Abstract
Aphis gossypii (Glover) is one of the major pests of melon crops as well as an efficient vector of non-persistent virus such as Cucumber Mosaic Virus and Zucchini Yellow Mosaic Virus among others. Host-plant resistance is one of the best strategies that can be used to control this pest. In this study 14 Tunisian melon accessions were screened to identify new sources of resistance/tolerance to Aphis gossypii using phenotypic and molecular approaches. Antixenosis, antibiosis and tolerance tests were carried out to phenotype those accessions which were also analyzed by molecular markers linked to the Vat gene which confers resistance to both A. gossypii colonization and virus transmission. Results evidenced that only the accession TUN-7 showed antixenosis, antibiosis and tolerance (no leaf curling), at a similar level to that of the resistant control PI414723. Although plants of the accession TUN-13 did not show leaf curling either, the presence of the Vat gene was only detected in TUN-7; its fruit characteristics, of Ananas type, makes this accession as a valuable source of resistance to this aphid that can be used in breeding programs to develop new aphid resistant melon cultivars.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Melon (Cucumis melo L., 2n = 24) is an outcrossing horticultural crop of economic importance that belongs to the family Cucurbitaceae. One of the main pests affecting this species around the world is Aphis gossypii Glover, the only aphid species able to colonize melon plants. Feeding on melon causes direct damage to the plant by removing photoassimilates producing stunting and severe leaf-curling, and heavy colonization can result in plant death. Aphids also excrete honeydew onto the leaves and fruits which acts as an ideal growth medium for sooty mold, which greatly decreases fruit quality (Dogimont et al. 2014). However the most dangerous damage produced by A. gossypii is indirect since this species is also an efficient vector of many non-persistent viruses such as Cucumber mosaic virus (CMV), Zucchini yellow mosaic virus (ZYMV), Papaya ring spot virus (PRSV) and Watermelon mosaic virus (WMV), causing important losses in melon crops (Sarria-Villada et al. 2009). Besides, this aphid species is also a vector of CABYV (Cucurbit aphid-borne yellows virus), a persistent virus affecting melon crops in many production areas (Lecoq and Desbiez 2012; Kassem et al. 2015).
To decrease the harmful effects of this pest, the most efficient and environmentally acceptable option is the use of aphid resistant melon varieties. A dominant gene, Ag (Aphis gossypii tolerance), was reported to control antixenosis and antibiosis under controlled no-choice tests and free-curling tolerance in LJ 90634, later called PI 414723 (Kishaba et al. 1971, 1976). Antixenosis or non-acceptance consists of disturbance in aphid behavior leading to the rejection of the plant for feeding. Antibiosis corresponds to a modification of the physiology of the insect, affecting demographic parameters (Panda and Khush 1995), resulting in a decrease in the intrinsic growth of the A. gossypii population (Boissot et al. 2010).
Later on, Pitrat and Lecoq (1980, 1986) reported resistance in PI 161375 and PI 414723 to several viruses when they are transmitted by A. gossypii and the gene controlling this character was named Vat (Virus aphid transmission resistance). The gene Vat controls the resistance to the aphid itself by non-acceptance and antibiosis mechanisms (Pitrat and Lecoq 1980; Chen et al. 1997) and, as a consequence, controls resistance to virus transmission by this aphid (Sarria-Villada et al. 2009).
The gene was shown to encode a cytoplasmic protein that has a nucleotide binding site and leucine-rich repeat domains CC-NBS-LRR (Pauquet et al. 2004; Dogimont et al. 2008, 2014), belonging to the R gene family, which is widely associated with active defence mechanisms against pathogens and insects (Shinoda 1993). Consequently, an active defence mechanism against A. gossypii would be present in melon genotypes that carry the Vat gene. Shinoda (1993) described early deposits of callose in leaf tissues of resistant melon plants in response to A. gossypii infestation. Also, Sarria-Villada et al. (2009) showed that the resistance conferred by this gene is associated with a microscopic hypersensitive response specific against A. gossypii. Soon after aphid infestation, phenol synthesis, deposits of callose and lignin in the cell walls, disturbing aphid behavior, were detected in genotypes carrying the Vat gene (Sarria-Villada et al. 2009). Four additive and two couples of epistatic QTLs affecting behaviour and biotic potential of A. gossypii were mapped in GL5 using a population of recombinant inbred lines derived from the cross Védrantais x PI 161375; among them, a major QTL, which affects both behavior and biotic potential of A. gossypii, corresponds to the Vat gene (Boissot et al. 2010). Several genomics studies focused on the region containing Vat showed that the density of genes conferring resistance to various pathogens in melon is highest in this region (Brotman et al. 2002; García-Mas et al. 2012).
The Vat gene has been sequenced and tightly linked markers have been reported (Dogimont et al. 2004, 2007) which allows their use in the selection of A. gossypii resistance. Dogimont et al. (2009) showed that these markers can be used in particular in the context of conventional techniques for varietal selection, in order to facilitate the identification of resistant varieties, and/or the monitoring of the introgression of the resistance characteristic in varieties of agronomic interest. These markers allow detecting the presence or the absence of an allele of the Vat gene that promotes resistance to colonization by the aphid Aphis gossypii and/or resistance to viral transmission by said aphid (Dogimont et al. 2009).
To date, several sources of resistance carrying the Vat gene have been used in melon breeding programs to prevent A. gossypii colonization and aphid virus transmission (Dogimont and Boissot 2014; Boissot et al. 2016); however, most of these sources are quite different from commercial types which, makes it difficult to eliminate undesirable melon characteristics introgressed into the commercial melon lines together with the Vat gene (Palomares-Rius et al. 2018). In spite of the difficulties, several melons varieties, mostly of Charentais type, carrying the Vat gene, have been released (Boissot et al. 2016). The identification of new sources of aphid resistance with agronomic traits and fruit characteristics similar to commercial types of melons is still of interest in breeding programs.
In this study, we screened Tunisian melon germplasm to identify new sources of resistance to aphid infestation using phenotypic screening (antixenosis, antibiosis and tolerance tests) and further molecular marker validation.
Material and methods
Insect and plant material
Aphis gossypii individuals collected from infested melon crops in a greenhouse (IHSM-La Mayora, Málaga, Spain) were kindly provided by Dr. de la Peña. One nymph from a female aphid was chosen to create the clone used in the bioassays. Aphid colonies were reared on plants of the aphid susceptible Spanish melon cultivar ‘ANC-57’ and maintained in a glasshouse at 25 °C (light) and 20 °C (dark) with a photoperiod of 16:8 h (light:dark).
The plant materials used in the experiments were the Tunisian accessions listed in Table 1 and the Indian melon line PI414723 in which resistance to A. gossypii is controlled by the Vat gene (Sarria et al. 2008); the Spanish cultivar ‘Bola de oro’ was used as susceptible control. The Korean melon line PI161375, carrying the Vat gene, was also used in the molecular marker validation.
Melon plants were grown in plastic pots (12 cm diameter) filled with soil-substrate composed of peat (60%), litonite (10%), and compost (30%) and placed in a glasshouse under insect-proof muslin net, in the same environmental conditions as aphid colonies.
Phenotypic evaluation
To assess the plant response to A. gossypii, five plants per accession at the 4th–5th true leaf stage were infested on the second expanded leaf from the apex with 10 apterous aphids (5 to 7 days old), which were previously starved for 2 h. Plants in pots were placed on dishes with water and carefully separated from each other to avoid contact between leaves and pots. Plants were then randomly distributed on the glasshouse bench.
Forty-eight and 72 h after infestation the numbers of aphids remaining on the plant were recorded to evaluate antixenosis. Since in previous experiments aphid disturbance behavior has been observed in resistant lines, which resulted in the aphids tending to leave the site of infestation (Sarria et al. 2008), the numbers of aphids that remained on the infested leaves 48 and 72 h after the infestation were also recorded. The average number of aphids on plants and on the infested leaves was considered as the phenotypic value of each accession.
Garzo et al. (2002) reported that the number of nymphs produced by aphids on resistant plants is lower than on susceptible plants and it could be considered as an antibiosis parameter; therefore the number of nymphs produced on the infested leaf 48 h after the release of the aphids was also scored.
Tolerance to A. gossypii was evaluated in young plants at the first true leaf stage under conditions of massive aphid infestation (10 adults/plantlet). Aphids used in the experiment were young wingless adults 5–7 days old. Tolerance was recorded seven days after infestation in accordance with Ivanoff (1945); plants with curled leaves were considered susceptible to aphids whereas plants with uncurled leaves were considered tolerant.
Molecular marker analysis
The presence of the Vat gene is associated with antixenosis, antibiosis and tolerance mechanisms. Therefore, if any resistant accession was found through the bioassays, molecular analysis based on molecular markers linked to Vat gene were also carried out. For PCR assays, two molecular markers D and E previously reported to be linked to the Vat gene were used (Dogimont et al. 2004). The sequences of the used primers and size fragment are shown in Table 2. The susceptible cultivar ‘Bola de oro’ and the resistant lines PI414723 and PI 161375 were used as controls.
DNA extraction and PCR amplification
Tunisian melon accessions were used to provide template DNA for PCR amplification. Fresh young leaf tissues, of 14-day old plantlets, was collected from each accession and stored at −20 °C till DNA extraction. The genomic DNA was extracted using the CTAB extraction protocol (Doyle and Doyle 1987) with some modifications (Oumouloud et al. 2012). The quality and quantity of extracted DNA was determined using Nanodrop Spectrophotometer ND-100 (Nanodrop Technologies, Delaware, USA). For PCR technique, the DNA concentration was adjusted to 10 ng/μl.
Polymerase chain reaction (PCR) for Marker-D and Marker-E of the Vat locus was carried out following the indications of Dogimont et al. (2007). PCR was performed in 50 μl solution containing 3.52 μl of distilled H2O, 5 μl 5xReaction buffer, 16.5 μl 15 mM MgCl2, 16.5 μl 5 mM dNTP, 3.3 μl of 0.5 μM primer, 10 ng of DNA and 1.37 μl of Taq Polymerase. PCR reactions were performed in a thermocycler (model 9700; Perkin-Elmer Corp., Norwalk, CT, USA). The optimization of PCR is done by a multiple choice of temperature for the annealing phase to set the required temperature.
The PCR products were separated in 2.2% agarose gel pre-stained with ethidium bromide and visualized using an UV transilluminator. The image was photographed and gel images were used for scoring.
Data analysis
To evaluate antixenosis the average number of aphids on both, plants and infested leaves 48 and 72 h after infestation was recorded; to estimate antibiosis, the average number of nymphs on the infested leaves 48 h after infestation was considered as the phenotypic value of each accession. In the tolerance test the occurrence or absence of leaf curling was used to evaluate the character. Data were subjected to analysis of variance (ANOVA) using SPSS for Windows (Version 20). The effect of different melon accession on different parameters was analyzed by one-way-ANOVA. For each parameter, differences among accessions were determined by Student- Newman- Keuls (SNK) test.
Results
Phenotypic evaluation
Most of the aphids remained on plants of all the accessions 48 and 72 h after aphid infestation except on plants of PI414723, where only 5–6 aphids were found (Table 3). The melon cultivar ‘Bola de oro’ was susceptible to aphids, as expected.
However, significant differences among accessions were observed for the number of remaining aphids on the infested leaf. PI414723 was clearly different from the rest since only 2–3 aphids were found on that leaf regardless of the time of evaluation. The number of aphids remaining on the infested leaf of the accession TUN-7 ranged from 5 to 6 at 72 and 48 h after infestation respectively; therefore, TUN-7 showed an antixenotic effect on A. gossypii. The remaining accessions were considered as susceptible as the control ‘Bola de oro’ (Table 3).
The number of nymphs on the infested leaf 48 h after infestation showed significant differences among accessions. Aphid reproduction was very high in most of the accessions and the highest number of nymphs was recorded in the accessions TUN-6 and TUN-12. PI414723 followed by TUN-7 showed the lowest values for this parameter (Table 3); therefore, these genotypes seem to have an antibiotic effect on the aphid.
Seven days after massive aphid infestation the accessions PI414723, TUN-7 and TUN-13 did not show any leaf curling symptoms; therefore, these Tunisian accessions could be considered as tolerant to the aphid. The susceptible cultivar ‘Bola de oro’ and the remaining Tunisian accessions showed leaf curling symptoms, which demonstrated their susceptibility to A. gossypii (Table 3).
Molecular marker analysis
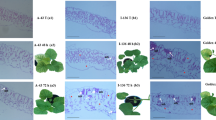
Tolerant melon accessions were further screened for presence of Vat gene conferring resistance to A. gossypii. The two markers (Marker-D and Marker-E) gave reproducible bands with known resistant and susceptible accessions (Fig. 1). The marker-D amplified, in the resistant controls (PI414723 and PI161375) and in TUN-7 a 1549 bp fragment corresponding to the Vat gene. In the susceptible ‘Bola de oro’, the marker amplified a fragment of approximately 750 bp. In TUN-13 only the susceptible allele was also amplified (Fig. 1).
Genotyping results of markers D and E linked to the Vat gene which confers resistance to both A. gossypii colonization and the viruses it transmits. The marker-D showed the resistance alleles of the Vat gene in PI161375, PI414723 and in TUN-7, whereas ‘Bola de oro’ (BO) and TUN-13 showed the susceptible alleles. The marker-E showed the resistance alleles of the Vat gene in PI161375, PI414723 and in TUN-7
The marker-E amplified a 1722 bp fragment corresponding to the Vat gene in PI161375, PI414723 and TUN-7. The amplification product obtained from the genomic DNA of the susceptible ‘Bola de oro’ had a length of approximately 1.3 kb, and the same amplification product was obtained for TUN-13 (Fig. 1).
The Vat gene was then identified in the resistant control melon lines (PI414723 and PI1611375) as well as in the Tunisian melon accession TUN-7.
Discussion
A. gossypii is an efficient vector for many non-persistent viruses, and also transmits CABYV in a persistent, non propagative manner, contributing to the spread of virus diseases. One of the most economically and environmentally options to control this pest is the use of resistant melon varieties. So, there is a need to screen germplasm accessions to identify strong sources of resistance to aphids that will serve as resistant donors in breeding programs. Selection of aphid resistance can be greatly improved by integrating molecular markers with the phenotypic screening.
In the antixenosis test, a high number of aphids remained on both resistant and susceptible plants at 48 h and 72 h after aphid infestation, except on plants of PI414723. These results showed that the methods described by Pitrat and Lecoq (1980) and Martín and Fereres (2003) did not enable us to discriminate between aphid resistance and susceptibility. Following their methodology, all the plants tested in our experiment should be considered susceptible to aphids. However, by scoring the number of aphids only on the infested leaves, significant differences between accessions were detected. Besides, scoring the number of aphids remaining on the infested leaf 72 h after infestation was a more accurate method for discrimination between resistant and susceptible plants than scoring after 48 h. This behavior was also observed by Sarria et al. (2008, 2010). The observed differences between these results and the results obtained by Pitrat and Lecoq (1980) and Martín and Fereres (2003) could be related to differences in the experimental conditions. For example, the age of the plants could influence the expression of resistance to aphids and other insect species, as reported for other plant species (Van Dam et al. 1999; Gurr and McGrath 2001; Leite et al. 2001; Nair et al. 2003; Goggin et al. 2004). In our experiments, the plants used were at the 4th–5th true leaf stage, whereas Pitrat and Lecoq (1980) and Martín and Fereres (2003) used plants at the first true leaf stage. In addition, the aphid clones used by Pitrat and Lecoq (1980) and Martín and Fereres (2003) and the clones employed in our experiments could exhibit different patterns of behavior upon exposure to resistant plants, which might influence the antixenosis and antibiosis results. As reported by Lombaert et al. (2009) and Boissot et al. (2016), different strains/clones of A. gossypii showed phenotypic variability for behavioral traits upon exposure to melon plants that carry the Vat gene.
Many accessions have been tested for their effect on the aphid life-parameters (Pitrat et al. 1996; Fergany et al. 2011). Those large screenings suggested that about 5% of accessions display resistance to colonization by A. gossypii. Among them, only a small number have been also tested for resistance to virus transmission. Up to now, the double phenotype (aphid tolerance and resistance to virus transmission) has been identified in several melon lines (Pitrat and Lecoq 1980; Soria et al. 2000; Thomas et al. 2012; Boissot et al. 2016) coming from Asia, Africa, America, and Europe. However, most of those accessions are quite different from commercial types, which make it difficult to eliminate the undesirable melon characteristics introgressed together with the Vat gene (Palomares-Rius et al. 2018).
Bioassays carried out in this work revealed that TUN-7 carried antixenotic and antibiotic resistance to A. gossypii. Seedlings of this accession grew normally without any curling symptoms after massive aphid infestation, which indicated its aphid tolerance. The accession TUN-13 showed no curling symptoms and also seemed to be tolerant to this aphid. The remaining Tunisian accessions were susceptible. On susceptible plants, aphids established a feeding site on the infested leaf; however, on resistant plants, as observed in previous experiments, most of the aphids left the infested leaf and moved to the apex. Similar findings had been reported by Martín and Fereres (2003).
Breeding programs, for pest resistance, are slow and laborious when based on conventional plant selection methods, since rearing healthy insect colonies and carrying out infestation tests are required. Therefore, the identification of molecular markers tightly linked to a resistance gene allows their use in marker assisted selection, making any breeding program much easier. Resistance to A. gossypii has been genetically characterized: the Vat gene controls antixenosis, antibiosis and A. gossypii tolerance and, resistance to virus transmission restricted to this aphid species (Dogimont et al. 2009; Sarria-Villada et al. 2009). The Vat gene has been sequenced and tightly linked markers have been reported (Dogimont et al. 2004, 2007). These markers allows detecting the presence or the absence of an allele of the Vat gene that promotes resistance to colonization by the aphid Aphis gossypii and resistance to viral transmission by said aphid (Dogimont et al. 2009). The usefulness of those markers in the selection of A. gossypii resistance has been demonstrated in the assays carried out herein. The results obtained support also their effective use in any breeding program leading to the introduction of the Vat gene in melons of commercial value.
Among all the Tunisian accessions the Vat gene was only identified in the TUN-7 accession. This accession belongs to the Ananas type, with ovate-elliptical fruits of yellow and netted skin, weighing around 1500 g, of yellow-orange flesh color. With these fruit characters, TUN-7 is a very interesting and suitable accession to be used as a source of aphid resistance to develop commercial melons of the Ananas type, which are highly appreciated not only in Tunisia but in other Mediterranean countries. The accession TUN-13 was considered as tolerant since non-curling of the infested leaf was observed 7 days after infestation; however, molecular analysis showed that this accession does not carry the Vat gene. To confirm the existence of any other genetic control different to the Vat gene in that accession a more careful study of its behavior against A. gossypii should be carried out. The confirmation of its tolerance by bioassays would lead to the DNA sequencing of TUN-13 looking for molecular polimorphisms that could be associated with aphid tolerance. The localization of resistance genes different from the Vat gene and their use in breeding could reinforce aphid resistance in melon crops.
Conclusion
In this work a new source of resistance to A. gossypii has been found. The TUN-7 accession carries antixenosis, antibiosis and tolerance to A. gossypii controlled by the Vat gene which could avoid the transmission of the virus transmitted by this aphid. Its fruit characteristics, of the Ananas type, make this accession an interesting donor of resistance to commercial melons. On the other hand, the TUN-13 accession should also be explored to clarify whether its A. gossypii tolerance is controlled by other gene different from Vat.
References
Boissot, N., Thomas, S., Sauvion, N., Marchal, C., Pavis, C., & Dogimont, C. (2010). Mapping and validation of QTLs for resistance to aphids and whiteflies in melon. Theoretical and Applied Genetics, 121, 117–125.
Boissot, N., Schoeny, A., & Vanlerberghe-Masutti, F. (2016). Vat, an amazing gene conferring resistance to aphids and viruses they carry: From molecular structure field effects. Frontiers in Plant Science, 7, 1420.
Brotman, Y., Silberstein, L., Kovalski, I., Périn, C., Dogimont, C., Pitrat, M., Klingler, J., Thompson, G. A., & Perl-Treves, R. (2002). Resistance gene homologues in melon are linked to genetic loci conferring disease and pest resistance. Theoretical and Applied Genetics, 104, 1055–1063.
Chen, J. Q., Martin, B., Rahl, M. Y., & Fereres, A. (1997). Early intracellular punctures by two aphid species on near-isogenic melon lines with and without the virus aphid transmission (Vat) resistance gene. European Journal of Plant Pathology, 103, 521–536.
Dogimont, C., & Boissot, N. (2014). Insect resistance in melon and its modification by molecular breeding techniques. In H. Ezura, T. Ariizumi, J. Garcia-Mas, & J. Rose (Eds.), Functional genomics and biotechnology in Solanaceae and Cucurbitaceae crops (pp. 199–299). Berlin: Springer.
Dogimont, C., Bendahmane, A., Pitrat, M., Burget-Bigeard, E., Hagen, L., LeMenn, A., Pauquet, J., Rousselle, P., Caboche, M. & Chovelon, V. (2004). New polynucleotide implicated in plant resistance, useful for producing transgenic plants resistant to Aphis gossypii and associated viral transmission, also encoded protein. World patent WO2004072109-A1.
Dogimont, C., Bendahmane, A., Pitrat, M., Burget-Bigeard, E., Hagen, L., LeMenn, A., Pauquet, J., Rouselle, P., Caboche, M. & Chovelon, V. (2007). Gene resistant to Aphis gossypii. US Patent Application US 2007/0016977 A1.
Dogimont, C., Chovelon, V., Tual, S., Boissot, N., Rittener, V., Giovinazzo, N. & Bendahmane, A. (2008). Molecular diversity at the Vat/Pm-W resistance locus in melon. In: Pitrat M (ed) Cucurbitaceae 2008, Proceedings of the IX th EUCARPIA Meeting on Cucurbit Genetics and Breeding. Avignon, 219–227.
Dogimont, C., Bendahmane, A., Pitrat, M., Burget-Bigeard, E., Hagen, L., LeMenn, A., Pauquet, J., Rouselle, P., Caboche, M. & Chovelon, V. (2009). Gene resistant to Aphis gossypii. US Patent Application US 7,576,264 B2.
Dogimont, C., Chovelon, V., Pauquet, J., Boualem, A., & Bendahmane, A. (2014). The vat locus encodes for a CC-NBS-LRR protein that confers resistance to Aphis gossypiiinfestation and A. gossypii-mediated virus resistance. Plant Journal, 80(6), 993–1004.
Doyle, J. J., & Doyle, J. L. (1987). A rapid isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bulletin, 19, 1–11.
Fergany, M., Kaur, B., Monforte, A. J., Pitrat, M., Rys, C., & Lecoq, H. (2011). Variation in melon (Cucumis melo L.) landraces adapted to the humid tropics of southern India. Genetic Resources and Crop Evolution, 58, 225–243.
García-Mas, J., Benjak, A., Sanseverino, W., Bourgeois, M., Mir, G., & Gonzalez, V. M. (2012). The genome of melon (Cucumis melo L.). Proceedings of the Natural Academy of Sciences, 109, 11872–11877.
Garzo, E., Soria, C., Gomez-Guillamon, M. L., & Fereres, A. (2002). Feeding behavior of Aphis gossypii on resistant genotypes of different melon genotypes (Cucumis melo). Phytoparasitica, 30, 129–140.
Goggin, F. L., Shah, G., Williamson, V. M., & Ullman, D. E. (2004). Developmental regulation of Mi-mediated aphid resistance is independent of Mi-1.2 transcript levels. Molecular Plant Microbe, 17, 532–536.
Gurr, G. M., & McGrath, D. (2001). Effect of plant variety, plant age and photoperiod on glandular pubescence and host-plant resistance to potato moth (Phthorimaea operculella) in Lycopersicon spp. Annals of Applied Biology, 138, 221–230.
Ivanoff, S. S. (1945). A seedling method for testing aphid resistance and its application to breeding and inheritance studies in cucurbits and other plants. Journal of Heredity, 36, 357–361.
Kassem, M. A., Gosalvez, B., Garzo, E., Fereres, A., Gómez-Guillamón, M. L., & Aranda, M. A. (2015). Resistance to Cucurbit aphid-borne yellows virus in melon accession TGR-1551. Phytopathology, 105(10), 1389–1396.
Kishaba, A. N., Bohn, G. W., & Toba, H. H. (1971). Resistance to Aphis gossypii in muskmelon. Journal of Economic Entomology, 64, 935–937.
Kishaba, A. N., Bohn, G. W., & Toba, H. H. (1976). Genetic aspects of antibiosis to Aphis gossypii in Cucumis melo from India. Horticultural Science, 101, 557–561.
Lecoq, H., & Desbiez, C. (2012). Viruses of cucurbit crops in the mediterranean region: An ever-changing picture. In G. Lebenstein & H. Lecoq (Eds.), Viruses and Virus Diseases of Vegetables in the Mediterranean Basin (pp. 68–114). Charleston: US Vegetable Laboratory.
Leite, G. L. D., Picanc, M., Guedes, R. N. C., & Zanuncio, J. C. (2001). Role of plant age in the resistance of Lycopersicon hirsutum f. glabratum to the tomato leaf miner Tuta absoluta (Lepidoptera: Gelechiidae). Scientia Horticulturae, 89, 103–113.
Lombaert, E., Carletto, J., Piotte, C., Fauvergue, X., Lecoq, H., Vanlerberghe-Masuttil, F., & Lapchin, L. (2009). Response of the melon aphid, Aphis gossypii, to host plant resistance: Evidence for high adaptive potential despite low genetic variability. Entomologia Experimentalis et Applicata, 133, 46–56.
Martín, B., & Fereres, A. (2003). Evaluation of a choice-test method to assess resistance melon to Aphis gossypii Glover (Homoptera: Aphididae) by comparison with conventional antibiosis and antixenosis trials. Applied Entomology and Zoology, 38(3), 405–411.
Nair, R. M., Craig, A. D., Auricht, G. C., Edwards, O. R., Robinson, S. S., Otterspoor, M. J., & Jones, J. A. (2003). Evaluating pasture legumes for resistance to aphids. Australian Journal of Experimental Agriculture, 43, 1345–1349.
Oumouloud, A., Mokhtari, M., Chikh-Rouhou, H., Arnedo-Andrés, M. S., Gonzalez-Torres, R., & Álvarez, J. M. (2012). Characterization of the Fusarium wilt resistance Fom-2 gene in melon. Molecular Breeding, 30, 325–334.
Palomares-Rius, F. J., Garcés-Claver, A., Picó, M. B., Esteras, C., Yuste-Lisbona, F. J., & Gómez-Guillamón, M. L. (2018). ‘Carmen’, a yellow-canary melon breeding line resistant to Podosphaera xanthii, Aphis gossypii and Cucurbit Yellow Stunting Disorder Virus. HortScience, 53(7), 1072–1075.
Panda, N., & Khush, G. S. (1995). Host plant resistance to insects. Wallingford: CAB International.
Pauquet, J., Burget, E., Hagen, L., Chovelon, V., Valot, N., Desloire, S., Caboche, M., Rousselle, P., Pitrat, M., Bendahmane, A. & Dogimont, C. (2004). Map-based cloning of the Vat gene from melon conferring resistance to both aphid colonization and aphid transmission of several viruses. In A. Lebeda, H. Paris (eds), Cucurbitaceae 2004, Proceedings of the VIII th Eucarpia Meeting on Cucurbit Genetics and Breeding. Olomouc, Czech Republic, 325–329.
Pitrat, M., & Lecoq, H. (1980). Inheritance of resistance to cucumber mosaic virus transmission by Aphis gossypii in Cucumis melo. Phytopathology, 70, 958–961.
Pitrat, M., & Lecoq, H. (1986). Relations génétiques entre les résistances par non-acceptation et par antibiose du melon Aphis gossypii. Recherche de liaisons avec d'autres gènes. Agronomie, 2, 503–508.
Pitrat, M., Risser, G., Blancard, D. & Lecoq, H. (1996). Evaluation of a melon collection for disease resistances. In M.L. Gómez-Guillamón et al. (eds), Cucurbits towards 2000, Proceedings of the VI th Eucarpia Meeting on Cucurbit Genetics and Breeding. Malaga, 49–58.
Pitrat, M., Hanelt, P., & Hammer, K. (2000). Some comments on infraspecific classification of cultivars of melon. Acta Horticulturae, 510, 29–36.
Sarria, E., Yuste-Lisbona, F.J., Palomares, F.J., López-Sesé, A.I. & Gómez-Guillamón, M.L. (2008). Inheritance of tolerance to Aphis gossypii in C. melo TGR-1551 and its relation with resistance to virus transmission. In: Pitrat M (ed) Cucurbitaceae 2008. Proceedings of the IX th EUCARPIA-Meeting on Cucurbit Genetics and Breeding, Avignon, 465–470.
Sarria, E., Palomares, F. J., López-Sesé, A. I., & Heredia, A. (2010). Role of leaf glandular trichomes of melon plants in deterrence of Aphis gossypii Glover. Plant Biology, 12(3), 503–511.
Sarria-Villada, E., Garzo, E., López-Sesé, A. I., Fereres, A., & Gómez-Guillamón, M. L. (2009). Hypersensitive response to Aphis gossypii Glover in melon genotypes carrying the Vat gene. Journal of Experimental Botany, 60(11), 3269–3277.
Shinoda, T. (1993). Callose reaction induced in melon leaves by feeding of melon aphid, Aphis gossypii GLOVER, as possible aphid-resistant factor. Japanese Journal of Applied Entomology and Zoology, 37(3), 145–152.
Soria, C., Diaz, J. A., Moriones, E., & Gomez-Guillamon, M. L. (2000). Resistance to Aphis gossypii and to virus transmission by this aphid in melon. Acta Horticulturae, 510, 305–312.
Thomas, S., Dogimont, C., & Boissot, N. (2012). Association between Aphis gossypii genotype and phenotype on melon accessions. Arthropod-Plant Interactions, 6, 93–101.
Van Dam, N. M., Hare, J. D., & Elle, E. (1999). Inheritance and distribution of trichome phenotypes in Datura wrightii. Journal of Heredity, 90, 220–227.
Acknowledgments
The authors thank L. Rodriguez and I. Díaz for their help in bioassays and molecular analysis and L. Currah for her help in language editing. This work was supported by a fellowship of one month provided to Ben Belgacem AM, by the Ministry of Higher Education and Scientific Research of Tunisia (MESRST) and partially financed by the Spanish National Research Project AGL2014-53398-C2-1-R.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chikh-Rouhou, H., Ben Belgacem, A.M., Sta-Baba, R. et al. New source of resistance to Aphis gossypii in Tunisian melon accessions using phenotypic and molecular marker approaches. Phytoparasitica 47, 405–413 (2019). https://doi.org/10.1007/s12600-019-00730-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12600-019-00730-5