Abstract
Estimating above-ground biomass (AGB) using machine learning (ML) algorithms and multi-sensor satellite data is a promising approach for monitoring and managing forest resources. This research integrated synthetic aperture radar (SAR) and multispectral imagery alongside in-field observations to accurately estimate above-ground biomass (AGB) in the Purna regional landscape of northern Western Ghats, India. The satellite data employed in the study included dual-polarization (VV + VH) imagery from Sentinel-1 and multi-spectral bands from Sentinel-2, processed and analysed using advanced ML algorithms. The ML algorithms, namely Random Forest (RF), Extreme Gradient Boosting (XGB), and Boosted Regression Trees (BRT), were strategically applied across different model scenarios to determine their effectiveness in AGB prediction. The XGB model displayed the highest accuracy with an R2 value of 0.61 and the lowest RMSE of 37.85 t/ha. The spatial distribution of AGB was successfully mapped, showing varied biomass concentrations throughout the study area. The study’s findings demonstrate the potential of integrating SAR and multispectral data for enhanced AGB estimation and suggest that ML models, specifically algorithms like RF, XGB, and BRT can address the complex relationships between AGB and satellite-derived variables more effectively than traditional methods.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Forests are vital in combating climate change, storing around 80% of terrestrial carbon (Liu et al., 2017). The carbon cycle and above-ground biomass (AGB) have been prioritized within the list of key biodiversity metrics to be monitored through satellite-based observations (Reddy et al., 2023). Accurate AGB measurement, particularly in spatial terms, supports initiatives like reducing emissions from deforestation and forest degradation (REDD +) and informs forest management plans to reduce carbon stock assessment uncertainties (Kaasalainen et al., 2015). The AGB of forests is typically assessed through conventional field measurements or remote sensing techniques (Sainuddin et al., 2023b; West, 2015). While for small forest stands, accurate AGB calculations are best achieved through direct field measurements (Lu, 2006), employing this method on a regional scale is impractical due to its high cost, labour intensity, and time demands (Lu, 2006; Henry, 2011).
Previous studies (Reddy et al., 2016; Saatchi et al., 2011) have demonstrated the effectiveness of remote sensing in quantifying and monitoring forest biomass on a regional level. Consequently, a range of remote sensors, encompassing both passive and active variants, have been employed to estimate AGB. The estimation of AGB through earth observation data requires the use of allometric equations and satellite-acquired structural or biophysical metrics (Boisvenue & White, 2019). Nonetheless, utilizing earth observation data for estimating AGB presents difficulties, such as choosing appropriate models and dealing with the constraints of data availability (Lu, 2006). Optical remote sensing data such as Landsat is frequently used due to its accessibility, extensive temporal coverage, and moderate spatial resolution (Dogru et al., 2020). Sentinel-2, part of the EU Copernicus program, offers improved forest monitoring in tropical regions with additional spectral bands, enhancing AGB estimation (Li et al., 2021; Mutanga et al., 2012). However, optical sensors face limitations, such as difficulty in penetrating dense canopies, susceptibility to cloud cover, and data saturation in areas with dense canopy cover (Lu et al., 2012; Powell et al., 2010). As Landsat-8, Sentinel-2 is less effective at estimating higher biomass levels. The challenge with saturation of biomass is a known problem with low- to medium-spatial-resolution multispectral data (Steininger, 2000). Synthetic aperture radar (SAR) has demonstrated greater efficiency in assessing medium- to high-stand-level biomass. Owing to regular cloud cover, SAR has proven to be a valuable instrument for evaluating AGB in tropical areas (Lu, 2006; Lu et al., 2016). SAR data offers the advantage of being collected during any weather and at all times of the day or night. Its capabilities include seeing through clouds and thick forest covers while also detecting variations in surface texture, dielectric properties, and water content. SAR can offer detailed insights into forest composition depending on the microwave bands (X-, C-, L-, and P-bands) utilized. Co-polarized and cross-polarized SAR data offer unique insights into the orientation and structural characteristics of forest canopies and tree stems, providing valuable information from the backscattered data (Ulaby et al., 1990a). Even though SAR systems don’t extract the vertical composition of vegetation as adeptly as airborne LiDAR, their wide orbital swath makes them advantageous for regional biomass monitoring.
There are three main approaches for estimating forest bio-physical parameters: Empirical data-driven relationships utilize ground measurements to predict variables using statistical regression but are limited by ground measurement quality and regional specificity (Fuchs et al., 2009; Lu et al., 2012; Næsset et al., 2013; Skowronski et al., 2014; Tian et al., 2012). Inverting physical models based on electromagnetic principles simulate a vegetation stand’s response to radiation interactions and require careful inversion due to simplifications of real-world phenomena (Ulaby et al., 1990b; Cartus et al., 2011, 2012; Santoro et al., 2011; Antropov et al., 2013; Sainuddin et al., 2021, 2023a). Non-parametric machine learning (ML) models, like random forest and gradient boosting, leverage complex relationships without assuming data distribution and integrate multiple sensor data for better estimations (Behera et al., 2023; Breidenbach et al., 2012; Jung et al., 2013; McRoberts et al., 2012; Mitchard et al., 2013; Mutanga et al., 2012; Saatchi et al., 2009). Previous research (Kellndorfer et al., 2010; Walker et al., 2007) has shown that integrating data from multiple sensors performs better than data from a single sensor in generating accurate biomass estimations. In the fusion of optical and radar data, numerous investigations (Li et al., 2020; Malhi et al., 2022) have incorporated multispectral bands, vegetation indices, and texture parameters from optical sensors, coupled with radar backscatter coefficients. Additionally, the textures generated from satellite imageries are known for their notable robust adaptability, and are leveraged in many previous studies (Dang et al., 2019; Dong et al., 2020; Eckert, 2012; Kelsey & Neff, 2014) and have confirmed the efficacy of these parameters in AGB assessment.
In this research, the AGB of tropical deciduous forests in the Purna regional forest landscape was estimated by integrating Sentinel 2 optical data with Sentinel-1 SAR data in association with topographical features from SRTM data and the GEDI canopy height product, as referenced in Potapov et al. (2021). Three ML models—random forest (RF), extreme gradient boosting (XGB), and boosted regression tree (BRT)—were methodically utilized in various modelling contexts to evaluate their performance in predicting AGB. The performance of these techniques in AGB prediction was rigorously evaluated by contrasting them against field-measured data, offering insights into their effectiveness and accuracy.
Materials and Methods
Study Area
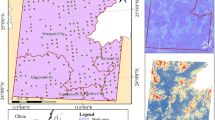
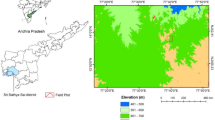
The selected study area is the Purna regional landscape, which includes the Purna Wildlife Sanctuary and surroundings (20° 51′—21° 21′N & 73° 32′—73° 48′ E) spanning the Dang district of Gujarat, India. The study area was outlined by generating a 2 km buffer extending from the boundaries of Purna Wildlife Sanctuary. The landscape spans around 324.88 km2, with 252.36 km2 of this area covered by forests, representing the northern region of the Western Ghats (Reddy et al., 2015). It is in the basins of the Purna and Gira rivers. The highest peak is Walu Dungar, rising to an altitude of 574 m. It experiences a predominantly dry climate. The Southwest Monsoon predominates from June to September. Purna features both moist and dry deciduous forests (Champion & Seth, 1968). The dominant tree species in the study area include Tectona grandis, Wrightia tinctoria, Terminalia alata, Haldina cordifolia, Acacia catechu, Butea monosperma, Desmodium oojeinense, and Mitragyna parvifolia. The study area was outlined by generating a 2 km buffer extending from the boundaries of Purna Wildlife Sanctuary (Fig. 1).
Field Sampling and AGB Estimation
The forest area was stratified based on the forest-type map from Reddy et al. (2015). Field inventory data was collected between 2019 and 2020 across 106 distinct 0.1 ha sample plots spread throughout the study area. This ensures the representation of the diversity of biomass within different forest types. A sampling intensity equivalent to 0.1% of the total forest area was selected due to practical feasibility. Stratified random sampling was utilised to establish these plots, and their coordinates were recorded using a global positioning system (GPS). For each plot, parameters such as height, diameter at breast height (DBH), number of individuals, and species names were documented. The AGB was estimated using an allometric equation (Eq. 1) that incorporated tree height and Diameter at Breast Height (DBH), with distinct coefficients specific to dry and moist deciduous forests proposed by Chave et al. (2005). In the sampled plots, 75.47% were located in the dry deciduous forests, and 24.53% were found in the moist deciduous forests.
Here, ρ signifies the wood density of the tree as suggested by the Forest Research Institute (Chowdhury & Ghosh, 1958), D stands for the diameter at breast height in centimetres, and H denotes the height of the tree, expressed in meters. Table 1 presents the unique coefficients for different forest types applied in the allometric equation. Figure 2 depicts the frequency distribution of the field-measured AGB. Table 2 shows the statistical overview of the field measured AGB in (t/ha) from the sampled plots.
Satellite Data and Predictor Variables
Sentinel-1 Data
The Sentinel-1 program features two satellites: Sentinel-1A (S1A; launched on April 3, 2014) and Sentinel-1B (S1B; launched on April 25, 2016). This satellite is designed with rapid revisit times, broad coverage, and rapid data distribution. Sentinel-1 operates a C-band imager at 5.405 GHz, with an incidence angle ranging from 200 to 450. The satellite maintains a Sun-synchronous, near-polar orbit at an altitude of 693 km. For this study, dual polarization (VV + VH) data from the Sentinel-1A interferometric wide (IW) ground range detection (GRD), acquired on May 3, 2019, was used. The data was accessed freely from the ESA Copernicus hub (https://sentinel.esa.int/web/sentinel/sentinel-data-access). The data preprocessing was conducted using the Sentinel Application Platform (SNAP) (version 8). Once the orbit was applied, the SAR data underwent radiometric calibration and then thermal noise removal. The data was resampled to a pixel size of 30 m to match the size of the sampled field plots. To mitigate the speckle noise in the image, a Gamma MAP filter with a 9 × 9 pixel window was employed.
Sentinel-2 Data
Sentinel-2 (S2A and S2B) has a powerful multispectral instrument (MSI) for advanced optical remote sensing. It offers 13 bands spanning various spectrums in a short 5-day revisit cycle. The spectral bands are divided into three separate spatial resolutions: 10 m, covering the blue, green, red, and near-infrared (NIR) bands; 20 m, including three vegetation red edge bands, a narrow NIR band, and two shortwave infrared (SWIR) bands; and 60 m, which capture the coastal aerosol, water vapor, and SWIR-cirrus bands. The data acquired from the ESA Copernicus hub for January 18, 2020 was used. The pre-processing of the data was primarily done with the Sen2cor tool in SNAP for atmospheric correction, and then the data was resampled to 30 m pixel spacing to align with the field plot dimensions. The data was then geocoded using the Shuttle Radar Topography Mission (SRTM) digital elevation model.
Predictor Variables
This study utilized the Sentinel-1 SAR as a key component in the analysis, using the VV and VH polarizations as predictor variables. The Principal Component Analysis (PCA) was applied to the multispectral bands of Sentinel-2 data to minimize dimensionality while preserving the variability between them. The initial two principal components, PC1 and PC2, accounted for 90% of the dataset variance and were selected for subsequent texture processing. The Gray-level Co-occurrence Matrix (GLCM) method (Haralick et al., 1973) was utilized, where eight GLCM elements were calculated within a 3 × 3 processing window using the SNAP toolbox. Additional predictor variables incorporated include vegetation indices from Sentinel-2 data, such as the Green Normalized Difference Vegetation Index (GNDVI) (Gitelson & Merzlyak, 1998), Green Red Vegetation Index (GRVI) (Tucker, 1979), and Normalized Difference Red Edge Index (NDRE1) (Gitelson and Merzlyak, 1996). These indices were chosen based on the correlation test with field-measured AGB, where GNDVI, GRVI, and NDRE1 emerged as the leading contributors, excluding other indices to prevent the impact of multicollinearity. The Leaf Area Index (LAI) was obtained through the biophysical processor available in the SNAP toolbox, serving as an indicator for biophysical parameters and aligning with the PROSAIL model (Jacquemoud et al., 2009). The assessment also integrated predictor variables like the global canopy height product (Potapov et al., 2021), elevation, slope, and aspects derived from SRTM data. All predictor variables were resampled to 30 m resolution to correspond with the field plots using the nearest neighbourhood method within the resample function of the SNAP toolbox. To mitigate the impacts of location inaccuracy, three neighbourhood statistics (minimum, maximum, and mean) for each variable were computed (Carreiras et al., 2013). This approach resulted in a one-pixel value at each field plot centre, supplemented by three neighbourhood statistical values for each plot, resulting in a total of four values per variable. Consequently, 112 predictor variables were available for modelling purposes. SAR polarizations, along with physical, spectral, biophysical, and texture parameters, were utilized in combinations as predictor variables within the models. For AGB estimation, four selected ML models were examined, each utilizing various variable combinations: (i) Model 1, which estimated AGB using polarizations and physical variables (27 in total), (ii) Model 2, which estimated AGB by combining both spectral and biophysical variables (40 in total), (iii) Model 3, which estimated AGB using only texture variables (68 in total), and (iv) Model 4, which estimated AGB by combining polarizations, physical, spectral, biophysical, and texture variables (112 in total). The choice of predictor variables was guided by findings from previous research, which suggest that integrating polarization channels, textural parameters, and spectral data often leads to reliable AGB estimates in different Indian forest ecosystems. Despite this, the exact combination mentioned in the previous studies was not adopted in the analysis. The predictor variables used for the study are listed in Table 3. A detailed list of employed predictor variables and their details is available in the Supplementary File.
Methods and Modelling
The workflow diagram (Fig. 3) provides a visual representation of the AGB estimation process and implementation of the ML models outlined in this study.
The procedure consists of the following phases:
-
Pre-processing the satellite images and deriving vital predictor variables
-
Training the selected ML models in distinct modeling scenarios
-
Evaluating the efficacy of the models against a test dataset
-
Generating the AGB map based on the best-performing model
This approach involved integrating data from Sentinel-1 and Sentinel-2 with terrain attributes from SRTM data and the canopy height product. The model’s performances were then cross-checked against ground truth data for accuracy. In this study, advanced ML algorithms, including RF, XGB, and BRT, were implemented to predict AGB in different modeling scenarios. Custom Python 3 scripts were utilized for both the modeling and validation processes.
Random Forest Model
The RF operates as an ensemble-learning algorithm, leveraging an extensive collection of decision trees for both regression and classification tasks. Decision trees, a widely recognized approach in machine learning, operate based on specified instructions or conditions for input variables, progressing from the tree’s root to its leaves (Quinlan, 2014). These trees utilize binary division to assign clusters of input variables to each node during the formulation of the regression tree. It’s essential to fine-tune both the number of regression trees and the quantity of input variables for each node. Predictions are then determined by averaging across all tree nodes. The underlying principle of RF centers on amplifying the reduction in variance, by minimizing the correlation among trees (Hastie et al., 2009). To achieve this, input variables are chosen at random during tree development phases.
Extreme Gradient Boosting Model
XGB (Chen et al., 2016) is an advanced ML algorithm that has garnered widespread recognition for its superior performance in Kaggle competitions. This model, which is an optimized version of gradient-boosted regression trees, is tailored for enhanced speed and efficiency. It leverages the second-order derivative of the loss function to hasten convergence and incorporates a regularization component to mitigate the risk of overfitting. As a result, XGB stands out as a versatile and scalable solution, especially adept at managing sparse datasets and achieving rapid convergence.
Boosted Regression Tree Model
The BRT model merges the principles of boosting with the decision tree algorithm to enhance predictive performance. Boosting contributes to reducing the risk of overfitting by selecting random subsets of the training data upon which to base the fitting of new trees. Unlike the RF model that apply bagging, BRTs employ a boosting approach, assigning varying weights to the input data for each successive tree (Biodiversity & Climate Change Virtual Laboratory, 2021). This method ensures that data points that were inadequately predicted by earlier trees are given a greater likelihood of influencing the formation of subsequent trees. Such a strategy increases the model’s precision by allowing it to correct for errors from previous trees when constructing the current one.
Tuning Process of ML Models
To identify the optimal settings, a series of tests employing various tuning parameter values were performed. Refining the tuning parameters for ML models revealed that the accuracy of RF models increased with the addition of trees until reaching a consistent level at a ‘ntree’ setting of 500. In the case of RF, the impact of the ‘mtry’ parameter was more pronounced with fewer trees, diminishing as tree numbers grew. Optimal performance for Model 1 was achieved with an ‘mtry’ of 10, where R2 slightly increased and RMSE remained stable as the tree count increased. Model 2 exhibited a more intricate R2 trend, yet RMSE was simpler to delineate, favoring an ‘mtry’ of 5. The optimal ‘mtry’ parameter for Model 3 was determined to be 3. For the Model 4, an ‘mtry’ of 10 delivered the best outcomes, with R2 and RMSE settling into a consistent range. XGB models showed less sensitivity to gamma, but required balance in child weight as tree depth increased to maintain accuracy. Lower learning rates were beneficial, preventing overfitting and necessitating more iterations for accuracy, with the optimal rate set at 0.01. The optimal subsample rates were below the default, set at 0.5, 0.7, 0.6, and 0.8 for the Model 1, Model 2, Model 3, and Model 4, respectively. The model performance also exhibited a positive correlation with lower ‘nrounds’, maintaining stability as boosting iterations increased. Choosing the right ‘nrounds’ was critical and differed from RF model selection. In the tuning of the BRT model, the learning rates were set within a spectrum from 0.001 to 0.03, specifically being 0.009 for Model 1, 0.001 for Model 2, 0.005 for Model 3, and 0.03 for Model 4 and ‘ntree’ was set to an optimal value of 500. This methodical approach of parameter adjustment led to the development of an optimal ML models.
Model Validation and AGB Estimation
To estimate AGB, four distinct modeling scenarios were evaluated, each employing different sets of variables. The field dataset was divided randomly, with 80% for model training and the remaining 20% for validation. To determine the most effective model for each variable combination, a five-fold cross-validation approach (Kuhn & Johnson, 2013) was employed on the training dataset. The coefficient of determination (R2), root mean square error (RMSE), and mean absolute error (MAE) were analyzed and compared across these models to identify and select the most effective model for mapping AGB. The AGB map was generated using the most accurately fitted model, with a spatial resolution of 30 m across the study area.
Results
This section reveals the findings derived from the study, which concentrates on estimating AGB through the application of multiple ML models using satellite data. The outcomes yielded from the application of advanced ML algorithms such as RF, XGB, and BRT with the combinations of different datasets have been thoroughly and strategically analyzed to identify the effectiveness and accuracy of each in estimating AGB. Figure 4 presents a categorical analysis of the importance of predictor variables. The spectral variables were identified as the most significant, whereas texture and polarization variables also exhibit substantial importance. Physical variables were found to be the least important in this analysis, as indicated by their lower median value and the presence of outliers in the data.
Predictive Modeling of AGB
Figure 5 presents the validation results of the predicted AGB against the observed values for Model 1, using the selected ML algorithms. For the RF model, a moderate correlation is observed with an R2 value of 0.52, an RMSE value of 42.25 t/ha, and a MAE value of 35.89 t/ha. The XGB model exhibits an R2 value of 0.51, an RMSE value of 41.64 t/ha, and a MAE value of 35.44 t/ha. Lastly, the BRT model presents comparable results with an R2 value of 0.47, an RMSE value of 43.02 t/ha, and a MAE value of 37.47 t/ha.
Figure 6 illustrates the validation results for the prediction of AGB in Model 2, employing various ML algorithms. For the RF model, the outcomes indicate a moderate correlation with an R2 value of 0.46, an RMSE value of 42.56 t/ha, and a MAE value of 37.07 t/ha. In the case of the XGB model, the results manifest a correlation with an R2 value of 0.51, an RMSE value of 42.56 t/ha, and a MAE value of 37.70 t/ha. Conversely, the BRT model demonstrated results with an R2 value of 0.44, an RMSE value equal to 40.55 t/ha, and a MAE value of 34.76 t/ha.
Figure 7 delineates the validation results of predicted AGB against observed values for Model 2 for the selected ML models. For the RF model, there’s a moderate correlation observed with an R2 value of 0.39, supplemented by an RMSE value of 44.99 t/ha and a MAE value of 37.40 t/ha. In the XGB model, the performance is slightly varied, with an R2 value of 0.44, an RMSE value of 42.37 t/ha, and a MAE value of 35.71 t/ha. Conversely, the BRT model showcased a moderate R2 value of 0.38, an RMSE value of 45.61 t/ha, and a MAE value of 36.52 t/ha.
Figure 8 showcases the validation results of AGB prediction in Model 4, utilizing the selected ML models. The RF model displays a modest correlation with an R2 value of 0.49, an RMSE value of 41.11 t/ha, and a MAE value of 35.27 t/ha. On the other hand, the XGB model results indicate a strong performance with an R2 value of 0.61, coupled with an RMSE value of 37.85 t/ha and a MAE value of 32.47 t/ha. The BRT model unveils outcomes with an R2 value of 0.41, an RMSE value of 41.81 t/ha, and a MAE value of 35.52 t/ha.
The various ML models applied in this study, namely RF, XGB, and BRT, exhibited a spectrum of performances in predicting AGB across different models. For instance, the RF model, showing variability in performance, managed to present reasonable outcomes in certain models. The XGB model consistently demonstrated moderate to strong correlations in the predictions across all the models. The BRT model displayed variability in its performance yet yielded satisfactory results in some of the tested models. The XGB algorithm showed its strongest performance in Model 4, yielding the highest R2 value. RF performed at its best in Model 4, where it demonstrated a relatively lower error in estimating AGB. The BRT algorithm showed its optimum performance in Model 2, showing a comparatively lower estimation error for AGB.
Figure 9 presents a comparison of the R2, RMSE, and MAE across the different models. The diversity in model performances underscores the importance of selecting an appropriate ML algorithm tailored to the specific characteristics and requirements of each dataset and model to enhance the accuracy and reliability of AGB predictions.
Spatial Mapping of AGB
The distribution map of AGB, depicted in Fig. 10, was produced at a 30 m spatial resolution derived using the XGB algorithm leveraging Model 4 variables. The XGB model incorporating Model 4 demonstrated the highest R2 and the lowest RMSE in comparison to other models. The mean AGB recorded in the field is 94.83 t/ha, while the mean for the predicted AGB is 41.45 t/ha. The predicted AGB within the study area spans from a minimum of 23.43 t/ha to a maximum of 176.61 t/ha. The AGB map utilizes a gradient colour scheme that transitions from yellow to a dark green, representing a range of AGB values from 23.43 to 176.61 t/ha. It depicts AGB density categorized into four distinct ranges, each represented by a colour on the legend. Most of the mapped area is dominated by AGB values in the range of 50–100 t/ha, as indicated by the prevalence of the lime colour. Following this, the next most extensive category is the 100–150 t/ha range, represented by olive shade, which corresponds to regions with relatively higher AGB. The map also shows substantial areas within the 23.43–50 t/ha category, highlighted in yellow, implying regions with a lower biomass density. The darkest green pixels on the map represent the areas with the highest AGB, ranging from 150 to 176.61 t/ha.
Discussions
The study’s findings indicate that multiple ML models such as RF, XGB, and BRT have varied performance in predicting AGB using the selected satellite data, with the RF model generally showing moderate to strong correlations. The strongest performance was observed in Model 4 using the XGB algorithm, achieved the highest R2 and lowest RMSE values, indicating its superior accuracy in AGB estimation. The spatial distribution of AGB was mapped at a 30 m resolution, with the majority of the area displaying AGB values in the range of 50–100 t/ha. This illustrates the proficiency of ML methods in precisely estimating AGB (Dube & Mutanga, 2015). Non-parametric models excel in managing the non-linear relationships between forest AGB and satellite data (Liu et al., 2017). Furthermore, the ability of the ML algorithms to manage non-linearity and assess the significance of predictor variables underscoring its effectiveness (Pandit et al., 2018). This research applied an allometric equation originally proposed by Chave et al. (2005) to estimate the AGB. This method takes into account both the DBH and the height of trees within the sample plots. In a study conducted by Lambert et al., (2005) found that adding tree height to allometric equations, alongside DBH, improves the accuracy of tree volume estimates and decreases the root mean squared error in predictions of total tree biomass. Furthermore, another research conducted by Frank et al., (2018) highlighted the importance of including tree height in models to better reflect variations across different locations.
Relationship Between Satellite Data and AGB
The integration of optical and SAR data marks an advancement in forest AGB estimation over the use of either data source in isolation. While optical imagery provides detailed information on the horizontal layout of forests, its penetrative capacity is limited, primarily capturing surface features rather than the full vertical profile (Myneni et al., 2001). SAR data, particularly at longer wavelengths such as L-band and P-band, can pierce through the canopy to reveal the crucial vertical structure indicative of AGB, which is predominantly composed of stem and branch biomass. The synergistic use of both optical and SAR data leverages the strengths of each. This combined approach, therefore, holds significant promise for enhancing the accuracy and reliability of AGB measurements. This study selected Sentinel-1 SAR data at C-band, because it was readily available for the geographic location of the study. The study examined the VV and VH polarization channels as the predictor variables of the SAR data. The accuracy of AGB estimation by SAR can be compromised by the terrain and can suffer from signal saturation in very dense or high-biomass areas (Imhoff, 1993; Le Toan et al., 1992; Luckman et al., 1997). It has been documented that C-band SAR backscatter typically reaches saturation at AGB levels ranging from 30 to 50 t/ha (Lucas et al., 2015). In the case of optical data, NDVI and EVI are commonly utilized vegetation indices, yet in this study, NDRE1 and GNDVI were found to be superior in estimating AGB in the correlation analysis. This aligns with Wang et al. (2007), who found GNDVI more precise than NDVI in LAI estimation across various conditions. Likewise, these findings are consistent with the research conducted by Otsu et al. (2019), who reported the superior performance of GNDVI in differentiating between broadleaf and needleleaf forests compared to NDVI. Supporting this, Yoder and Waring (1994) identified the green spectral band as more correlational with photosynthetic activity in the tree canopies of miniature Douglas-firs than the red spectral band. The difference in efficacy between NDVI and GNDVI can be attributed to NDVI being more sensitive to lower chlorophyll concentrations, while GNDVI is more effective at detecting higher chlorophyll levels, thereby providing greater accuracy in assessing chlorophyll concentration in tree crowns (Gitelson et al., 1996). In this study, NDRE1 also emerged as a superior predictor for estimating AGB primarily due to its sensitivity in capturing chlorophyll content. The sensitivity of the red-edge bands is particularly crucial, as the reflectance in these bands is influenced by the thickness of the tree canopy layers. Research conducted by Horler et al. (1983), and Eitel et al. (2011), has shown that the red-edge spectral band is particularly adept at estimating AGB in areas of dense canopy coverage, providing a more accurate measurement than traditional vegetation indices through its ability to detect chlorophyll absorption and reflection in leaves. This finding is supported by Mutanga et al. (2012) and Laurin et al. (2018), who have also reported a relationship between the reflectance of red-edge bands and factors such as canopy density and biomass. Since NDRE1 effectively captures variations in these red-edge bands, it serves as a more accurate indicator of the chlorophyll content and, by extension, the overall health and biomass of the canopy. This sensitivity makes NDRE1 particularly effective in environments with dense vegetation, where traditional indices like NDVI might be less responsive due to saturation. NDRE1’s ability to detect subtle changes in chlorophyll content in these dense canopy layers provides a more nuanced and accurate estimation of biomass, distinguishing it from other vegetation indices and explaining its superior performance in the study. In compliance with the previous studies (Ali et al., 2015; Ghosh & Behera, 2018; Liu et al., 2019; Sinha et al., 2015), this study has also demonstrated that by integrating SAR parameters with optical (particularly the red-edge (B5) spectral band) and terrain parameters in ML models, the saturation threshold for biomass density measurements increases, extending up to a higher value.
Efficacy of Machine Learning Approaches in AGB Estimation
Earlier studies on biomass estimation predominantly employed standard statistical regression techniques, for instance, linear regression, which implied a direct linear correlation between independent and dependent variables (Dong, et al., 2003; Le Toan et al., 1992). However, the complexity of the relationship between AGB and satellite data is not adequately addressed by these classical methods. Advanced ML approaches, like RF and XGB, are adept at delineating the intricate non-linear relationships present within heterogeneous data distributions and effectively integrating diverse data sources to enhance the accuracy of biomass estimations. Many previous studies revealed that combining ML algorithms with multi-sensor RS data helps in preventing overfitting and significantly enhances estimation accuracy. For instance, a study conducted by Behera et al. employed a combination of 71 spectral and texture variables, derived from Sentinel-2 in the RF model for estimating AGB in the regional landscape of Eastern Ghats (Behera et al., 2023). Another study conducted by David et al. combined Sentinel-1 SAR and Sentinel-2 multispectral imagery in the RF model to assess AGB of dryland forests of Southern Africa (David et al., 2022). In a related study, Singh and the team compared the efficacy of RF and Artificial Neural Network (ANN) models to estimate the AGB of dry deciduous forests using Sentinel-2 data of different seasons (Singh et al., 2022). In their study, Ghosh and Behera used RF and stochastic gradient Boosting modelling to assess the AGB of dense tropical forests by harnessing 70 predictor variables derived from Sentinel-1 and Sentinel-2 data (Ghosh & Behera, 2018). Similarly, the present study incorporated 112 predictive variables from Sentinel-1, Sentinel-2 data along with variables derived from elevation data and the height product (Supplementary File). Among the three modelling approaches analysed in this study, XGB achieved the best results, exhibiting the highest R2 and the lowest RMSE, outperforming both the RF and BRT models. The superior performance of XGB in this study can be primarily attributed to its inherent algorithmic strengths. XGB represents an enhanced gradient boosting framework known for its flexibility and ability to adjust residuals in the process of developing new trees from existing ones, unlike the RF model where trees are constructed independently (Chen & Guestrin, 2016; Friedman, 2002). XGB represents a more refined version of gradient boosting systems, excelling in processing a regularized learning objective, a feature instrumental in mitigating overfitting (Chen & Guestrin, 2016). However, it’s important to note that challenges like overestimation and underestimation, a common issue in ML algorithms for AGB estimation, were not entirely resolved (Stelmaszczuk-Górska et al., 2015). A key limitation of the decision trees, fundamental to both RF and XGB methods, is their inability to extrapolate beyond the data present in the training set. Moreover, when employing remote sensing datasets for biomass estimation, issues of data saturation can arise (Mutanga & Skidmore, 2004). Additionally, the limited number of plots used in this study restricted the opportunity for a more stratified estimation approach, which might be based on different biomass levels or forest types. Such an approach could potentially reduce estimation errors further. Li et al. (2021) observed that XGB surpassed RF in performance, and another comparison by Li et al. (2020) revealed that XGB excelled beyond both RF and linear regression. The findings of this study are also in concordance with the research done by Zhang et al. (2021) and Luo et al. (2022), which have shown that XGB tends to surpass RF in the performance of regression models. The RF algorithm demonstrated greater ease of calibration and resilience against overfitting compared to BRT, an advantage linked to the bagging technique, which lessens the prediction model’s variance. This aligns with the literature indicating superior performance of the RF model over BRT (Wang et al., 2018).
Multi-Sensor Earth Observation Studies in Indian Forests
Studies have employed remote sensing methods to investigate the biomass of Indian forests, adopting either single or combined use of optical, SAR, and LiDAR data. Reddy et al. (2016) explored the spatial distribution of biomass carbon density in Indian forests from 1930 to 2013 using satellite remote sensing data, historical archives, and collateral data. The study estimated the total aboveground carbon stock (3070.27 Tg C) in 2013, with notable variations observed through different periods. In a study carried out by Ghosh and Behera (2018), they investigated AGB estimation in dense tropical forests using multi-sensor data from Sentinel-1A and Sentinel-2A satellites, combined with machine learning algorithms like RF and stochastic gradient boosting. Their research, focused on Shorea robusta and Tectona grandis species in Katerniaghat Wildlife Sanctuary, Uttar Pradesh, demonstrates the efficacy of integrating SAR data, texture images, and vegetation indices in enhancing AGB estimation accuracy, highlighting the potential of Sentinel satellite data and machine learning in forest biomass assessments. Singh et al. (2022) applied a methodology employing open-source satellite data and ML techniques to monitor AGB at finer scales in Tundi Reserved Forest, Jharkhand. Their case study in the dry deciduous tropical forest of Tundi forest highlighted the superior performance of RF and ANN models using wet season Sentinel-2 data, while dry season data proved challenging for AGB estimation, underscoring the potential of the methodology in enhancing forest carbon stock monitoring. Bhandari and Nandy (2023) conducted research that utilized terrestrial laser scanning (TLS) and satellite-derived forest canopy density (FCD) and spectral indices to predict AGB in the Barkot Reserve Forest in Uttarakhand, demonstrating a strong correlation between TLS measurements and field data. Their approach, combining TLS data with FCD classifications from Landsat-8 OLI, proved effective in estimating the study area’s AGB with high precision. Another study conducted by Singh et al. (2023) Barkot Reserve Forest focused on integrating TLS and ALOS PALSAR L-band SAR data for AGB estimation using machine learning algorithms. The research combined various SAR-derived parameters with TLS measurements of tree dimensions, finding that the RF algorithm outperformed the ANN in AGB prediction, demonstrating the potential of SAR and LiDAR data fusion in enhancing forest biomass assessments. In research conducted by Behera et al. (2023) on estimating regional forest landscape AGB integrated textural and spectral variables from Sentinel-2 with ancillary data, effectively overcoming optical remote sensing saturation effects. Utilizing an RF model, the study achieved a significant correlation in AGB variability, demonstrating the potential of this integrated approach for enhancing AGB mapping accuracy and its applicability in developing generalized AGB models. Sainuddin et al. (2023a) investigated the use of multifrequency SAR data in estimating AGB in the tropical forests of the Western Ghats region of Kerala by applying a vector radiative transfer (VRT) theory-based scattering model. The study utilized dual-pol SAR data from L-band ALOS-2, S-band NovaSAR, and C-band Sentinel-1 to retrieve biophysical parameters like tree height and trunk radius, which were then used to estimate AGB using a general allometric equation. Validation with ground truth data showed the L-band data provided the most accurate AGB estimates, demonstrating its superior potential in biomass estimation over S- and C-band data. In a study conducted by Ayushi et al. (2024), they addressed the complexity of estimating AGB in tropical biodiversity hotspots by employing seven machine learning algorithms to analyse multisource datasets, including Sentinel-1 and -2, topography, soil, and climate. Their findings highlight the effectiveness of an ensemble stacking approach, which integrates these diverse datasets for AGB prediction, showcasing high accuracy and the importance of environmental variables in enhancing estimation precision.
Conclusion
This research has integrated SAR and multispectral imagery from satellites along with physical parameters to map AGB across the deciduous forests of the Purna regional landscape in the Western Ghats. The findings of this study indicate that the enhanced accuracy in AGB estimation can be achieved through the synergy of different data types—both SAR and multispectral sensors. By meticulously applying and comparing models like RF, XGB, and BRT, the study has unveiled their unique advantages when used in synergy with satellite data. The models demonstrated their capability to handle the complex, non-linear relationships that exist between the satellite-derived variables and AGB, with XGB consistently surpassing the performance of RF and BRT in accuracy. Model 4, leveraging XGB, emerged as the most precise, with its superior performance being reflected in the highest R2 of 0.61 and the lowest RMSE of 37.85 t/ha. The spatial analysis at a 30 m resolution highlighted the distribution of AGB across the landscape, revealing the effectiveness of ML methods in capturing the gradations of biomass densities, from low to high AGB ranges. The study demonstrates that the fusion of freely accessible SAR and multispectral data (from Sentinel-1 and Sentinel-2) has the capacity to enhance the accuracy of AGB estimation. SAR backscatter data, when combined with selected optical band data, particularly from red-edge wavelengths, markedly improved the efficacy of the estimation process and mitigated the saturation phenomena usually seen in high biomass areas. Indices such as NDRE1 and GNDVI exhibited stronger linear correlations with AGB than traditional indices like NDVI, with GRVI and EVI. The precision and timeliness provided by these methods are vital for a deeper comprehension of tropical forest ecosystems and for the effective management of forest resources within protected areas. Moving forward, using new technologies and methods could make the estimation of AGB even more accurate. Advancements in sensor technology, including the arrival of higher-resolution satellite imagery, promise to provide data with greater detail, facilitating a more accurate analysis of AGB. The next generation of sensors, including LiDAR profilers like ICESat-2, GEDI, and MOLI, along with SAR sensors such as NISAR, BIOMASS, and ALOS-2, are poised to deliver unparalleled precision and resolution in AGB measurements. Exploring the potential of convolutional neural networks and other deep learning frameworks might reveal patterns and correlations in environmental data that are currently underutilized. As the accuracy of AGB estimation continues to improve, these methodologies hold great promise for better informed and more effective environmental policy and resource management decisions.
References
Ali, I., Greifeneder, F., Stamenkovic, J., Neumann, M., & Notarnicola, C. (2015). Review of machine learning approaches for biomass and soil moisture retrievals from remote sensing data. Remote Sensing, 7(12), 16398–16421. https://doi.org/10.3390/rs71215841
Antropov, O., Rauste, Y., Ahola, H., & Hame, T. (2013). Stand-level stem volume of boreal forests from spaceborne SAR imagery at L-band. IEEE Journal of Selected Topics in Applied Earth Observations and Remote Sensing, 6(1), 35–44. https://doi.org/10.1109/JSTARS.2013.2241018
Ayushi, K., Babu, K. N., Ayyappan, N., Nair, J. R., Kakkara, A., & Reddy, C. S. (2024). A comparative analysis of machine learning techniques for aboveground biomass estimation: A case study of the Western Ghats India. Ecological Informatics, 20, 102479. https://doi.org/10.1016/j.ecoinf.2024.102479
Behera, D., Kumar, V. A., Rao, J. P., Padal, S. B., Ayyappan, N., & Reddy, C. S. (2023). Estimating aboveground biomass of a regional forest landscape by integrating textural and spectral variables of sentinel-2 along with ancillary data. Journal of the Indian Society of Remote Sensing, 14, 1–13. https://doi.org/10.1007/s12524-023-01740-x
Bhandari, S. K., & Nandy, S. (2023). Forest aboveground biomass prediction by integrating terrestrial laser scanning data, Landsat 8 OLI-derived forest canopy density and spectral indices. Journal of the Indian Society of Remote Sensing, 18, 1–12. https://doi.org/10.1007/s12524-023-01687-z
Biodiversity and Climate Change Virtual Laboratory. (2021). Boosted Regression Tree. Retrieved March 17, 2023, from https://support.bccvl.org.au/support/solutions/articles/6000083202-boosted-regression-tree
Boisvenue, C., & White, J. C. (2019). Information needs of next-generation forest carbon models: Opportunities for remote sensing science. Remote Sensing, 11(4), 463. https://doi.org/10.3390/rs11040463
Breidenbach, J., Næsset, E., & Gobakken, T. (2012). Improving k-nearest neighbor predictions in forest inventories by combining high and low density airborne laser scanning data. Remote Sensing of Environment, 117, 358–365. https://doi.org/10.1016/j.rse.2011.10.010
Carreiras, J., Melo, J., & Vasconcelos, M. (2013). Estimating the above-ground biomass in miombo savanna woodlands (Mozambique, East Africa) using L-band synthetic aperture radar data. Remote Sensing, 5(4), 1524–1548. https://doi.org/10.3390/rs5041524
Cartus, O., Santoro, M., & Kellndorfer, J. (2012). Mapping forest aboveground biomass in the Northeastern United States with ALOS PALSAR dual-polarization L-band. Remote Sensing of Environment, 124, 466–478. https://doi.org/10.1016/j.rse.2012.05.029
Cartus, O., Santoro, M., Schmullius, C. C., & Li, Z. (2011). Large area forest stem volume mapping in the boreal zone using synergy of ERS-1/2 tandem coherence and MODIS vegetation continuous fields. Remote Sensing of Environment, 115, 931–943. https://doi.org/10.1016/j.rse.2010.12.003
Champion, H. G., & Seth, S. K. (1968). A Revised Survey of the Forest Types of India, Government of India, New Delhi.
Chave, J., Andalo, C., Brown, S., Cairns, M. A., Chambers, J. Q., Eamus, D., Folster, H., Fromard, F., Higuchi, N., Kira, T., Lescure, J. P., Nelson, B. W., Ogawa, H., Puig, H., Riera, B., & Yamakura, V. (2005). Tree allometry and improved estimation of carbon stocks and balance in tropical forests. Oecologia, 145(1), 87–99. https://doi.org/10.1007/s00442-005-0100-x
Chen, T., & Guestrin, C. (2016, August). Xgboost: A scalable tree boosting system, in proceedings of the 22nd acm sigkdd international conference on knowledge discovery and data mining (pp. 785–794). https://doi.org/10.1145/2939672.2939785
Chowdhury, K. A., & Ghosh, S. S. (1958). Indian Wood their Identification, Properties and Uses (Vols. 1–6). Forest Research Institute, Dehradun
Dang, A. T. N., Nandy, S., Srinet, R., Luong, N. V., Ghosh, S., & Kumar, A. S. (2019). Forest aboveground biomass estimation using machine learning regression algorithm in Yok Don National Park. Vietnam. Ecological Informatics, 50, 24–32. https://doi.org/10.1016/j.ecoinf.2018.12.010
David, R. M., Rosser, N. J., & Donoghue, D. N. (2022). Improving above ground biomass estimates of Southern Africa dryland forests by combining Sentinel-1 SAR and Sentinel-2 multispectral imagery. Remote Sensing of Environment, 282, 113232. https://doi.org/10.1016/j.rse.2022.113232
Dogru, A. O., Goksel, C., David, R. M., Tolunay, D., Sözen, S., & Orhon, D. (2020). Detrimental environmental impact of large scale land use through deforestation and deterioration of carbon balance in Istanbul Northern Forest Area. Environmental Earth Sciences, 79, 1–13. https://doi.org/10.1007/s12665-020-08996-3
Dong, J., Kaufmann, R. K., Myneni, R. B., Tucker, C. J., Kauppi, P. E., Liski, J., Buermann, W., Alexeyev, V., & Hughes, M. K. (2003). Remote sensing estimates of boreal and temperate forest woody biomass: Carbon pools, sources, and sinks. Remote Sensing of Environment, 84(3), 393–410. https://doi.org/10.1016/S0034-4257(02)00130-X
Dong, L., Du, H., Han, N., Li, X., Zhu, D. E., Mao, F., Zhang, M., Zheng, J., Liu, H., Huang, Z., & He, S. (2020). Application of convolutional neural network on lei bamboo above-ground-biomass (AGB) estimation using Worldview-2. Remote Sensing, 12(6), 958. https://doi.org/10.3390/rs12060958
Dube, T., & Mutanga, O. (2015). Evaluating the utility of the medium-spatial resolution Landsat 8 multispectral sensor in quantifying aboveground biomass in uMgeni catchment, South Africa. ISPRS Journal of Photogrammetry and Remote Sensing, 101, 36–46. https://doi.org/10.1016/j.isprsjprs.2014.11.001
Eckert, S. (2012). Improved forest biomass and carbon estimations using texture measures from WorldView-2 satellite data. Remote Sensing, 4(4), 810–829. https://doi.org/10.3390/rs4040810
Eitel, J. U., Vierling, L. A., Litvak, M. E., Long, D. S., Schulthess, U., Ager, A. A., Krofcheck, D. J., & Stoscheck, L. (2011). Broadband, red-edge information from satellites improves early stress detection in a New Mexico conifer woodland. Remote Sensing of Environment, 115(12), 3640–3646. https://doi.org/10.1016/j.rse.2011.09.002
Frank, J., Castle, M., Westfall, J. A., Weiskittel, A. R., MacFarlane, D. W., Baral, S., Radtke, P. J., & Pelletier, G. (2018). Variation in occurrence and extent of internal stem decay in standing trees across the eastern US and Canada: Evaluation of alternative modelling approaches and influential factors. Forestry: An International Journal of Forest Research, 91(3), 382–399. https://doi.org/10.1093/forestry/cpx054
Friedman, J. H. (2002). Stochastic gradient boosting. Computational Statistics & Data Analysis, 38(4), 367–378. https://doi.org/10.1016/S0167-9473(01)00065-2
Fuchs, H., Magdon, P., Kleinn, C., & Flessa, H. (2009). Estimating aboveground carbon in a catchment of the Siberian forest tundra: Combining satellite imagery and field inventory. Remote Sensing of Environment, 113(3), 518–531. https://doi.org/10.1016/j.rse.2008.07.017
Ghosh, S. M., & Behera, M. D. (2018). Aboveground biomass estimation using multi-sensor data synergy and machine learning algorithms in a dense tropical forest. Applied Geography, 96, 29–40. https://doi.org/10.1016/j.apgeog.2018.05.011
Gitelson, A. A., Kaufman, Y. J., & Merzlyak, M. N. (1996). Use of a green channel in remote sensing of global vegetation from EOS-MODIS. Remote Sensing of Environment, 58(3), 289–298. https://doi.org/10.1016/S0034-4257(96)00072-7
Gitelson, A. A., & Merzlyak, M. N. (1998). Remote sensing of chlorophyll concentration in higher plant leaves. Advances in Space Research, 22(5), 689–692. https://doi.org/10.1016/S0273-1177(97)01133-2
Haralick, R. M., Shanmugam, K., & Dinstein, I. H. (1973). Textural features for image classification. IEEE Transactions on Systems, Man, and Cybernetics, 6, 610–621. https://doi.org/10.1109/TSMC.1973.4309314
Hastie, T., Tibshirani, R., Friedman, J. H., & Friedman, J. H. (2009). The elements of statistical learning: data mining, inference, and prediction (pp. 1–758). Springer.
Henry, M., Picard, N., Trotta, C., Manlay, R., Valentini, R., Bernoux, M., & Saint André, L. (2011). Estimating tree biomass of sub-Saharan African forests: A review of available allometric equations.
Horler, D. N. H., Dockray, M., Barber, J., & Barringer, A. R. (1983). Red edge measurements for remotely sensing plant chlorophyll content. Advances in Space Research, 3(2), 273–277. https://doi.org/10.1016/0273-1177(83)90130-8
Imhoff, M. L. (1993, August). Radar backscatter/biomass saturation: Observations and implications for global biomass assessment. In Proceedings of IGARSS’93-IEEE International Geoscience and Remote Sensing Symposium (pp. 43–45). IEEE. https://doi.org/10.1109/IGARSS.1993.322465
Jacquemoud, S., Verhoef, W., Baret, F., Bacour, C., Zarco-Tejada, P. J., Asner, G. P., François, C., & Ustin, S. L. (2009). PROSPECT+ SAIL models: A review of use for vegetation characterization. Remote Sensing of Environment, 113, S56–S66. https://doi.org/10.1016/j.rse.2008.01.026
Jung, J., Kim, S., Hong, S., Kim, K., Kim, E., Im, J., & Heo, J. (2013). Effects of national forest inventory plot location error on forest carbon stock estimation using k-nearest neighbor algorithm. ISPRS Journal of Photogrammetry and Remote Sensing, 81, 82–92. https://doi.org/10.1016/j.isprsjprs.2013.04.008
Kaasalainen, S., Holopainen, M., Karjalainen, M., Vastaranta, M., Kankare, V., Karila, K., & Osmanoglu, B. (2015). Combining lidar and synthetic aperture radar data to estimate forest biomass: Status and prospects. Forests, 6(1), 252–270. https://doi.org/10.3390/f6010252
Kellndorfer, J. M., Walker, W. S., LaPoint, E., Kirsch, K., Bishop, J., & Fiske, G. (2010). Statistical fusion of Lidar, InSAR, and optical remote sensing data for forest stand height characterization: A regional-scale method based on LVIS, SRTM, Landsat ETM+, and ancillary data sets. Journal of Geophysical Research: Biogeosciences, 115(G2), 997. https://doi.org/10.1029/2009JG000997
Kelsey, K. C., & Neff, J. C. (2014). Estimates of aboveground biomass from texture analysis of Landsat imagery. Remote Sensing, 6(7), 6407–6422. https://doi.org/10.3390/rs6076407
Kuhn, M., & Johnson, K. (2013). Applied predictive modeling (p. 13). Springer.
Lambert, M. C., Ung, C.H., & Raulier, F. (2005). Canadian national tree aboveground biomass models. Canadian Journal of Forest Research, 35(8), 1996–2018. https://doi.org/10.1139/x05-112
Laurin, G. V., Balling, J., Corona, P., Mattioli, W., Papale, D., Puletti, N., Rizzo, M., Truckenbrodt, J., & Urban, M. (2018). Above-ground biomass prediction by Sentinel-1 multitemporal data in central Italy with integration of ALOS2 and Sentinel-2 data. Journal of Applied Remote Sensing, 12(1), 016008–016008. https://doi.org/10.1117/1.JRS.12.016008
Le Toan, T., Beaudoin, A., Riom, J., & Guyon, D. (1992). Relating forest biomass to SAR data. IEEE Transactions on Geoscience and Remote Sensing, 30(2), 403–411. https://doi.org/10.1109/36.134089
Li, C., Zhou, L., & Xu, W. (2021). Estimating aboveground biomass using sentinel-2 MSI data and ensemble algorithms for grassland in the Shengjin Lake Wetland. China. Remote Sensing, 13(8), 1595. https://doi.org/10.3390/rs13081595
Li, Y., Li, M., Li, C., & Liu, Z. (2020). Forest aboveground biomass estimation using Landsat 8 and Sentinel-1A data with machine learning algorithms. Scientific Reports, 10(1), 9952. https://doi.org/10.1117/1.JRS.9.097696
Liu, K., Wang, J., Zeng, W., & Song, J. (2017). Comparison and evaluation of three methods for estimating forest above ground biomass using TM and GLAS data. Remote Sensing, 9(4), 341. https://doi.org/10.3390/rs9040341
Liu, Y., Gong, W., Xing, Y., Hu, X., & Gong, J. (2019). Estimation of the forest stand mean height and aboveground biomass in Northeast China using SAR Sentinel-1B, multispectral Sentinel-2A, and DEM imagery. ISPRS Journal of Photogrammetry and Remote Sensing, 151, 277–289. https://doi.org/10.1016/j.isprsjprs.2019.03.016
Lu, D. (2006). The potential and challenge of remote sensing-based biomass estimation. International Journal of Remote Sensing, 27(7), 1297–1328. https://doi.org/10.1080/01431160500486732
Lu, D., Chen, Q., Wang, G., Liu, L., Li, G., & Moran, E. (2016). A survey of remote sensing-based aboveground biomass estimation methods in forest ecosystems. International Journal of Digital Earth, 9(1), 63–105. https://doi.org/10.1080/17538947.2014.990526
Lu, D., Chen, Q., Wang, G., Moran, E., Batistella, M., Zhang, M., Vaglio Laurin, G., & Saah, D. (2012). Aboveground forest biomass estimation with Landsat and LiDAR data and uncertainty analysis of the estimates. International Journal of Forestry Research. https://doi.org/10.1155/2012/436537
Lucas, R. M., Mitchell, A. L., & Armston, J. (2015). Measurement of forest above-ground biomass using active and passive remote sensing at large (subnational to global) scales. Current Forestry Reports, 1, 162–177. https://doi.org/10.1007/s40725-015-0021-9
Luckman, A., Baker, J., Kuplich, T. M., Yanasse, C. C. F., & Frery, A. C. (1997). A study of the relationship between radar backscatter and regenerating tropical forest biomass for spaceborne SAR instruments. Remote Sensing of Environment, 60(1), 1–13. https://doi.org/10.1016/S0034-4257(96)00121-6
Luo, K., Wei, Y., Du, J., Liu, L., Luo, X., Shi, Y., Pei, X., Lei, N., Song, C., Li, J., & Tang, X. (2022). Machine learning-based estimates of aboveground biomass of subalpine forests using Landsat 8 OLI and Sentinel-2B images in the Jiuzhaigou National Nature Reserve, Eastern Tibet Plateau. Journal of Forestry Research, 10, 1–12. https://doi.org/10.1007/s11676-021-01421-w
Malhi, R. K. M., Anand, A., Srivastava, P. K., Chaudhary, S. K., Pandey, M. K., Behera, M. D., Kumar, A., Singh, P., & Kiran, G. S. (2022). Synergistic evaluation of Sentinel 1 and 2 for biomass estimation in a tropical forest of India. Advances in Space Research, 69(4), 1752–1767. https://doi.org/10.1016/j.asr.2021.03.035
McRoberts, R. E., Gobakken, T., & Næsset, E. (2012). Post-stratified estimation of forest area and growing stock volume using lidar-based stratifications. Remote Sensing of Environment, 125, 157–166. https://doi.org/10.1016/j.rse.2012.07.002
Mitchard, E. T., Saatchi, S. S., Baccini, A., Asner, G. P., Goetz, S. J., Harris, N. L., & Brown, S. (2013). Uncertainty in the spatial distribution of tropical forest biomass: A comparison of pan-tropical maps. Carbon Balance and Management, 8, 1–13. https://doi.org/10.1016/j.rse.2010.05.010
Mutanga, O., Adam, E., & Cho, M. A. (2012). High density biomass estimation for wetland vegetation using WorldView-2 imagery and random forest regression algorithm. International Journal of Applied Earth Observation and Geoinformation, 18, 399–406. https://doi.org/10.1016/j.jag.2012.03.012
Mutanga, O., & Skidmore, A. K. (2004). Narrow band vegetation indices overcome the saturation problem in biomass estimation. International Journal of Remote Sensing, 25(19), 3999–4014. https://doi.org/10.1080/01431160310001654923
Myneni, R. B., Dong, J., Tucker, C. J., Kaufmann, R. K., Kauppi, P. E., Liski, J., Zhou, L., Alexeyev, V., & Hughes, M. K. (2001). A large carbon sink in the woody biomass of Northern forests. Proceedings of the National Academy of Sciences, 98(26), 14784–14789. https://doi.org/10.1073/pnas.261555198
Næsset, E., Gobakken, T., Bollandsås, O. M., Gregoire, T. G., Nelson, R., & Ståhl, G. (2013). Comparison of precision of biomass estimates in regional field sample surveys and airborne LiDAR-assisted surveys in Hedmark County, Norway. Remote Sensing of Environment, 130, 108–120. https://doi.org/10.1016/j.rse.2012.11.010
Otsu, K., Pla, M., Duane, A., Cardil, A., & Brotons, L. (2019). Estimating the threshold of detection on tree crown defoliation using vegetation indices from UAS multispectral imagery. Drones, 3(4), 80. https://doi.org/10.3390/drones3040080
Pandit, S., Tsuyuki, S., & Dube, T. (2018). Estimating above-ground biomass in sub-tropical buffer zone community forests, Nepal, using sentinel 2 data. Remote Sensing, 10(4), 601. https://doi.org/10.3390/rs10040601
Potapov, P., Li, X., Hernandez-Serna, A., Tyukavina, A., Hansen, M. C., Kommareddy, A., & Hofton, M. (2021). Mapping global forest canopy height through integration of GEDI and Landsat data. Remote Sensing of Environment, 253, 112165. https://doi.org/10.1016/j.rse.2020.112165
Powell, S. L., Cohen, W. B., Healey, S. P., Kennedy, R. E., Moisen, G. G., Pierce, K. B., & Ohmann, J. L. (2010). Quantification of live aboveground forest biomass dynamics with Landsat time-series and field inventory data: A comparison of empirical modeling approaches. Remote Sensing of Environment, 114(5), 1053–1068. https://doi.org/10.1016/j.rse.2009.12.018
Quinlan, J. R. (2014). C4. 5: programs for machine learning. Elsevier.
Reddy, C. S., Jha, C. S., Diwakar, P. G., & Dadhwal, V. K. (2015). Nationwide classification of forest types of India using remote sensing and GIS. Environmental Monitoring and Assessment, 187, 1–30. https://doi.org/10.1007/s10661-015-4990-8
Reddy, C. S., Rakesh, F., Jha, C. S., Athira, K., Singh, S., Alekhya, V. P., Rajashekar, G., Diwakar, P. G., & Dadhwal, V. K. (2016). Geospatial assessment of long-term changes in carbon stocks and fluxes in forests of India (1930–2013). Global and Planetary Change, 143, 50–65. https://doi.org/10.1016/j.gloplacha.2016.05.011
Reddy, C. S., Satish, K. V., Saranya, K. R. L., Sri Surya, N. N., Neha, P. A., & Rajashekar, G. (2023). Harnessing essential biodiversity variables and remote sensing of earth observations-Synthesizing biodiversity insights. Spatial Information Research, 21, 1–12. https://doi.org/10.1007/s41324-023-00558-6
Saatchi, S. S., Harris, N. L., Brown, S., Lefsky, M., Mitchard, E. T., Salas, W., Zutta, B. R., Buermann, W., Lewis, S. L., Hagen, S., & Morel, A. (2011). Benchmark map of forest carbon stocks in tropical regions across three continents. Proceedings of the National Academy of Sciences, 108(24), 9899–9904. https://doi.org/10.1073/pnas.1019576108
Saatchi, S., Malhi, Y., Zutta, B., Buermann, W., Anderson, L. O., Araujo, A. M., Phillips, O. L., Peacock, J., Ter Steege, H., Lopez Gonzalez, G., & Ramírez, H. A. (2009). Mapping landscape scale variations of forest structure, biomass, and productivity in Amazonia. Biogeosciences Discussions, 6(3), 5461–5505. https://doi.org/10.5194/bgd-6-5461-2009
Sainuddin, F. V., Chirakkal, S., Asok, S. V., & Putrevu, D. (2021). Forest stand height estimation by inversion of Polarimetric canopy scattering models. In 2021 IEEE International India Geoscience and Remote Sensing Symposium (InGARSS) (pp. 532–536). IEEE. https://doi.org/10.1109/InGARSS51564.2021.9791867
Sainuddin, F. V., Chirakkal, S., Asok, S. V., Das, A. K., & Putrevu, D. (2023a). Evaluation of multifrequency SAR data for estimating tropical above-ground biomass by employing radiative transfer modeling. Environmental Monitoring and Assessment, 195(9), 1–21. https://doi.org/10.1007/s10661-023-11715-7
Sainuddin, F. V., Mathew, S. M., Saranya, K. R. L., Suthari, S., Smitha, V. A., & Reddy, C. S. (2023). From field to map: a review of state-of-the-art approaches to estimate above-ground biomass integrating remote sensing techniques. In S. Suthari (Ed.), Exploring emerging techniques in plant sciences (pp. 168–181). Vaagdevi Colleges.
Santoro, M., Beer, C., Cartus, O., Schmullius, C., Shvidenko, A., McCallum, I., Wegmüller, U., & Wiesmann, A. (2011). Retrieval of growing stock volume in boreal forest using hyper-temporal series of Envisat ASAR ScanSAR backscatter measurements. Remote Sensing of Environment, 115(2), 490–507. https://doi.org/10.1016/j.rse.2010.09.018
Singh, A., Kushwaha, S. K. P., Nandy, S., Padalia, H., Ghosh, S., Srivastava, A., & Kumari, N. (2023). Aboveground forest biomass estimation by the integration of TLS and ALOS PALSAR data using machine learning. Remote Sensing, 15(4), 1143. https://doi.org/10.3390/rs15041143
Singh, C., Karan, S. K., Sardar, P., & Samadder, S. R. (2022). Remote sensing-based biomass estimation of dry deciduous tropical forest using machine learning and ensemble analysis. Journal of Environmental Management, 308, 114639. https://doi.org/10.1016/j.jenvman.2022.114639
Sinha, S., Jeganathan, C., Sharma, L. K., & Nathawat, M. S. (2015). A review of radar remote sensing for biomass estimation. International Journal of Environmental Science and Technology, 12, 1779–1792. https://doi.org/10.1007/s13762-015-0750-0
Skowronski, N. S., Clark, K. L., Gallagher, M., Birdsey, R. A., & Hom, J. L. (2014). Airborne laser scanner-assisted estimation of aboveground biomass change in a temperate oak–pine forest. Remote Sensing of Environment, 151, 166–174. https://doi.org/10.1016/j.rse.2013.12.015
Steininger, M. K. (2000). Satellite estimation of tropical secondary forest above-ground biomass: Data from Brazil and Bolivia. International Journal of Remote Sensing, 21(6–7), 1139–1157. https://doi.org/10.1080/014311600210119
Stelmaszczuk-Górska, M. A., Rodriguez-Veiga, P., Ackermann, N., Thiel, C., Balzter, H., & Schmullius, C. (2015). Non-parametric retrieval of aboveground biomass in Siberian Boreal Forests with ALOS PALSAR interferometric coherence and backscatter intensity. Journal of Imaging, 2(1), 1. https://doi.org/10.3390/jimaging2010001
Tian, X., Su, Z., Chen, E., Li, Z., van der Tol, C., Guo, J., & He, Q. (2012). Reprint of: Estimation of forest above-ground biomass using multi-parameter remote sensing data over a cold and arid area. International Journal of Applied Earth Observation and Geoinformation, 17, 102–110. https://doi.org/10.1016/j.jag.2012.03.007
Tucker, C. J. (1979). Red and photographic infrared linear combinations for monitoring vegetation. Remote Sensing of Environment, 8(2), 127–150. https://doi.org/10.1016/0034-4257(79)90013-0
Ulaby, F. T., & Elachi, C. (1990). Radar polaritnetry for geoscience applications. Taylor & Francis.
Ulaby, F. T., Sarabandi, K., Mcdonald, K. Y. L. E., Whitt, M., & Dobson, M. C. (1990). Michigan microwave canopy scattering model. International Journal of Remote Sensing, 11(7), 1223–1253. https://doi.org/10.1080/01431169008955090
Walker, W. S., Kellndorfer, J. M., LaPoint, E., Hoppus, M., & Westfall, J. (2007). An empirical InSAR-optical fusion approach to mapping vegetation canopy height. Remote Sensing of Environment, 109(4), 482–499. https://doi.org/10.1016/j.rse.2007.02.001
Wang, B., Waters, C., Orgill, S., Cowie, A., Clark, A., Li Liu, D., Simpson, M., McGowen, I., & Sides, T. (2018). Estimating soil organic carbon stocks using different modelling techniques in the semi-arid rangelands of eastern Australia. Ecological Indicators, 88, 425–438. https://doi.org/10.1016/j.ecolind.2018.01.049
Wang, F. M., Huang, J. F., Tang, Y. L., & Wang, X. Z. (2007). New vegetation index and its application in estimating leaf area index of rice. Rice Science, 14(3), 195–203. https://doi.org/10.1016/S1672-6308(07)60027-4
West, P. W. (2015). Tree and forest measurement (Vol. 20). Springer. https://doi.org/10.1007/978-3-319-14708-6
Yoder, B. J., & Waring, R. H. (1994). The normalized difference vegetation index of small Douglas-fir canopies with varying chlorophyll concentrations. Remote Sensing of Environment, 49(1), 81–91. https://doi.org/10.1016/0034-4257(94)90061-2
Zhang, Y., Xia, C., Zhang, X., Cheng, X., Feng, G., Wang, Y., & Gao, Q. (2021). Estimating the maize biomass by crop height and narrowband vegetation indices derived from UAV-based hyperspectral images. Ecological Indicators, 129, 107985. https://doi.org/10.1016/j.ecolind.2021.107985
Acknowledgements
The authors express their gratitude for the financial support provided to Faseela V. Sainuddin by the Department of Environment and Climate Change (DoECC), Kerala, as part of the Paristhithi Poshini Fellowship scheme. Gratitude is extended to the Director of NRSC, the Principal of All Saints’ College, Thiruvananthapuram, and the Head of the Department of Botany at The Maharaja Sayajirao University, Vadodara, for providing facilities and encouragement.
Funding
This work has been carried out as part of a project on ‘Biodiversity Characterization at Community level in India using Earth Observation Data’ through the Department of Biotechnology and the Department of Space, Government of India.
Author information
Authors and Affiliations
Contributions
FVS: Conceptualization, Methodology, Data curation, Software, Writing—original draft, Visualization. GM: Data curation, Writing—review & editing, AR: Data curation, Writing—review & editing. PSN: Data curation, Validation, Writing—review & editing, Supervision. SVA: Writing—review & editing, Supervision. CSR: Data curation, Writing—review & editing, Supervision, Project administration, funding acquisition.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
About this article
Cite this article
Sainuddin, F.V., Malek, G., Rajwadi, A. et al. Estimating Above-Ground Biomass of the Regional Forest Landscape of Northern Western Ghats Using Machine Learning Algorithms and Multi-sensor Remote Sensing Data. J Indian Soc Remote Sens 52, 885–902 (2024). https://doi.org/10.1007/s12524-024-01836-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12524-024-01836-y