Abstract
Chromium is a carcinogenic toxicant widely used in many industries. The concentration of chromium is increasing in various areas of the world causing threat to living beings. Bioremediation is an inexpensive and eco-friendly approach to detoxify such contaminants. Many bacteria can live both as free cells (planktonic) and as biofilms (sessile), and the pattern of chromium reduction is different in these live forms. In the present study, chromium reduction by planktonic cells and biofilms of bacteria was determined in a comparative manner. Chromium-resistant bacteria capable of biofilm formation were isolated from contaminated soil and wastewater. The bacteria were characterized morphologically and biochemically, and identified by 16S rRNA gene sequencing. Cr(VI) reduction by planktonic cells and biofilms was determined over different periods of incubation. The wastewater isolates showed significant resistance against Cr(VI). Minimum inhibitory concentration (MIC) of Cr (VI) was found to be 900 μg ml−1 for Staphylococcus simulans KW1 and Staphylococcus hominis KW2. Whereas, Bacillus cereus KW4 and Staphylococcus equorum KS1 showed MIC values of 800 μg ml−1 and 850 μg ml−1 respectively. Among all the isolates, maximum biofilm formation was shown by Staphylococcus equorum KS1 both qualitatively and quantitatively. Planktonic cells of these bacteria were more efficient in Cr(VI) reduction as compared to their biofilms in 24 h. Maximum Cr(VI) reduction in planktonic form was shown by Staphylococcus simulans KW1 (27.9 μg ml−1, 55%), whereas in biofilm mode of growth the maximum Cr(VI) reduction was shown by Staphylococcus equorum KS1 (4.6 μg ml−1, 9.2%). The results show that for faster Cr(VI) reduction in wastewater the planktonic form of these bacteria is more suited.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Unchecked industrialization and unwise use of natural resources are the most important reasons for the deterioration of natural environment around the globe (Bhargava et al. 2012; Dhal et al. 2013). Heavy metals are of great concern among all the pollutants. These heavy metals can accumulate in food chains and can badly affect living beings as these are toxic in nature (Dixit et al. 2015). Chromium (Quintelas et al. 2013) is one such toxic heavy metal. Though Cr in the form of trivalent ion (Cr III) is required as a micronutrient by many living organisms (Lewicki et al. 2014), however, the elevated concentration of this metal can cause serious hazards to life as it is carcinogenic and mutagenic.
Cr in different forms such as metallic Cr, Cr (III) and Cr (VI) is used in different industrial processes such as leather tanning, electroplating, manufacturing of alloys, in chemical and pigment synthesis, mining procedures and petroleum refining. There are two stable forms of this metal: Cr(VI) and Cr(III). The former is highly soluble and toxic as compared to the latter which is less soluble and less toxic (Oliveira 2012; Mishra and Bharagava 2016). Cr VI containing solid and liquid wastes are produced from these industries on regular basis and are discharged into the environment in many areas of the world, thus contaminating water and soil (Avudainayagam et al. 2003; Oliveira 2012; Tripathi et al. 2012; Mishra and Bharagava 2016). This contamination of water and soil causes toxicity and carcinogenicity for living beings (Dhal et al. 2013).
Remediation efforts focus on reduction-mediated immobilization of Cr. When Cr(VI) is converted to Cr(III), its mobility decreases. Conventional methods such as precipitation by the use of chemicals, electrochemical processing, membrane filtration, ion exchange mechanisms, redox processes, reverse osmosis and extraction through solvents for the reduction/removal of Cr are neither cost effective nor eco-friendly and are difficult to operate (Agarwal et al. 2006; Ahluwalia and Goyal 2007). This difficulty can be overcome by developing some innovative, cost effective and eco-friendly chromium remediation methods. Microbial reduction of Cr is a cost-effective and environment friendly strategy. Bioremediation is the deliberate use of microorganisms for the detoxification/removal of contaminants from the environment and has several advantages over the conventional methods (Wadood and Sabri 2013). The major advantage of using this technology includes no or minimum harm to the environment and the living organisms. Moreover, no toxic chemicals are used in this approach.
Both planktonic and sessile (biofilms) bacteria can be used for bioremediation. Biofilm is a sessile bacterial community formed on abiotic or biotic surfaces and encapsulated in a self-produced matrix of extracellular polymeric substance consisting majorly of different polysaccharides and proteins. This sessile mode of bacterial life provides them extra benefits to withstand harsh environment and tolerate environmental pollutants. Biofilm-based bioremediation is considered as an efficient and advantageous strategy because in this mode of growth the bacteria have the ability to absorb, adsorb and immobilize the pollutants (Quintelas et al. 2013; Hoh et al. 2016; Mitra and Mukhopadhyay 2016).
The aim of the present study was to compare chromium reduction ability of bacteria in planktonic and biofilm modes of growth. The major objectives to achieve this aim were to isolate and characterize Cr-resistant bacteria from indigenous environment, assess their biofilm forming ability and estimate chromium reduction by planktonic cells and biofilms of these bacteria in a comparative manner.
Materials & methods
Sample collection and isolation of Cr-resistant bacteria
Wastewater and soil samples were collected from wastewater treatment plant Kasur, Punjab, in sterilized screw capped bottles and plastic zipper bags, respectively. For bacterial isolation, 50 μl wastewater samples and 50 μl of 10−4 dilution of soil samples were plated on Cr(VI)-supplemented LB agar plates (K2CrO4, 100 μg ml−1). The petri plates were then incubated aerobically at 37 °C for a period of 24 h. After incubation, the colonies were selected based on morphological differences and purified on Cr(VI)-supplemented LB agar using quadrant streaking method. Morphological and biochemical analysis of Cr-resistant bacteria was performed as given by Cappuccino and Sherman (2007).
Optimization of bacterial growth
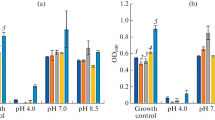
The physiological characterization included the determination of pH and temperature optima for the growth of isolated bacteria. LB broth flasks maintained at pH 5, 6, 7 and 8 were prepared and autoclaved. For the determination of temperature optima, three sets of LB broth medium were prepared and sterilized. For each set (both for pH and temperature), blank LB medium without inoculum was used as control. Media flasks were inoculated with 100 μl of overnight cultures set at 0.1 OD600. The flasks for pH optima determination were incubated at 37 °C at 100 rpm for 24 h, whereas the flasks for the determination of optimum temperature were incubated at 25, 37 and 45 °C at 100 rpm for 24 h. After the completion of the incubation, the OD600nm was measured with the help of a spectrophotometer (IRMECO UV-VIS).
Determination of Cr resistance by the bacterial isolates
Resistance against Cr (VI) was checked by broth dilution method as reported by Randrianarivelo et al. (2009). LB broth (5 ml culture media per test tube) was prepared and supplemented with varying concentrations of Cr(VI) (0–1000 μg ml−1, with 50 μg ml−1 intervals) and autoclaved. McFarland standards were prepared at OD600=0.1 using overnight bacterial cultures, and 50 μl of these was inoculated in the Cr(VI)-supplemented media tubes. The inoculated tubes were put on 24-h incubation at 100 rpm and 37 °C. The tubes were then visualized for presence or absence of growth.
Determination of biofilm forming ability of the bacterial isolates
Initially, the biofilm forming ability of the isolated bacteria was checked by a qualitative ring-test assay. Bacterial isolates were inoculated in LB broth with (50 μg ml−1) and without Cr(VI). These tubes were then incubated under static conditions at a temperature of 37 °C for a period of 24 h. Following the incubation, the liquid culture was decanted, and the tubes were air dried and stained with 0.1% crystal violet at room temperature for 20 min. Excess stain was removed, and the tubes were washed with autoclaved deionized water and air dried at room temperature. The tubes were then observed for the presence of a purple ring (Qurashi and Sabri 2012).
The quantitative biofilm formation was studied in sterile 96-well microtitre plates. Overnight bacterial cultures were standardized (OD600 = 0.1) for the inoculation. Two sets of sterilized LB broth medium, one with (50 μg ml−1) and one without Cr (VI), were prepared and dispensed (250 μl/well) in microtitre plates. Before inoculation, the wells were labeled accordingly. Each experimental well was inoculated with 20 μl of bacterial cultures (OD600 = 0.1) accordingly. The wells for negative control contained the broth without inoculum. The plates were incubated at 37 °C for 24 h. After the completion of the incubation period, the contents of each well were decanted, and wells were washed with sterile distilled water (300 μl/well) thrice. The attached bacteria (biofilms) were fixed using methanol (200 μl/well) for 15 min. After that, the methanol was discarded, and plates were left to air dry. The biofilms were stained for 5 min with 160 μl of crystal violet per well. The plates were washed with tap water to remove extra stain and then air dried. The optical density (OD) of microtitre plates was measured at 570 nm by an ELISA plate reader, after solubilizing the crystal violet dye with 33% glacial acetic acid (160 μl/well) (Yassien and Khardori 2001; Stepanović et al. 2003).
The bacterial isolates can be classified into three categories according to the criteria given by Christensen et al. (1985): absorbance < 0.125 = non biofilm formers, absorbance between 0.125 and 0.25 = weak biofilm formers and absorbance > 0.25 = strong biofilm formers.
Determination of bacterial reduction of Cr in planktonic and biofilm modes of growth
Cr(VI) reduction was determined both by planktonic cells and biofilms of the bacteria. Planktonic cells were prepared by inoculating sterilized LB broth flasks and incubating them at 37 °C, 120 rpm shaking, overnight. To immobilize bacterial cells onto the glass slides (bacterial biofilms), clean glass slides were placed in the flasks containing LB broth and autoclaved. These flasks were then inoculated with overnight bacterial cultures (OD600 = 0.1) and incubated at 37 °C for a period of 48 h under static conditions.
Two sets of LB broth medium were prepared and autoclaved (50 ml medium per flask). The flasks were supplemented with 50 μg ml−1 Cr(VI). One of the sets designated for planktonic cultures was inoculated with overnight bacterial cultures (OD600=0.1). The second set of flasks was inoculated with glass slides containing immobilized bacterial cells (biofilms). Planktonic culture flasks were incubated at 37 °C and 100 rpm for 24 h, whereas the flasks with biofilms of bacteria were incubated at 37 °C under static conditions for 24 h. Uninoculated flasks were used as control.
After incubation, samples were drawn from each flask (both planktonic and biofilms), centrifuged at 10,000 rpm and supernatants were separated. One hundred microliters of diphenylcarbazide solution (0.25 g diphenylcarbazide, 100 ml acetone and 1.0 drop H3PO4) was added to 1 ml of appropriately diluted (usually 1/200) supernatant sample and placed at room temperature for 10 min for the development of purple-colored complex. OD540nm was measured using an IRMECO UV-VIS spectrophotometer (Desjardin et al. 2003).
DNA isolation, 16S rDNA amplification and sequencing
Genomic DNA of four bacterial isolates showing best results was isolated using the method described by Wilson (2001). Universal primers (27F and 1492R) were used to amplify the 16S rRNA genes of the bacterial isolates.. Polymerase chain reaction (PCR) was carried out according to the conditions described by Mohsin et al. (2019). The purity of the PCR products was analyzed by agarose gel electrophoresis. The PCR products were got sequenced from Macrogen Inc. (South Korea) through dideoxy sequencing. The quality of the obtained sequences was checked using Finch TV (Geospiza, Inc. Seattle, WA), and the sequences were cleaned. Contigs were made using NCBI 2-sequence BLAST followed by classification through NCBI-BLAST using 16S rRNA database. Nearest homologous sequences were downloaded, and phylogenetic trees were made using Mega 5.0 (Tamura et al. 2011). The sequences were submitted in GenBank, and accession numbers were obtained.
Statistical analysis
All experiments were performed in triplicates or mentioned otherwise. Mean values were plotted where error bars represent the standard deviation. Microsoft Excel 2016 was used for the analysis.
Results
Isolation of Cr-resistant bacteria
Six morphologically diverse bacterial colonies were selected from the two types of samples plated on LB agar plates containing 100 μg ml−1 Cr(VI). Four bacterial isolates were from wastewater sample, while two of them were from the soil sample. These selected bacteria were purified and labeled as KW1, KW2, KW3 and KW4 (from wastewater) and KS1 and KS2 (from soil sample). The results of morphological and biochemical characterization have been shown in Table 1. The optimum temperature and pH were found to be 37 °C and 7 respectively for all the bacterial isolates (Figs. 1 and 2).
Identification of the isolated bacteria
Analysis of 16S rRNA gene sequences of the bacterial isolates showed that the bacteria belonged to Staphylococcus and Bacillus genera. KS1, KW1, KW2 and KW4 were identified as Staphylococcus equorum, Staphylococcus simulans, Staphylococcus hominis and Bacillus cereus, respectively. The accession numbers assigned to the sequences submitted to GenBank are given in Table 2. The phylogenetic tree of these sequences constructed along with nearest homologues from NCBI BLAST results also confirmed the identity of these isolates (Fig. 3).
Cr resistance by the bacterial isolates
Resistance against Cr(VI) by the bacterial isolates ranged from 650 to 900 μg ml−1. The bacterial isolates KW3 and KS2 resisted Cr(VI) up to 650 μg ml−1, Bacillus cereus KW4 resisted Cr(VI) up to 800 μg ml−1, Staphylococcus equorum KS1 resisted Cr(VI) up to 850 μg ml−1 whereas Staphylococcus simulans KW1 and Staphylococcus hominis KW2 resisted Cr(VI) up to 900 μg ml−1 (Fig. 4).
Biofilm formation by the bacterial isolates
Both qualitative and quantitative assays showed that all the isolated bacteria were biofilm formers in LB broth medium in the absence of Cr(VI). Qualitatively, biofilm was seen as purple rings on the walls of the test tubes visible after staining with 0.1% crystal violet following the incubation (Fig. 5). Quantitatively, the results of biofilm formation were shown as the values of optical density (Fig. 6). When given 50 μg ml−1 Cr(VI) in the media, only four bacterial isolates (Staphylococcus simulans KW1, Staphylococcus hominis KW2, Bacillus cereus KW4 and Staphylococcus equorum KS1) were able to form biofilms (Table 3). Staphylococcus equorum KS1 showed maximum biofilm formation both qualitatively and quantitatively (Figs. 5 and 6). These four bacterial isolates were selected for further experiments.
Cr reduction by planktonic cells and biofilms of the bacteria
In planktonic form, highest concentration of Cr(VI) was reduced by Staphylococcus simulans KW1 (27.9 μg ml−1, 55%), followed by Staphylococcus hominis KW2 (23.7 μg ml−1, 47.4%) and Bacillus cereus KW4 (6.16 μg ml−1, 12.3%). No Cr(VI) reduction was detected by Staphylococcus equorum KS1 planktonic cells (Fig. 7). In case of biofilms, Staphylococcus equorum KS1 showed 4.6 μg ml−1 (9.2%) Cr(VI) reduction, whereas no detectable Cr(VI) reduction was observed by biofilms of the other bacterial isolates.
Discussion
Cr is a carcinogenic and mutagenic pollutant which is discharged by a number of industries in our environment and causes a great threat to living organisms (Ahemad 2014). Attention has been lately focused on bioremediation processes which involve microbes to detoxify toxicants. A large number of bacterial species are known to reduce Cr(VI) to its less toxic form by using different cellular mechanisms (Samuel et al. 2013; Ahemad 2015). In this study, most of the bacteria (KW1, KW2 and KW4) that resisted highest concentration of Cr(VI) belonged to the wastewater sample. Maximum Cr(VI) reduction, in planktonic mode of growth, was also shown by bacteria isolated from wastewater (Staphylococcus simulans KW1, Staphylococcus hominis KW2 and Bacillus cereus KW4) as compared to the soil-borne bacteria. In wastewater environment, mostly bacteria are in planktonic mode of life because of the moving nature of the water, and microorganisms are more readily exposed to toxic substances which enables them to work efficiently in this mode of life (Elahi and Rehman 2019; Liu et al. 2020). On the contrary, soil, being a complex matrix, offers many microsites where microbes can keep their selves safe from the exposure and hazardous effects of the toxicants, and this enables bacteria to exist mostly in biofilm mode which increases their efficiency in this mode of life for various processes including Cr reduction (Almås et al. 2005; SU et al. 2019). Thus, the microbes surviving in wastewater can have more resistance to the toxicants as compared to the microbes present in contaminated soil (Qamar et al. 2017). In this study, soil-borne bacteria showed more biofilm formation as compared to the other bacteria even in the presence of Cr(VI). Out of all the bacterial isolates, Staphylococcus equorum KS1 showed most firm and thick biofilm formation both with and without Cr(VI). Majority of the soil-borne bacteria are known to have biofilm forming character (Foster 1981; Lünsdorf et al. 2000; Burmølle et al. 2007). The key of Cr reduction is their survival in such harsh conditions so that microorganisms (bacteria) reduce toxic hexavalent chromium into less toxic chromium to make the environment better for their survival. The bacteria in this study belonged to two different genera namely Staphylococcus and Bacillus. Cr-resistant Staphylococcus and Bacillus species have also been reported by other researchers (Ilias et al. 2011; Upadhyay et al. 2017).
Many researchers have used biofilms as tools for bioremediation and have reported increased resistance to metals by biofilms (Priester et al. 2006; Harrison et al. 2007; Nancharaiah et al. 2010). However, many researchers have also reported that planktonic bacterial cells are more efficient in reducing Cr(VI) as compared to their biofilm mode of growth (Chen et al. 2012; Pan et al. 2014). Toxicants are more readily available to planktonic bacteria due to larger surface area as compared to biofilms. This could be a reason of higher Cr(VI) reduction by planktonic bacteria as compared to biofilms. This is especially true for shorter incubation times, as in our study. For instance, Saba et al. (2018) reported 3.73 mmol l−1 As(V) reduction in 48 h by planktonic cells and 3.6 mmol l−1 As(V) reduction in 72 h by biofilms of the same bacterial species. This is a clear indication of fast reduction by planktonic cells as compared to biofilms. Biofilms, being more resilient as compared to planktonic cells, might be more suited for longer incubation times. Pan et al. (2014) also reported higher Cr reduction by planktonic cells as compared to the biofilms. They also suggested that the unavailability of the active sites during the biofilm formation could be a reason behind it. In biofilms, most of the cells are hidden inside the EPS matrix, and only the cells exposed to the outside can reduce chromium. This can lead to lower chromium reduction by biofilms as compared to planktonic cells. Cr (VI) reduction by bacteria can be enzymatic or non enzymatic process. Cr(VI) reduction under aerobic conditions commonly uses soluble cytoplasmic reductases which use NADH/NADPH as coenzymes for Cr (VI) reduction (Zhu et al. 2019). Nonenzymatic Cr(VI) reduction usually uses intracellular or extracellular chemical species like organic acids, glutathione or other microbial metabolites (Ahemad 2014). It was also observed in this study, the planktonic mode of growth showed higher Cr reduction as compared to the biofilm mode of growth.
Conclusion
In this study, we compared Cr reduction by planktonic cells and biofilms of Cr-resistant bacteria. The bacteria were isolated from indigenous soil and wastewater. It was found that in planktonic form, the bacteria isolated from wastewater showed maximum Cr reduction, whereas in biofilm mode the bacteria isolated from soil showed maximum Cr reduction. However, irrespective of the isolation source, the Cr reduction was rapid and higher by planktonic cells. Therefore, the planktonic form of these bacteria should be preferred for Cr(VI) reduction for the bioremediation of wastewater. The use of “indigenous environment” means that microorganisms (bacteria) are adapted to the environment and are more capable of cleaning the environment laden with such toxic metal ions. So these isolated bacteria can be used to remediate the indigenous environment (from where they have been isolated) using shorter incubation times as revealed by the study.
Data availability
Not applicable.
References
Agarwal A, Kumar V, Pandey B (2006) Remediation options for the treatment of electroplating and leather tanning effluent containing chromium—a review. Miner Process Extr Metall Rev 27:99–130
Ahemad M (2014) Bacterial mechanisms for Cr (VI) resistance and reduction: an overview and recent advances. Folia Microbiol 59:321–332
Ahemad M (2015) Enhancing phytoremediation of chromium-stressed soils through plant-growth-promoting bacteria. J Genet Eng Biotechnol 13:51–58
Ahluwalia SS, Goyal D (2007) Microbial and plant derived biomass for removal of heavy metals from wastewater. Bioresour Technol 98:2243–2257
Almås AR, Mulder J, Bakken LR (2005) Trace metal exposure of soil bacteria depends on their position in the soil matrix. Environ Sci Technol 39:5927–5932
Avudainayagam S, Megharaj M, Owens G, Kookana R, Chittleborough D, Naidu R (2003) Chemistry of chromium in soils with emphasis on tannery waste sites. In: Ware GW (ed) Reviews of environmental contamination and toxicology, vol 178. Springer, New York, pp 53–91
Bhargava A, Carmona FF, Bhargava M, Srivastava S (2012) Approaches for enhanced phytoextraction of heavy metals. J Environ Manag 105:103–120
Burmølle M, Hansen LH, Sørensen SJ (2007) Establishment and early succession of a multispecies biofilm composed of soil bacteria. Microb Ecol 54:352–362
Cappuccino JG, Sherman N (2007) Microbiology: a laboratory manual. Pearson Education, London, UK
Chen Z, Huang Z, Cheng Y, Pan D, Pan X, Yu M, Pan Z, Lin Z, Guan X, Wu Z (2012) Cr (VI) uptake mechanism of Bacillus cereus. Chemosphere 87:211–216
Christensen GD, Simpson WA, Younger J, Baddour L, Barrett F, Melton D, Beachey E (1985) Adherence of coagulase-negative staphylococci to plastic tissue culture plates: a quantitative model for the adherence of staphylococci to medical devices. J Clin Microbiol 22:996–1006
Desjardin V, Bayard R, Lejeune P, Gourdon R (2003) Utilisation of supernatants of pure cultures of Streptomyces thermocarboxydus NH50 to reduce chromium toxicity and mobility in contaminated soils. Water Air Soil Pollut 3:153–160
Dhal B, Thatoi H, Das N, Pandey B (2013) Chemical and microbial remediation of hexavalent chromium from contaminated soil and mining/metallurgical solid waste: a review. J Hazard Mater 250:272–291
Dixit R, Malaviya D, Pandiyan K, Singh U, Sahu A, Shukla R, Singh B, Rai J, Sharma P, Lade H (2015) Bioremediation of heavy metals from soil and aquatic environment: an overview of principles and criteria of fundamental processes. Sustainability 7:2189–2212
Elahi A, Rehman A (2019) Comparative behavior of two gram positive Cr6+ resistant bacterial strains Bacillus aerius S1 and Brevibacterium iodinum S2 under hexavalent chromium stress. Biotechnol Rep 21:e00307
Foster RC (1981) Polysaccharides in Soil Fabrics. Science 214:665–667
Harrison JJ, Ceri H, Turner RJ (2007) Multimetal resistance and tolerance in microbial biofilms. Nat Rev Microbiol 5:928–938
Hoh D, Watson S, Kan E (2016) Algal biofilm reactors for integrated wastewater treatment and biofuel production: a review. Chem Eng J 287:466–473
Ilias M, Rafiqullah IM, Debnath BC, Mannan KSB, Mozammel Hoq M (2011) Isolation and characterization of chromium(VI)-reducing bacteria from tannery effluents. Indian J Microbiol 51:76–81
Lewicki S, Zdanowski R, Krzyzowska M, Lewicka A, Debski B, Niemcewicz M, Goniewicz M (2014) The role of Chromium III in the organism and its possible use in diabetes and obesity treatment. Ann Agric Environ Med 21:331–335
Liu L, Liu X, Wang D, Lin H, Huang L (2020) Removal and reduction of Cr(VI) in simulated wastewater using magnetic biochar prepared by co-pyrolysis of nano-zero-valent iron and sewage sludge. J Clean Prod 257:120562
Lünsdorf H, Erb RW, Abraham WR, Timmis KN (2000) ‘Clay hutches’: a novel interaction between bacteria and clay minerals. Environ Microbiol 2:161–168
Mishra S, Bharagava RN (2016) Toxic and genotoxic effects of hexavalent chromium in environment and its bioremediation strategies. J Environ Sci Health C 34:1–32
Mitra A, Mukhopadhyay S (2016) Biofilm mediated decontamination of pollutants from the environment. AIMS Bioeng 3:44–59
Mohsin H, Asif A, Rehman Y (2019) Anoxic growth optimization for metal respiration and photobiological hydrogen production by arsenic-resistant Rhodopseudomonas and Rhodobacter species. J Basic Microbiol 59:1208–1216
Nancharaiah Y, Dodge C, Venugopalan V, Narasimhan S, Francis A (2010) Immobilization of Cr (VI) and its reduction to Cr (III) phosphate by granular biofilms comprising a mixture of microbes. Appl Environ Microbiol 76:2433–2438
Oliveira H (2012) Chromium as an environmental pollutant: insights on induced plant toxicity. Aust J Bot 2012:1–8
Pan X, Liu Z, Chen Z, Cheng Y, Pan D, Shao J, Lin Z, Guan X (2014) Investigation of Cr (VI) reduction and Cr (III) immobilization mechanism by planktonic cells and biofilms of Bacillus subtilis ATCC-6633. Water Res 55:21–29
Priester JH, Olson SG, Webb SM, Neu MP, Hersman LE, Holden PA (2006) Enhanced exopolymer production and chromium stabilization in Pseudomonas putida unsaturated biofilms. Appl Environ Microbiol 72:1988–1996
Qamar N, Rehman Y, Hasnain S (2017) Arsenic-resistant and plant growth-promoting Firmicutes and γ-Proteobacteria species from industrially polluted irrigation water and corresponding cropland. J Appl Microbiol 123:748–758
Quintelas C, Pereira R, Kaplan E, Tavares T (2013) Removal of Ni (II) from aqueous solutions by an Arthrobacter viscosus biofilm supported on zeolite: from laboratory to pilot scale. Bioresour Technol 142:368–374
Qurashi AW, Sabri AN (2012) Bacterial exopolysaccharide and biofilm formation stimulate chickpea growth and soil aggregation under salt stress. Braz J Microbiol 43:1183–1191
Randrianarivelo R, Sarter S, Odoux E, Brat P, Lebrun M, Romestand B, Menut C, Andrianoelisoa HS, Raherimandimby M, Danthu P (2009) Composition and antimicrobial activity of essential oils of Cinnamosma fragrans. Food Chem 114:680–684
Saba AR, Li Y, Rehman Y, Ahmed M, Meyer RL, Sabri AN (2018) Prospective role of indigenous Exiguobacterium profundum PT2 in arsenic biotransformation and biosorption by planktonic cultures and biofilms. J Appl Microbiol 124:431–443
Samuel J, Paul ML, Ravishankar H, Mathur A, Saha DP, Natarajan C, Mukherjee A (2013) The differential stress response of adapted chromite mine isolates Bacillus subtilis and Escherichia coli and its impact on bioremediation potential. Biodegradation 24:829–842
Stepanović S, Djukić V, Djordjević V, Djukić S (2003) Influence of the incubation atmosphere on the production of biofilm by staphylococci. Clin Microbiol Infect 9:955–958
SU C-Q, L-Q LI, Zhi-hui YZ-H, Chai L-ZY, Liao Q, Shi Y, Li J-W (2019) Cr(VI) reduction in chromium-contaminated soil by indigenous microorganisms under aerobic condition. Trans Nonferrous Metals Soc China 29:1304–1311
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tripathi DK, Singh VP, Kumar D, Chauhan DK (2012) Impact of exogenous silicon addition on chromium uptake, growth, mineral elements, oxidative stress, antioxidant capacity, and leaf and root structures in rice seedlings exposed to hexavalent chromium. Acta Physiol Plant 34:279–289
Upadhyay N, Vishwakarma K, Singh J, Mishra M, Kumar V, Rani R, Mishra RK, Chauhan DK, Tripathi DK, Sharma S (2017) Tolerance and reduction of chromium (VI) by Bacillus sp. MNU16 isolated from contaminated coal mining soil. Front Plant Sci 8:778
Wadood HZ, Sabri AN (2013) Screening, characterization and biofilm formation of nickel resistant bacteria isolated from indigenous environment. Pol J Microbiol 62:411–418
Wilson K (2001) Preparation of genomic DNA from bacteria. In: Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K (eds) Current protocols in molecular biology, vol 56. John Wiley & Sons, New York
Yassien M, Khardori N (2001) Interaction between biofilms formed by Staphylococcus epidermidis and quinolones. Diagn Microbiol Infect Dis 40:79–89
Zhu Y, Yan J, Xia L, Zhang X, Luo L (2019) Mechanisms of Cr(VI) reduction by Bacillus sp. CRB-1, a novel Cr(VI)- reducing bacterium isolated from tannery activated sludge. Ecotoxicol Environ Saf 186:109792
Code availability
Not applicable.
Funding
This research project was funded by the Higher Education Commission of Pakistan under Start-up Research Grant for PhDs (SRGP) project (21-1455/SRGP/R&D/HEC).
Author information
Authors and Affiliations
Contributions
AL, HM and MJ performed the experimental part of the manuscript including isolation, selection and screening of the Cr-resistant bacteria including the performance of Cr reduction experiments. HZW and YR supervised the overall research project. YR and HM provided the facilities in their research labs to carry out this research project.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Responsible Editor: Haroun Chenchouni
Rights and permissions
About this article
Cite this article
Wadood, H.Z., Latif, A., Mukhtar, H. et al. Planktonic cells of Staphylococcus and Bacillus species capable of faster chromium reduction in short incubation times as compared to their biofilms. Arab J Geosci 14, 1797 (2021). https://doi.org/10.1007/s12517-021-08226-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12517-021-08226-5