Abstract
Bovine mastitis causes economic losses on dairy farms worldwide. Lactic acid bacteria (LAB) in animal health are an alternative tool to avoid antibiotic therapy on the prevention of bovine mastitis. In previous studies, 12 LAB isolated from bovine milk were selected taking into account some of the following characteristics: hydrophobicity, auto aggregative capability, inhibition of indicator pathogens, hydrogen peroxide, and capsular polysaccharide production. These LAB were considered because of their beneficial properties. In this work, we also analyzed the antimicrobial activity and the co-aggregation against mastitis causing bacteria, auto-inhibition, adhesion to bovine teat canal epithelial cells (BTCEC), and growth kinetic curves for the 12 LAB. Two of them, Lactococcus lactis subsp. lactis CRL 1655 and Lactobacillus perolens CRL 1724, were selected because they had an interesting pattern of adhesion to BTEC, the inhibition of pathogens and the co-aggregation with the 100% of the assayed pathogens. They showed a predictable difference in the PFGE genomic pattern bands. The kinetic growth of these two strains was similar between them and with the rest of the assayed LAB. The strains selected in the present study showed indispensable characteristics for their inclusion in a probiotic formulation to be used at dry-off period for the prevention of bovine mastitis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
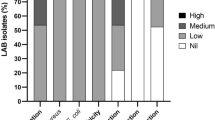
Bovine mastitis is defined as the inflammation of the mammary gland where the tissue of the udder is severely affected. It is considered as the major endemic and prevalent disease of dairy cattle [1,2,3] and is generally caused by microorganisms [4,5,6]. In approximately 70% of mastitis cases, microorganisms can be isolated. In the most cases, the infection occurs when bacteria get into the udder through the teat canal and multiply causing an inflammation with or without clinical signs (clinical or subclinical mastitis, respectively) [1]. The disease causes considerable distress on the animal, a decreased milk production, and major economic losses on dairy farms worldwide [7]. Different levels of economic losses have been reported in different countries [8,9,10]. The conventional methods for the control of bovine mastitis include preventive strategies such as diagnosis, segregation of the animals and improved hygiene [11, 12]. These practices diminish the appearance of the disease, but the control of bovine mastitis still relies heavily on the use of therapeutic and preventive protocols with antibiotics [13]. Antibiotics are usually applied during lactation and also in dry cow therapies. The susceptibility of cows to mastitis increases significantly during the early days of the dry-off period, as well as in the peripartum, due to the immunosuppression that the animals develop during this period. Even though antibiotics improve cows’ health and their consequent milk production, the risk of bacterial antibiotic resistance still exists and the presence of residues in milk makes their use questionable for human consumption [14, 15]. Due to these weaknesses, the developments of complementary and natural alternatives are very appealing to prevent the disease [1]. During the last years, the use of probiotics, which are “live microorganisms administered in adequate amounts conferring a health benefit on the host” [16], constitute an alternative tool for the prevention of bovine mastitis. The use of probiotic bacteria has been widely studied as a novel approach to prevent infections in animals, especially in the gastrointestinal and vaginal tract [17, 18], but few studies are directed to investigate the application of probiotics in the mammary gland [19,20,21,22,23]. Probiotics may exert their beneficial effects on the host health through several mechanisms: adhesion to epithelial cells, colonization, production of biosurfactants, auto-aggregation and co-aggregation of phatogens, production of antagonistic metabolites (organic acids, hydrogen peroxide, bacteriocins), competition for nutrients and production of enzymes, and/or immunomodulation [17]. Lactic acid bacteria (LAB) are the main components of the indigenous microbiota of the teat canal and because of this they are optimal candidates to design a species-specific probiotic product to prevent mastitis [24]. In previous studies [25, 26], LAB isolated from bovine milk were selected taking into account at least one of the following characteristics: high or medium degree of hydrophobicity, high or medium auto aggregative capability, inhibition of some of the indicator pathogens, high hydrogen peroxide production, and exopolysaccharides or capsular polysaccharide production. In the present work, 12 selected LAB were analyzed with the aim of studying their adhesion to bovine teat canal epithelial cells and their capability to bind to other bacteria of the same species (co-aggregate) and to mastitis pathogens (immobilization) to exert the antimicrobial effect.
These properties are very valuable to select potentially beneficial strains to be included in a probiotic product to prevent bovine mastitis.
Materials and Methods
Bacterial Strains and Culture Conditions
LAB strains analyzed in this study were previously isolated from the milk of healthy Holstein quarters from Córdoba and Tucumán, Argentina and were genetically identified by 16S rRNA gene sequencing as Lactobacillus (Lact.) perolens CRL 1724 (Centro de Referencia para Lactobacilos, Culture Collection), Lact. plantarum CRL 1716, Lactococcus (L.) lactis subsp. lactis CRL 1655, Enterococcus (Ent.) mundtii CRL 1656, Pediococcus (Ped.) pentosaceus CRL 1831, Ped. pentosaceus CRL 1832, Weissella (W.) cibaria CRL 1840, W. cibaria CRL 1833, Ent. hirae CRL 1834, Ent. hirae CRL 1835, Ent. hirae CRL 1837, and Ent. hirae 7-3. These strains were selected being as potentially probiotics because of their hydrophobicity index, auto-aggregation, and ability to produce antagonistic metabolites (organic acids, hydrogen peroxide and bacteriocins) [25, 26]. LAB strains were grown in de Man, Rogosa, and Sharpe (MRS, Britania) broth and incubated and stored as described by Frola et al. [27]. Before performing additional studies, bacteria were subcultured three times, every 12–14 h at 37 °C in MRS broth. The following mastitis-causing bacteria (MCB) were used to assess antagonistic activity and to assess co-aggregation assays: Streptococcus (Strep.) agalactiae ATCC27956, Strep. dysgalactiae ATCC27957, Strep. uberis 102, Strep. uberis ATCC27958, Strep. bovis ATCC27960, Staphylococcus (Staph.) hyicus 112,249, Staph. epidermidis ATCC14990, Staph. aureus ATCC25923, Ent. faecalis 19,433, Ent. faecium 35,667, Escherichia (E.) coli ATCC35218, and Klebsiella (Kl.) pneumoniae ATCC10031. These bacteria were kindly provided by Dr. Odierno (Universidad Nacional de Río Cuarto, Argentina). Staph. aureus RC108, E. coli 345, and Pseudomonas spp. were isolated from the milk of cows with bovine mastitis and identified in our laboratory. All these strains were cultivated and stored as described by Frola et al. [28].
Antimicrobial Activity Against Mastitis Causing Bacteria (MCB)
The in vitro antimicrobial activity of LAB strains against MCB was assayed through the streak line method [29]. The inhibitory effect was estimated as the width of the inhibition zone and ranked as high (> 25 mm, +++), intermediate (13–25 mm, ++), low (1–12 mm, +), and no inhibition (0 mm, −). The assay was performed in triplicate.
Co-aggregation Assay
To assess the interaction between LAB strains and MCB, the method described by Reid et al. [30] was followed. A suspension of each LAB adjusted to a concentration of 109 cfu mL−1 in 1 M of phosphate-buffered saline (PBS) (pH 6.2) was mixed with 500 μL of 108 cfu mL−1 of each MCB and incubated at 37 °C in an orbital shaker at 2 g for 4 h. After incubation, suspensions were Gram-stained and observed under an optical microscope. Pure cultures were used as negative controls and assays were done in duplicate. Positive co-aggregation was considered when LAB strains were attached or close to assayed MCB. This was reported as negative (−) when there was no co-aggregation in five observed areas and one positive rood (+), two positive roods (++), and three positive roods (+++) were reported when co-aggregation was observed in one to two, three to four, and five of the five observed areas.
Adhesion to BTCEC
The adhesion of LAB strains to BTCEC was determined through the methodology described previously by Otero and Nader-Macías [31] with the modifications described by Frola et al. [27]. Bacterial binding to BTCEC was examined by optical microscopy (Gram stain) and results expressed as (1) percentage of adhesion (PA): (number of BTCEC with bacteria adhered/total number of BTCEC) × 100 and (2) adhesion index (AI): (total number of bacteria attached to BTCEC/total number of cells with bacteria adhered). The application of the index allowed us to evaluate the efficiency of adhesion. The experiment was done in triplicate.
Auto-inhibition Assay
The antimicrobial activity between LAB strains was assayed through the streak line method [29]. The inhibitory effect was estimated as the width of the inhibition zone and ranked as high (> 25 mm, +++), intermediate (13–25 mm, ++), low (1–12 mm, +), and no inhibition (0 mm, −). The assay was performed in triplicate.
LAB Growth Curves
One milliliter of an overnight culture of each LAB strains was inoculated in 20 mL of sterile MRS. The cultures were incubated at 37 °C, and the optical density at 600 nm (OD600) and colony forming units (cfu) per milliliter were determined immediately after inoculation and 2, 4, 6, 8, 10, and 24 h after inoculation. The cfu per milliliter was determined by plating serial dilutions of each culture in MRS agar and incubated 24 h at 37 °C. The experiment was done in triplicate. The specific growth rate (μ) was calculated from the exponential phase of the growth curve using the Malthus model [32].
Genomic Profile of Selected LAB Strains by PFGE
Preparation of Genomic DNA in Agarose Blocks
The method used to prepare genomic DNA in situ in agarose blocks was adapted from the one described in previous works [33]. An overnight culture of strains was diluted (1:25) in fresh MRS broth and grown at 37 °C to an A600 of 0.6 (approximately 3 h). The cells were harvested by centrifugation at 1200×g for 5 min and washed once with 1 mL of 1 M NaCl-100 mM EDTA. The pellet was suspended in 100 μL of lysozyme (Promega) solution (10 mg mL−1), mixed with 100 μL of 2% low melting point agarose at 50 °C, and the mixture was allowed to solidify in appropriate molds at 4 °C. The embedded cells in agarose blocks were treated with 0.6 mL of NaCl-EDTA containing 5 mg mL−1 of lysozyme for 17 h at 37 °C, and 500 μL of 7.5% SDS solution was added. The incubation continued for 5 h. The blocks were washed twice with SL buffer (10 mM Tris-ClH, 1 M NaCl, pH = 7.6) and incubated overnight at 50 °C in 1 mL of BR buffer (500 mM sodium EDTA (pH = 9), 1% Sarkozyl, proteinase K 1 mg mL−1). Then, the blocks were washed twice with BL buffer (20 mM Tris-ClH, 50 mM sodium EDTA (pH = 8), treated for 1 h with BL buffer supplemented with 1 mM phenylmethylsulfonyl fluoride (PMSF) and finally washed four times, 30 min each in TE buffer (10 mM Tris-ClH [pH 8.0], 1 mM EDTA [pH 8.0]).
Digestion of DNA
Agarose blocks containing DNA were washed three times in TE buffer for 30 min and cut into slices (3 × 3 mm). Prior to use, they were equilibrated in multicore buffer. The DNAs were restricted with SmaI (10 U μL−1, Sigma-Aldrich) for 18 h at 25 °C. The buffer was discarded and the agarose blocks kept in TE buffer for 30 min, and then they were kept in TBE buffer 0.5× (0.045 M Tris-borate, 0.001 M EDTA, pH 8.3) for 15 min.
Pulsed-Field Gel Electrophoresis
Samples were electrophoresed through 1% (wt vol−1) agarose gels in 0.5× TBE buffer at 200 V for 22 h with a 2–20-s pulse time at 14 °C in a Bio-Rad CHEFF DRIII electrophoresis cell. Gels were visualized by staining them with ethidium bromide (5 μg mL−1) for 20 min, and DNA was detected under UV light.
Statistical Analysis
The statistical analysis of the data was performed using the SAS software (Institute, Cary, NC, USA). Differences between specific growth rates (μ) of the LAB were analyzed by an ANOVA test, and comparison between means was performed using Tukey’s post-hoc test. A p value < 0.05 was considered as being indicative of a statistically significant difference.
Results
Antimicrobial Activity
In the present work, the antagonistic activity of 12 LAB against 15 MCB was assayed. Ten LAB strains (83.3%) were able to inhibit all MCB assayed (Table 1). Ent. faecalis 19433 and Ent. faecium 35667 were not inhibited by W. cibaria CRL 1833, and Lact. plantarum CRL 1716 was not able to inhibit Staph. aureus ATCC25923, Staph. aureus RC108, Staph. epidermidis ATCC14990, Strep. agalactiae ATCC27956, Strep. uberis 102, Strep. uberis ATCC27958, Strep. bovis ATCC27960, Ent. faecalis 19433, and Ent. faecium 35667. Ent. hirae CRL 1835, Ent. hirae CRL 1837, L. lactis subsp. lactis CRL 1655, and Ent. mundtii CRL 1656 showed the highest inhibitory effect against Strep. dysgalactiae ATCC27957, inhibiting it completely. One of the most prevalent pathogens of mastitis, Staph. aureus, was inhibited by 11 of the 12 tested LAB. E. coli, which causes acute clinical mastitis, was inhibited for the entire LAB assayed.
Co-Aggregation
Ten of the twelve LAB strains (83%) showed co-aggregation with all of the MCB assayed (Table 2). Lact. plantarum CRL 1716 and W. cibaria CRL 1833 did not show co-aggregation with Pseudomonas spp. Moreover, Lact. plantarum CRL 1716 was not able to co-aggregate with E. coli and W. cibaria CRL 1833 with Strep. epidermidis ATCC14990. Staph. aureus and Streptococcus spp. co-aggregated with all LAB included into the study.
Adhesion Capacity
A high number of epithelial cells could be isolated from the bovine teat canal and no bacterial contaminants were observed after Gram staining as seen in previous reports [27]. All strains were able to adhere to BTCEC. The PA and the AI were different between strains (Table 3). Ent. hirae CRL 1835 and L. lactis subsp. lactis CRL 1655 showed the highest AI (36.7 and 27.9, respectively). Ped. pentosaceus CRL 1831 showed the highest PA (99%). Ped. pentosaceus CRL 1831 and Lact. perolens CRL 1724 adhered to BTCEC as can be seen in Fig. 1a. Figure 1b, c shows different numbers of bacteria adhered to the surface of epithelial cells, showing an irregular pattern of distribution on the cell surface. Figure 1d demonstrates an auto-aggregative pattern and an adherence of lactobacilli as clusters on the cell surface.
Light photomicrographs showing Gram-stained lactic acid bacteria strains adhered to bovine teat canal epithelial cells. Bacterial adhered to the surface of epithelial cells showing an irregular pattern of distribution on the cell surface. a Control (cells without bacteria). b Pediococcus pentosaceus CRL 1831. c Lactobacillus perolens CRL 1724. d Autoagreggative pattern and adherence of lactobacilli as clusters on the cell surface (× 1000)
Auto-inhibition
Three LAB strains showed a capacity for auto-inhibition. Enterococcus mundtii CRL 1656 was able to inhibit Ent. hirae 7-3 and W. cibaria CRL 1833. These strains were discarded to perform the probiotic formulation.
LAB Growth Curves
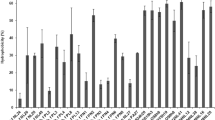
In general, the kinetic growth curves of the 12 selected LAB strains were similar (Fig. 2). They showed a latency phase of 2 h and an exponential phase that lasted until 8 h of growth. L. lactis subsp. lactis CRL 1655 and Ped. pentosaceus CRL 1831 showed the highest biomass yield (3 × 109 cfu mL−1) at 24 h of incubation, whereas W. cibaria CRL 1833 and W. cibaria CRL 1840 only reached a concentration around 6.5 × 108 cfu mL−1. No significant differences (p = 0.3476) were found between the specific growth rates of the selected LAB. Ent. hirae 7-3 and Lact. perolens CRL 1724 showed the highest specific growth rate, whereas Ped. pentosaceus CRL 1832, Ent. hirae CRL 1837, and Lact. plantarum CRL 1716 presented the lowest.
Selection of Strains
For the selection of LAB strains to be included in the intramammary formulation, the following criteria were applied: high IA and PA of adherence to BTCEC, high percentage of co-aggregation, and inhibition against MCP, inability to auto-inhibit and middle to high specific growth rate. Based on this, L. lactis subsp. lactis CRL 1655 and Lact. perolens CRL 1724 were selected. Moreover, L. lactis subsp. lactis CRL 1655 showed an interesting pattern of adhesion to the bovine teat canal epithelial cells and produced nisin Z. Lact. perolens CRL 1724 showed some surface properties and produced organic acid [25]. Even though Ent. hirae 7-3 presented good probiotic characteristics, especially elevated PA and AI and high co-aggregation, it was discarded because it encodes a potential vancomycin resistance gene undesirable for a biological product [26]. The selected LAB inhibits completely the growth of Staph. aureus ATCC25923 and Staph. aureus RC108, a bovine mastitis strain, after 24 h of co-incubation (Table 1). L. lactis subsp. lactis CRL 1655 and Lact. perolens CRL 1724 were able to decrease the pH of the culture from 6 to 3 after 24 h of incubation with Staph. aureus ATCC25923. Furthermore, the selected LAB showed an in vitro synergistic inhibitory effect against Staph. aureus ATCC25923 (data not shown).
Genomic Profile of Selected LAB Strains
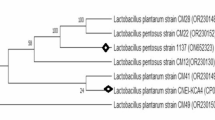
In addition to the 16s RNA genetic identification, a further PFGE restriction profile characterization of the selected LAB strains was performed. The PFGE profiles obtained from chromosomal DNA restricted with SmaI allowed to differentiate easily between L. lactis subsp. lactis CRL 1655 and Lact. perolens CRL 1724. The analysis of genetic profile showed the presence of 6 bands between 55 and 96 kb for Lact. perolens CRL 1724 and a pattern of 11 bands between 76 and 286 kb for L. lactis subsp. lactis CRL 1655 (Fig. 3).
Discussion
In Argentina, as in many other countries, Staph. aureus is still one of the major causes of clinical and subclinical mastitis, especially due to resistance to antibiotic treatment and its ability to persist in a herd in an undetected form. Even though current management practices such as proper milking hygiene and reduced exposure to environmental pathogens contribute to a decrease in the occurrence of the disease, the treatment for bovine mastitis relies heavily on the use of antibiotics, both for prophylaxis and therapy. This has proved to be ineffective in controlling Staph. aureus infections, causing a persistent bacterial reservoir within a herd with chronic and recurrent infections [1, 3,4,5]. The increasing importance of Staph. aureus in veterinary medicine, the problems related to its antimicrobial resistance and the need to have effective tools to fight bovine mastitis, strongly support the ongoing research on the development of safe preventive alternative approaches [1, 34]. This will permit having new and more powerful preventive and therapeutic tools to reduce its impact on animal health. The antimicrobial activity produced by the LAB strains against different pathogens seems to be an interesting research field and has two essential roles: to prevent the invasion of different bacterial strains considered as causal agents of diseases and to prevent new infections by MCB [35]. Several researchers have studied the ability of LAB to inhibit the pathogenic strains that cause bovine mastitis, attributing this ability to the production of inhibitory substances. Juarez Tomas et al. [36] established that vaginal Lactobacillus spp. has the capacity to inhibit urogenital pathogens (such as Staph. aureus, Strep. agalactiae, and Enterococcus spp.) throughout the production of lactic acid and through the reduction of medium pH. Maršálková et al. [37] and Hütt et al. [29] demonstrated the inhibitory effect of Lact. plantarum against mastitis (Staph. aureus, Strep. uberis, and Strep. agalactiae) and human (enteric and urinary) pathogens. Çon and Gökalp [38] attributed the inhibitory effect of LAB to different metabolites like lactic acid, acetic acid, diacetyl, hydrogen peroxide, and bacteriocins. Recently, Bouchard et al. [23] demonstrated the ability of LAB isolated from bovine milk to inhibit the growth of representative strains of E. coli, Strep. uberis, and Staph. aureus. They reported that bacterial inhibition lies on acidification, since neutralization of the supernatants totally relieves the inhibitory effect/activity. In this work, it was found that 83.3% of the isolated LAB were able to inhibit all the assayed MCB. Based on previous results, we can attribute this inhibitory effect to the ability of strains to produce hydrogen peroxide, lactic acid, acetic acid, organic acid, and/or bacteriocins [26]. L. lactis subsp. lactis CRL 1655 produces hydrogen peroxide, lactic acid (6 g L−1), acetic acid (0.5 g mL−1), and nisin Z. Lact. perolens CRL 1724 produces lactic acid (10 g L−1.), acetic acid (0.6 g mL−1) and organic acid. LAB exert their beneficial effects as a consequence of one or more mechanisms. Adhesion to epithelial cells, aggregation, and inhibition of pathogens were some of them. The degree of auto-aggregation and hydrophobicity predict the ability of a strain to adhere to epithelial cells, and it is a specific property of each microorganism [39,40,41]. The LAB strains described in this work were selected previously adopting recognized criteria [26] like no pathogens of mastitis, good health status of the quarters where the isolation was performed, high hydrophobicity and auto-aggregation phenotype, and absence of antibiotic resistance genes. Co-aggregation is an important method for assessing the close interaction between LAB and pathogenic bacteria, since LAB present surface binding proteins related to environmental surfaces and bacteria [42]. Some authors [43] propose that co-aggregation may be beneficial to these LAB because they produce an area around the pathogen where the concentration of antimicrobial substances increases. All LAB assayed were able to co-aggregate with all the MCB, except Lact. plantarum CRL 1716 and W. cibaria CRL 1833. Staph. aureus and Streptococcus spp., the major bovine mastitis pathogens, were co-agregated by all LAB strains. There are few publications where in vitro co-aggregation is determined. Soleimani et al. [42] reported co-aggregation between four strains of Lactobacillus (Lact. acidophilus, Lact. plantarum, Lact. casei and Lact. reuteri) and two strains of Staph. aureus (one isolated from a bovine mastitis case and other one belonging to a collection strain, Staph. aureus ATCC25923). Boris et al. [44] suggested that the ability of LAB to enclose pathogenic bacteria would be related to inhibiting the adherence of pathogens to tissue receptors. This effect would prevent the colonization of the epithelial cells by pathogenic bacteria, thus favoring their removal.
The adhesion of LAB to the epithelium is the first step in the formation of a barrier to prevent the invasion of photogenic bacteria and it is an essential characteristic when selecting probiotic strains [45, 46]. In this work, we obtained PA between 37 and 99% and AI between 6.9 and 36.7. Ent. hirae CRL 1835 and L. lactis subsp. lactis CRL 1655 showed the highest AI (36.7 and 27.9, respectively). Ped. pentosaceus CRL 1831 showed the highest PA (99%). Most of the published works of adherence are carried out in bovine mammary epithelial cells (MAC-T). Ocaña and Nader-Macías [47] and Otero and Nader-Macías [31] isolated bovine vaginal cells and confronted them with different Lactobacillus strains (Lact. crispatus CRL 1266, Lact. gasseri CRL 1412, Lact. gasseri CRL 1421, and Lact. delbrueckii). The results were similar to those obtained in this work: PA between 61 and 100% and AI between 10 and 22. In a recent study [48], where the technique described by Frola et al. [27] was used, four LAB strains (Lact. paracasei subsp. paracasei 78/37, Lact. plantarum 118/37, L. lactis subsp. lactis ATCC11454 and Lact. rhamnosus ATCC7469) were tested for their capacity of adhesion. The highest PA was shown by L. lactis subsp. lactis ATCC 11454 (100%) with an AI of 14.2. The lowest PA was shown by Lact. paracasei subsp. paracasei 78/37 with a PA of 56.1% and an AI of 7.8. Some studies suggested that Lactobacillus adherence is mediated by proteins associated to the external protein S-layer [49,50,51], while others have suggested a role for lipoteichoic acid and carbohydrates [52]. Further studies need to be conducted to determine the chemical nature of the structures involved in the adhesion of assayed LAB to BTCEC. Taking into account the results obtained in this work and in previous reports [25,26,27,28], we selected two LAB to be included in a future product for the prevention of bovine mastitis. L. lactis subsp. lactis CRL 1655 showed an interesting pattern of adhesion to the BTCEC (PA 93% and AI 27.9) and produced hydrogen peroxide, lactic acid, acetic acid, and nisin Z as inhibitory substance. Lact. perolens CRL 1724 has an adhesion capacity of 75% of PA and 14.4 of AI and produces lactic acid, acetic acid, and organic acid as inhibitory substances [25]. The two LAB strains inhibited and co-aggregated with 100% of the assayed pathogens, especially Staphylococcus spp. and did not show capacity for auto-inhibition. They completely inhibited the growth of Staph. aureus ATCC25923 and Staph. aureus RC108 after 24 h of co-incubation. Furthermore, the kinetic growth of these strains was similar among them and similar to the rest of the assayed LAB and showed a considerable and specific growth rate. These constitute an important feature for the industrial production of an intrammamary formulation. The selected strains show a predictable difference in the pattern of bands. SmaI is a restriction enzyme that recognizes and cuts in a specific sequence (5′ CCCGGG 3′) rich in G and C bases. It is expected that genomes with low G + C content, such as L. lactis (35% of G + C), have a few recognition sites, resulting in few large DNA fragments [53]. On the other hand, in Lact. perolens, the percentage of G + C is close to 50% and the cut sites are more frequent [54]. Because of this, a larger number of small size fragments are obtained, which explains the low molecular weight of the bands observed in the profile of Lact. perolens. The different genetic profiles of bands obtained by PFGE for the selected LAB also constitute a very important tool to ensure and to monitor the integrity of a veterinary product used to prevent bovine mastitis.
Conclusion
The development of non-antibiotic formulations for the prevention of bovine mastitis has the potential to reduce the dependence on antibiotics for prophylactic therapies in the future. Some studies indicate that the use of LAB is an interesting alternative for the prevention or treatment of bovine mastitis, especially in the dry-off period. The capacity of the two selected strains to inhibit and co-aggregate with the main bovine mastitis pathogens and to adhere to BTCEC, described in this paper, constitutes a fundamental characteristic for its inclusion in the development of a probiotic formulation for the prevention of bovine mastitis at dry-off period.
References
Bogni C, Odierno L, Raspanti C, Giraudo J, Larriestra A, Reinoso E et al (2011) War against mastitis: current concepts on controlling bovine mastitis pathogens. In: Méndez-Vilas A (ed) Science against microbial pathogens: communicating current research and technological advances. FORMATEX Research Center Inc., Spain, pp 483–494
Schukken H, Günthe J, Fitzpatrick J, Fontaine MC, Goetze L, Holst O et al (2011) Host-response patterns of intramammary infections in dairy cows. Vet Immunol Immunop 144(3):270–289. https://doi.org/10.1016/j.vetimm.2011.08.022
De Vliegher S, Fox L, Piepers S, McDougall S, Barkema HW (2012) Invited review: mastitis in dairy heifers: nature of the disease, potential impact, prevention, and control. J Dairy Sci 95(3):1025–1040. https://doi.org/10.3168/jds.2010-4074
Dieser S, Vissio C, Lasagno M, Bogni CI, Larriestra AJ, Odierno LM (2014) Prevalence of pathogens causing subclinical mastitis in Argentinean dairy herds. Pak Vet J 34:124–126
Petrovski K, Grinberg A, Williamson N, Abdalla ME, Lopez-Villalobos N, Parkinson TJ et al (2015) Susceptibility to antimicrobials of mastitis-causing Staphylococcus aureus, Streptococcus uberis and Str. dysgalactiae from New Zealand and the USA as assessed by the disk diffusion test. Aust Vet J 93(7):227–233. https://doi.org/10.1111/avj.12340
Raspanti C, Bonetto C, Vissio C, Pellegrino MS, Reinoso EB, Dieser SA et al (2016) Prevalence and antibiotic susceptibility of coagulase-negative Staphylococcus species from bovine subclinical mastitis in dairy herds in the central region of Argentina. Rev Arg Microbiol 48(1):50–56
Thompson-Crispi K, Atalla H, Miglior F, Mallard B (2014) Bovine mastitis: frontiers in immunogenetics. Front Immunol 5:493
Dua K (2001) Incidence, etiology and estimated loss due to mastitis in India—an update. Indian Dairyman 53:41–48
Halasa T, Nielen M, De Roos A, Van Hoorne R, De Jong G, Lam TJGM et al (2009) Production loss due to new subclinical mastitis in Dutch dairy cows estimated with a test-day model. J Dairy Sci 92(2):599–606. https://doi.org/10.3168/jds.2008-1564
Vissio C, Dieser S, Agnelli H, Odierno LM, Larriestra A (2014) Accuracy of the composite somatic cell count to detect intra-mammary infection in dairy cows using latent class analysis. Prev Vet Med 113(4):547–555. https://doi.org/10.1016/j.prevetmed.2013.11.016
Bradley A (2002) Bovine mastitis: an evolving disease. Vet J 164(2):116–128. https://doi.org/10.1053/tvjl.2002.0724
Dufour S, Dohoo I (2012) Monitoring dry period intramammary infection incidence and elimination rates using somatic cell count measurements. J Dairy Sci 95(12):7173–7185. https://doi.org/10.3168/jds.2012-5839
McDougall S, Bryan M, Tiddy R (2009) Effect of treatment with the nonsteroidal antiinflammatory meloxicam on milk production, somatic cell count, probability of re-treatment, and culling of dairy cows with mild clinical mastitis. J Dairy Sci 92(9):4421–4431. https://doi.org/10.3168/jds.2009-2284
Song E, Yu M, Wang Y, Hu W, Cheng D, Swihart MT, Song Y (2015) Multi-color quantum dot-based fluorescence immunoassay array for simultaneous visual detection of multiple antibiotic residues in milk. Biosens Bioelectron 72:320–325. https://doi.org/10.1016/j.bios.2015.05.018
Wang D, Wang Z, Yan Z, Wu J, Ali T, Li J, Lv Y, Han B (2015) Bovine mastitis Staphylococcus aureus: antibiotic susceptibility profile, resistance genes and molecular typing of methicillin-resistant and methicillin-sensitive strains in China. Infect Genet Evol 31:9–16. https://doi.org/10.1016/j.meegid.2014.12.039
FAO, WHO (2015) Guidelines for the evaluation of probiotics in food. Food and Agriculture Organization of the United Nations and World Health Organization Working Group, Geneva
Nader-Macías MEF, Bogni C, Sesma F, Espeche MC, Pellegrino M, Saavedra L et al (2011) Alternative approaches for the prevention of bovine mastitis. In: Bitterlich A, Fisch S (eds) Probiotics, bioactive compounds and vaccines: bioactives compounds. Nova Science Publishers Inc., New York, pp 1–34
Walsh M, Gardiner G, Hart O, Lawlor PG, Daly M, Lynch B et al (2008) Predominance of a bacteriocin-producing Lactobacillus salivarius component of a five-strain probiotic in the porcine ileum and effects on host immune phenotype. FEMS Microbiol Ecol 64(2):317–327. https://doi.org/10.1111/j.1574-6941.2008.00454.x
Ryan M, Flynn J, Hill C, Ross RP, Meaney WJ (1999a) The natural food grade inhibitor lacticin 3147 can prevent mastitis in non-lactating dairy cows. J. Dairy Sci 82(12):2625–2631. https://doi.org/10.3168/jds.S0022-0302(99)75519-0
Twomey D, Wheelock A, Flynn J, Meaney WJ, Hill C, Ross R (2000) Protection against Staphylococcus aureus mastitis in dairy cows using a bismuth-based teat seal containing the bacteriocin, lacticin 3147. J Dairy Sci 83(9):1981–1988. https://doi.org/10.3168/jds.S0022-0302(00)75075-2
Crispie F, Alonso-Gómez M, O’Loughlin C, Klostermann K, Flynn J, Arkins S et al (2008) Intramammary infusion of a live culture for treatment of bovine mastitis: effect of live lactococci on the mammary immune response. J Dairy Res 75(3):374–384. https://doi.org/10.1017/S0022029908003385
Klostermann K, Crispie F, Flynn J, Ross RP, Hill C, Meaney W (2008) Intramammary infusion of a live culture of Lactococcus lactis for treatment of bovine mastitis: comparison with antibiotic treatment in field trials. J Dairy Res 75(3):365–373. https://doi.org/10.1017/S0022029908003373
Bouchard D, Seridan B, Saraoui T, Rault L, Germon P, Gonzalez-Moreno C et al (2015) Lactic acid bacteria isolated from bovine mammary microbiota: potential allies against bovine mastitis. PLoS One 10(12):e0144831. https://doi.org/10.1371/journal.pone.0144831
Giannino M, Aliprandi M, Feligini M, Vanoni L, Brasca M, Fracchetti F (2009) A DNA array based assay for the characterization of microbial community in raw milk. J Microbiol meth 78(2):181–188. https://doi.org/10.1016/j.mimet.2009.05.015
Espeche M, Otero M, Sesma F, Nader-Macias MEF (2009) Screening of surface properties and antagonistic substances production by lactic acid bacteria isolated from the mammary gland of healthy and mastitic cows. Vet Microbiol 135(3):346–357. https://doi.org/10.1016/j.vetmic.2008.09.078
Espeche MC, Pellegrino M, Frola I, Larriestra A, Bogni C, Nader-Macías MEF (2012) Lactic acid bacteria from raw milk as potentially beneficial strains to prevent bovine mastitis. Anaerobe 18(1):103–109. https://doi.org/10.1016/j.anaerobe.2012.01.002
Frola I, Pellegrino M, Espeche MC, Giraudo JA, Nader-Macias MEF, Bogni C (2011) Effects of intramammary inoculation of lactobacillus perolens CRL 1724 in lactating cows’ udders. J Dairy Res 79(1):84–92. https://doi.org/10.1017/S0022029911000835
Frola I, Pellegrino M, Espeche M, Giraudo JA, Nader-Macias MEF, Bogni CI (2012) Effects of intramammary inoculation of Lactobacillus perolens CRL 1724 in lactating cows’ udders. J Dairy Res 79(01):84–92. https://doi.org/10.1017/S0022029911000835
Hütt P, Shchepetova J, Lõivukene K, Kullisaar T, Mikelsaar M (2006) Antagonistic activity of probiotic lactobacilli and bifidobacteria against entero- and uropathogens. J Appl Microb 100(6):1324–1332. https://doi.org/10.1111/j.1365-2672.2006.02857.x
Reid G, Mac Groarty J, Chow A, Chow AW, Bruce AW, Eisen A et al (1990) Coaggregation of urogenital bacteria in vitro and in vivo. Curr Microbiol 20(1):47–52. https://doi.org/10.1007/BF02094024
Otero M, Nader-Macías MEF (2007) Lactobacillus adhesion to epithelial cells from bovine vagina. In: Méndez-Vilas A (ed) Communicating current research and educational topics and trends in applied microbiology, FORMATEX Research Center Inc., Spain, pp 749–757
Malthus TR (1826) An essay on the principle of population. 6th. John Murray, London
Tanskanen E, Tulloch D, Hillier A, Davidson B (1990) Pulsed-field gel electrophoresis of SmaI digests of lactococcal genomic DNA, a novel method of strain identification. Appl Environ Microb 56(10):3105–3111
Pellegrino M, Rodriguez N, Vivas A, Giraudo J, Bogni C (2016) Staphylococcus aureus avirulent mutant vaccine induces humoral and cellular immune responses on pregnant heifers. Vaccine 34(29):3356–3362. https://doi.org/10.1016/j.vaccine.2016.05.014
Perea-Vélez M, Hermans K, Verhoeven T, Lebeer SE, Vanderleyden J, De Keersmaecker SCJ (2007) Identification and characterization of starter lactic acid bacteria and probiotics from Columbian dairy products. J appl microb 103(3):666–674. https://doi.org/10.1111/j.1365-2672.2007.03294.x
Juárez Tomás M, Ocaña V, Wies B, Nader-Macías MEF (2003) Growth and lactic production by vaginal Lactobacillus acidophilus CRL 1259, and inhibition of uropathogenic Escherichia coli. J Med Microbiol 52(12):1117–1124. https://doi.org/10.1099/jmm.0.05155-0
Maršálková S, Cizek M, Vasil M, Bomba A, Nad P, Datelinka I et al (2003) Testing two Lactobacillus plantarum and Lactobacillus acidophilus strains for their suitability as a lipoid probiotic. Berl Munch Tierarztl 117(3–4):145–147
Çon A, Gökalp H (2000) Production of bacteriocin-like metabolites by lactic acid cultures isolated from sucuk samples. Meat Sci 55(1):89–96. https://doi.org/10.1016/S0309-1740(99)00129-1
Saarela M, Mogensen G, Fondén R, Mättö J, Mattila-Sandholm T (2000) Probiotic bacteria: safety, functional and technological properties. J Biotechnol 84(3):197–215. https://doi.org/10.1016/S0168-1656(00)00375-8
Ouwehand A, Salminen S, Isolauri E (2002) Probiotics: an overview of beneficial effects. A Van Leeuw J Microb 82(1/4):279–289. https://doi.org/10.1023/A:1020620607611
Gueimonde M, Salminen S (2006) New methods for selecting and evaluating probiotics. Dig Liver Dis 38:S242–S247. https://doi.org/10.1016/S1590-8658(07)60003-6
Soleimani N, Kermanshahi R, Yakhchali B, Sattari T (2010) Antagonistic activity of probiotic lactobacilli against Staphylococcus aureus isolated from bovine mastitis. Afric J Microbiol Res 4(20):2169–2173
Reid G, McGroarty J, Angotti R, Cook RL (1988) Lactobacillus inhibitor production against Escherichia coli and co-aggregation ability with uropathogens. Can J Microbol 34(3):344–351. https://doi.org/10.1139/m88-063
Boris S, Suarez J, Velazquez F, Barbés C (1998) Adherence of human lactobacilli to vaginal epithelial cells and interaction with uropathogens. Infect Immun 66(5):1985–1989
Havenaar R, Brink BT, Huis In’t Veld J (1992) Selection of strains for probiotics use. In: Fuller R (ed) Probiotics. The scientific basis. Chapman and Hall Inc., London, pp 209–223
Reid G, Jass J, Sebulsky M, McCormick J (2003) Potential use of probiotics in clinical practice. Clin Microbial Rev 16:652–658
Ocaña V, Nader-Macías MEF (2001) Adhesión of Lactobacillus vaginal strains with probiotic properties to vaginal epithelial cells. Biocell 25(3):265–273
Diepers A, Krömker V, Zinke C, Wente N, Pan L, Paulsen K et al (2017) In vitro ability of lactic acid bacteria to inhibit mastitis-causing pathogens. Sust Chem Pharm 5:84–92
Wadstroum T, Andersson K, Sydow M, Axelsson L, Lindgren S, Gullmar B (1987) Surface properties of lactobacilli isolated from the small intestine of pigs. J Appl Bacteriol 62(6):513–520. https://doi.org/10.1111/j.1365-2672.1987.tb02683.x
Henriksson A, Szewzyk R, Conway P (1991) Characteristics of the adhesive determinants of Lactobacillus fermentum 104. Appl Environ Microb 57(2):499–502
Frece J, Kos B, Svetec I, Zgaga Z, Mrša V, Šušković J (2005) Importance of S-layer proteins in probiotic activity of Lactobacillus acidophilus M92. J Appl Microb 98(2):285–292. https://doi.org/10.1111/j.1365-2672.2004.02473.x
Fuller R (1975) Nature of the determinant responsible for the adhesion of lactobacilli to chicken crop epithelial cells. Microbiology 87(2):245–250
Siezen R, Bayjanov J, Felis G, van der Sijde M, Starrenburg M, Molenaar D et al (2011) Genome-scale diversity and niche adaptation analysis of Lactococcus lactis by comparative genome hybridization using multi-strain arrays. Microb Biotechnol 4(3):383–402. https://doi.org/10.1111/j.1751-7915.2011.00247.x
Sun Z, Harris H, McCann A, Guo C, Argimo’n S, Zhang W et al (2015) Expanding the biotechnology potential of lactobacilli through comparative genomics of 213 strains and associated genera. Nat Commun 6:8322. https://doi.org/10.1038/ncomms9322
Acknowledgements
These are the results obtained from the project called “Design of a probiotic product for bovine mastitis prevention” signed between Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET) and Universidad Nacional de Río Cuarto. M. Pellegrino and M.E.F. Nader-Macías are career investigators of the CONICET, and N. Berardo is recipient of a fellowship from CONICET.
Funding
This work was supported by grants of Secretaría de Ciencia y Técnica, Universidad Nacional de Río Cuarto (SECYT-UNRC) and Agencia Nacional de Promoción Científica y Tecnológica (ANPCYT- PICT 543).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This article does not contain any studies with human or animal subjects.
Conflict of Interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Pellegrino, M.S., Frola, I.D., Natanael, B. et al. In Vitro Characterization of Lactic Acid Bacteria Isolated from Bovine Milk as Potential Probiotic Strains to Prevent Bovine Mastitis. Probiotics & Antimicro. Prot. 11, 74–84 (2019). https://doi.org/10.1007/s12602-017-9383-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12602-017-9383-6