (Noseworthy 2017; Breuil et al. 2019; van den Burg and Weissgold 2020). Overall, the identified allelic variation (Fig. 4D) will aid in the discovery of non-native and hybrid iguanas and the geographic origin of these as well as of iguanas in the global commercial hobbyist trade.
In conclusion, this manuscript provides a reference for Iguana microsatellite loci in order to allow standardization and comparison among studies, aiding broader assessment of research and conservation hypotheses. Importantly, this work provides an essential tool for conservation of the Critically Endangered I. delicatissima, and taxa within the I. iguana complex. To ensure future comparability, we strongly urge that future studies utilize: (1) the same allele size ladder, (2) positive controls from localities with fixed alleles, and/or (3) samples scored in prior studies.
Data Availability
Data are freely available on figshare, https://doi.org/10.6084/m9.figshare.13584923.v1).
References
ANT/ATE/STENAPA (2018) Lesser Antillean Iguana Iguana delicatissima Conservation Strategy and Action Plan for the Northern Caribbean Sub-region (Anguilla, St. Barthélemy, St. Eustatius), 2018–2023. Anguilla National Trust, Agence Territoriale de l’Environnement and St. Eustatius National Parks Foundation
Angin B (2017) Plan National d’Actions pour le rétablissement de l’iguane des petites Antilles, Iguana delicatissima, 2018–2022. 69p + annexes
Bochaton C, Bailon S, Ineich I, Breuil M, Tresset A, Grouard S (2016) From a thriving past to an uncertain future: Zooarchaeological evidence of two millenia of human impact on a large emblematic lizard (Iguana delicatissima) on the Guadeloupe Islands (French West Indies). Quat Sci Rev 150:172–183
Breuil M (2003) In the footsteps of French Naturalists, a Battle of Iguanas and Improvements in Biodiversity, Islands and the Sea: Essays on Herpetological Exploration in the West Indies. Henderson RW, Powell R (eds.). Society for the Study of Amphibians and Reptiles, Ithaca (New York). Contrib to Herpetol 20:255–270.
Breuil M (2013) Caractérisation morphologique de l’iguane commun Iguana iguana (Linnaeus, 1758), de l’iguane des Petites Antilles Iguana delicatissima Laurenti, 1768 et de leurs hybrides. Bull Soc Herpetol Fr 147:309–346
Breuil M (2021) Les iguanes des Petites Antilles. Les espèces endémiques sur le déclin. Le Courrier De La Nature 326:27–33
Breuil M et al. (submitted) Iguana insularis (Iguanidae) from the Southern Lesser Antilles: an endemic partially hybridized species. Conserv Genet.
Breuil M, Vuillaume B, Schikorski D, Krauss U, Morton MN, Haynes P, Daltry JC, Corry E, Gaymes G, Gaymes J, Bech N, Jelic M, Grandjean F (2019) A story of nasal horns: Two new subspecies of Iguana Laurenti, 1768 (Squamata, Iguanidae) in Saint Lucia, St Vincent & the Grenadines, and Grenada (southern Lesser Antilles). Zootaxa 4608:201–232
Breuil M, Schikorski D, Vuillaume B, Krauss U, Morton MN, Corry E, Bech N, Jelic M, Grandjean F (2020) Painted black: Iguana melanoderma (Reptilia, Squamata, Iguanidae) a new melanistic endemic species from Saba and Montserrat islands (Lesser Antilles). ZooKeys 926:95–131
Bryan JJ, Gerber GP, Welch ME, Stephen CL (2007) Re-evaluating the taxonomic status of the Booby Cay Iguana, Cyclura carinata bartschi. Copeia 3:734–739
Bock B, Malone CL, Knapp C, Aparicio J, Avila-Pires TCS, Cacciali P, Caicedo JR, Chaves G, Cisneros-Heredia DF, Gutiérrez-Cárdenas P, Lamar W, Moravec J, Perez P, Porras LW, Rivas G, Scott N, Solórzano A, Sunyer J (2018) Iguana iguana. The IUCN Red List of Threatened Species. https://doi.org/10.2305/IUCN.UK.2018-1.RLTS.T174481A1414646.en
Carsten BC, Pelletier TA, Reid NM, Satler JD (2013) How to fail at species delimitation. Mol Ecol 22:4369–4383
Censky EJ, Hodge K, Dudley J (1998) Over-water dispersal of lizards due to hurricanes. Nature 395:556
Coetzer WG, Downs CT, Perrin MR, Willows-Munro S (2017) Testing of microsatellite multiplexes for individual identification of Cape Parrots (Poicephalus robustus): paternity testing and monitoring trade. PeerJ 5:e2900
Colosimo G, Knapp CR, Wallace LE, Welch ME (2014) Inferred vs realized patterns of gene flow: An analysis of population structure in the Andros Island Rock Iguana. PLoS ONE 9:e106963
Dabrowski MJ, Pilot M, Kruczyk M, Zmihorski M, Umer HM, Gliwicz J (2014) Reliability assessment of null allele detection: inconsistencies between and within different methods. Mol Ecol Res 14:361–373
De Jesús Villanueva CN, Falcón W, Velez-Zuazo X, Papa R, Malone CL (2021) Origin of the green iguana (Iguana iguana) invasion in the greater Caribbean Region and Fiji. Biol Invasions. https://doi.org/10.1007/s10530-021-02524-5
Earl DA, von Holdt BM (2012) Structure Harvester: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Engelhardt A, Muniz L, Perwitasari-Farajallah D, Widdig A (2017) Highly polymorphic microsatellite markers for the assessment of male teproductive skew and genetic variation in Critically Endangered Crested Macaques (Macaca nigra). Int J Primatol 38:672–691
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Falcón W, Ackerman JD, Daehler CC (2012) Non-native distribution and predicted geographic range of Iguana iguana in the Greater Caribbean region. IRCF Reptiles & Amphibians 19:150–160
Hedges SB, Powell R, Henderson RW, Hanson S, Murphy JC (2019) Definition of the Caribbean Islands biogeographic region, with checklist and recommendations for standardized common names of amphibians and reptiles. Caribb Herpetol 67:1–53
Iguana Taxonomy Working Group (2016) A checklist of the iguanas of the world (Iguanidae; Iguaninae), pages 4–46. In: Iguanas: Biology, Systematics, and Conservation, Iverson JB, Grant TD, Knapp CR, Pasachnik SA, editors. Herpetol Conserv Biol 11, Monograph 6.
IUCN (2020) The IUCN Red List of Threatened Species. Version 2020-1. https://www.iucnredlist.org
Janes JK, Miller JM, Dupuis JR, Malenfant RM, Gorrell JC, Cullingham CI, Andrew RL (2017) The K = 2 conundrum. Mol Ecol 26:3594–3602
Jombart T (2008) adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24:1403–1405
Knapp CR, Breuil M, Rodrigues C, Iverson JB (2014) Lesser Antillean Iguana, Iguana delicatissima: conservation Action Plan, 2014–2016. Gland (Switzerland): IUCN SSC Iguana Specialist Group.
Knapp CR, Grant TD, Pasachnik S, Angin B, Boman E, Brisbane J, Buckner SD, Haakonsson JE, Harlow PS, Mukhida F, Thomas-Moko N, van den Burg MP, Wasilewski JA (2020) The global need to address threats from invasive alien iguanas. Anim Conserv. https://doi.org/10.1111/acv.12660
Kuehn R, Hindenlang KE, Holzgang O, Senn J, Stoeckle B, Sperisen C (2007) Genetic effect of transportation infrastructure on Roe deer populations (Capreolus capreolus). J Hered 98:13–22
Luís C, Cothran EG, Mar Oom M (2007) Inbreeding and genetic structure in the endangered sorraia horse breed: implications for its conservation and management. J Hered 98:232–237
Martin Judson JL, Knapp CR, Welch ME (2018) Age-dependent, negative heterozygosity-fitness correlations and local effects in an endangered Caribbean reptile, Iguana delicatissima. Ecol Evol 8:2088–2096
Martin JL, Knapp CR, Gerber GP, Thorpe RS, Welch ME (2015) Phylogeography of the endangered lesser antillean iguana, Iguana delicatissima: A recent diaspora in an archipelago known for ancient herpetological endemism. J Hered 106:315–321
Meirmans PG (2020) GENODIVE version 3.0: Easy-to-use software for the analysis of genetic data of diploids and polyploids. Mol Ecol Resour 20:1126–1131
Moss JB, Welch ME, Burton FJ, Vallee MV, Houlcroft EW, Laaser T, Gerber GP (2017) First evidence for crossbreeding between invasive Iguana iguana and the native rock iguana (Genus Cyclura) on Little Cayman Island. Biol Invasions 20:817–823
Noseworthy J (2017) Cold-blooded conflict: Tackling the illegal trade in endemic Caribbean island reptiles. Unpublished M.SC. Thesis (pp. 1–106). Cambridge: University of Cambridge.
Pounder KC, Mukhida F, Brown RP, Carter D, Daltry JC, Fleming T, Goetz M, Halsey LG, Hughes G, Questel K, Saccheri IJ, Williams R, Soanes LM (2020) Testing for hybridisation of the Critically Endangered Iguana delicatissima on Anguilla to inform conservation efforts. Conserv Genet 21:405–420
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
R Core Team (2019) A Language and Environment for Statistical Computing. Version 3.5.1. Available at www.R-project.org/.
Raymond M, Rousset F (1995) GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Rowe G, Beebee TJC, Burke T (2003) A microsatellite analysis of natterjack toad, Bufo calamita. Oikos 88:641–651
Schlötterer C (2000) Evolutionary dynamics of microsatellite DNA. Chromosoma 109:365–371
Scott PA, Allison LJ, Field KJ, Averill-Murray RC, Bradley Shaffer H (2020) Individual heterozygosity predicts translocation success in threatened deser tortoises. Science 370:1086–1089
Stephen CL, Pasachnik S, Reuter A, Mosig P, Ruyle L, Fitzgerald L (2011) Survey of status, trade, and exploitation of Central American iguanas. Report, Department of Interior, United States Fish and Wildlife Service, Washington, DC. Available at https://www.cites.org/sites/default/files/common/com/ac/26/E26-04i.pdf
Stephen CL, Reynoso VH, Collett WS, Hasbun CR, Breinholt JW (2013) Geographical structure and cryptic lineages within common Green Iguanas, Iguana iguana. J Biogeogr 40:50–62
Valette V, Filipova L, Vuillaume B, Cherbonnel C, Risterucci AM, Delaunay C, Breuil M, Grandjean F (2013) Isolation and characterization of microsatellite loci from Iguana delicatissima (Reptilia: Iguanidae), new perspectives for investigation of hybridization events with Iguana iguana. Conserv Genet Resour 5:173–175
van den Burg MP, Meirmans PG, van Wagensveld TP, Kluskens B, Madden H, Welch ME, Breeuwer JAJ (2018b) The Lesser Antillean Iguana (Iguana delicatissima) on St. Eustatius: genetically depauperate and threatened by ongoing hybridization. J Hered 109:426–437
van den Burg M, Breuil M, Knapp C (2018a) Iguana delicatissima. The IUCN Red List of Threatened Species. https://doi.org/10.2305/IUCN.UK.2018-1.RLTS.T10800A122936983.en
van den Burg MP, Weissgold BJ (2020) Illegal trade of morphologically distinct populations prior to taxonomic assessment and elevation, with recommendations for future prevention. J Nat Conserv 57:125887
van den Burg MP, Brisbane JLK, Knapp CR (2020a) Post-hurricane relief facilitates invasion and establishment of two invasive alien vertebrate species in the Commonwealth of Dominica, West Indies. Biol Invasions 22:195–203
van den Burg MP, Van Belleghem SM, De Jesús Villanueva CN (2020b) The continuing march of Common Green Iguanas: Arrival on mainland Asia. J Nat Conserv 57:125888
van den Burg MP, Grandjean F, Schikorski D, Breuil M, Malone CL (2021a) IguanaBase: a live microsatellite database to guide research and conservation in the genus Iguana (Reptilia: Iguanidae). figshare. https://doi.org/10.6084/m9.figshare.13584923.v1
van den Burg MP, Daltry JD, Angin B, Boman E, Brisbane JLK, Collins K, Haaksonsson JE, Hill A, Horrocks JA, Mukhida F, ProvidenceF Questel K, Ramnanan N, Steele S, Vique Bosquet IM, Knapp CR (2021b) Biosecurity for humanitarian aid. Science 372(6542):581–582. https://doi.org/10.1126/science.abj0449
van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) MICROCHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
van Wagensveld TP, van den Burg MP (2018) First record on fecundity of an Iguana hybrid and its implications for conservation: evidence for genetic swamping by non-native iguanas. Herpetology Notes 11:1079–1082
Vashistha G, Deepika S, Dhakate PM, Khudsar FA, Kothamasi D (2020) The effectiveness of microsatellite DNA as a genetic tool in crocodilian conservation. Conserv Genet Resour 12:733–744
Vuillaume B, Valette V, Lepais O, Grandjean F, Breuil M (2015) Genetic evidence of hybridization between the endangered native species Iguana delicatissima and the invasive Iguana iguana (Reptilia, Iguanidae) in the Lesser Antilles: management implications. PLoS ONE 10:e0127575
Welch ME, Colosimo G, Pasachnik PA, Malone CL, Hilton J, Long GJ, Getz AH, Alberts AC, Gerber GP (2017) Molecular variation and population structure in critically endangered Turks and Caicos rock iguanas: identifying intraspecific conservation units and revising subspecific taxonomy. Conserv Genet 18:479–493
Wright S (1931) Evolution in Mendelian populations. Bull Math Biol 52:241–295
Acknowledgements
We thank Ty Park and all donators to IguanaFest 2019 for financial support. We also want to acknowledge the commitment of all conservation practioners working to protect these endangered reptiles. We are extremely grateful to Joanne Justo-Gaymes, Glenroy Gaymes (Forestry Department, Ministry of Agriculture, Industry, Forestry, Fisheries and Rural Transformation, Richmond Hill, Kingstown, St Vincent & the Grenadines), and Jennifer Daltry (Fauna & Flora International, Cambridge) for allowing us to include preliminary data generated for a forthcoming manuscript on Iguana insularis insularis in St. Vincent and the Grenadines to better understand genetic variability in the genus Iguana in order to better identify geographic origins and hybridization phenomena throughout the Caribbean.
Funding
We thank Ty Park and all donators to IguanaFest 2019 for financial support.
Author information
Authors and Affiliations
Contributions
Project design: MPB, CLM; Sample collection: MPB, MB, CLM; Laboratory work: DS; Data collation: MB; Data analysis: MPB, DS; Paper writing: MPB, CLM; Funding acquisition: MPB, CLM; Commenting on manuscript: All authors.
Corresponding author
Ethics declarations
Conflict of interest
Authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
van den Burg, M.P., Grandjean, F., Schikorski, D. et al. A genus-wide analysis of genetic variation to guide population management, hybrid identification, and monitoring of invasions and illegal trade in Iguana (Reptilia: Iguanidae). Conservation Genet Resour 13, 435–445 (2021). https://doi.org/10.1007/s12686-021-01216-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12686-021-01216-5