Abstract
Purpose of Review
Clinical management of triple negative breast cancer (TNBC) is challenging as patients have heterogeneous responses to systemic therapy, and there is no established therapeutic target. Gene expression profiling and genomic sequencing analysis are among the first steps in understanding the biology of TNBC. In this paper, we review the molecular classification of TNBC and discuss the implications for systemic therapy.
Recent Findings
Clonal and mutational spectrum analyses of TNBC show that it is highly heterogeneous with a diverse mutational pattern and can be clustered into different subtypes including basal-like, luminal androgen receptor, and mesenchymal, based on gene expression profiling. Although knowledge of these subtypes is not used in routine clinical practice, studies have shown that patient outcomes differ according to subtype, with higher pathological complete response rates to chemotherapy reported in basal-like subtypes. Clinical trials with targeted agents are now starting to incorporate molecular subtypes into eligibility criteria.
Summary
TNBC is molecular heterogeneous, and therefore, a wide spectrum of patients’ clinical outcomes exists. Incorporating molecular subtypes into treatment algorithms may offer clinicians greater precision in managing TNBC patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
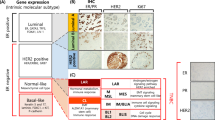
Breast cancer is the most commonly diagnosed cancer in women with an estimated incidence of over 250,000 new cases in the USA in 2017 [1]. It is the second most common cause of cancer death in women, after lung cancer, and accounts for approximately 6.8% of all cancer deaths in women [1]. The clinical classification of breast cancer is discernable by the presence of certain markers, including the estrogen receptor (ER), the progesterone receptor (PR), and overexpression and/or amplification of the human epidermal growth factor receptor 2 (HER2). The knowledge of expression of these three markers guides management, allowing rational and tailored management for breast cancer patients. Attempts to provide further insights into breast cancer heterogeneity based on gene expression patterns has resulted in the development of the molecular classification of breast cancer. The description of the so-called intrinsic subtypes based on gene cluster analysis led to the classification into luminal A, luminal B, Her2-enriched, basal-like, and normal subtypes [2]. Since then, other studies have also described additional breast cancer molecular subtypes [3, 4].
The subset of breast cancer that lacks expression of ER, PR, and HER2 is termed “triple negative breast cancer” (TNBC). Therefore, TNBC is a diagnosis of exclusion. It accounts for approximately 10–20% of all breast cancers [5, 6] and is more commonly diagnosed in younger women, under the age of 50 years [7]. Racial disparities have also been noted in the incidence of TNBC; women of African-American descent have a higher attributable risk than Caucasian women (odds ratio [OR] 2.41, 95% CI 1.81–3.21) [7]. TNBC is also more common in individuals with a germline BRCA mutation, especially BRCA1 [8, 9]. As such, the National Comprehensive Cancer Network (NCCN) recommends genetic risk evaluation for individuals diagnosed with TNBC before the age of 60 years [10]. TNBC is also associated with more biologically aggressive disease at presentation than ER positive breast cancer [11]. As TNBC lacks therapeutic targets, the standard approach for systemic treatment is cytotoxic chemotherapy. Although a subset of TNBC is chemosensitive and carries a good prognosis, resistance to chemotherapy is common and is associated with a much higher risk of early disease recurrence compared to other breast cancer subtypes [11]. Although generally an aggressive subtype, there have been descriptions of low-grade TNBC variants, including adenoid cystic and secretory carcinomas [12].
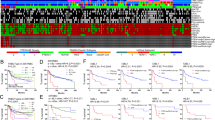
Clonal and mutational spectrum analyses of TNBC suggest that it has a higher mutational frequency than other breast cancer subtypes [13,14,15]. The only frequently recurrent somatic mutation identified is in TP53, present in over 80% of patients [13,14,15,16]. Mutations seen in this gene are commonly frameshift or nonsense mutations versus missense mutations seen in TP53 in patients with luminal breast cancers [17]. The Cancer Genome Atlas study lists PIK3CA as the second most common somatic mutated gene (9%) [17]. Although aberrations in these genes are clonally dominant compared with others, the clonal frequencies in some TNBC patients are not consistent with a founder status, also supporting the observed mutational heterogeneity [13]. In this review, we present two clinical cases to highlight differences in TNBC outcome and discuss molecular classifications of TNBC.
Clinical Case Studies
Patient 1
A healthy 56-year-old postmenopausal woman discovered a palpable lump in the upper outer quadrant of her left breast. Physical examination was remarkable for a 3.5-cm firm mass at the 2 o’clock position of the left breast, with multiple bulky left axillary lymph nodes. Diagnostic breast imaging revealed a 3-cm hyperdense irregular mass at the 2 o’clock position in the left breast, with several markedly enlarged left axillary lymph nodes suspicious for metastatic disease. Her staging CT of the chest, abdomen, and pelvis, and bone scan did not show distant metastatic disease. After initial breast and lymph node biopsies confirmed high-grade TNBC with nodal involvement, she underwent neoadjuvant carboplatin and docetaxel chemotherapy on a clinical trial, followed by mastectomy and axillary node dissection. Pathology review indicated a residual focus of invasive ductal cancer measuring 1.5 cm, with one of three nodes involved with a 3-mm focus of carcinoma. All surgical margins were negative. Genetic testing did not reveal a deleterious mutation in BRCA1 or BRCA2. She received adjuvant post-mastectomy radiation therapy. Six months after therapy completion, she developed abdominal pain and was found to have widespread recurrence in the liver, lungs, lymph nodes, and bones. A liver biopsy confirmed recurrent TNBC. She received palliative treatment with eribulin and pembrolizumab on a clinical trial but died from liver failure due to disease progression within 2 months.
Patient 2
A healthy 54-year-old postmenopausal woman discovered a palpable lump in the upper outer quadrant of her right breast. Physical examination revealed a 4-cm firm mass in the upper right breast, with no palpable right axillary lymph nodes. Diagnostic breast imaging revealed a 4.2-cm upper outer quadrant right breast mass, with a 7-mm satellite mass located 1.8 cm anterior to the dominant mass. Her staging CT showed two indeterminate non-calcified subcentimeter left upper lobe lung nodules. After initial breast biopsy confirmed high-grade TNBC, she also underwent neoadjuvant carboplatin and docetaxel chemotherapy on a clinical trial, followed by lumpectomy and sentinel lymph node biopsy. Pathology review indicated residual invasive ductal cancer measuring 2.4 cm, with one of three nodes involved with macro-metastatic carcinoma. All margins were negative. Genetic testing did not reveal a deleterious mutation in BRCA1 or BRCA2. She received adjuvant post-lumpectomy radiation therapy. She remains disease-free 4 years after therapy completion.
Both patients were managed by the same multidisciplinary team and received the same chemotherapy regimen. The disease course experienced by each patient highlights the diversity in clinical outcomes seen in TNBC, even in individuals who present at similar stages, and are managed in similar ways. This suggests that other unmeasured factors such as tumor biology influence long-term clinical outcomes.
Molecular Heterogeneity of TNBC
In order to deepen our understanding of TNBC biology, several attempts have been made to subclassify TNBC. Studies by Lehmann et al. identified six TNBC subtypes (“TNBCtype”) using gene expression profiles of 587 TNBC cases: (i) basal-like 1 (BL1) subtype enriched in genes involved in cell cycle and proliferation; (ii) basal-like 2 (BL2) involving growth factor signaling; (iii) immunomodulatory (IM) associated with immune cell and cytokine signal transduction pathway; (iv) mesenchymal like (M) genes involved in cell motility, growth, and differentiation; (v) mesenchymal stem-like (MSL), similar to M subtype, however, with low level of genes associated with proliferation; and (vi) luminal androgen receptor (LAR) enriched with genes involved in steroid synthesis metabolism [18, 19]. This subclassification was recently refined as “TNBCtype-4” (BL1, BL2, M, and LAR) based on studies that showed that the gene expression patterns for the previously defined IM and MSL subtypes were heavily influenced by tumor associated stromal cells and infiltrating lymphocytes [20]. Similarly, Burstein et al. identified four distinct TNBC subtypes using mRNA and DNA profiling: (i) basal-like immunosuppressed (BLIS) characterized by downregulation of immune cell and cytokines pathways; (ii) basal-like immune activated (BLIA) with upregulation of genes associated with B, T, and NK cell functions; (iii) mesenchymal (MES) enriched in pathways associated with cell cycle, mismatch repair, and growth factor; and (iv) LAR exhibiting androgen, ER, and ErbB4 signaling, with negative ER staining on IHC [21]. Another group defined three distinct subtypes of TNBC using analysis of microarray gene-expression profiles of 107 TNBC patients: (i) basal-like with low immune response and high M2-like macrophages, (ii) basal enriched with high immune response and low M2-like macrophages, and (iii) LAR [22]. Subsequently, Liu et al. proposed a new classification system by integrating gene expression profiles of mRNAs and lncRNAs of TNBC into four subtypes (i) BLIS, (ii) IM, (iii) LAR, and (iv) MES [23].
All classification systems and the significant overlap between them not only support the remarkable degree of heterogeneity in TNBC tumors but also emphasize the need for a more uniform and standardized classification system for the eventual translation to patient care (Table 1). The overlap suggests at least four distinct subtypes (basal-like, immunomodulatory, mesenchymal, and LAR) with potential clinical implications.
Basal-like
This subtype has the most controversy in its description by the classification systems. It was initially first coined by the “intrinsic subtype” model to describe a subset of breast cancers lacking ER and ErbB2 expression but associated with a unique gene expression profile similar to that expressed by the basal epithelial cells. This group was initially thought to involve all TNBC tumors; however, research has shown that not all TNBC tumors are basal-like, with reported concordance of 70% [24]. Lehman et al. described two basal-like groups (BL1 and BL2) in their original and refined classification system [18]. BL1 is enriched in genes that were associated with cell cycle and cell division (AURK, MYC, NRAS, PLK1, BIRC5), and genes associated with DNA damage repair (RAD5, FANC, MSH2, MDC1). It is also associated with mutations in DNA damage repair genes (BRCA1, ATR) [18, 20,21,22]. Breast cancers in patients with germline BRCA1 mutation have been described as basal-like and have a TNBC phenotype. Interestingly, although most basal-like TNBC patients do not have germline BRCA1 mutation, a high degree of BRCA1 dysfunction and low levels of BRCA1 mRNA expression have been reported in this subtype [25]. The proposed mechanisms of decreased BRCA1 expression include loss of 17q21 (the BRCA1 locus), increased expression of ID4, regulating BRCA1 transcription, and BRCA1 promoter hypermethylation [26, 27]. These provide the basis of the BRCAness of basal-like breast cancer, and the rationale for the use of PARP inhibitors in this subtype. BL1 is also associated with a high Ki-67 mRNA expression supporting its proliferative nature and the observed chemosensitivity [28]. Masuda et al. described varying rates of pathological complete response (pCR) to neoadjuvant chemotherapy among the different subtypes, with BL1 having the highest pCR rate of 52% (M 31%, IM 30%, MSL 23%, LAR 10%, and BL2 0%) [29]. BL2 is associated with genes involved in growth factor signaling (EGFR, MET, Wnt/β-catenin) [18].
Other groups have proposed a substratification of the basal-like subtype based on the immune signature and tumor niche. Two separate groups proposed two basal-like subtypes, BLIS and BLIA (with low and high immune response), while Liu et al. described BLIS as the basal-like subtype [21,22,23]. BLIS (basal-like immunosuppressed) represent tumors with basal-like gene expression profile but exhibit downregulation of B cells, T cells, NK cells, cytokine, and complement pathways, but with the unique expression of several SOX transcription factors. BLIA (basal-like immune activated) as its name implies shows upregulation of genes associated with B, T, and NK cell functions and high expression of STAT family transcription factors [21,22,23]. Jézéquel reported that the subtype associated with downregulated immune response was associated with high M2 macrophages implicated with tumor invasion and metastases and associated with a poorer prognosis [22].
LAR
Androgen receptors (AR) belong to the steroid hormone group of nuclear receptor family, which is encoded on the long arm of X chromosome (Xq12) [30]. This intracellular binding of testosterone or 5-α dihydrotestosterone results in translocation of this complex into the nucleus, with binding to promoter regions of target genes associated with cell growth and survival [30]. AR is more commonly expressed in ER positive breast cancer (~ 80%); however, 10–35% of TNBC have expression of AR. Those tumors are described as LAR given similarity of gene expression profiles to those ER positive breast cancers [31,32,33,34]. Lehman et al. described that in the LAR subtype, AR mRNA was overexpressed, on average at 9-fold greater than the other TNBC subtypes [18]. LAR subtype displays a unique gene expression profile enriched in genes associated with hormone regulation and steroid synthesis. It involves expression of downstream androgen receptor target genes (APOD, FASN, SPDEF, CLDN8), estrogen receptor (ESR1), and genes associated with estrogen signaling pathways (FOXA1, XBP1, GATA-3) [18]. This demonstrates that although the LAR subtype is immunohistochemically negative for ER, it exhibits increased molecular activity of estrogen-mediated pathways. Taken together, these suggest the possibility of therapeutic benefit with anti-estrogens and anti-androgens.
There is also increased frequency of PIK3CA activating mutations in the LAR subtype, suggesting a potential for therapeutic targeting with PI3K/AKT/mTOR inhibition [18, 21].
Mesenchymal
This subtype has a gene expression profile that exhibits increased expression of pathways associated with cell motility and cellular differentiation. Several groups have described increased expression of genes associated with TGF-β, mTOR, Wnt/β-catenin, and ALK signaling pathways [18]. The initial proposed classification by Lehman et al. described two different subtypes including M and the MSL, with similar gene expression profiles. The major differences between both are increased growth factor signaling (EGFR, PDGFR), and low levels of proliferation genes in MSL compared to the M subtype [20]. The refined classification system proposed by Lehman et al. “TNBCtype-4” excludes the MSL subtype as the gene expression profile suggests an interaction between tumor cells and the microenvironment. However, subtypes proposed by others groups describe a mesenchymal type with gene expression profile enriched in growth factor signaling (PGDGFR, VEGF, IGF) and low levels of genes associated with cell proliferation, similar to the previously proposed MSL subtype. This suggests possible opportunities for therapeutic targeting with growth factor inhibitors.
Immunomodulatory
It remains unclear if this can be truly identified as a subtype. This was first proposed in the initial classification system proposed by Lehman et al. as enriched in genes involved in immune cell signaling [18]. The refined TNBCtype-4 classification system proposed that this subtype was associated with low tumor cellularity, with gene expression profile driven by the tumor infiltrating lymphocytes (TILs) [20]. The IM subtype has the highest amount of associated lymphocytes when compared to other subtypes, and high levels of immune checkpoint regulatory genes encoding for PD-L1 and PD1 [20], making this subtype a potential target for immune checkpoint inhibition. This subtype has been corroborated by the classification schema proposed by Liu et al. [23]; however, there seems overlap between this group and the BLIA/basal-like with high immune response.
Potential Therapeutic Applications
Although chemotherapy remains the mainstay of systemic therapy for TNBC patients, the molecular classification systems may allow for personalized and targeted therapy with the goal of improving clinical outcomes in TNBC.
Dysregulation of BRCA1 in basal-like tumors leads to impaired homologous recombinant-dependent DNA repair pathways which are a mechanism of TNBC tumorigenesis [26, 27]. The BRCAness of basal-like breast cancer is the basis of the use of drugs that engage DNA-repair mechanisms. PARP1 inhibitors are approved for treatment of breast cancer patients with germline BRCA1 and BRCA2 mutations. In vitro studies demonstrate that the combination of PARP1 inhibitors and chemotherapeutic agents that induce DNA damage augment the cytotoxic and anti-proliferative effects of chemotherapy in TNBC cell lines [35, 36]. Clinical trials with PARP1 inhibitors in patients with BRCA-associated advanced breast cancer show improved clinical outcomes [37, 38]. Currently, there are several ongoing clinical trials investigating PARP1 inhibitors in TNBC, alone or in combination with cytotoxic chemotherapy (NCT00516724, NCT03205761, NCT01445418).
EGFR overexpression has been demonstrated in basal-like tumors and is reported as a predictor of worse survival independent of tumor stage [39, 40]. Interestingly, basal-like breast cancer cell lines are more sensitive to EGFR inhibition compared to luminal cell lines [39]. A phase II study of cetuximab, an EGFR inhibitor, and cisplatin doubled the overall response rate in patients with advanced TNBC [41]. Larger studies are needed to define patient subtypes with EGFR activating mutations that may benefit from EGFR targeted therapies.
TNBC cell lines with AR overexpression demonstrate increased proliferation in response to androgens and estrogens, but no response to treatment with estrogen antagonists, suggesting an AR-dependent, ER-independent growth response [42]. Anti-androgens have been investigated alone and in combination with chemotherapy in several clinical trials in TNBC with varying outcomes. A phase II trial of bicalutamide in AR-positive ER-negative breast cancer showed a 6-month clinical benefit rate (CBR) of 19% and a median progression-free survival (PFS) of 12 weeks [43]. A recently published phase II trial with enzalutamide in 118 AR-positive TNBC patients showed a 16-week CBR of 25%, a median PFS of 2.9 months, and a median overall survival of 12.7 months all in the intent to treat population [44]. Enzalutamide was very well tolerated with fatigue being the only treatment-related grade 3 or higher adverse event. In vitro studies have also demonstrated that the LAR subtype is much more sensitive to CDK4/6 inhibition compared to basal-like subtypes [45]. CDK4/6 inhibitors are approved in combination with aromatase inhibitors and fulvestrant for advanced ER positive breast cancer. Clinical trials evaluating the combination of anti-androgens and CDK4/6 inhibitors for AR-positive TNBC patients are ongoing (NCT02605486, NCT03090165). The increased frequency of activating PIK3CA mutations in TNBC also suggests a possible role for therapeutic targeting with PI3k/mTOR pathway inhibition [17, 18]. Taselisib, an oral PI3K inhibitor, is being studied in combination with enzalutamide for AR-positive TNBC patients (NCT02457910). Table 2 lists clinical trials with targeted agents in TNBC.
Role of Immunotherapy
TILs are more prevalent in TNBC compared to non TNBC, and their presence is a prognostic factor for disease-free survival, overall survival, and chemotherapy response in TNBC [46,47,48]. PD-L1 expression in TNBC ranges from 20 to 26% [49, 50]. Furthermore, the higher mutational frequency in TNBC compared to other breast cancer subtypes suggests immunogenicity in TNBC. These findings suggest a potential role of immunotherapy in TNBC. Pembrolizumab, an anti-PD1 monoclonal antibody, has been investigated in heavily pretreated PD-L1 positive metastatic TNBC with varying response rates of 5–18.5% (NCT01848834, NCT02447003).
Currently, there are several ongoing clinical trials using anti-PD1 or anti-PD-L1 monoclonal antibodies as monotherapy or in combination with other systemic therapies in the treatment of metastatic TNBC.
Conclusion
Various attempts to classify TNBC demonstrate its marked heterogeneity. There is no standard uniformly accepted molecular classification system. Several clinical studies investigating targeted therapies in TNBC patients have had varying results, which can be attributed to lack of proper patient selection. Although TNBC has a higher mutational frequency compared to other breast cancer subtypes, there is no approved targeted therapy for TNBC. To help address this gap, research should focus on the standardization of TNBC classification, as this may enable proper patient selection in future clinical trials with targeted therapies.
References
Perou CM, Parker JS, Prat A, Ellis MJ, Bernard PS. Clinical implementation of the intrinsic subtypes of breast cancer. Lancet Oncol. 2010;11(8):718–9 author reply 20-1.
Perou CM, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, et al. Molecular portraits of human breast tumours. Nature. 2000;406(6797):747–52.
Prat A, Parker JS, Karginova O, Fan C, Livasy C, Herschkowitz JI, et al. Phenotypic and molecular characterization of the claudin-low intrinsic subtype of breast cancer. Breast Cancer Res. 2010;12(5):R68.
Farmer P, Bonnefoi H, Becette V, Tubiana-Hulin M, Fumoleau P, Larsimont D, et al. Identification of molecular apocrine breast tumours by microarray analysis. Oncogene. 2005;24(29):4660–71.
Diaz LK, Cryns VL, Symmans WF, Sneige N. Triple negative breast carcinoma and the basal phenotype: from expression profiling to clinical practice. Adv Anat Pathol. 2007;14(6):419–30.
Stover DG, Winer EP. Tailoring adjuvant chemotherapy regimens for patients with triple negative breast cancer. Breast. 2015;24:S132–S5.
Trivers KF, Lund MJ, Porter PL, Liff JM, Flagg EW, Coates RJ, et al. The epidemiology of triple-negative breast cancer, including race. Cancer Causes Control. 2009;20(7):1071–82.
Rakha EA, Reis-Filho JS, Ellis IO. Basal-like breast cancer: a critical review. J Clin Oncol. 2008;26(15):2568–81.
Reis-Filho JS, Tutt AN. Triple negative tumours: a critical review. Histopathology. 2008;52(1):108–18.
National Comprehensive Cancer Network. Breast Cancer (Version 1.2018). https://www.nccn.org/professionals/physician_gls/pdf/breast_blocks.pdf.
Dent R, Trudeau M, Pritchard KI, Hanna WM, Kahn HK, Sawka CA, et al. Triple-negative breast cancer: clinical features and patterns of recurrence. Clin Cancer Res. 2007;13(15 Pt 1):4429–34.
Pareja F, Geyer FC, Marchio C, Burke KA, Weigelt B, Reis-Filho JS. Triple-negative breast cancer: the importance of molecular and histologic subtyping, and recognition of low-grade variants. NPJ Breast Cancer. 2016;2:16036.
Shah SP, Roth A, Goya R, Oloumi A, Ha G, Zhao Y, et al. The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature. 2012;486(7403):395–9.
Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature. 2012;490(7418):61–70.
Ma CX, Luo J, Ellis MJ. Molecular profiling of triple negative breast cancer. Breast Dis. 2010;32(1–2):73–84.
Kandoth C, McLellan MD, Vandin F, Ye K, Niu B, Lu C, et al. Mutational landscape and significance across 12 major cancer types. Nature. 2013;502(7471):333–9.
Cancer Genome Atlas N. Comprehensive molecular portraits of human breast tumours. Nature. 2012;490(7418):61–70.
Lehmann BD, Bauer JA, Chen X, Sanders ME, Chakravarthy AB, Shyr Y, et al. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Invest. 2011;121(7):2750–67.
Lehmann BD, Pietenpol JA. Identification and use of biomarkers in treatment strategies for triple-negative breast cancer subtypes. J Pathol. 2014;232(2):142–50.
Lehmann BD, Jovanovic B, Chen X, Estrada MV, Johnson KN, Shyr Y, et al. Refinement of triple-negative breast cancer molecular subtypes: implications for neoadjuvant chemotherapy selection. PLoS One. 2016;11(6):e0157368.
Burstein MD, Tsimelzon A, Poage GM, Covington KR, Contreras A, Fuqua SA, et al. Comprehensive genomic analysis identifies novel subtypes and targets of triple-negative breast cancer. Clin Cancer Res. 2015;21(7):1688–98.
Jezequel P, Loussouarn D, Guerin-Charbonnel C, Campion L, Vanier A, Gouraud W, et al. Gene-expression molecular subtyping of triple-negative breast cancer tumours: importance of immune response. Breast Cancer Res. 2015;17:43.
Liu YR, Jiang YZ, Xu XE, Yu KD, Jin X, Hu X, et al. Comprehensive transcriptome analysis identifies novel molecular subtypes and subtype-specific RNAs of triple-negative breast cancer. Breast Cancer Res. 2016;18(1):33.
Bertucci F, Finetti P, Cervera N, Esterni B, Hermitte F, Viens P, et al. How basal are triple-negative breast cancers? Int J Cancer. 2008;123(1):236–40.
Foulkes WD, Stefansson IM, Chappuis PO, Begin LR, Goffin JR, Wong N, et al. Germline BRCA1 mutations and a basal epithelial phenotype in breast cancer. J Natl Cancer Inst. 2003;95(19):1482–5.
Esteller M, Silva JM, Dominguez G, Bonilla F, Matias-Guiu X, Lerma E, et al. Promoter hypermethylation and BRCA1 inactivation in sporadic breast and ovarian tumors. J Natl Cancer Inst. 2000;92(7):564–9.
Turner NC, Reis-Filho JS, Russell AM, Springall RJ, Ryder K, Steele D, et al. BRCA1 dysfunction in sporadic basal-like breast cancer. Oncogene. 2006;26:2126.
Echavarria I, López-Tarruella S, Picornell A, García-Saenz JÁ, Jerez Y, Hoadley K, et al. Pathological response in a triple-negative breast cancer cohort treated with neoadjuvant carboplatin and docetaxel according to Lehmann’s refined classification. In: Clin Cancer Res, vol. 24; 2018. p. 1845–52.
Masuda H, Baggerly KA, Wang Y, Zhang Y, Gonzalez-Angulo AM, Meric-Bernstam F, et al. Differential response to neoadjuvant chemotherapy among 7 triple-negative breast cancer molecular subtypes. Clin Cancer Res. 2013;19(19):5533–40.
Gao W, Bohl CE, Dalton JT. Chemistry and structural biology of androgen receptor. Chem Rev. 2005;105(9):3352–70.
Alshenawy HA. Prevalence of androgen receptors in invasive breast carcinoma and its relation with estrogen receptor, progesterone receptor and Her2/neu expression. J Egypt Natl Canc Inst. 2012;24(2):77–83.
Loibl S, Muller BM, von Minckwitz G, Schwabe M, Roller M, Darb-Esfahani S, et al. Androgen receptor expression in primary breast cancer and its predictive and prognostic value in patients treated with neoadjuvant chemotherapy. Breast Cancer Res Treat. 2011;130(2):477–87.
Mina A, Yoder R, Sharma P. Targeting the androgen receptor in triple-negative breast cancer: current perspectives. Onco Targets Ther. 2017;10:4675–85.
Park S, Koo J, Park HS, Kim JH, Choi SY, Lee JH, et al. Expression of androgen receptors in primary breast cancer. Ann Oncol. 2010;21(3):488–92.
Alli E, Sharma VB, Sunderesakumar P, Ford JM. Defective repair of oxidative dna damage in triple-negative breast cancer confers sensitivity to inhibition of poly(ADP-ribose) polymerase. Cancer Res. 2009;69(8):3589–96.
Hastak K, Alli E, Ford JM. Synergistic chemosensitivity of triple-negative breast cancer cell lines to poly(ADP-Ribose) polymerase inhibition, gemcitabine, and cisplatin. Cancer Res. 2010;70(20):7970–80.
Robson M, Im SA, Senkus E, Xu B, Domchek SM, Masuda N, et al. Olaparib for metastatic breast cancer in patients with a germline BRCA mutation. N Engl J Med. 2017;377(6):523–33.
A study evaluating talazoparib (BMN 673), a PARP inhibitor, in advanced and/or metastatic breast cancer patients with BRCA mutation (EMBRACA Study) 2018. Available from: https://clinicaltrials.gov/ct2/show/NCT01945775.
Cheang MC, Voduc D, Bajdik C, Leung S, McKinney S, Chia SK, et al. Basal-like breast cancer defined by five biomarkers has superior prognostic value than triple-negative phenotype. Clin Cancer Res. 2008;14(5):1368–76.
Stratford AL, Habibi G, Astanehe A, Jiang H, Hu K, Park E, et al. Epidermal growth factor receptor (EGFR) is transcriptionally induced by the Y-box binding protein-1 (YB-1) and can be inhibited with Iressa in basal-like breast cancer, providing a potential target for therapy. Breast Cancer Res. 2007;9(5):R61.
Baselga J, Gomez P, Greil R, Braga S, Climent MA, Wardley AM, et al. Randomized phase II study of the anti-epidermal growth factor receptor monoclonal antibody cetuximab with cisplatin versus cisplatin alone in patients with metastatic triple-negative breast cancer. J Clin Oncol. 2013;31(20):2586–92.
Doane AS, Danso M, Lal P, Donaton M, Zhang L, Hudis C, et al. An estrogen receptor-negative breast cancer subset characterized by a hormonally regulated transcriptional program and response to androgen. Oncogene. 2006;25:3994–4008.
Gucalp A, Tolaney S, Isakoff SJ, Ingle JN, Liu MC, Carey LA, et al. Phase II trial of bicalutamide in patients with androgen receptor–positive, estrogen receptor–negative metastatic breast cancer. Clin Cancer Res. 2013;19(19):5505–12.
Traina TA, Miller K, Yardley DA, Eakle J, Schwartzberg LS, O’Shaughnessy J, et al. Enzalutamide for the treatment of androgen receptor-expressing triple-negative breast cancer. J Clin Oncol. 2018;36(9):884–90.
Asghar U, Barr A, Cutts R, Beaney M, Sampath D, Giltnane J, et al. 44PUnravelling mechanisms of resistance to CDK4/6 inhibitors using triple negative breast cancer (TNBC). Ann Oncol. 2017;28(suppl_1):mdx145-mdx.
Adams S, Gray RJ, Demaria S, Goldstein L, Perez EA, Shulman LN, et al. Prognostic value of tumor-infiltrating lymphocytes in triple-negative breast cancers from two phase III randomized adjuvant breast cancer trials: ECOG 2197 and ECOG 1199. J Clin Oncol. 2014;32(27):2959–66.
Denkert C, Loibl S, Noske A, Roller M, Muller BM, Komor M, et al. Tumor-associated lymphocytes as an independent predictor of response to neoadjuvant chemotherapy in breast cancer. J Clin Oncol. 2010;28(1):105–13.
Denkert C, von Minckwitz G, Darb-Esfahani S, Lederer B, Heppner BI, Weber KE, et al. Tumour-infiltrating lymphocytes and prognosis in different subtypes of breast cancer: a pooled analysis of 3771 patients treated with neoadjuvant therapy. Lancet Oncol. 2018;19(1):40–50.
Mittendorf EA, Philips AV, Meric-Bernstam F, Qiao N, Wu Y, Harrington S, et al. PD-L1 expression in triple-negative breast cancer. Cancer Immunol Res. 2014;2(4):361–70.
Tung N, Garber JE, Hacker MR, Torous V, Freeman GJ, Poles E, et al. Prevalence and predictors of androgen receptor and programmed death-ligand 1 in BRCA1-associated and sporadic triple-negative breast cancer. NPJ Breast Cancer. 2016;2:16002.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Cynthia X. Ma has served on the advisory board for Merck, Pfizer, Novartis, Eli Lily, and AstraZeneca and has received a grant from Eisai.
Nkiruka Ezenwajiaku and Foluso O. Ademuyiwa declare that they have no competing interests.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
This article is part of the Topical Collection on Clinical Trials
Rights and permissions
About this article
Cite this article
Ezenwajiaku, N., Ma, C.X. & Ademuyiwa, F.O. Updates on Molecular Classification of Triple Negative Breast Cancer. Curr Breast Cancer Rep 10, 289–295 (2018). https://doi.org/10.1007/s12609-018-0292-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12609-018-0292-9