Abstract
Probiotics represent a possible strategy for controlling intestinal infections in livestock. Members of the Weissella genus are increasingly being studied for health-related applications in animals and humans. Here we investigated the functional properties of two Weissella cibaria strains isolated from cows reared in Kuwait breeding facilities by combining phenotypic with genomic analyses. W. cibaria SP7 and SP19 exhibited good growth in vitro under acidic conditions and in the presence of bile salts compared to the reference probiotic Lacticaseibacillus (formerly Lactobacillus) rhamnosus GG. Both strains were able to adhere to Caco-2 and HT-29 cell lines, as well as to mucin. The cell-free supernatants of the two isolates exhibited inhibitory activity towards Escherichia coli ATCC 25,922 and Salmonella enterica UC3605, which was ultimately due to the low pH of supernatants. W. cibaria SP19 showed a co-aggregation ability similar to that of L. rhamnosus GG when incubated with S. enterica. Whole genome sequencing and analysis revealed that both strains harbored several genes involved in carbohydrate metabolism and general stress responses, indicating bacterial adaptation to the gastrointestinal environment. We also detected genes involved in the adhesion to host epithelial cells or extracellular matrix. No evidence of acquired antibiotic resistance or hemolytic activity was found in either strain. These findings shed light on the potential of W. cibaria for probiotic use in livestock and on the mechanisms underlying host-microbe interaction in the gut. W. cibaria` strain SP19 exhibited the best combination of in vitro probiotic properties and genetic markers, and is a promising candidate for further investigation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Over the last decade, there has been a considerable increase in the use of probiotics in animal feeding, with the aim of improving animal performance, digestion, productivity, and resistance to infectious diseases [1, 2]. Infection control is a serious problem for cattle facilities in Kuwait, where dairy farms are managed in an intensive system due to the extremely arid and harsh environment. The type of housing puts stress on animals and, as a consequence, dairy farmers are faced with a high calf mortality rate that is mainly attributed to enterotoxaemia, diarrhea, and pneumonia caused by Pasteurella.
Among lactic acid bacteria, the probiotic potential of Weissella spp. has only recently been investigated in humans as well as in animals. Several studies have reported Weissella strains, mostly belonging to the species Weissella cibaria, Weissella confusa and Weissella paramesenteroides, to possess some functional properties relevant to their use as probiotics [3,4,5,6]. Notably, W. cibaria JW15 was found to possess anti-oxidant and anti-inflammatory properties in vitro [7, 8], and to exert immune-modulatory effects in mice [9] and in human subjects [10]. W. cibaria WIKIM28 had also important immune effects in mice, suppressing allergic Th2 responses and inducing Treg responses in atopic dermatitis [11]. In addition, W. cibaria has been suggested as oral care probiotics: strain CMU has shown significant antimicrobial activity against oral pathogens [12] and a beneficial impact on halitosis and periodontitis in animals and humans [13,14,15].
The development of efficient probiotics relies on a proper selection of bacterial strains to secure the desired benefit in the target host. This implies the absence of pathogenic traits, the capacity to survive during passage and colonize the gastrointestinal tract, as well as to produce antimicrobial substances against pathogens. In the light of recent findings showing that colonization of the intestine may be highly dependent on the host genotype and gut microbiome [16], it is reasonable to assume that better performing strains may be identified if they are specifically selected for the target population. Weissella spp. seem to be well adapted to cattle, in that they have been often identified in cow feces [17, 18], vagina [19], skin [20], and milk [21, 22]. Recently, Weissella spp. were isolated from hosts that have a high capacity for adaptation to survive in arid and desert areas characterized by poor nutrients, high temperatures, and desiccation, such as camels [23]. The authors of the study hypothesized that organisms living in arid lands may select specific gut commensals with special metabolic characteristics contributing to the adaptation of their hosts to harsh environmental conditions.
In this study we evaluated the probiotic potential of two W. cibaria strains selected from among a range of lactic acid bacteria isolated from dairy cows in Kuwait; we sought to make a preliminary assessment from the perspective of developing a tailored treatment strategy to support calf intestinal health. We used conventional in vitro assays to determine W. cibaria resistance to acid and bile stress, adhesion ability, and antimicrobial activity against pathogens. Whole genome sequencing and sequence analysis was used as a complementary approach to deeply investigate the safety and probiotic potential of these promising candidates, as well as to gain insight into the genome evolutionary strategies adopted by these two strains to adapt to the gastrointestinal tract and interact with the host.
Material and Methods
Sample Collection, Strain Isolation, and Identification
Sampling was carried out on two dairy farms (Al-Adwani and Al-Dhbabi) located in the Kabd area of Kuwait. A sampling schedule was prepared to cover the four different seasons: winter, spring, summer, and autumn. Fecal samples were collected from healthy adult cows and calves, preserved in an ice box, and transported to the laboratory for analysis. Milk samples were collected in sterile glass bottles, refrigerated, and transported to the laboratory where they were analyzed immediately. A total of 26 samples were collected. Fecal samples were obtained with informed written consent from the owners of the animals. All efforts were made to minimize animal discomfort according to high standards of veterinary care. No experimentation was done on the animals.
Each 10 g sample was mixed with 90 mL of sterile saline solution (0.85% w/v NaCl) and homogenized using a Stomacher machine (400 Circulator, International PBI, Milan, Italy) at 260 rpm for 1.5 min. One milliliter aliquots of the homogenate were serially diluted in 9 mL of saline and used to isolate strains on MRS agar medium (Merck, Darmstad, Germany) incubated at 37 °C for 48 h in anaerobic jars. No antifungal agents were added in order to avoid any possible interference with LAB growth. Different bacterial colonies were selected on the basis of morphological characteristics (small greyish white colonies) and subjected to three further purification steps under the same anaerobic conditions. Gram-positive and catalase-negative isolates were retained as putative LAB and their identities were confirmed by 16S rRNA amplicon sequencing. Two W. cibaria strains, identified as SP7 and SP19, were selected for further analysis from among all LAB cultures recovered from cow feces. The known probiotic Lacticaseibacillus (formerly Lactobacillus) rhamnosus GG was included as a control strain.
In Vitro Phenotypic Assays
pH and Bile Salt Tolerance
The acid tolerance of W. cibaria strains and L. rhamnosus GG was determined by exposing bacterial cells to low pH. The strains were grown in MRS broth at 37 °C overnight and centrifuged at 3000 rpm for 10 min at 4 °C; the pellets were washed in sterile phosphate-buffered saline (PBS, pH 7) and then suspended in PBS [optical density at 600 nm (OD600nm) = 0.8]. An aliquot (0.15 mL) of each washed cell suspension was added to 1.485 mL of PBS at pH 2, 3, 4, and 7 (control). The mixture was then vortexed at maximum speed for 10 s and incubated at 37 °C. An aliquot of 0.1 mL was removed after 1, 2, and 4 h, serially diluted in 0.85% sterile saline, and plated on MRS agar to determine the total viable count [24].
Bile salt tolerance was evaluated by inoculating 1 mL of bacterial suspension (OD600nm = 0.8) in 9 mL of MRS broth supplemented with 0 (control), 0.5, and 2% (w/v) Ox-bile dehydrated purified (Merck KGaA, Germany). After thorough mixing, the test tubes were incubated at 37 °C. One milliliter of culture was taken from each tube immediately (0 h) and after 2 h incubation, and a series of tenfold dilutions were prepared. Dilutions were plated on MRS agar and incubated at 37 °C for 48 h [25]. Experiments were performed in triplicate.
Aggregation and Co-Aggregation Tests
Auto-aggregation assays were performed according to Kos et al. [26]. Pellets of bacterial cultures grown overnight in MRS broth were collected by centrifugation at 5000 rpm for 15 min, washed twice in PBS (pH 7), and re-suspended in PBS after adjusting the OD600nm to 1 (approximately 107 CFU mL−1). Cell suspensions (4 mL) were mixed by vortexing for 10 s and auto-aggregation measured immediately (t = 0) and after 1, 2, 3, and 4 h of incubation at 37 °C. For every hour interval, 100 µL from the upper suspension was transferred to another tube with 3.9 mL of PBS and the absorbance (A) measured at 600 nm. The auto-aggregation was expressed as a percentage determined by 1-(At/A0) × 100, where At represents the absorbance at time (t) = 1, 2, 3, or 4 h and A0 the absorbance at t = 0.
For co-aggregation tests, bacterial strains were grown anaerobically at 37 °C for 18 h in MRS broth. Salmonella enterica UC3605 from the Università Cattolica del Sacro Cuore Bacteria Culture Collection and Escherichia coli ATCC 25,922 were used as test pathogens. Suspensions of W. cibaria strains and test pathogens were adjusted to an OD600nm of 0.5. Two milliliters of each LAB strain suspension were mixed with the same volume of test pathogen, blended on a vortex mixer for 10 s, and incubated at 37 °C. Control tubes contained 4.0 mL of the bacterial suspension of each strain alone. The OD600nm of the resulting suspensions was measured immediately (t = 0) and after 1, 2, 3, and 4 h by transferring 100 µL from the upper suspension to another tube with 3.9 mL of PBS as described previously. Co-aggregation was calculated as a percentage according to the equation described by Handley et al. [27]. Experiments were performed in triplicate.
Adhesion to Cell Lines and Porcine Gastric Mucin
The mucin and eukaryotic cell adhesion assays were carried out for all strains using porcine gastric mucin, type II (Sigma-Aldrich, Germany) and Caco-2 and HT-29 cells obtained from European Collection of Authenticated Cell Cultures (ECACC). Culturing, maintenance, and adhesion tests were performed as described previously [28, 29] with minor modifications. Strains were inoculated in MRS broth and incubated for 24 h at 37 °C under anaerobic conditions. Broth cultures were centrifuged and pellets washed twice with Hank's balanced salt solution (HBSS). Washed cells were resuspended and diluted to 1.5 × 108 CFU mL−1. The bacterial suspensions were further diluted 1:10 in high glucose Dulbecco's Modified Eagle Medium (DMEM), and 125 µL of these dilutions were used to inoculate the wells seeded with Caco-2 or HT-29 cells, or mucin-coated and the other control wells in order to obtain a multiplicity of infection (MOI) of 5:1.
Cell lines or mucin plus probiotic strains were incubated for 1 h at 37 °C with 5% of CO2. After incubation, the volume in the control wells was harvested and serially diluted and plated on MRS agar according to Rapporti Istisan 08/36 [30]. The medium was removed from the seeded wells and the Caco-2 or HT-29 cell monolayers and mucin coating washed three times with 1 mL of HBSS for 5 min. The wells seeded with eukaryotic cells were inoculated with 100 µL trypsin, and the mucin-coated wells with 100 µL of a 0.5% solution of Triton-X and incubated for 5 min at 37 °C to break the cell monolayers and mucus coating, respectively. The homogenates composed of cell lines or mucin and probiotics were recovered with 900 µL of Maximum Recovery Diluent (MRD, BD Difco, England), serially diluted, and plated onto MRS agar. Count plates were incubated for 72 h at 37 °C under anaerobic conditions.
The adhesion percentages were obtained by referring to the following formula: P = (µ/M) × 100, where P represents the percentage of tested strains adhering to the human cells or mucin, µ represents the vital count of analyzed strains bonded to intestinal cell lines or mucin expressed as a logarithmic value, and M represents the vital count of analyzed strains transformed as a logarithmic value in the wells without human cells or mucin coating.
Antagonistic Activity Against Pathogens
The method in Choi et al. [31] was used with minor modifications. After 24 h at 37 °C, W. cibaria MRS broth cultures were centrifuged at 4000 rpm for 15 min and filtered using a 0.22 µm filter (Jet Biofil, Guangzhou, China) to obtain the cell-free supernatant (CFS). S. enterica UC3605 and E. coli ATCC 25,922 were incubated in Luria Broth for 24 h, diluted in 2X Luria Broth to an OD600nm of 0.06, corresponding to McFarland standard 0.5, and then 50 μL was added to each well of a microtiter plate. The same volume of CFS from W. cibaria strains was added to each well and incubated at 37 °C for 24 h; filtered CFS was tested either after neutralization to pH 6.5 or without pH neutralization. In the control trials, sterile and neutralized MRS broth was used instead of the supernatant. The growth of pathogens was determined by measuring the OD600nm after 24 h. Tests were conducted in duplicate.
Carbohydrate Fermentation Patterns and Exopolysaccharide (EPS) Production
Carbohydrate utilization profiles of W. cibaria SP7 and SP19 were assessed using API 50 CH strips (bioMérieux, Marcy l’Etoile, France) in API 50 CHL Lactobacilli broth as indicated by the manufacturer. The inoculated strips were incubated at 37 °C in anaerobiosis and the reactions observed after 48 h.
The W. cibaria strains were screened for their ability to produce EPS using modified MRS (mod-MRS) agar medium (Proteose Peptone No. 3 10 gL−1, Beef extract 10 gL−1, Yeast extract 5 gL−1, Polysorbate 80 1 gL−1, Ammonium Citrate 2 gL−1, Sodium Acetate 5 gL−1, Magnesium Sulfate 0.1 gL−1, Manganese Sulfate 0.05 gL−1, Dipotassium Phosphate 2 gL−1, Agar 15 gL−1), testing individual sugars that had a positive fermentation output in the carbohydrate fermentation pattern assessment. Mod-MRS agar plates containing 20 g L−1 of glucose, fructose, D-maltose, sucrose, galactose, L-arabinose, D-ribose, D-xylose, D-mannose, D-cellobiose, or gentiobiose (Sigma-Aldrich, Germany) as the sole source of carbohydrate were streaked with the W. cibaria isolates and incubated at 37 °C for 2 to 3 days. The detection of EPS production was facilitated by the addition of Ruthenium Red [0.08% (w/v); Sigma-Aldrich, Germany] [32]. Tests were conducted in duplicate.
Safety Evaluation of W. cibaria Strains
The antibiotic susceptibility profiles of the two W. cibaria strains were determined by broth microdilution according to the ISO 10,932/IDF 233 standard [33] and EFSA guidelines [34] with VetMIC Lact-1 and Lact-2 plates (SVA National Veterinary Institute, Uppsala, Sweden). Briefly, bacterial colonies were suspended in 5 mL of sterile saline solution (0.9% NaCl) to obtain a density corresponding to McFarland standard 1. Suspensions were diluted 1:1000 in LAB susceptibility test medium (LSM); 100 μL of this dilution was added to each well of the VetMIC plate. The plates were incubated at 37 °C for 48 h in anaerobiosis. The minimum inhibitory concentrations (MICs) were determined by spectrophotometric analysis of the plates at 620 nm and compared to the breakpoints proposed by EFSA for obligate heterofermentative Lactobacillus spp.[34].
To evaluate the hemolytic properties, W. cibaria SP7 and SP19 were anaerobically cultured on Columbia agar plates (Oxoid, Altrincham, England) containing 5% (w/v) defibrinated sheep blood (Biolife, Milano, Italy) at 37 °C for 48 h. Staphylococcus aureus ATCC 6538 was used as a positive control for hemolysis. The presence of β- or α-hemolysis was indicated by a clear or greenish zone around the colonies, respectively.
Whole Genome Sequencing and Analysis
Genomic DNA Extraction and Sequencing
Genomic DNA was extracted from exponential phase MRS broth culture of W. cibaria SP7 and W. cibaria SP19 using the NucleoSpin Tissue kit according to the manufacturer’s instructions (Macherey–Nagel, Düren, Germany). Genomic DNA was sequenced at Fasteris (Geneve, Switzerland) using an Illumina MiSeq operating with V3 chemistry in 300X2 bp paired-reads. Basecalling was performed with MiSeq Control Software 2.4.1.3, RTA 1.18.54.0, and CASAVA-1.8.2.
Genome Assembly and Annotation
Raw Fastq reads obtained in Illumina were screened for quality by FastQC software (https://www.bioinformatics.bbsrc.ac.uk/projects/fastqc), retaining the reads with Phred score > 30. Genome assembly was performed using SPAdes (v3.11.1) [35], resulting in 64 contigs for W. cibaria SP7 and 25 contigs for W. cibaria SP19. Genome annotation was performed with Prokka (v1.13.3) [36]. Putative virulence factors were assessed by BLASTn with the VFDB [37] and MvirDB [38] databases. Putative antibiotic resistance genes were identified by BLASTn using ARDB database 1.1 [39], and by the Resistance Gene Identifier of the CARD database [40]. ResFinder 2.1 [41] was used to detect acquired antimicrobial resistance genes. Bacteriocin-coding genes were searched using BAGEL4 [42]. The presence of plasmids was assessed by PlasmidFinder 1.3 [43]. The presence of CRISPR-Cas systems was evaluated using the online tool CRISPRCasFinder [44].
To evaluate the pan- and core-genome of W. cibaria, all publicly available complete genomes (nine strains) and the two newly sequenced strains were analyzed using Roary (v3.12.0) [45]. The choice to consider only complete genome sequences was made since it has been previously evidenced that incomplete genomes highly affect the core-genome identification within a species [46]. The core gene alignment obtained with Roary was then used in RAxML (v8.2.12) to build a maximum-likelihood phylogenetic tree. The tree was visualized using iTOL [47]. Orthogroups (OGs) were inferred with OrthoFinder (v2.2.3) [48], and the gene content analysis was performed by mapping the OGs to the eggNOG bacterial database using eggNOG-mapper (v1.0.3) [49]. The gene content per OG for each strain (considering COG categories) was visualized with MeV [50].
Statistical Analysis
Statistical analyses were performed using GraphPad Prism version 5 (GraphPad Software, San Diego, CA, USA). Phenotypic data were analyzed by two-way ANOVA with Bonferroni post-hoc test, with P < 0.05 considered significant.
Results
Phenotypic Characterization of W. cibaria SP7 and SP19
Resistance to Low pH and Bile Salts
The results of the acid tolerance test are shown in Fig. 1. No viable bacterial cells were detected after the first hour of exposure to pH 2, suggesting that all three strains were severely impaired upon acid shock. Exposure to pH 3 resulted in a decrease in viability for all strains; after 2 h, a decrease of 2 log and 1 log was detected for W. cibaria SP7 (P < 0.001) and SP19 (P < 0.01), respectively, as compared to pH 7. In contrast, for L. rhamnosus GG, significant growth reduction was observed only after 4 h (P < 0.001). At this time point, the concentration of viable W. cibaria SP19 cells declined to undetectable levels. All tested strains had a high survival rate when exposed to pH 4; only the viability of L. rhamnosus GG decreased significantly after 4 h incubation (P < 0.01) when compared to control.
The results of the bile tolerance test are shown in Fig. 2. All tested strains demonstrated good survival over the 2 h of exposure to concentrations of bile salts as high as 2%, with only a reduction in microbial counts as compared to 0% bile salts detected for SP19 (P < 0.01).
Aggregation and Co-Aggregation Tests
As shown in Fig. 3, the aggregation of W. cibaria SP7 was very similar to the aggregation of L. rhamnosus GG over the 4 h incubation, whereas W. cibaria SP19 exhibited significantly less aggregation. Neither the two W. cibaria isolates nor L. rhamnosus GG exhibited co-aggregative abilities with E. coli ATCC 25,922. Co-aggregation with S. enterica UC3605 was not observed for W. cibaria SP19 up to 2 h of incubation; afterwards, the percentage of co-aggregation was comparable to L. rhamnosus GG. In contrast, co-aggregation values for W. cibaria SP7 reached a maximum after 1 h of incubation and then progressively declined, becoming barely detectable after 4 h of incubation.
Adhesion to Cells and Porcine Gastric Mucin
All tested strains demonstrated a capacity to adhere to porcine gastric mucin at varying levels. L. rhamnosus GG exhibited the highest adhesion (77 ± 0.85%), followed by W. cibaria SP7 (72 ± 0.0%) and W. cibaria SP19 (53 ± 1.2%). Significant differences were observed between the adhesion of SP19 (P < 0.001) and SP7 (P < 0.01) compared to L. rhamnosus GG. W. cibaria SP7 and L. rhamnosus GG exhibited the same adhesion to Caco-2 cells (78 ± 1%), whereas W. cibaria SP19 exhibited less adhesion (69 ± 0.28%;i P< 0.001). In regards to HT-29 cells, L. rhamnosus GG displayed a percentage of adhesion (76 ± 2.0%) that was significantly higher than that of W. cibaria SP19 (64 ± 1.3%) and of W. cibaria SP7 (70 ± 0.8%) (Fig. 4). No significant differences were found between strains considering the percentage of adhesion to Caco-2 and HT-29 cells.
Antagonistic Activity Against Pathogens
The CFSs of both W. cibaria strains and L. rhamnosus GG exhibited antagonistic activity against reference E. coli and S. enterica strains. After 24 h of incubation with the W. cibaria SP7 supernatant, the growth of E. coli and S. enterica decreased 96 ± 1.3% (P < 0.001) and 94 ± 0.8% (P < 0.001), respectively. Similar percentages of inhibition were observed in E. coli (98 ± 1.2%; P < 0.001) and S. enterica (96 ± 0.0%; P < 0.001) when exposed to W. cibaria SP19 CFS. Analogously, L. rhamnosus GG significantly inhibited the growth of E. coli and S. enterica (97 ± 1.2% and 95 ± 0.8%, respectively; P < 0.001). Neutralization of CFS resulted in complete reversal of the inhibitory effects (data not shown), suggesting a major role of organic acids in supporting the antagonistic activity of the three tested bacteria against pathogens.
Carbohydrate Fermentation Patterns
The results of the carbohydrate fermentation test are shown in Table 1. Both W. cibaria SP19 and SP7 were able to ferment a number of sugars, including L-arabinose, D-ribose, D-glucose, D-fructose, D-mannose, N-acetylglucosamine, D-maltose, and potassium gluconate. In addition, W. cibaria SP19 was able to ferment D-xylose, D-galactose, and D-saccharose, as well as β-glucosides D-cellobiose, amygdalin, arbutin, esculin, gentobiose, and salicin.
The production of EPS was assessed by evaluating the color of colonies grown on mod-MRS agar plates containing various carbohydrates. W. cibaria SP19 resulted in slimy colonies, indicating EPS synthesis and secretion only in the presence of sucrose, whereas W. cibaria SP7 grew as red colonies on the same plates, showing no ability to produce EPS (data not shown).
Safety Assessment
The MICs of the tested antibiotics are listed in Table 2. For both W. cibaria strains, the MICs did not exceed the MIC cut-off values proposed by EFSA for obligate heterofermentative Lactobacillus spp.
Neither β-hemolytic nor α-hemolytic phenotypes were detected for W. cibaria SP7 and SP19 grown on blood agar plates. In contrast, the positive control S. aureus ATCC 6538 exhibited the expected β-hemolysin activity (data not shown).
Features of the W. Cibaria SP7 and SP19 Genomes
Genes Associated with Persistence in the Host and Probiotic Function
The general genomic properties of W. cibaria SP7 and W. cibaria SP19 are presented in Table 3 and Fig. S1. In the preliminary RAST annotation, both the SP7 and SP19 genomes contained a number of genes encoding enzymes required for the metabolism of mannose, D-ribose, xylose, D-gluconate and ketogluconates, and L-arabinose. The SP19 genome also included genes encoding enzymes required for the utilization of fructose, D-galacturonate, and D-glucuronate. Liver-produced β-D-glucuronides are detoxification products excreted into the mammalian gut, and several symbiotic microorganisms have adapted to exploit host-derived glucuronides as a source of carbon for growth. Both strains harbored putative PTS transporter and permease genes for sucrose, lactose, trehalose, and galactose utilization. Genome mining with Prokka annotation identified genes involved in chitin and N-acetylglucosamine utilization, coding for N-acetylglucosamine-6-phosphate deacetylase (NagA), N-acetylglucosamine-6-phosphate deaminase (NagB), and N-acetylglucosamine repressor (NagC). Notably, two beta-glucoside operons were found in both genomes, the bglABFHK operon and licABCT operon (with SP7 lacking bglB and licBC in the two operons, respectively), which are responsible for the catabolism of polysaccharides present in the plant cell wall, such as cellobiose, salicin, and arbutin. W. cibaria SP19 possessed putative enzymes required for glycogen synthesis in bacteria, including the glycogen synthase (GlgA), 1,4-alpha-glucan branching enzyme (GlgB), glucose-1-phosphate adenylyltransferase (GlgC), glycogen biosynthesis protein glucose-1-phosphate adenylyltransferase (GlgD), and glycogen phosphorylase (GlgP), responsible for glycogen and starch breakdown.
We also found several genes associated with stress resistance in SP7 and SP19, including genes encoding for resistance to osmotic stress (aquaporin Z, glycerol uptake facilitator protein, and OpuABC and GbuA gene clusters related to the uptake and biosynthesis of osmo-protectants choline and betaine). Bile stress has been shown to induce oxidative stress, and a number of genes involved in resistance to oxidative stress were observed in the two Weissella isolates. These genes include proteins with ferroxidase activity, redox-sensitive transcriptional regulator (AT-rich DNA-binding protein), and genes involved in the biosynthesis and reduction of the cellular antioxidant glutathione. Both strains harbored hflX and hflK, which are associated with the 50S ribosomal subunit and may play a role during protein synthesis or ribosome biogenesis. Finally, genes that confer resistance to cold (CspLA and CspB proteins) and 11 genes conferring resistance to heat shock were found in both genomes. They belong to the heat shock dnaK gene cluster extended subsystem and include the DnaK (HrcA-GrpE-DnaK-DnaJ) operon. Other genes are involved in protecting ribosomal function during heat shock and other stresses (lepA, prmA, rdgB, smpAB, rsmABCDEFGHI). No bile salt hydrolase (bsh) genes were present.
Gene prediction with Prokka identified genes specifically associated with adhesion. Specifically, the presence of genes coding for capsular proteins (cps12A and cap8A) and fibronectin-binding protein (fbpA) was demonstrated, as well as three and two copies of mucin-binding protein (mucBP) in SP7 and SP19, respectively. Moreover, SP7 and SP19 harbored two and three copies of the LPXTG specific sortase gene, respectively, along with three copies of the gene coding for the biofilm regulatory protein A precursor (brpA). EPS clusters, namely EpsDFHJLE, were identified in the two strains, with both lacking EpsA. EPS may contribute to cell adhesion and exert a protective function in the bacterial cells against the environment. Notably, both strains harbored genes encoding enzymes predicted to be involved in N-acetylglucosamine catabolism and synthesis (nagA, nagB, nagC, glmS, glmM, glmU, manX). However, the SP19 genome also harbored genes involved in dTDP-rhamnose biosynthesis (rfbC, rmlD), whereas SP7 does not harbor these genes. Moreover, multiple copies of glycosyltransferase genes were found in both strains (gtf1, gtfC), as well as maltose metabolism genes (maa, malP, malL).
Gene prediction by Prokka highlighted the presence of a putative gene coding for the bacteriocin Colicin V, in both strains of W. cibaria. However, a more in-depth characterization with BAGEL4 did not detect a related gene cluster for the bacteriocin production.
CRISPRCasFinder identified one CRISPR-Cas system in W. cibaria SP19, including CAS-TypeIIA proteins (Cas1, Cas2, Cas9, Csn2).
A summary of the main genes identified in the two newly sequenced strains of W. cibaria, that can be potentially associated to probiotic functions, are reported in Table 4.
Genes Associated with Safety
Analysis of the SP7 and SP9 genomes by comparisons to the VFDB and MvirDB databases indicated that no putative virulence-associated genes were present. The Antibiotic Resistance Database (ARDB) identified no gene above the cut-off (70% identity, P < 0.0001). Analysis by ResFinder (selected % ID threshold: 98.00%, selected minimum length: 60%) identified no acquired antimicrobial resistance genes in the genome of either strain. Finally, interrogation of the comprehensive Antibiotic Resistance Database (CARD) resulted in no positive hits when selecting perfect and strict hits, or hits below a percentage identity of 45% when allowing loose hits. No plasmids were found by PlasmidFinder.
Two genes coding for hemolysin A (TlyA) and hemolysin-III-related protein were found in both SP7 and SP19.
Pan- and Core-Genome Analysis
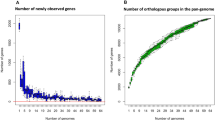
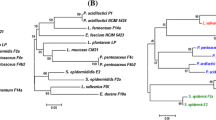
Comparative genomics of the newly sequenced strains with complete W. cibaria genomes revealed the presence of 1323 core genes (99% ≤ strains ≤ 100%), 1360 shell genes (15% ≤ strains < 95%), and 2008 cloud genes (0% ≤ strains < 15%), resulting in a pan-genome of 4691 genes (Fig. S2a). Variations in the content of unique and new genes in the W. cibaria pan-genomes, as a function of the number of analyzed strains, are shown in Fig. S2b. The phylogenetic tree based on the core gene alignment of the 11 strains was re-rooted on strain SP7, serving as an outgroup (Fig. S2c). Interestingly, the two newly sequenced strains clustered differently, even if the isolation source was the same. Excluding the outgroup (SP7 strain), two main clades were highlighted: one including strains M2, CBA3612, and CH2, and the other including BM2, SRCM103448, SP19, CMS2, CMS3, and the proposed oral care probiotic strains CMS1 and CMU [51,52,53] (Fig. S2b).
Evaluating the pan-genome, 97% of the genes identified in all genomes were clustered into 2509 OGs, and 1724 were defined as core OGs (i.e., containing all species). OGs were then mapped to the bacterial eggNOG database to evaluate the main functional COG categories related to the pan-genome considering each strain (Fig. S3). The major fraction of OGs (24%) was grouped under the “S” category of “unknown function” (not included in Fig. S3). From the hierarchical clustering based on the Euclidean distance, two main clusters were shown, grouping the more and less abundant COG categories, respectively. Similarly, two main clades were formed by separating the two strains: CBA3612 and CH2. Considering the relative abundance profiles, no differences were highlighted between the strains when considering the most abundant COG categories, which were related to amino acid and carbohydrate transport and metabolism, and DNA replication and translation activities (E, G, J, L). However, some differences were found within the categories of “Cell envelope biogenesis, outer membrane” (M), “Transcription” (K), “Inorganic ion transport and metabolism” (P), and “Defense mechanisms” (V) (Fig. S3).
Discussion
Probiotics are gaining interest from research and the food industry as a possible strategy for controlling intestinal infections in livestock and to reduce antibiotic use in animal feed, thus helping contain the risks associated with the spread of antibiotic-resistance genes or the contamination of animal-derived products with antibiotic residues. In addition, the inclusion of probiotics in cattle diets has the potential to improve gut function and increase the utilization of feed, ultimately enhancing the health and productivity of ruminant animals. In this study, the potential of two W. cibaria strains newly isolated from cattle in Kuwait as animal probiotics was investigated based on their whole genome sequences and corresponding phenotypes. The phenotypic assessment of W. cibaria SP7 and SP19 showed that both strains possess good tolerance to low pH and bile salts in vitro to varying degrees. The comparison with well-known probiotic L. rhamnosus GG indicated similar levels of pH and bile resistance. Although data are available only for a limited number of strains, resistance of W. cibaria strains to a pH as low as 2.5 and 0.3% bile salts has been described in some previous studies on fermented food and intestinal isolates of human and animal origin [6, 54,55,56,57]. The bile salt resistance of Weissella spp. is likely to be considered a strain-dependent property similar to what has been observed in Lactobacillus and Bifidobacterium spp.[58]. A W. cibaria WD2 isolate obtained from fermenting cassava mash survived in simulated gastric and intestinal transit, tolerating acid (pH 2) and bile salt (1%) [59], whereas W. cibaria KCTC 3746 isolated from kimchi was not able to retain its viability when exposed to 0.3% bile salts in the presence of pancreatin [31]. At the genomic level, both strains under investigation seem to carry a good set of genes to counteract stress; some of them are frequently found in lactic acid bacteria, such as the chaperone system DnaK, which has been demonstrated to play a role in the acid resistance response [60, 61]. In contrast, little is known about the HflX GTPase, but proteomic studies in probiotic Lactobacillus gasseri ATCC 33323 suggest that HflX may act as a key regulator during stress [62]. Furthermore, it must be stressed that the occurrence of a coding gene not always implies the formation of a functional gene product or a direct involvement in a specific biological activity. We found no evidence of bile salt hydrolases in the W. cibaria strains under study. In addition to bile acid modification activity, bile tolerance develops through multiple mechanisms that can be bile-specific, such as efflux pumps and cell wall and membrane modification, as well as mechanisms underlying the overall bacterial capacity to maintain intracellular homeostasis [58]. Notably, recent studies relying on omics technologies have allowed the detection of numerous proteins for which their expression is influenced by bile salt stress, including proteins involved in carbohydrate metabolism, cell envelope and lipid metabolism, efflux pumps, and proteins related to the general stress response [63].
Phenotypic test results indicated the ability of both tested strains to ferment carbohydrates such as L-arabinose, D-ribose, D-glucose, D-fructose, D-mannose, N-acetylglucosamine, D-maltose; D-saccharose, D-galactose, and gentiobiose were fermented only by W. cibaria SP19. More than 12% and 15% of the identified genes in the two bovine isolates, W. cibaria SP7 and SP19, respectively, were involved in carbohydrate metabolism. In this context, Lynch et al. [64] reported that the sourdough isolate W. cibaria MG1 has the potential to metabolize galactose, maltose, fructose, ribose, xylose, sucrose, and gluconate. They also identified putative phosphotransferase systems (PTSs) for N-acetyl-D-glucosamine, cellobiose, mannose, b-glucosides, and fructose. In contrast to W. cibaria SP7, W. cibaria SP19 exhibited an ability to utilize sugars widely distributed in plants, namely D-xylose and β-glucosides, including arbutin, esculin, and salicin, suggesting some level of adaptation to the plant-based diet of ruminants.
Interestingly, W. cibaria strain SP19 also possessed putative enzymes conferring an ability to synthesize glycogen and a glycogen phosphorylase involved in glycogen catabolism. The glycogen metabolic pathway has been detected in probiotic strains or intestinal isolates belonging to the Lactobacillus genus, suggesting potential involvement of this pathway in the persistence and probiotic functionalities of lactobacilli in the gut [65, 66].
At the phenotypic level, both W. cibaria strains exhibited adhesion properties to epithelial cell lines and mucin in vitro, though significantly lower percentages were detected compared to L. rhamnosus GG. These results are consistent with previous findings [6, 55] indicating the in vitro adhesion capacity of W. cibaria strains to Caco-2 cells. W. cibaria KCTC 3746 exhibited a high percentage of auto-aggregation after 24 h incubation and effectively adhered to human intestinal epithelial HT-29 cells [31]. At the genomic level, the two isolates harbored genes encoding proteins specifically associated with adhesion, such as fibronectin-binding protein [67] and mucin-binding protein. No mucus-binding proteins were detected in W. cibaria KACC 11,862 in a previous study [68]. The presence of sortases, enzymes anchoring surface proteins to the cell wall of Gram-positive bacteria, could also play an important role. Deletion of the sortase-encoding gene reduced adherence of Ligilactobacillus (formerly Lactobacillus) salivarius UCC118 to epithelial cells [69].
Antagonistic activity and the production of antimicrobial compounds is another very important characteristic needed to inhibit the growth of pathogenic bacteria. The ability of W. cibaria SP19 and SP7 to exert effective antimicrobial activity against E. coli and Salmonella spp., which are major causes of calf enteric disorders in Kuwait dairy herds [70], was associated with the production of organic acids and the resulting decrease in pH. W. cibaria is able to produce several antimicrobial substances; recently, the antibacterial activity of the oral probiotic W. cibaria CMU was linked to its ability to produce H2O2, organic acids, oleic acid, and specific proteins, such as N-acetylmuramidase [53]. The production of bacteriocins was described previously in a limited number of Weissella strains belonging to the species W. cibaria, W. paramesenteroides, and Weissella hellenica [71,72,73,74,75]. Unfortunately, we were not able to highlight any production of antimicrobial substances other than organic acids in the growth medium or putative bacteriocin gene clusters in the genomes of the strains under investigation.
However, the production of antimicrobial compounds is only one of several mechanisms through which probiotics can prevent or reduce the colonization and infection of livestock and humans. In our experimental conditions, co-aggregation of W. cibaria SP19 with the gut pathogen S. enterica, though expressed at a low level, was comparable to that of L. rhamnosus GG after 4 h incubation, and the ability of probiotic strains to co-aggregate with pathogens is thought to play an important role in their protective function [76].
In regards to safety, we exploited whole genome sequencing data analysis in combination with in vitro testing to conduct a comprehensive assessment of the antibiotic resistance and pathogenic potential of W. cibaria isolates, as recommended by Hill et al. [77]. We found no evidence of phenotypic antibiotic resistance characteristics in the two W. cibaria strains we investigated. Few studies are available on the antibiotic resistance profiles of Weissella species and, unlike other lactic acid bacteria genera, no specific MIC breakpoint values have been defined by EFSA for the assessment of antimicrobial susceptibility. Jeong and Lee [78] found that the W. cibaria, W. confusa, and W. paramesenteroides strains are resistant to streptomycin, and W. cibaria strains to penicillin G. Low rates of resistance to aminoglycosides have been reported in the Leuconostoc/Weissella group [79, 80], and Leuconostoc strains of food origin have been susceptible to beta-lactams in other studies [80,81,82]. In regards to the resistance to penicillin G, Jeong and Lee [78] reported that the results from the disk diffusion test were inconsistent with those from the microdilution broth assay, indicating the need to define specific cut-off values for Weissella spp. However, in our study, the search for antibiotic resistance determinants identified no specific gene in either strains, confirming the antimicrobial susceptibility testing results.
In line with previous findings [68], both W. cibaria strains harbored genes for hemolysin or hemolysin-like proteins, but the presence of such genes was not associated with the expression of a hemolytic phenotype, further confirming the safety profiles of the isolates.
Comparative genomic analysis showed that the W. cibaria pan-genome is still “open”, as nearly 25 new genes are included for each additional genome considered. This is also due to the low number of genomes currently available for W. cibaria. However, the presence of variable genetic content that relies on strain-specific genes (unique) is suggested.
Although a lot of commercial probiotic preparations are currently available for a wide range of conditions, it is clear that not all organisms are well suited for all applications. Notably, species-specificity [83, 84] and intestinal origin [85, 86] of bacterial strains have been reported as relevant factors influencing probiotics efficacy.
The results of this study add to our understanding of the genomics and metabolic capabilities of gut-associated W. cibaria species, and shed light on potential mechanisms of host-microbe interaction that represent a crucial input for the probiotics development process. Since functional attributes of specific strains can be related to their evolution and the mutualistic relationship established with different hosts [87], our findings could contribute to select probiotics which are more likely to colonize and maintain in the intestine of cow, and to exert specific physiological effects on the host. Future investigations are necessary to ascertain whether the in vitro characteristics of our W. cibaria strains may reflect the in vivo behavior of the isolates when administered to control infection and reduce the use of antibiotics under field conditions.
In conclusion, the integration of whole-genome sequencing and phenotyping provides insight into the genomics and physiology of W. cibaria strains of animal origin and demonstrated that specific strains belonging to this species have relevant features that make them suitable for probiotic applications. To the best of our knowledge, this study is the first to report the draft genome sequences of W. cibaria strains derived from bovine intestine. Our results indicate that W. cibaria strain SP19, and to a lesser extent strain SP7, possesses several in vitro probiotic capacities, such as good tolerance to acidic pH and bile salts, adherence to intestinal cells, and antagonistic activity against pathogens. These properties, together with genetic characteristics, such as the lack of transferable antibiotic resistance determinants and pathogenicity factors, and the presence of genes revealing adaptation to the gut environment, suggest a potential use of these strains as novel probiotics tailored to prevent intestinal infections in cattle production. In vivo trials are mandatory to confirm the in vitro results of this study.
Data Availability
The whole genome sequencing projects for W. cibaria strains SP7 and SP19 have been deposited at DDBJ/ENA/GenBank under accession numbers SDGJ00000000 and SDGK00000000, and in the NCBI Sequence Read Archive (SRA) database with accession numbers SRR8486238 and SRR8494494
References
Markowiak P, Śliżewska K (2018) The role of probiotics, prebiotics and synbiotics in animal nutrition. Gut Pathog 10. https://doi.org/10.1186/s13099-018-0250-0
Uyeno Y, Shigemori S, Shimosato T (2015) Effect of probiotics/prebiotics on cattle health and productivity. Microbes Environ 30:126–132. https://doi.org/10.1264/jsme2.me14176
Xiong L, Ni X, Niu L et al (2019) Isolation and preliminary screening of a Weissella confusa strain from Giant Panda (Ailuropoda melanoleuca). Probiotics Antimicrob Proteins 11:535–544. https://doi.org/10.1007/s12602-018-9402-2
Pabari K, Pithva S, Kothari C et al (2020) Evaluation of probiotic properties and prebiotic utilization potential of Weissella paramesenteroides isolated from fruits. Probiotics Antimicrob Proteins. https://doi.org/10.1007/s12602-019-09630-w
Xia Y, Qin S, Shen Y (2019) Probiotic potential of Weissella strains isolated from horse feces. Microb Pathog 132:117–123. https://doi.org/10.1016/j.micpath.2019.04.032
Lee KW, Park JY, Jeong HR et al (2012) Probiotic properties of Weissella strains isolated from human faeces. Anaerobe 18:96–102. https://doi.org/10.1016/j.anaerobe.2011.12.015
Yu H-S, Lee N-K, Choi A-J et al (2019a) Antagonistic and antioxidant effect of probiotic Weissella cibaria JW15. Food Sci Biotechnol 28:851–855. https://doi.org/10.1007/s10068-018-0519-6
Yu H-S, Lee N-K, Choi A-J et al (2019b) Anti-inflammatory potential of probiotic strain Weissella cibaria JW15 Isolated from kimchi through regulation of NF-κB and MAPKs pathways in LPS-induced RAW 264.7 Cells. J Microbiol Biotechnol 29:1022–1032. https://doi.org/10.4014/jmb.1903.03014
Park H-E, Kang K-W, Kim B-S et al (2017) Immunomodulatory potential of Weissella cibaria in aged C57BL/6J mice. J Microbiol Biotechnol 27:2094–2103. https://doi.org/10.4014/jmb.1708.08016
Lee YJ, Lee A, Yoo HJ et al (2018) Supplementation with the probiotic strain Weissella cibaria JW15 enhances natural killer cell activity in nondiabetic subjects. J Funct Foods 48:153–158. https://doi.org/10.1016/j.jff.2018.07.009
Lim SK, Kwon M-S, Lee J et al (2017) Weissella cibaria WIKIM28 ameliorates atopic dermatitis-like skin lesions by inducing tolerogenic dendritic cells and regulatory T cells in BALB/c mice. Sci Rep 7:40040. https://doi.org/10.1038/srep40040
Jang H-J, Kang M-S, Yi S-H et al (2016) Comparative study on the characteristics of Weissella cibaria CMU and probiotic strains for oral care. Molecules 21:1752. https://doi.org/10.3390/molecules21121752
Do K-H, Park H-E, Kang M-S et al (2019) Effects of Weissella cibaria CMU on halitosis and calculus, plaque, and gingivitis indices in Beagles. J Vet Dent 36:135–142. https://doi.org/10.1177/0898756419872562
Kim J-W, Jung BH, Lee JH et al (2020) Effect of Weissella cibaria on the reduction of periodontal tissue destruction in mice. J Periodontol. https://doi.org/10.1002/JPER.19-0288
Lee D-S, Lee S-A, Kim M et al (2020) Reduction of halitosis by a tablet containing Weissella cibaria CMU: randomized, double-blind, placebo-controlled study. J Med Food. https://doi.org/10.1089/jmf.2019.4603
Zmora N, Zilberman-Schapira G, Suez J et al (2018) Personalized gut mucosal colonization resistance to empiric probiotics is sssociated with unique host and microbiome features. Cell 174:1388-1405.e21. https://doi.org/10.1016/j.cell.2018.08.041
Adeniyi BA, Adetoye A, Ayeni FA (2015) Antibacterial activities of lactic acid bacteria isolated from cow faeces against potential enteric pathogens. Afr Health Sci 15:888. https://doi.org/10.4314/ahs.v15i3.24
Ayeni FA, Sánchez B, Adeniyi BA et al (2011) Evaluation of the functional potential of Weissella and Lactobacillus isolates obtained from Nigerian traditional fermented foods and cow’s intestine. Int J Food Microbiol 147:97–104. https://doi.org/10.1016/j.ijfoodmicro.2011.03.014
Wang J, Sun C, Liu C et al (2016) Comparison of vaginal microbial community structure in healthy and endometritis dairy cows by PCR-DGGE and real-time PCR. Anaerobe 38:1–6. https://doi.org/10.1016/j.anaerobe.2015.11.004
Braem G, De Vliegher S, Verbist B et al (2012) Culture-independent exploration of the teat apex microbiota of dairy cows reveals a wide bacterial species diversity. Vet Microbiol 157:383–390. https://doi.org/10.1016/j.vetmic.2011.12.031
Masoud W, Vogensen FK, Lillevang S et al (2012) The fate of indigenous microbiota, starter cultures, Escherichia coli, Listeria innocua and Staphylococcus aureus in Danish raw milk and cheeses determined by pyrosequencing and quantitative real time (qRT)-PCR. Int J Food Microbiol 153:192–202. https://doi.org/10.1016/j.ijfoodmicro.2011.11.014
Espeche MC, Otero MC, Sesma F, Nader-Macias MEF (2009) Screening of surface properties and antagonistic substances production by lactic acid bacteria isolated from the mammary gland of healthy and mastitic cows. Vet Microbiol 135:346–357. https://doi.org/10.1016/j.vetmic.2008.09.078
Fhoula I, Rehaiem A, Najjari A et al (2018) Functional probiotic assessment and in vivo cholesterol-lowering efficacy of Weissella sp. associated with arid lands living-hosts. Biomed Res Int 2018:1–11. https://doi.org/10.1155/2018/1654151
Conway PL, Gorbach SL, Goldin BR (1987) Survival of lactic acid bacteria in the human stomach and adhesion to intestinal cells. J Dairy Sci 70:1–12. https://doi.org/10.3168/jds.s0022-0302(87)79974-3
Mishra V, Pradad D (2005) Application of in vitro methods for selection of strains as potential probiotics. Int J Food Microbiol 103:109–115. https://doi.org/10.1016/j.ijfoodmicro.2004.10.047
Kos B, Šušković J, Vuković S et al (2003) Adhesion and aggregation ability of probiotic strain Lactobacillus acidophilus M92. J Appl Microbiol 94:981–987. https://doi.org/10.1046/j.1365-2672.2003.01915.x
Handley PS, Harty DWS, Wyatt JE et al (1987) A comparison of the adhesion, coaggregation and cell-surface hydrophobicity properties of fibrillar and fimbriate strains of Streptococcus salivarius. Microbiology 133:3207–3217. https://doi.org/10.1099/00221287-133-11-3207
Letourneau J, Levesque C, Berthiaume F et al (2011) “In vitro”assay of bacterial adhesion onto mammalian epithelial cells. J Vis Exp. https://doi.org/10.3791/2783
Lukić J, Strahinić I, Jovčić B et al (2012) Different roles for lactococcal aggregation factor and mucin binding protein in adhesion to gastrointestinal mucosa. Appl Environ Microbiol 78:7993–8000. https://doi.org/10.1128/aem.02141-12
Aureli P, Fiore A, Scalfaro C, Franciosa G (2008) Metodi microbiologici tradizionali e metodi molecolari per l’analisi degli integratori alimentari a base di o con probiotici per uso umano. Rapp ISTISAN 08:36
Choi AR, Patra JK, Kim WJ, Kang SS (2018) Antagonistic activities and probiotic potential of lactic acid bacteria derived from a plant-based fermented food Front. Microbiol 9:1963. https://doi.org/10.3389/fmicb.2018.01963
Mora D, Fortina MG, Parini C et al (2002) Genetic diversity and technological properties of Streptococcus thermophilus strains isolated from dairy products. J Appl Microbiol 93:278–287. https://doi.org/10.1046/j.1365-2672.2002.01696.x
International Organization for Standardization (2010) ISO 10932:2010 - Milk and milk products - Determination of the minimal inhibitory concentration (MIC) of antibiotics applicable to bifidobacteria and non-enterococcal lactic acid bacteria (LAB). ISO 10932/IDF 233 Stand
EFSA Panel on Additives and Products or Substances used in Animal Feeds (2012) Guidance on the assessment of bacterial susceptibility to antimicrobials of human and veterinary importance. EFSA J 10(6):2740. https://doi.org/10.2903/j.efsa.2012.2740
Bankevich A, Nurk S, Antipov D et al (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477. https://doi.org/10.1089/cmb.2012.0021
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069. https://doi.org/10.1093/bioinformatics/btu153
Chen L, Yang J, Yu J et al (2005) VFDB: a reference database for bacterial virulence factors. Nucleic Acids Res 33:D325–D328. https://doi.org/10.1093/nar/gki008
Zhou CE, Smith J, Lam M et al (2007) MvirDB–a microbial database of protein toxins, virulence factors and antibiotic resistance genes for bio-defence applications. Nucleic Acids Res 35:D391–D394. https://doi.org/10.1093/nar/gkl791
Liu B, Pop M (2009) ARDB–Antibiotic Resistance Genes Database. Nucleic Acids Res 37:D443–D447. https://doi.org/10.1093/nar/gkn656
McArthur AG, Waglechner N, Nizam F et al (2013) The comprehensive antibiotic resistance database. Antimicrob Agents Chemother 57:3348–3357. https://doi.org/10.1128/aac.00419-13
Zankari E, Hasman H, Cosentino S et al (2012) Identification of acquired antimicrobial resistance genes. J Antimicrob Chemother 67:2640–2644. https://doi.org/10.1093/jac/dks261
de Jong A, van Hijum SAFT, Bijlsma JJE et al (2006) BAGEL: a web-based bacteriocin genome mining tool. Nucleic Acids Res 34:W273–W279. https://doi.org/10.1093/nar/gkl237
Carattoli A, Zankari E, García-Fernández A et al (2014) In silico detection and typing of plasmids using PlasmidFinder and Plasmid Multilocus Sequence Typing. Antimicrob Agents Chemother 58:3895–3903. https://doi.org/10.1128/aac.02412-14
Couvin D, Bernheim A, Toffano-Nioche C et al (2018) CRISPRCasFinder, an update of CRISRFinder, includes a portable version, enhanced performance and integrates search for Cas proteins. Nucleic Acids Res 46:W246–W251. https://doi.org/10.1093/nar/gky425
Page AJ, Cummins CA, Hunt M et al (2015) Roary: rapid large-scale prokaryote pan genome analysis. Bioinformatics 31:3691–3693. https://doi.org/10.1093/bioinformatics/btv421
Inglin RC, Meile L, Stevens MJA (2018) Clustering of pan- and core-genome of Lactobacillus provides novel evolutionary insights for differentiation. BMC Genomics 19:284. https://doi.org/10.1186/s12864-018-4601-5
Letunic I, Bork P (2006) Interactive Tree Of Life (iTOL): an online tool for phylogenetic tree display and annotation. Bioinformatics 23:127–128. https://doi.org/10.1093/bioinformatics/btl529
Emms DM, Kelly S (2015) OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biol 16:157. https://doi.org/10.1186/s13059-015-0721-2
Huerta-Cepas J, Szklarczyk D, Forslund K et al (2015) eggNOG 4.5: a hierarchical orthology framework with improved functional annotations for eukaryotic, prokaryotic and viral sequences. Nucleic Acids Res 44:D286–D293. https://doi.org/10.1093/nar/gkv1248
Saeed AI, Sharov V, White J et al (2003) TM4: A Free, open-source system for microarray data management and analysis. Biotechniques 34:374–378. https://doi.org/10.2144/03342mt01
Kang M-S, Chung J, Kim S-M et al (2006) Effect of Weissella cibaria isolates on the formation of Streptococcus mutans biofilm. Caries Res 40:418–425. https://doi.org/10.1159/000094288
Kang M-S, Kim B-G, Chung J et al (2006) Inhibitory effect of Weissella cibaria isolates on the production of volatile sulphur compounds. J Clin Periodontol 33:226–232. https://doi.org/10.1111/j.1600-051x.2006.00893.x
Lim H-S, Yeu J-E, Hong S-P, Kang M-S (2018) Characterization of antibacterial cell-free supernatant from oral care probiotic Weissella cibaria. CMU Molecules 23:1984. https://doi.org/10.3390/molecules23081984
Patel A, Prajapati JB, Holst O, Ljungh A (2014) Determining probiotic potential of exopolysaccharide producing lactic acid bacteria isolated from vegetables and traditional Indian fermented food products. Food Biosci 5:27–33. https://doi.org/10.1016/j.fbio.2013.10.002
Singh S, Bhatia R, Singh A et al (2018) Probiotic attributes and prevention of LPS-induced pro-inflammatory stress in RAW264.7 macrophages and human intestinal epithelial cell line (Caco-2) by newly isolated Weissella cibaria strains. Food Funct 9:1254–1264. https://doi.org/10.1039/c7fo00469a
Divya JB, Varsha KK, Nampoothiri KM (2012) Newly isolated lactic acid bacteria with probiotic features for potential application in food industry. Appl Biochem Biotechnol 167:1314–1324. https://doi.org/10.1007/s12010-012-9561-7
García-Hernández Y, Pérez-Sánchez T, Boucourt R et al (2016) Isolation, characterization and evaluation of probiotic lactic acid bacteria for potential use in animal production. Res Vet Sci 108:125–132. https://doi.org/10.1016/j.rvsc.2016.08.009
Ruiz L, Margolles A, Sánchez B (2013) Bile resistance mechanisms in Lactobacillus and Bifidobacterium. Front Microbiol 4:396. https://doi.org/10.3389/fmicb.2013.00396
Ojekunle O, Banwo K, Sanni AI (2017) In vitro and in vivo evaluation of Weissella cibaria and Lactobacillus plantarum for their protective effect against cadmium and lead toxicities. Lett Appl Microbiol 64:379–385. https://doi.org/10.1111/lam.12731
Lorca GL, Font de Valdez G, Ljungh A (2002) Characterization of the protein-synthesis dependent adaptive acid tolerance response in Lactobacillus acidophilus. J Mol Microbiol Biotechnol 4:525–532
Weiss G, Jespersen L (2010) Transcriptional analysis of genes associated with stress and adhesion in Lactobacillus acidophilus NCFM during the passage through an in vitro gastrointestinal tract model. J Mol Microbiol Biotechnol 18:206–214. https://doi.org/10.1159/000316421
Suokko A, Poutanen M, Savijoki K et al (2008) ClpL is essential for induction of thermotolerance and is potentially part of the HrcA regulon in Lactobacillus gasseri. Proteomics 8:1029–1041. https://doi.org/10.1002/pmic.200700925
Bustos AY, Font de Valdez G, Fadda S, Taranto MP (2018) New insights into bacterial bile resistance mechanisms: the role of bile salt hydrolase and its impact on human health. Food Res Int 112:250–262. https://doi.org/10.1016/j.foodres.2018.06.035
Lynch KM, Arendt EK, Lucid A et al (2015) Genomics of Weissella cibaria with an examination of its metabolic traits. Microbiology 161:914–930. https://doi.org/10.1099/mic.0.000053
Goh YJ, Klaenhammer TR (2013) A functional glycogen biosynthesis pathway in Lactobacillus acidophilus : expression and analysis of the glg operon. Mol Microbiol 89:1187–1200. https://doi.org/10.1111/mmi.12338
Goh YJ, Klaenhammer TR (2014) Insights into glycogen metabolism in Lactobacillus acidophilus: impact on carbohydrate metabolism, stress tolerance and gut retention. Microb Cell Fact 13:1–12. https://doi.org/10.1186/s12934-014-0094-3
Wang L, Si W, Xue H, Zhao X (2017) A fibronectin-binding protein (FbpA) of Weissella cibaria inhibits colonization and infection of Staphylococcus aureus in mammary glands. Cell Microbiol 19:e12731. https://doi.org/10.1111/cmi.12731
Abriouel H, Lerma LL, del Casado Muñoz MC et al (2015) The controversial nature of the Weissella genus: technological and functional aspects versus whole genome analysis-based pathogenic potential for their application in food and health. Front Microbiol 6:1197. https://doi.org/10.3389/fmicb.2015.01197
van Pijkeren J-P, Canchaya C, Ryan KA et al (2006) Comparative and functional analysis of sortase-dependent proteins in the predicted secretome of Lactobacillus salivarius UCC118. Appl Environ Microbiol 72:4143–4153. https://doi.org/10.1128/AEM.03023-05
Razzaque MA, Mohammed SA, Al- Mutawa T, Bedair M (2009) Improving performance of replacement heifers in hot arid environment under intensive management. Am J Anim Vet Sci 4:85–91. https://doi.org/10.3844/ajavsp.2009.85.91
Srionnual S, Yanagida F, Lin L-H et al (2007) Weissellicin 110, a newly discovered bacteriocin from Weissella cibaria 110, isolated from plaa-som, a fermented fish product from Thailand. Appl Environ Microbiol 73:2247–2250. https://doi.org/10.1128/AEM.02484-06
Papagianni M, Papamichael EM (2011) Purification, amino acid sequence and characterization of the class IIa bacteriocin weissellin A, produced by Weissella paramesenteroides DX. Bioresour Technol 102:6730–6734. https://doi.org/10.1016/j.biortech.2011.03.106
Masuda Y, Zendo T, Sawa N et al (2011) Characterization and identification of weissellicin Y and weissellicin M, novel bacteriocins produced by Weissella hellenica QU 13. J Appl Microbiol 112:99–108. https://doi.org/10.1111/j.1365-2672.2011.05180.x
Leong K-H, Chen Y-S, Lin Y-H et al (2013) Weissellicin L, a novel bacteriocin from sian-sianzih -isolated Weissella hellenica 4–7. J Appl Microbiol 115:70–76. https://doi.org/10.1111/jam.12218
Chen C, Chen X, Jiang M et al (2014) A newly discovered bacteriocin from Weissella hellenica D1501 associated with Chinese Dong fermented meat (Nanx Wudl). Food Control 42:116–124. https://doi.org/10.1016/j.foodcont.2014.01.031
Tuo Y, Yu H, Ai L et al (2013) Aggregation and adhesion properties of 22 Lactobacillus strains. J Dairy Sci 96:4252–4257. https://doi.org/10.3168/jds.2013-6547
Hill C, Guarner F, Reid G et al (2014) The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat Rev Gastroenterol Hepatol 11:506–514. https://doi.org/10.1038/nrgastro.2014.66
Jeong D-W, Lee J-H (2015) Antibiotic resistance, hemolysis and biogenic amine production assessments of Leuconostoc and Weissella isolates for kimchi starter development. LWT - Food Sci Technol 64:1078–1084. https://doi.org/10.1016/j.lwt.2015.07.031
Rodríguez-Alonso P, Fernández-Otero C, Centeno JA, Garabal JI (2009) Antibiotic resistance in lactic acid bacteria and Micrococcaceae/Staphylococcaceae isolates from artisanal raw milk cheeses, and potential implications on cheese making. J Food Sci 74:M284–M293. https://doi.org/10.1111/j.1750-3841.2009.01217.x
Morandi S, Cremonesi P, Silvetti T, Brasca M (2013) Technological characterisation, antibiotic susceptibility and antimicrobial activity of wild-type Leuconostoc strains isolated from north Italian traditional cheeses. J Dairy Res 80:457–466. https://doi.org/10.1017/s0022029913000447
Ammor MS, Belén Flórez A, Mayo B (2007) Antibiotic resistance in non-enterococcal lactic acid bacteria and bifidobacteria. Food Microbiol 24:559–570. https://doi.org/10.1016/j.fm.2006.11.001
Flórez AB, Campedelli I, Delgado S et al (2016) Antibiotic susceptibility profiles of dairy Leuconostoc, analysis of the genetic basis of atypical resistances and transfer of genes in vitro and in a food matrix. PLoS ONE 11:e0145203. https://doi.org/10.1371/journal.pone.0145203
Ripamonti B, Agazzi A, Bersani C et al (2011) Screening of species-specific lactic acid bacteria for veal calves multi-strain probiotic adjuncts. Anaerobe 17:97–105. https://doi.org/10.1016/j.anaerobe.2011.05.001
Dowarah R, Verma AK, Agarwal N et al (2018) Selection and characterization of probiotic lactic acid bacteria and its impact on growth, nutrient digestibility, health and antioxidant status in weaned piglets. PLoS ONE 13:e0192978. https://doi.org/10.1371/journal.pone.0192978
De Angelis M, Siragusa S, Berloco M et al (2006) Selection of potential probiotic lactobacilli from pig feces to be used as additives in pelleted feeding. Res Microbiol 157:792–801. https://doi.org/10.1016/j.resmic.2006.05.003
Balasingham K, Valli C, Radhakrishnan L, Balasuramanyam D (2017) Probiotic characterization of lactic acid bacteria isolated from swine intestine. Vet World 10:825–829. https://doi.org/10.14202/vetworld.2017.825-829
Oh PL, Benson AK, Peterson DA et al (2009) Diversification of the gut symbiont Lactobacillus reuteri as a result of host-driven evolution. ISME J 4:377–387. https://doi.org/10.1038/ismej.2009.123
Funding
Open access funding provided by Università Cattolica del Sacro Cuore within the CRUI-CARE Agreement. The study was supported by the Kuwait Foundation for the Advancement of Sciences (KFAS) and the Kuwait Institute for Scientific Research (KISR) through project FB075C, and by Regione Lombardia and Fondazione Cariplo within the framework of the project “Cremona Agri-food technologies (CRAFT)”.
Author information
Authors and Affiliations
Contributions
Vania Patrone: Writing—original draft, Methodology, Data curation, Investigation, Visualization. Tahani Al-Surrayai: Writing—original draft, Methodology, Data curation, Investigation, Visualization. Francesco Romaniello: Methodology, Investigation. Alessandra Fontana: Writing—original draft, Methodology, Investigation, Visualization. Giovanni Milani: Methodology, Investigation. Edoardo Puglisi: Writing—original draft, Methodology, Data curation, Investigation, Visualization. Maria Luisa Callegari: Writing—original draft, Methodology, Data curation, Investigation, Visualization. Hamad Al-Mansour: Conceptualization, Supervision, Writing – review & editing; Mohamed Waheed Kishk: Conceptualization, Supervision, Writing – review & editing; Lorenzo Morelli: Conceptualization, Supervision, Writing – review & editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
Not applicable
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Patrone, V., Al-Surrayai, T., Romaniello, F. et al. Integrated Phenotypic-Genotypic Analysis of Candidate Probiotic Weissella Cibaria Strains Isolated from Dairy Cows in Kuwait. Probiotics & Antimicro. Prot. 13, 809–823 (2021). https://doi.org/10.1007/s12602-020-09715-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12602-020-09715-x