Abstract
Discovery of a novel bacteriocin is always an event in sciences, since cultivation of most bacterial species is a general problem in microbiology. This statement is reflected by the fact that number of bacteriocins is smaller for tenfold comparing to known antimicrobial peptides. We cultivated Enterococcus faecium on simplified medium to reduce amount of purification steps. This approach allows to purify the novel heavy weight bacteriocin produced by E. faecium ICIS 7. The novelty of this bacteriocin, named enterocin-7, was confirmed by N-terminal sequencing and by comparing the structural-functional properties with available data. Purified enterocin-7 is characterized by a sequence of amino acid residues having no homology in UniProt/SwissProt/TrEMBL databases: NH2 - Asp - Ala - His - Leu - Ser - Glu - Val - Ala - Glu - Arg - Phe - Glu - Asp - Leu - Gly. Isolated thermostable protein has a molecular mass of 65 kDa, which allows it to be classified into class III in bacteriocin classification schemes. Enterocin-7 displayed a broad spectrum of activity against some Gram-positive and Gram-negative microorganisms. Fluorescent microscopy and spectroscopy showed the permeabilizing mechanism of the action of enterocin-7, which is realized within a few minutes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
During evolution, microorganisms acquire an effective chemical arsenal that regulates inter-microbial relationships in communities. Bacteriocins are biologically active protein moieties, which determine antagonistic activity of bacteria and play a significant role in the ability to widespread [1]. According to their structural and functional characteristics, bacteriocins can be related to antimicrobial peptides (AMPs) produced by eukaryotes. Meanwhile bacteriocins are a relatively small group compared to other AMPs, since they are included in about 10% of the total known AMPss [2]. This disparity is due to the fact that most bacterial species are uncultivated in laboratory conditions [3]; in addition, separation of bacteriocins from microbial metabolites is an another problem [4]. Partial solution of stated problems is provided by searching for new bacteriocins using genome database mining approach [5, 6]. Nevertheless, purification of novel bacteriocins is a significant event in microbiology and related sciences. Every finding expands the knowledge about the microbial world and can be used in biotechnological or medical sectors [7].
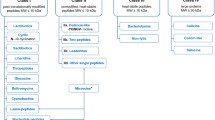
The classification of bacteriocins undergoes changing and updating, which led to the existence of several classification schemes [8]. In our opinion, the most objective classification includes three groups of bacteriocins [9]. The first group includes bacteriocins that are subjected to enzymatic modifications during biosynthesis, as the mature peptide contains unusual amino acids. The second group of bacteriocins is the most diverse, characterized by molecular mass of less than 10 kDa. The third group is represented by proteins with molecular mass of more than 30 kDa. Significant numbers of novel bacteriocins were classified to the first and second groups, but few bacteriocins were classified in the third group [8].
We report on the purification and structural-functional features of enterocin-7, which is a novel bacteriocin produced by Enterococcus faecium ICIS 7 strain isolated from human feces. This strain is characterized by the presence of antagonistic activity against Gram-positive and Gram-negative strains.
Achievement of the stated goal was realized through complex approaches. Initially, we performed optimization of medium composition to reduce chromatographically steps. At the next stage, the one-step liquid high-performance chromatography was successfully used. The mode of action was assessed using the fluorescent spectroscopy and routine bacteriological methods.
Materials and Methods
Cultivation Condition
Investigations of growth and bacteriocin production were performed using the bacteriocin-producing strain E. faecium ICIS 7. Bacteria were incubated in 50-ml flacons at 37 °C for 24–36 h on a simplified media (Table 1). The dynamic of bacterial growth was assessed by reading and plotting the absorbance data at 620 nm obtained by spectrophotometer IEMS MF (Labsystems, Finland). Evaluation of the bacteriocin content was performed by isolating an aliquot of the culture medium followed by centrifugation (20 min, 4 °C, 9000 g) and then filtering through a Durapore PVDF filter with a pore size of 0.22 μm (Millipore). The activity of the supernatant was evaluated by agar diffusion assay (described below) against Listeria monocytogenes EGDe and was expressed in arbitrary units per milliliter, calculated according to formula (1) [10]:
where,
- V:
-
supernatant volume
- D:
-
dilution factor
Purification of Bacteriocin
The chromatography manipulation was performed as described earlier [11]. In the next step, desalting of the culture medium was performed using reversed-phase low-pressure chromatography on an FPLC column completed with a Synchroprep RP-P C18 phase (SynChrom Inc., USA) equilibrated with solvent A (10% acetonitrile; Merck, Germany) in ultra-purified water (18 Mohms, Milli-Q, Millipore) with 0.1% trifluoroacetic acid (TFA). Elution was performed using solvent B (80% acetonitrile in ultra-purified water with 0.1% TFA) followed by lyophilization to withdraw a residual quantity of TFA and concentration. The obtained desalted extract was tested to reveal antibacterial properties. The lyophilized mass was resuspended in ultrapure water and applied to semi-preparative column C18 Luna (250 × 10 mm, Phenomenex, USA) integrated into a high-performance liquid chromatography (HPLC) system (Knauer SmartLine 200, Knauer, Germany). Elution was performed using solvent B (80% acetonitrile in ultrapure water with 0.1% TFA) in a linear gradient according to the following scheme: 5–80% for 60 min at a flow rate of 2.0 ml/min. Absorbance was detected at 214 nm.
Structural Analysis of Enterocin-7
SDS-PAGE of the Active Fraction
The electrophoretic separation of the active fraction obtained by RP-HPLC was conducted according to the protocol described [12]. To an analyzed sample an equal volume of buffer containing 0.125 M Tris-HCl, pH 6.8, 4% SDS, 20% glycerol, 10% 2-mercaptoethanol was added. 1 ml of a 0.2% solution of bromophenol blue was mixed with the sample and applied to the wells of the polyacrylamide plate consisting of 4% in the stacking gel and 10% in the separating gel. The phosphorylase B (97 kDa), bovine serum albumin (66 kDa), ovalbumin (45 kDa), carbonic anhydrase (30 kDa), trypsin inhibitor (20.1 kDa), and α-lactalbumin (14.4 kDa) (Amersham Biosciences, USA) were used as the markers of molecular mass of the proteins.
After the electrophoresis (1.5–2.0 h) gel was transferred to PVDF membrane “Immobilone,” the membrane was stained with 0.1% solution of Coomassie R-250 in 50% aqueous methanol for 3–5 min, and then washed with water.
N-Terminal Sequencing
The proteins were sequenced by automated Edman degradation on a model PPSQ-33A protein sequencer (Shimadzu Corp., Japan) according to the manufacturer’s protocol. Cysteine residues were determined as pyridilethylated derivatives. Homology search was performed using databases with a BLASTP algorithm.
Effect of Enzymes and Temperature Treatment on Activity
The protease susceptibility was carryout by incubation of enterocin-7 with 10 mg mL of protease K and 20 mg mL of trypsin (biotechnology grade, Sigma-Aldrich, USA) at 37 °C for 1 h in buffer (5 mM HEPES, pH 7.5) as described in [13]. Enterocin-7 in buffer without enzymes was used as positive control; enzyme in buffer solution and buffer alone were used as negative controls. Evaluation of thermostability of enterocin-7 was performed by heating in the solid-state thermostat “Thermit” designed for microfuge tubes (“DNK-Technologiya,” Russia). The inhibitory activity of active fraction after treatments were determined using the agar diffusion assay.
Determination of Antimicrobial Activity by Agar Diffusion Assay
Determination of bacterial metabolites’ bactericidal properties, HPLC fractions, and purified enterocin-7 was performed using the agar diffusion assay [14]. Briefly, the microorganisms (Table 3) were cultured for 18 h in LB broth (HiMedia, India), after which 50 μL of the bacterial suspension (containing ~ 107 CFU) was mixed with 10 mL of soft (0.6%) LB agar and placed immediately over a Petri dish, which was previously overlaid with a 1.5% agar plate. The test samples were transferred on the upper layer of soft agar. After incubation at 37 °C overnight, inhibitory areas were observed.
Characterization of Mode of Action of Enterocin-7
Determination of Minimal Inhibitory Concentration
To evaluate the minimal inhibitory concentration of enterocin-7, L. monocytogenes EGDe was incubated in 96-wells microtiter plate with twofold dilutions of the enterocin-7 according to protocol [15]. Dynamic of bacterial growth was assessed by reading and plotting the absorbance data at 620 nm obtained by spectrophotometer IEMS MF (Labsystems, Finland). Antimicrobial activity was expressed by the minimal inhibitory concentration (MIC), which was defined as the lowest peptide dose at which no visible growth were detected.
Permeabilization of the Plasma Membrane
The LIVE/DEAD BacLight bacterial viability kit (Molecular Probes, USA) was used to evaluate the cytoplasmatic membrane integrity of L. monocytogenes EGDe according to the manufacture’s protocol. The fluorescence microscopy of stained bacteria was performed using a Mikromed-3-Lum fluorescent microscope (Lomo, Russia) equipped with filter sets useful for simultaneous viewing of SYTO 9 and PI stains.
Measurement of SYTO 9 fluorescence kinetic was performed using a plate reader Infinite F200 pro (Tecan, Austria) at 485 nm emission and 535 nm of excitation wavelength as described earlier [16]. Indolicidin and pure water were used as positive and negative controls, respectively.
Statistical Analysis
All experiments were preformed using three independent series.
Results
Purification of Enterocin-7 and Structure Determination
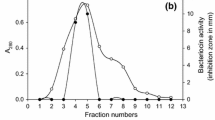
E. faecium ICIS 7 was quickly adapted to the simplified nutrient medium (Table 1) since the exponential phase was finished at 6 h of cultivation. Bacteriocin production was observed in the middle of the stationary phase of bacterial growth (Fig. 1). After cultivation, the growth medium was desalted by low-pressure liquid chromatography and, after evaporation of acetonitrile, applied on a semi-preparative C18 reverse-phase HPLC column. The chromatographic profile obtained was characterized by 23 main peaks (Fig. 2a) among which antimicrobial activity was demonstrated for the most hydrophobic fractionwith a retention time of 34–37 min. The active fraction demonstrates thermostability when heated at 90 °C for 30 min (Table 1). Enzymatic treatments with proteinase K and trypsin led to complete loss of activity, which is preliminarily suggested about a proteinaceous nature of the active substance (Table 2).
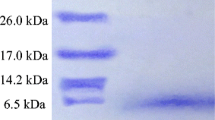
PAAG electrophoresis of the concentrated fraction in native denaturated and denaturated/reduced conditions revealed two separated stained bands characterized by similar mobility in polyacrylamide gel (Fig. 2b). The upper band have comparable electrophoretic mobility with the bovine serum albumin, and its molecular mass was determined to be about 65 kDa, whereas the bottom band was approximately 50–55 kDa. A preparative PAAG electrophoresis, followed by electroblotting to Immobilon PVDF hydrophobic membrane, was carried out for structure identification of detected proteins (named Ent-1 and Ent-2 according to Fig. 1b). Both bands were cut and analyzed separately by Edman sequencing. As a result, for Ent-1, we could estimate 15 N-terminal amino acid residues: NH2 - Asp - Ala - His - Leu - Ser - Glu - Val - Ala - Glu - Arg - Phe - Glu - Asp - Leu-Gly; for Ent-2, only eight amino acid residues were identical to Ent-1. Thus, both isolated proteins are homologous, but differences in molecular mass hypothetically can be explained by partial absence of amino acid residue at the C-terminal region of the Ent-2. Concerning homology analysis using a BLASTP algorithm, there was no identity among E. faecium deposited in UniProt/SwissProt/TrEMBL primary structure databases; the same result was obtained for the whole Enterococcus genus. When result had analyzed, no translated amino acid sequences (including ESTs) were found. We tentatively designated this bacteriocin enterocin-7.
Biological Activity of Enterocin-7
Qualitative evaluation revealed the ability of enterocin-7 to inhibit growth of some Gram-positive and Gram-negative microorganisms (Table 3). Agar diffusion assay revealed a more pronounced inhibitory effect of enterocin-7 on Gram-positive than on Gram-negative strains. The most sensitive strain of L. monocytogenes EGDe was selected for further studies as a model. The addition of enterocin-7 to the culture medium inoculated by L. monocytogenes EGDe allowed to record of the minimal inhibitory concentration (MIC), which was 1.75 μg mL (Fig. 3).
Effects of enterocin-7 treatment on bacterial membranes were evaluated with fluorescent spectroscopy and microscopy imaging. Using a commercial staining kit containing DNA-intercalating dyes, we visualized treated bacterial cells with a permeabilized membrane (Fig. 4b). Moreover, we traced the kinetic of fluorophore accumulation within bacterial cells and found that enterocin-7 was able to disturb cell’ membranes very quickly (Fig. 4с). The lightweight green fluorescence SYTO 9 (~ 400 Da) freely penetrates even through integral bacterial membrane, in contrast heavier red fluorescence PI (668 Da) is accumulated inside only through disordered plasma membrane. Membrane disordering was spectroscopically recorded as quenching of the green fluorescence since the SYTO 9 replaces by PI.
The fluorescence microscopy images of intact (a) cells of L. monocytogenes EGDe and treated with enterocin-7 (b). Dynamics of permeation of SYTO 9 into bacterial cells treated with AMPs (c): enterocin-7 at 2 MIC; indolicidin at 2 MIC (positive control). Arrows show the time when test substances were added. If bacterial membranes are permeabilized, PI penetrates into the cell after displacing the SYTO 9 from DNA, which leads to decreasing fluorescence intensity in a green region of the spectrum (emission at 535 nm). The scale bar - 10 µm
Discussion
Enterococci are among of the most important intestinal microbiota of humans and animals [17]. The significance of Enterococci for human is controversial, because while known to cause infectious disease, but they are still actively used in the food industry. Benefits which are provided by enterococci partially are determined through bacteriocin’s production. For example, bacteriocin-producing enterococcus introduced in the gut can able to replace indigenous enterococci including vancomycin-resistant strains, which are lacking bacteriocins [18].
Unlike peptides of animal origin, bacteriocins are much more difficult to study, because bacteria usually synthesize bacteriocins in small amounts, while their extraction and purification from microbial metabolites is a non-trivial task. The main problems of purification are related to the fact that bacteria grow and produce bacteriocins in multi-component media saturated with various proteins and peptides (for example, beta-casein hydrolyzate). To obtain a homogeneous form of bacteriocins, a variety of isolation and purification methods needs to be used such as precipitation, ion-exchange chromatography, and multiple reversed-phase HPLC (semi-preparative and analytical) [8, 19].
Effectiveness of liquid chromatography may be increased due to cultivation on media with a minimum content of proteins and other hydrophobic components that impede purification of bacteriocins [20, 21]. For preparation of the simplified medium, we used only inorganic salts and yeast extract. According to the manufacturer (Becton Dickenson, BBL), the used yeast extract consisted of 70% peptides with molecular masses of about 250 Da and 20% with 0.5–2.0 kDa. Thus, using the yeast extract as a one component for cultivation medium reduces loading of chromatographically column by proteins and peptides. Respectively, this approach decreases number of purification steps and increases the efficiency of bacteriocin purification from E. faecium metabolites. Of course, not all strains grow and produce bacteriocin on media with a minimal nutrient content. Moreover, using of specialized rich media such as MRS or Shaedler increase production of bacteriocins, but ultimately it complicates purification and reduces the bacteriocin content at each step of chromatography.
Today, performed enquiry to the antimicrobial peptide database (APD) resulted in 36, entries, which are corresponded to various annotated enterocins [2]. The greatest number of enterocins belongs to class II, only a few large proteins related to class III have been revealed [22, 23]. According to the existing classification of bacteriocins, large-molecular- mass and heat-labile antimicrobial proteins are classified as class III [8, 9]. If compared with existing representatives, enterocin-7 has significant differences. Apparently, purified active fraction is represented by large protein with a molecular mass of about 65 kDa. Currently, only bacteriocin 41 (Bac41), produced by E. faecalis have a similar molecular mass > 60 kDa (GenBank ID: BAH02371.1) [22]. It is important to note that enterocin-7 had no homologue with previously isolated bacteriocins and predicted sequences encoded with potential bacteriocin-like proteins. Comparison of the first 15 amino acids of enterocin-7 with existing protein sequences from the Enterococcus genus and Bacteria kingdom revealed no sequence similarity to any known proteins or peptides.
Secondly, enterocin-7 is a heat-stable protein. Only a few high-molecular-mass bacteriocins are known, but they are characterized by heat-labile property [22, 24,25,26,27,28]. Previously, we isolated a novel heat-stable bacteriocin-like protein, which is secreted by the E. faecium ICIS 8 into culture liquid. This protein has a molecular mass of about 14 kDa, and consists two Cys residues located in the N-terminal part of the molecule [11].
In addition, enterocin-7 is active against some Gram-negative bacteria, which is intrinsic to bacteriocins of low molecular mass, for example enterocin E-760 [19]. The ability to inhibit bacteria with a Gram-negative cell wall type is a property of a relatively small fraction of known bacteriocins. According to the APD database, enterocins which are inhibited both Gram-positive and Gram-negative bacteria constitute the minor group (< 15%). It is difficult to identify any physico-chemical features of enterocins that determine their ability to kill Gram-negative bacteria. Whereas anti-Gram-negative activity revealed for various enterocins including lantibiothics [29], circular [30], and two-chain peptide [31].
Another feature of enterocin-7 is rapid permeabilization of bacterial membranes like peptides, but not proteins. For example, interaction of lacticin 3147 with Gram-positive bacteria resulted in potassium efflux immediately after addition [32]; enterococcal cytolysin interact in a similar way [33]. In turn, known proteins related to class III have endopeptidase enzymatic activity that is realized within several hours [24, 34]. The time-killing dynamic similar with enterocin-7 was described for zoocin A that stopped growth of S. mutans within 10 min [35].
Thus, the isolated enterocin-7 has some differences from known enterocins; moreover, it differs from other bacteriocins too.
Conclusion
In the present work, according to the results of amino acid sequencing, we report firstly about a novel bacteriocin. On the one hand, a molecular mass of 65 kDa allows to classify enterocin-7 as a class III bacteriocin. On the other hand, a number of features are not in accordance with existing data. However, the small number of bacteriocins of class III does not allow us to speak unambiguously about distinctive or non-distinctive features. Further study will be focused on revealing the complete primary structure of enterocin-7 and detailed molecular characteristics of interaction with cellular targets.
References
Riley MA, Wertz JE (2002) Bacteriocins: evolution, ecology, and application. Annu Rev Microbiol 56(1):117–137. https://doi.org/10.1146/annurev.micro.56.012302.161024
Wang G, Li X, Wang Z (2016) APD3: the antimicrobial peptide database as a tool for research and education. Nucleic Acids Res 44(D1):D1087–D1093. https://doi.org/10.1093/nar/gkv1278
Epstein SS (2009) General model of microbial uncultivability. In: Epstein SS (ed) Uncultivated microorganisms. Springer, Heidelberg, Germany, pp 131–159
Pingitore VE, Salvucci E, Sesma F, Nader-Macías ME (2007) Different strategies for purification of antimicrobial peptides from lactic acid bacteria (LAB). Communicating current research and educational topics and trends in applied microbiology a. Méndez-Vilas(Ed). Formatex, 557-568
Carson DA, Barkema HW, Naushad S, De Buck J (2017) Bacteriocins of non-aureus staphylococci isolated from bovine milk. Appl Environ Microbiol 83(17):e01015–e01017. https://doi.org/10.1128/AEM.01015-17
Sandiford SK (2017) Genome database mining for the discovery of novel lantibiotics. Expert Opin Drug Discov 12(5):489–495. https://doi.org/10.1080/17460441.2017.1303475
Chikindas ML, Weeks R, Drider D, Chistyakov VA, Dicks LM (2018) Functions and emerging applications of bacteriocins. Curr Opin Biotechnol 49:23–28. https://doi.org/10.1016/j.copbio.2017.07.011
Kaškonienė V, Stankevičius M, Bimbiraitė-Survilienė K, Naujokaitytė G, Šernienė L, Mulkytė K, Malakauskas M, Maruška A (2017) Current state of purification, isolation and analysis of bacteriocins produced by lactic acid bacteria. Appl Microbiol Biotechnol 101(4):1323–1335. https://doi.org/10.1007/s00253-017-8088-9
Alvarez-Sieiro P, Montalbán-López M, Mu D, Kuipers OP (2016) Bacteriocins of lactic acid bacteria: extending the family. Appl Microbiol Biotechnol 100(7):2939–2951. https://doi.org/10.1007/s00253-016-7343-9
Yamamoto Y, Togawa Y, Shimosaka M, Okazaki M (2003) Purification and characterization of a novel bacteriocin produced by Enterococcus faecalis strain RJ-11. Appl Environ Microbiol 69(10):5746–5753. https://doi.org/10.1128/AEM.69.10.5546-5553.2003
Vasilchenko AS, Rogozhin EA, Valyshev AV (2017) Purification of a novel bacteriocin-like inhibitory substance produced by Enterococcus faecium ICIS 8 and characterization of its mode of action. Microb Drug Resist 23(4):447–456. https://doi.org/10.1089/mdr.2016.0069
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227(5259):680–685. https://doi.org/10.1038/227680a0
Gupta A, Tiwari SK, Netrebov V, Chikindas ML (2016) Biochemical properties and mechanism of action of enterocin LD3 purified from Enterococcus hirae LD3. Probiotics Antimicrob Proteins 8(3):161–169. https://doi.org/10.1007/s12602-016-9217-y
Bell SC, Grundy WE (1968) Preparation of agar wells for antibiotic assay. Appl Microbiol 16(10):1611–1612
Wiegand I, Hilpert K, Hancock REW (2008) Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat Protoc 3(2):163–175. https://doi.org/10.1038/nprot.2007.521
Vasilchenko AS, Vasilchenko AV, Pashkova TM, Smirnova MP, Kolodkin NI, Manukhov IV, Zavilgelsky GB, Sizova EA, Kartashova OL, Simbirtsev AS, Rogozhin EA, Duskaev GK, Sycheva MV (2017) Antimicrobial activity of the indolicidin-derived novel synthetic peptide In-58. J Pept Sci 23(12):855–863. https://doi.org/10.1002/psc.3049
Fisher K, Phillips C (2009) The ecology, epidemiology and virulence of Enterococcus. Microbiology 155(6):1749–1757. https://doi.org/10.1099/mic.0.026385-0
Kommineni S, Bretl DJ, Lam V, Chakraborty R, Hayward M, Simpson P, Cao Y, Bousounis P, Kristich CJ, Salzman NH (2015) Bacteriocin production augments niche competition by enterococci in the mammalian GI tract. Nature 526(7575):719–722. https://doi.org/10.1038/nature15524
Garsa AK, Kumariya R, Sood SK, Kumar A, Kapila S (2014) Bacteriocin Production and Different Strategies for Their Recovery and Purification. Probiotics Antimicrob Proteins 6(1):47–58.
Khan H, Flint SH, Yu PL (2013) Development of a chemically defined medium for the production of enterolysin A from Enterococcus faecalis B9510. J Appl Microbiol 114(4):1092–1102. https://doi.org/10.1111/jam.12115
Liu W, Zhang L, Yi H (2017) Development of a chemically defined medium for better yield and purification of enterocin Y31 from Enterococcus faecium Y31. Journal of Food Quality vol. 2017, Article ID 9017452, 8 pages, 2017. doi:https://doi.org/10.1155/2017/9017452
Tomita H, Kamei E, Ike Y (2008) Cloning and genetic analyses of the bacteriocin 41 determinant encoded on the Enterococcus faecalis pheromone-responsive conjugative plasmid pYI14: a novel bacteriocin complemented by two extracellular components (lysin and activator). J Bacteriol 190(6):2075–2085. https://doi.org/10.1128/JB.01056-07
Nes IF, Diep DB, Ike Y (2014) Enterococcal bacteriocins and antimicrobial proteins that contribute to niche control, In Enterococci: from commensals to Leading Causes of Drug Resistant Infection Boston: Massachusetts Eye and Ear Infirmary
Nilsen T, Nes IF, Holo H (2003) Enterolysin A, a cell wall-degrading bacteriocin from Enterococcus faecalis LMG 2333. Appl Environ Microbiol 69(5):2975–2984. https://doi.org/10.1128/AEM.69.5.2975-2984.2003
Joerger MC, Klaenhammer TR (1986) Characterization and purification of helveticin J and evidence for a chromosomally determined bacteriocin produced by Lactobacillus helveticus 481. J Bacteriol 167(2):439–446. https://doi.org/10.1128/jb.167.2.439-446.1986
Beukes M, Bierbaum G, Sahl H-G, Hastings JW (2000) Purification and partial characterization of a murein hydrolase, millericin B, produced by Streptococcus milleri NMSCC 061. Appl Environ Microbiol 66(1):23–28. https://doi.org/10.1128/AEM.66.1.23-28.2000
Simmonds RS, Naidoo J, Jones CL, Tagg JR (1995) The streptococcal bacteriocin-like inhibitory substance, zoocin A, reduces the proportion of Streptococcus mutans in an artificial plaque. Microb Ecol Health Dis 8(6):281–292. https://doi.org/10.3109/08910609509140107
Schindler CA, Schuhardt VT (1964) Lysostaphin: a new bacteriolytic agent for the staphylococcus. Proc Natl Acad Sci U S A 51(3):414–421. https://doi.org/10.1073/pnas.51.3.414
Booth MC, Bogie CP, Sahl HG, Siezen RJ, Hatter KL, Gilmore MS (1996) Structural analysis and proteolytic activation of Enterococcus faecalis cytolysin, a novel lantibiotic. Mol Microbiol 21(6):1175–1184. https://doi.org/10.1046/j.1365-2958.1996.831449.x
Gálvez A, Maqueda M, Valdivia E, Quesada A, Montoya E (1986) Characterization and partial purification of a broad spectrum antibiotic AS-48 produced by Streptococcus faecalis. Can J Microbiol 32(10):765–771. https://doi.org/10.1139/m86-141
Franz CM, Grube A, Herrmann A, Abriouel H, Stärke J, Lombardi A, Tauscher B, Holzapfel WH (2002) Biochemical and genetic characterization of the two-peptide bacteriocin enterocin 1071 produced by Enterococcus faecalis FAIR-E 309. Appl Environ Microbiol 68(5):2550–2554. https://doi.org/10.1128/AEM.68.5.2550-2554.2002
Wiedemann I, Böttiger T, Bonelli RR, Wiese A, Hagge SO, Gutsmann T, Seydel U, Deegan L, Hill C, Ross P, Sahl HG (2006) The mode of action of the lantibiotic lacticin 3147- a complex mechanism involving specific interaction of two peptides and the cell wall precursor lipid II. Mol Microbiol 61(2):285–296. https://doi.org/10.1111/j.1365-2958.2006.05223.x
Van Tyne D, Martin MJ, Gilmore MS (2013) Structure, function, and biology of the Enterococcus faecalis cytolysin. Toxins 5(5):895–911. https://doi.org/10.3390/toxins5050895
Kurushima J, Nakane D, Nishizaka T, Tomita H (2015) Bacteriocin protein BacL 1 of Enterococcus faecalis targets cell division loci and specifically recognizes L-Ala 2-cross-bridged peptidoglycan. J Bacteriol 197(2):286–295. https://doi.org/10.1128/JB.02203-14
Simmonds RS, Pearson L, Kennedy RC, Tagg JR (1996) Mode of action of a lysostaphin-like bacteriolytic agent produced by Streptococcus zooepidemicus 4881. Appl Environ Microbiol 62(12):4536–4541
Funding
This work was partially supported by the Program of fundamental research of Ural branch of RAS, project No. 15-4-4-28 “Analysis of the resistom, its phenotypic manifestations and QS-systems of bacterial populations in order to develop a new methods for overcoming the antibiotic resistance of microorganisms.”
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interests
The authors declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Vasilchenko, A.S., Vasilchenko, A., Valyshev, A. et al. A Novel High-Molecular-Mass Bacteriocin Produced by Enterococcus faecium: Biochemical Features and Mode of Action. Probiotics & Antimicro. Prot. 10, 427–434 (2018). https://doi.org/10.1007/s12602-018-9392-0
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12602-018-9392-0