Abstract
The present study aimed to isolate an optimal lactic acid bacterial strain from the feces of healthy giant pandas. The strain exhibited good stability at low pH and high bile salt concentrations, activity against pathogens relevant to pandas, and antibiotic susceptibility. In the current study, 25 isolates were obtained from de Man, Rogosa, and Sharpe agar. Two (E21 and G83) and eight (E1, E2, E16, E18, E21, E69, E70, and G83) isolates demonstrated good performance at pH 2.0 and bile 2% (w/v), respectively. Three isolates (G83, G88, and G90) possessed better antimicrobial effect on enterotoxigenic Escherichia coli CVCC196 (ETEC) than the rest. One isolate (G83) strongly affected Salmonella, whereas three (G83, G87, and G88) exhibited inhibitory activity against Staphylococcus aureus. All isolates were multi-drug resistant. These isolates were identified as Lactobacillus (5 isolates) and Enterococcus (20 isolates) by 16S rRNA sequencing. Virulence genes were detected in Enterococcus isolates. Isolate G83 was identified as Lactobacillus plantarum and was considered as the best probiotic candidate among all of the experimental isolates. This study provided necessary and important theoretical guidance for further experiments on G83 in vivo.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The giant panda, Ailuropoda melanoleuca, is famous as a living fossil. It is a vulnerable, endemic species that is extremely popular worldwide. Conservation strategies have resulted in the survival of about 1800 giant pandas worldwide. The giant panda is a herbivore that has retained a typical carnivorous digestive system. It is easily afflicted by various intestinal diseases when the feed structure changes. Under conditions of captivity, the prevention and treatment of diseases rely on antibiotics. However, scholars sought alternative treatments because of the disadvantages of antibiotics. One of the best choices is probiotics, which maintain or restore normal gut microbiota, inhibit pathogen invasion, and prevent inflammation [1,2,3].
Studies on the effects of intestinal probiotics on giant pandas focused mostly on cellulolytic bacterium and bacillus [4,5,6]. The present work is the first to investigate lactic acid bacteria (LAB) isolated from the giant panda feces, thereby providing crucial data to guide further study. We analyzed the intestinal microflora structure of captive giant pandas of different ages and focused on the survival in extreme acid and bile condition, antagonistic activity, and antibiotic susceptibility of the strains [7]. The pathogenic strains used in the antagonistic test were hyperendemic enteric pathogens of giant pandas [8, 9]. The present study is the first step in the search for probiotics that prevent and treat gut diseases in giant pandas. The next steps must involve in vivo safety test and the extensive assessment of LAB. We aim to develop new probiotics for giant pandas in the future.
Materials and Methods
Strains and Feces Samples
The feces samples were collected from seven healthy giant pandas (Chengdu Research Base of Giant Panda Breeding); ETEC (O8:H19:F4ac+, LT+, STa−, STb+), Escherichia coli ATCC25922, and Lactobacillus rhamnosus GG ATCC53103 (LGG) were purchased from Chinese Veterinary Drug Control; Salmonella and Staphylococcus aureus were received from Laboratory of Animal Infectious Disease and Microarray (Sichuan Agricultural University).
Isolation of LAB
Feces samples (10 g) were homogenized in sterile saline (90 mL), serially diluted and plated onto MRS agar and incubated at 37 °C for 24~48 h in constant anaerobic environment. Colonies which showed different morphologies were selected and purified by restreaking three times or more on MRS agar. The pure isolates which exhibited Gram-positive were selected and subcultured in MRS broth for further study. Biochemical reaction method (Hangzhou Microbial Reagent co., Ltd.) was the first step of screening and referenced with Bergey’s Manual Of Systematic Bacteriology and Isolation And Identification And Test Methods Of Lactic Acid Bacteria.
Acid and Bile Tolerance Test
The isolates were subcultured in MRS broth for 24 h. The equal volume of suspension was added to MRS broth which was adjusted to pH 1.0, 2.0, and 3.0 with 1 M HCl and contained bile 0.3, 1.0, and 2.0% (w/v), respectively. Broth was incubated at 37 °C and then viable count was conducted at 0 and 3 h for acid tolerance test and 0 and 4 h for bile test.
Antimicrobial Activity Test
Antimicrobial activity of isolates, for ETEC, Staphylococcus aureus, and Salmonella, was assessed using the Oxford cup method [10]. The isolates were inoculated to MRS broth at 37 °C for 24 h. The cell-free supernatant was collected (15,600×g, 10 min). Freshly grown pathogen cultures (100 μL, 107 CFU/mL) were spread on LB agar plate and allowed to dry. Oxford cups were placed on plates. A 100 μL cell-free supernatant was poured into a cup on plates. The zone of inhibition was measured and recorded after inoculating at 37 °C for 24 h.
Antibiotic Susceptibility Test
Antibiotic susceptibility of the isolates was assessed using disk diffusion method [11]. Isolates and paper disk (Beijing Tiantan Biological Products co., Ltd.) were placed onto the surface of MRS agar plates. The zone of inhibition was measured and recorded after inoculating at 37 °C for 24 h. Results were compared with interpretative zone diameters described by Performance Standards for Antimicrobial Disk Susceptibility Tests [12]. The antibiotics tested were kanamycin (30 μg), gentamicin (10 μg), amikacin (30 μg), streptomycin (10 μg), sulfamethoxazole (25 μg), tetracycline (30 μg), doxycycline (30 μg), florfenicol (30 μg), chloramphenicol (30 μg), cefotaxime (30 μg), cephradine (30 μg), ceftriaxone (30 μg), cefoperazone (75 μg), and ciprofloxacin (5 μg).
Molecular Identification
The isolates were inoculated at 10 ml MRS broth at 37 °C overnight and the culture was centrifuged (4000 rev/min) to harvest the cells and wash 2–3 times by sterile saline. The genomic DNA was extracted by using E.Z.N.A.® Stool DNA kit (Omega Biotechnology, USA). The primers are 27F and 1492R. The amplified DNA fragment was separated on a 2% agarose gel. The fragment was used directly for DNA sequencing (Beijing BGI Sequencing). The resulting sequences were compared with the sequences in the GenBank database using the BLAST program available on the National Center for Biotechnology Information (NCBI) website. The criterion used to identify an isolate to the species level was identity greater than 99% in the 16S rRNA gene sequence.
Statistical Analysis
All data were expressed as means and standard deviations and analyzed using SPSS version 19.0. The difference was evaluated by one-way ANOVA and statistical significance was set at P < 0.05.
Result
Morphological and Phenotypic Characteristics
A total of 207 isolates were obtained and initially screened. The isolates were observed for their morphological and phenotypic characteristics. Only 25 isolates were Gram-positive. G83 exhibited a rod-shaped morphology, whereas that of G87, G88, G89, and G90 was bent. The morphology of the remaining isolates was ellipsoidal (Table 1). Among the isolates, 25 were oyster white and facultative anaerobes, arranged singly, in pairs, or in short chains. Catalase, xylose, motility, nitrate reduction, and H2S production tests showed negative results.
Acid and Bile Resistance
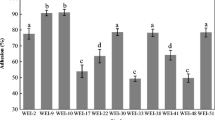
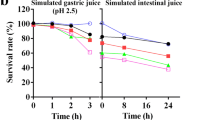
The majority of the isolates showed high resistance to acid after inoculation at pH 3.0 for 3 h. Isolates E21 and G83 revealed good survival ability after inoculation at pH 2.0 for 3 h. No isolates could survive at pH 1.0 (Table 2). Viable cell counts of isolates after 4 h culture at different bile concentrations are shown in Table 3. At 0.3% bile, all isolates except E3, E9, E23, E33, and E57 showed high resistance, with concentrations higher than 107 CFU/mL. Under 2% bile, the viable cell count of E1, E2, E16, E18, E21, E69, E70, and G83 were higher than that of 106 CFU/mL, whereas E3, E9, E23, E33, and E57 did not survive. Out of 25, only 20 isolates exhibited a good ability to resist low pH and high bile salts and were selected for further analysis.
Antimicrobial Activity
The isolates exhibited significant antimicrobial effects on enterotoxigenic Escherichia coli CVCC196 (ETEC), Staphylococcus aureus, and Salmonella (Table 4). Isolates G83, G88, and G90 demonstrated better antimicrobial effect on ETEC (> 20 mm) than the other isolates. Isolate G83 showed a stronger effect on Salmonella (> 20 mm) than the others, and isolates G83, G87, and G88 possessed inhibitory activity against S. aureus (> 17 mm). The inhibition zones of G83 were longer with respect to LGG on each pathogen. G83 showed excellent antimicrobial ability. Antimicrobial activity was not detected after excluding the interference of acid materials by adjusting the supernatant to pH 6.5 using NaOH.
Antibiotic Susceptibility
Selected isolates exhibited multi-drug resistance. However, the antibiotic resistance ratio of isolates was not high, except that of E1 (Table 5). All isolates were sensitive to florfenicol, chloramphenicol, and doxycycline. Isolate G83 was only resistant to amikacin, streptomycin, cotrimoxazole, and ciprofloxacin.
Phylogenetic Analysis
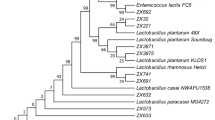
All isolates were further identified by 16S rRNA sequencing and phylogenetic analysis. The five isolates were species of Lactobacillus, whereas the rest of the isolates were species of Enterococcus. G83 showed the highest sequence similarity (99%) to L. plantarum based on BLASTn. Its sequence was uploaded to NCBI (GenBank accession number KU291268). A phylogenetic tree was built using DNAMAN5.2 and MEGA6.0 (Fig. 1).
Detection of Virulence Genes
Enterococcus is one of the dominant bacteria in the gut of giant pandas [7]. Enterococcus can be used as probiotics without virulence genes [13, 14]. We performed virulence gene detection on the isolates for security. Some common virulence genes were detected via PCR (Table 6). A total of 15 isolates of Enterococcus exhibited different virulence genes, whereas 5 isolates of Lactobacillus did not. Eight kinds of virulence genes were detected, namely sex pheromone (ccf), gelatinase gene from Enterococcus (gelE), accessory colonization factor (ace), cytolysin A (cylA), aggregation substance gene (agg), enterococcal surface protein gene (esp), endocarditis antigen in E. faecalis (efaAfs), and E. faecium (efaAfm), with ratios fluctuating from 37.5 to 62.5%. The detection rates of ccf and efaAfs reached 100%, whereas those of gelE, ace, and efaAfm were 66.7, 86.7, and 33.3%, respectively. Overall, G83 was selected for further analysis.
Discussion
Probiotics, such as Lactobacillus, Bifidobacterium, and Bacillus, can prevent gut bacterial disease, maintain or restore the normal microbiota, and maintain intestinal integrity [19,20,21,22]. The current work aimed to screen potential probiotics from giant panda feces that exhibit outstanding abilities. We aimed to use the probiotics as an alternative for antibiotics, as demonstrated in giant panda. Mimicking gastrointestinal tract conditions, the acid and bile tolerance of isolates were initially screened [23]. Acid and bile tolerance are important selection criteria for probiotic strains. Gastric acidity and bile conditions are relevant for digestion and metabolic activity [24]. The isolates must withstand acidic fluid with a pH of 1.5 to 3.0 and bile concentrations that fluctuate between 0.5 and 2.0% [25]. Isolate G83 grew well at pH 2.0 and under 2.0% bile conditions, thereby showing its ability to survive the harsh conditions in the stomach and small intestines. The LAB isolated from canine feces could live in broth with a pH level of 2.0 for 4 h [26]. In addition, nitrate and H2S reactions were negative, thereby indicating that isolates can enhance the safety of the host. Carbohydrates were also produced, and these provide energy for host and gut microflora.
Captive giant pandas are easily afflicted by gut bacterial diseases. These gut diseases are always caused by E. coli, Salmonella, S. aureus, Shigella, Klebsiella, and Proteus [8, 9]. In the current study, selected isolates were assessed for antimicrobial activity against hyperendemic enteric pathogens of giant pandas. Isolate G83 showed maximum inhibition zones against ETEC, Salmonella, and S. aureus. The antagonistic activity of the acknowledged probiotic LGG was inferior to that of G83. Other studies reported the antimicrobial activity of probiotic strains against some common pathogens, such as E. coli, Salmonella, Listeria, and S. aureus [23, 27, 28].
Susceptibility to antibiotics is species- and strain-specific [29]. All isolates (except the indicator strain ATCC25922) showed different levels of antibiotic resistance. Consistent with previous studies, isolate G83 was resistant to amikacin, streptomycin, cotrimoxazole, and ciprofloxacin. Strains of L. reuteri and L. rhamnosus were resistant to tetracycline [30, 31]. Corynebacterium vitaeruminis MRU4 isolated from cow rumen was resistant to oxacillin, gentamicin, erythromycin, clindamycin, sulfa/trimethoprim, and rifampicin [32]. Antibiotic use in animals can result in the presence of antibiotic residues in their meat and milk. Antibiotic-resistant bacteria and resistance genes can transfer between animals and people [33]. However, this situation is not applicable to giant pandas. Antibiotic resistance was not detected in the strains isolated from giant panda feces.
Enterococcus is a natural flora present in traditional food. It is used for food ripening and flavor improvement [34]. Some Enterococcus strains have been successfully developed as probiotics to improve the health of human and animals [34, 35]. However, most Enterococcus strains carry various virulence factors and can cause many diseases, including urethral infections, bacteremia, endocarditis, peritonitis, and wound infections [36, 37]. We detected virulence factors in our 20 isolates to determine their safety. Five kinds of virulence genes were detected from 20 Enterococcus isolates. The use of the strains relies on various indexes that indicate their safety as probiotics in the Korean market [38]. These indexes not only include virulence genes but also enterotoxin genes, which are carried by some Enterococcus strains. Several factors have been implicated as potential virulence determinants that cause serious human diseases. Virulent strains harm the host through their adherence to host tissue, invasion and abscess formation, host inflammatory response modulation, and toxin secretion [13]. For security purposes, we selected G83 as a potential probiotic.
As previously discussed, some strains in this study showed good acid and bile resistance, activity against pathogens, and sensitivity to most antibiotics, with Lactobacillus G83 exhibiting the best activity. In future studies, G83 may show some interesting probiotic traits. Further sequential trials of its effects, as well as animal trials to test its in vivo effects, are required. The safety of these isolates must also be evaluated in vivo.
References
Geier MS, Mikkelsen LL, Torok VA, Allison GE, Olnood CG, Boulianne M, Hughes RJ, Choct M (2010) Comparison of alternatives to in-feed antimicrobials for the prevention of clinical necrotic enteritis. J Appl Microbiol 109(4):1329–1338. https://doi.org/10.1111/j.1365-2672.2010.04758.x
Archambaud C, Nahori MA, Soubigou G, Bécavin C, Laval L, Lechat P, Smokvina T, Langella P, Lecuit M, Cossart P (2012) Impact of lactobacilli on orally acquired listeriosis. Proc Natl Acad Sci 109(41):16684–16689. https://doi.org/10.1073/pnas.1212809109
Yu Q, Yuan L, Deng J, Qian Y (2015) Lactobacillus protects the integrity of intestinal epithelial barrier damaged by pathogenic bacteria. Front Cell Infect Microbiol 5:26. https://doi.org/10.3389/fcimb.2015.00026
Zhou Z, Zhou X, Zhong Z, Wang C, Zhang H, Li D, He T, Li C, Liu X, Yuan H (2014) Investigation of antibacterial activity of Bacillus spp. isolated from the feces of Giant Panda and characterization of their antimicrobial gene distributions. World J Microbiol Biotechnol 30(12):3129–3136. https://doi.org/10.1007/s11274-014-1740-y
Wu C, Wu L, Zhang L, Gelbič I, Xu L, Guan X (2014) Characterization of eight Bacillus thuringiensis isolates originated from fecal samples of Fuzhou Zoo and Fuzhou Panda Center. J Asia Pac Entomol 17(3):395–397. https://doi.org/10.1016/j.aspen.2014.02.009
Chen F, Shuang L, Cheng L, Shuang M, Li Z, Qi W (2012) Isolation, identification and cellulase production of a cellulolytic bacterium from intestines of giant panda (in Chinese). Acta Microbiol Sin 52(9):1113–1121 (樊程, 李双江, 李成磊, 马双, 邹立扣, 吴琦 (2012) 大熊猫肠道纤维素分解菌的分离鉴定及产酶性质. 微生物学报 52 (9):1113-1121)
Peng Z, Dong Z, Qiang W, Niu L, Ni X, Zou F, Yang M, Hao S, Yi Z, Qian L (2016) Decreased microbial diversity and Lactobacillus group in the intestine of geriatric giant pandas (Ailuropoda melanoleuca). World J Microbiol Biotechnol 32(5):79. https://doi.org/10.1007/s11274-016-2034-3
Chen X, Yin M, Wang X, Pu Z, Ma Y, Chen Z (2015). Isolation and preliminary identification of intestinal pathogens of captive giant panda (In Chinese). J Mianyang Normal Univ (2):1–7. (陈希文, 尹苗, 王雄清, 蒲中慧, 马缨, 陈紫娟 (2015) 圈养大熊猫肠道致病菌的分离与初步鉴定. 绵阳师范学院学报 (2):1–7). https://doi.org/10.3969/j.issn.1672-612x.2015.02.001
Fei S, Jing L, Dan X, Wan W, Geng G, Ning F, Shui Y (2002) Pathogens of intestinal disease in giant panda (in Chinese). J Economic Animal 6(2):20–23 (孙飞龙, 刘敬贤, 席丹, 王万云, 高更更, 冯宁, 杨水云 (2002) 大熊猫肠道疾病致病菌. 经济动物学报 6 (2):20-23). https://doi.org/10.3969/j.issn.1007-7448.2002.02.006
Fontana C, Cocconcelli PS, Vignolo G, Saavedra L (2015) Occurrence of antilisterial structural bacteriocins genes in meat borne lactic acid bacteria. Food Control 47:53–59. https://doi.org/10.1016/j.foodcont.2014.06.021
Ghosh K, Ray M, Adak A, Halder SK, Das A, Jana A, Parua S, Vágvölgyi C, Mohapatra PKD, Pati BR (2015) Role of probiotic Lactobacillus fermentum KKL1 in the preparation of a rice based fermented beverage. Bioresour Technol 188:161–168. https://doi.org/10.1016/j.biortech.2015.01.130
Cockerill FR (2013) Performance standards for antimicrobial susceptibility testing : twenty-first informational supplement. Clinical and Laboratory Standards Institute
Eaton TJ, Gasson MJ (2001) Molecular screening of Enterococcus virulence determinants and potential for genetic exchange between food and medical isolates. Appl Environ Microbiol 67(4):1628–1635. https://doi.org/10.1128/AEM.67.4.1628-1635.2001
Bellomo G, Mangiagle A, Nicastro L, Frigerio G (1980) A controlled double-blind study of SF68 strain as a new biological preparation for the treatment of diarrhoea in pediatrics. Curr Ther Res Clin 28(6):927–936
Shankar V, Baghdayan AS, Huycke MM, Lindahl G, Gilmore MS (1999) Infection-derived Enterococcus faecalis strains are enriched in esp, a gene encoding a novel surface protein. Infect Immun 67(1):193–200
Su YA, Sulavik MC, He P, Makinen KK, Makinen PL, Fiedler S, Wirth R, Clewell DB (1991) Nucleotide sequence of the gelatinase gene (gelE) from Enterococcus faecalis subsp. liquefaciens. Infect Immun 59(1):415–420
Gilmore MS, Segarra RA, Booth MC, Bogie CP, Hall LR, Clewell DB (1994) Genetic structure of the Enterococcus faecalis plasmid pAD1-encoded cytolytic toxin system and its relationship to lantibiotic determinants. J Bacteriol 176(23):7335–7344. https://doi.org/10.1128/jb.176.23.7335-7344.1994
Mannu L, Paba A, Daga E, Comunian R, Zanetti S, Duprè I, Sechi LA (2003) Comparison of the incidence of virulence determinants and antibiotic resistance between Enterococcus faecium strains of dairy, animal and clinical origin. Int J Food Microbiol 88(2):291–304. https://doi.org/10.1016/S0168-1605(03)00191-0
Barbavidal E, Castillejos L, Lópezcolom P, Rivero UM, Moreno Muñoz JA, Martínorúe SM (2017) Evaluation of the probiotic strain Bifidobacterium longum subsp. Infantis CECT 7210 capacities to improve health status and fight digestive pathogens in a piglet model. Front Microbiol 8:533. https://doi.org/10.3389/fmicb.2017.00533
Nishida S, Ishii M, Nishiyama Y, Abe S, Ono Y, Sekimizu K (2017) Lactobacillus paraplantarum 11-1 isolated from rice bran pickles activated innate immunity and improved survival in a silkworm bacterial infection model. Front Microbiol 8:436. https://doi.org/10.3389/fmicb.2017.00436
Wu Y, Wang Y, Zhou H, Wang B, Sun Q, Fu A, Wang Y, Wang Y, Xu X, Li W (2017) Probiotic Bacillus amyloliquefaciens SC06 induces autophagy to protect against pathogens in macrophages. Front Microbiol 8:469. https://doi.org/10.3389/fmicb.2017.00469
Gareau MG, Sherman PM, Walker WA (2010) Probiotics and the gut microbiota in intestinal health and disease. Nat Rev Gastroenterol Hepatol 7(9):503–514. https://doi.org/10.1038/nrgastro.2010.117
Yadav R, Puniya AK, Shukla P (2016) Probiotic properties of Lactobacillus plantarum RYPR1 from an indigenous fermented beverage Raabadi. Front Microbiol 7:1683. https://doi.org/10.3389/fmicb.2016.01683
Havenaar R, Brink BT, Veld JHJHI (1992) Selection of strains for probiotic use. Springer Netherlands:209–224. https://doi.org/10.1007/978-94-011-2364-8_9
Klopper KB, Deane SM, Lmt D (2017) Aciduric strains of Lactobacillus reuteri and Lactobacillus rhamnosus, isolated from human feces, have strong adhesion and aggregation properties. Probiotics Antimicro 2:1–9. https://doi.org/10.1007/s12602-017-9307-5
Beasley SS, Manninen TJK, Saris PEJ (2006) Lactic acid bacteria isolated from canine faeces. J Appl Microbiol 101(1):131–138. https://doi.org/10.1111/j.1365-2672.2006.02884.x
Blazenka K, Jagoda Š, Jasna B, Krešimir G, Jadranka F, Carlo I, Francesco C (2008) Characterization of the three selected probiotic strains for the application in food industry. World J Microbiol Biotechnol 24(5):699–707
Verso LL, Lessard M, Talbot G, Fernandez B, Fliss I (2017) Isolation and selection of potential probiotic bacteria from the pig gastrointestinal tract. Probiotics Antimicro 5:1–14. https://doi.org/10.1007/s12602-017-9309-3
Sharma P, Tomar SK, Sangwan V, Goswami P, Singh R (2016) Antibiotic resistance of Lactobacillus sp. isolated from commercial probiotic preparations. J Food Saf 36(1):38–51. https://doi.org/10.1111/jfs.12211
Egervärn M, Lindmark H, Olsson J, Roos S (2010) Transferability of a tetracycline resistance gene from probiotic Lactobacillus reuteri to bacteria in the gastrointestinal tract of humans. Anton Leeuw Int J G 97(2):189–200. https://doi.org/10.1007/s10482-009-9401-0
Korhonen JM, Hoek AHAMV, Saarela M, Huys G, Tosi L, Mayrhofer S, Wright AV (2010) Antimicrobial susceptibility of Lactobacillus rhamnosus. Benefic Microbes 1(1):75–80. https://doi.org/10.3920/BM2009.0002
Colombo M, Castilho NP, Todorov SD, Nero LA (2017) Beneficial and safety properties of a Corynebacterium vitaeruminis strain isolated from the cow rumen. Probiotics Antimicro 9(2):157–162. https://doi.org/10.1007/s12602-017-9263-0
Barton MD (2000) Antibiotics or probiotics: reducing antibiotic resistance. J Magn Reson Imaging 13:352–355
Franz CMAP, Melanie H, Hikmate A, Wilhelm H, Antonio G (2011) Enterococci as probiotics and their implications in food safety. Int J Food Microbiol 151(2):125–140. https://doi.org/10.1016/j.ijfoodmicro.2011.08.014
Kreuzer S, Machnowska P, Aßmus J, Sieber M, Pieper R, Schmidt MF, Brockmann GA, Scharek-Tedin L, Johne R (2012) Feeding of the probiotic bacterium Enterococcus faecium NCIMB 10415 differentially affects shedding of enteric viruses in pigs. Vet Res 43(1):95–96. https://doi.org/10.1186/1297-9716-43-58
Hunt CP (1998) The emergence of enterococci as a cause of nosocomial infection. Brit J Biomed Sci 55(2):149–156
Jett BD, Huycke MM, Gilmore MS (1994) Virulence of enterococci. Clin Microbiol Rev 7(4):462–478. https://doi.org/10.1128/CMR.7.4.462
Lee K, Lee M, Lee Y (2008) Safety assessment of commercial Enterococcus probiotics in Korea. J Microbiol Biotechnol 18(5):942–945
Funding
The present study was supported by the National Natural Science Foundation of China (31672318) and the Funded Project of Chengdu Giant Panda Breeding Research Foundation (CPF2014-15, CPF2015-06).
Author information
Authors and Affiliations
Author notes
Xueqin Ni, Qiang Wang and Zhirong Peng are joint first authors.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Liu, Q., Ni, X., Wang, Q. et al. Investigation of Lactic Acid Bacteria Isolated from Giant Panda Feces for Potential Probiotics In Vitro. Probiotics & Antimicro. Prot. 11, 85–91 (2019). https://doi.org/10.1007/s12602-017-9381-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12602-017-9381-8