Abstract
Phosphatidylinositol-specific phospholipase C (PI-PLC) catalyzes the hydrolysis of phosphatidylinositol-4,5-bisphosphate into diacylglycerol and inositol 1,4,5-trisphosphate. It can play an essential role in plant stress response and signaling. However, the functions of PLCs remain unclear in wooden plants. This study carried out the bioinformatics analysis of the PLC gene family in poplar. The expression pattern analysis suggested the transient up-regulation of PsnPLC in salt stress conditions. The transgenic tobacco plants overexpressing PsnPLC were generated by the Agrobacterium-mediated leaf disc method. The transgenic lines showed a significant increase in plant height, SOD and POD activity, and proline content under salt stress. By contrast, transgenic lines showed a decreased malondialdehyde content under salt stress. Comparative transcriptome analysis indicated that the overexpression of PsnPLC mainly affects membrane-associated GO terms and signaling pathways, and the differential expression of salt-responsive genes may contribute to the enhanced salt tolerance. Overall, the results indicated that PsnPLC plays an essential role in modulating salt tolerance in transgenic tobacco.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Salt stress is one of the primary factors which can severely affect plant growth and development. Plants have evolved multiple cellular events to mitigate the damage caused by high salinity, including sensing salt stimuli and converting them into internal signals (Chinnusamy et al. 2006). As a crucial part of the cell membrane, phospholipids play a critical role in the signal transduction system. They can act as the substrates for phospholipases to convert the external signals into cellular responses (Kadamur and Ross. 2013; Musille et al. 2013; Pokotylo et al. 2014). As an important class of phospholipases, phospholipase C (PLC) can be divided into two families, phosphatidylinositol-specific phospholipase C (PI-PLC, hereafter referred to as PLC) and non-specific PLC (NPC), depending on their substrates. PI-PLC preferentially catalyzes the hydrolysis of phosphoinositide (4,5) bisphosphate (PIP2) (Dickson et al. 2014), releasing the secondary messenger inositol 1,4,5-trisphosphate (IP3) and diacylglycerol (DAG) (Meijer and Munnik 2003). In animal cells, IP3 triggers the release of Ca2+, and DAG is involved in the PKC signal pathways together with Ca2+ (Kadamur and Ross 2013). However, plant cells not only lack the genes encoding IP3 receptors and PKC family protein but possess a different conserved structure compared to animal PLCs (Wang, 2004). In this context, the phosphorylation of IP3 and DAG into IP6 and PA play a vital role in mediating the PLC pathways in plants (Stevenson-Paulik and Phillippy 2010).
Many plant PLCs have been demonstrated to regulate plant growth and development, abiotic stress, and biotic stress response (Wang 2004; Xue et al. 2007; Singh et al. 2015; Abd-El-Haliem and Joosten 2017). However, the function of PLC in wooden plants remains unclear. In this study, the full-length cDNA of PLC was cloned from Populus simonii x P. nigra. Bioinformatic analysis was carried out for poplar PLCs, including phylogenetic analysis, motif prediction, chromosome distribution, gene duplication, and synteny analysis. The expression pattern of PsnPLC in salt stress condition was further analyzed by qPCR. The transgenic tobacco overexpressing PsnPLC was then generated via Agrobacterium tumefaciens-mediated transformation. Phenotypic, physiological traits and transcriptomic data analysis were carried out to understand the functions of PsnPLC in salt stress response. This study will provide experimental evidence for the application of PsnPLC to improve salt tolerance in molecular breeding and outline the relation between salt response and phospholipid signals.
Materials and Methods
Plant Materials
Populus simonii x P. nigra were grown in pots containing water at 26 °C/22 °C (day/night) under a 16 h/8 h light/dark cycle. The tobacco (Nicotiana tabacum L.cv. Petit Havana SR-1) seeds were sterilized in 70% ethanol for 30 s, washed twice with sterile water, then treated with 10% NaClO for 10 min, and washed 4 times with sterile water. The sterilized seeds were transferred to MS medium and cultured at 25–26 °C for 30 days under 14 h light/10 h dark conditions, and the seedlings were used for transformation.
Cloning and Sequence Analysis of PsnPLC
The total RNA was extracted from 2-month-old leaves of Populus simonii x P. nigra. Then, PsnPLC was amplified by RT-PCR. The primers were designed according to the transcript sequence of Potri.010g188800.1. The PLC sequences of Populus trichocarpa, Arabidopsis thaliana, Oryza sativa L., and other plant species were obtained from NCBI. The phylogenetic tree was constructed using the Neighbor-Joining method in MEGA 7.0. The reliability was tested using bootstrapping with 1000 replicates (Tamura et al. 2011). The phylogenetic tree was displayed using iTOL (http://itol.embl.de/) (Letunic and Bork 2021). Motif analysis was performed with MEME (Bailey et al. 2009). Chromosomal distribution and local collinearity of PLC genes in poplar were visualized using TBtools (Chen et al. 2020).
Expression Patterns of PsnPLC
2-month-old seedlings with new roots and leaves were subject to the treatment of 200 mM NaCl. Leaves were harvested at 0, 6, 12, 24, 48, and 72 h with three biological replicates and quickly frozen in liquid nitrogen for RNA extraction. 1 µg of total RNA was reverse-transcribed into cDNA in a 20 µl volume using a PrimeScript™ RT reagent kit with gDNA Eraser (Takara). cDNAs were used as templates for real-time PCR, and the primers were shown in Table S1. Poplar ubiquitin (UBQ) gene (Potri.001G418500) was used as the internal control. Real-time PCR was performed using the SYBR Premix Ex Taq™ (Takara) on a CFX connect real-time system (Bio-RAD). The amplification was performed under the following conditions: 94 °C for 30 s, 40 cycles of 94 °C for 12 s, 54 °C for 30 s, 72 °C for 30 s, and 1 s at 81 °C for plate reading. Three replicates were carried out for each assay, and the relative expression levels were calculated using the 2−ΔΔCt method.
Tobacco Transformation
The PsnPLC sequence was inserted into the Sma I and Sac I sites of the pROKII vector. The recombinant plasmid was transformed into tobacco using Agrobacterium tumefaciens-mediated transformation (Gallois and Marinho 1995). The regenerated shoots were transferred to the rooting medium containing 100 mg/L kanamycin, 600 mg/L ceftriaxone, and 0.02 mg/L 1-naphthaleneacetic acid (NAA) for root induction. The putative transgenic lines were subject to PCR assays. The T2 homozygous transgenic lines were selected for salt stress tolerance analysis.
Evaluation of Salt Stress Tolerance
The seeds of T2 homozygous transgenic lines and WT were sown on the MS medium containing 100 mg/L kanamycin and MS medium without antibiotics, respectively. The 2-leaf stage seedlings of transgenic lines and WT were then transferred to the MS medium containing 0 mM and 150 mM NaCl. After 30-day treatment, the leaves of three T2 homozygous transgenic lines and WT were harvested for physiological traits analysis. POD activities were measured by the Peroxidase assay kit using the guaiacol method (Jiancheng Bioengineering Institute, Nanjing, China). Proline content was estimated by the Pro content assay kit using the acidic-ninhydrin reaction method (Cominbio, Suzhou, China). MDA contents were measured by the “lipid peroxidation MDA assay kit” using the thiobarbituric acid (TBA) method (Beyotime Biotechnology). SOD activities were measured by the “total superoxide dismutase assay kit with NBT” (Beyotime Biotechnology). For all the above assays, six replicates were performed for each sample.
NBT Staining
NBT staining was applied to detect the accumulated ROS. The leaves of T2 homozygous transgenic lines and WT were infiltrated in 0.1 mM NBT solution (Coolaber) in the dark for 10 h. The staining leaves were then transferred to 95% ethanol and incubated in 70 °C water bath.
RNA-Seq Analysis of Transgenic Plants
Wild type and transgenic tobacco lines overexpressing PsnPLC were treated with 150 mM NaCl for 30 days. Total RNA was extracted from each sample using TRIzol Reagent (Invitrogen). The RNA-seq experiments were carried out by Genewiz (Suzhou, China) using an Illumina HiSeq 2500 sequencing platform. Gene expressions were quantified using the FPKM (Fragments Per Kilo bases per Million reads). The differentially expressed genes were defined as those with the false discovery rate < 0.05 and |log2FC|> 1. GO terms and KEGG pathways with adjusted P value < 0.05 were considered significantly enriched.
Statistical Analysis
The experimental data were analyzed using SPSS22.0. The results were presented as mean ± standard deviation. One-way analysis of variance (ANOVA) based on least significant difference was used for statistical analysis. The threshold of significance was P < 0.05 (*).
Results
Cloning and Sequence Analysis of PsnPLC
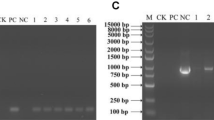
The cDNA of PsnPLC was cloned by RT-PCR based on its partial sequence identified in the previous study. PsnPLC shares high sequence similarity (98.8%) with PtPLC6 (Potri.010G188800.1) (Fig. S1), and contains a 1764 bp open reading frame encoding 587 amino acid residues. A phylogenetic analysis based on the amino acid sequences of PsnPLC and PLCs from poplar, Arabidopsis, rice, and other plant species revealed a close evolutionary relationship between PsnPLC and PtPLC6, PtPLC4 (Potri.008G068400.1), and PePLC (XP_011035945.1) from Populus euphratica (Fig. 1). In addition, 30 PLC sequences were submitted to MEME to predict the conserved motifs. The motif prediction result indicated that most of the input sequences consist of the conserved domains of PLC (EF-hand, X domain, Y domain, and C2 domain), which were indicated by colored bars in Fig. 2. In addition, AtPLC8 and AtPLC9 lack the EF-hand domain (Fig. S2) and are, therefore, more distantly related to other PLC family members phylogenetically.
Phylogenetic tree between poplar, rice, Arabidopsis and other plant species. Phylogenetic analysis was carried out based on the amino acid sequences of PLCs. The red block represents PsnPLC and its closest relatives in poplar. The green block indicated by blue lines represents the PLCs lacking the EF-hand domain. Accession numbers of genes are showed in Table S2
MEME motif prediction result for PLC family proteins. The colored rectangles on the sequence labeling line indicate the conserved domain of PLC, that is, EF-hand domain (purple), X domain (light blue), Y domain (red), and C2 domain (green) from left to right. Accession numbers of genes are showed in Table S2
Poplar genome annotation was downloaded from NCBI to study the evolutionary relationship of the PLC family genes in poplar, and the chromosomal distribution of poplar PLCs was visualized using TBtools. As shown in (Fig. 3a), 8 PLC family members were assigned to chromosome 1, 8, 9, and 10, respectively. Among them, four tandem duplication events were observed, half of which occurred on chromosome 1. In addition, local collinearity analysis of PLCs was also carried out by TBtools using BLASTP and MCScanX methods. Segmental duplication events were found between two PLC gene pairs, Potri.001g252500/Potri.008g068400 and Potri.008g068300/Potri.010g188900 (Fig. 3b). These results suggested that tandem duplication events occur mainly on chromosome 1. In addition, the tandem duplication region on chromosome 8 is also critical for segmental duplication during the evolution of PLC family genes.
Gene duplication analysis for poplar PLCs. a Chromosomal distribution and tandem duplication analysis for poplar PLCs. Only the chromosomes harboring PLC (chromosome 1, 8, 9, and 10) are displayed. The genes involving tandem duplication events are marked. b Collinearity analysis for poplar PLCs. The pale yellow rectangles indicate chromosome 1–19 in poplar. The intensity of the genes on the corresponding chromosome is reflected by the heatmap and line graph. Red curves represent the segmental duplication events between poplar PLCs, and blue curves represent the segmental duplication events among gene pairs encoding the partial PLC-X domains such as Potri.018G109200/Potri.006G186300 and Potri.004G172000/Potri.009G131600
PsnPLC Expression in Response to Salt Stress
To investigate the expression patterns of PsnPLC under abiotic stress conditions, the leaves of Populus simonii x P. nigra were harvested at different time points after stress treatment and used for RNA extraction. Real-time PCR results showed a significant transient up-regulation of PsnPLC at 6 h (P < 0.01). In contrast to the transient up-regulation at 6 h, a down-regulation of PsnPLC was observed under salt condition, with elevated expression after 48 h (Fig. 4). These results suggested that PsnPLC expression is sensitive to salt stress.
Expression patterns of PsnPLC Expression analysis of PsnPLC in response to salt stress (200 mM NaCl) in poplar leaves. The expression level of PsnPLC was calculated relative to its expression level at 0 h. The results were displayed as the mean ± SD. Double and single asterisk indicate significant difference at P < 0.01 and P < 0.05 (t-test)
Overexpression of PsnPLC Improves Salt Tolerance in Transgenic Tobacco
To further determine the function of PsnPLC, the recombinant vector containing 35S::PsnPLC (pROKII-PsnPLC) was constructed and introduced into tobacco via Agrobacterium tumefaciens-mediated transformation. The transgenic lines were screened by kanamycin-resistance selection and further validated by PCR. After disinfection, the seeds of three T2 homozygous transgenic lines (TL 1–3) and wild type were sown on the medium containing 100 mg/L kanamycin and MS medium without antibiotics, respectively. The 2-leaf stage seedlings of the same size were transferred to the subculture medium containing 0 mM and 150 mM NaCl. After culturing for 1 month, the growth of transgenic lines and WT were approximately the same under normal condition; however, after 30-day 150 mM NaCl treatment, the transgenic tobacco appeared a better morphological condition than WT (Fig. 5a). Meanwhile, the growth condition may also be reflected by the plant height measurement, and no significant difference was shown between transgenic lines and WT under normal condition. However, under 150 mM NaCl, the plant height of T-2 (P < 0.01) and T-3 (P < 0.05) were significantly higher than WT (Fig. 5b). These results suggested that transgenic tobacco maintained better growth under salt condition.
Phenotype and physiological traits analysis. a Phenotype pictures of transgenic and WT tobacco plants under normal conditions and 150 mM NaCl. b–e Plant height, POD and SOD activity, and proline content between WT and transgenic lines under normal condition and 150 mM NaCl. WT: wild type. T-1, T-2, T-3: transgenic tobacco lines. The data were expressed as the mean ± SD calculated from six replicate experiments. Double and single asterisks indicate significant difference at P < 0.01 and P < 0.05, respectively. f. NBT staining assay of transgenic tobacco plants and WT under 150 mM NaCl. g MDA content between WT and transgenic lines under normal condition and 150 mM NaCl
SOD and POD can effectively scavenge the accumulated ROS, mitigating oxidative damage caused by salt stress. The POD activity assays showed no significant difference between transgenic lines and WT under normal condition, but a significant increase in POD activity was observed in transgenic lines under 150 mM NaCl (Fig. 5c). Compared with POD, PsnPLC might mediate SOD signaling in a different manner. As shown in the result of SOD assay, higher SOD activity was observed in T-1 and T-2 than WT under normal condition. Meanwhile, the SOD activity of transgenic lines was still higher under 150 mM NaCl (Fig. 5d). These results suggested that overexpression of PsnPLC could maintain the cellular ROS homeostasis by regulating POD and SOD activities.
Small molecules, such as proline, can function as osmo-regulator to main the stability of the cell membrane and keep the plant from dehydration in the presence of osmotic imbalance. Measurements of proline content showed no difference between transgenic lines and WT under normal conditions. However, the proline content was much higher in transgenic lines, particularly in T-2 and T-3 under 150 mM NaCl (Fig. 5e). These results implied that PsnPLC could maintain a stable osmotic pressure by regulating the cellular proline content.
The oxidative damage degree of cell membrane can be estimated by NBT staining assay and the concentration of MDA. The NBT staining assay displayed that more ROS accumulated in WT than TL1-3 under NaCl treatment (Fig. 5f). Under normal condition, there was no significant difference in MDA level between WT and TL1-3, whereas under salt stress, the MDA concentration in transgenic lines was significantly lower than the WT (Fig. 5g). Overall, these data suggested transgenic lines suffered less oxidative damage under salt stress.
Differentially Expressed Genes in Transgenic Tobacco
To investigate the interactions between PsnPLC and other signaling pathways, we carried out a comparative transcriptome analysis of leaves in transgenic lines against WT. A total of 1676 DEGs were identified, including 1147 up-regulated genes (URGs) and 529 down-regulated genes (DRGs) (Fig. S3). The functions of DEGs could be predicted by GO and KEGG annotations. The significantly enriched GO terms and KEGG pathways are shown in Fig. 6. The most notable GO terms included integral component of membrane (cellular component, CC), metal ion binding (molecular function, MF), carbohydrate metabolic process (biological process, BP), and metabolic process (BP) (Fig. 6a); KEGG analysis showed that the DEGs mainly clustered in 6 pathways, including plant hormone signal transduction (34), toll-like receptor signaling pathway (20), starch and sucrose metabolism (17), glutathione metabolism (14), alanine, aspartate and glutamate metabolism (10) and butanoate metabolism (6) (Fig. 6b). These results showed that PsnPLC might directly or indirectly affect the above signaling pathways.
The enriched Gene ontology terms and KEGG pathways of transgenic tobacco compared to WT. a Classification of the enriched GO terms. Different colors are used to distinguish the GO terms (molecular function, cellular component, and biological process). b Top clustered KEGG pathways with the most DEGs. The total number of DEGs is indicated by the stacked red (up-regulated) and blue (down-regulated) bars. The rich factor, which represents the percentage of DEGs among the genes in the corresponding pathways, is indicated by green lines
To investigate the salinity tolerance mechanism of the transgenic lines overexpressing PsnPLC, we studied the expression profiles of stress-responsive genes based on transcriptome analysis. The expression level of potassium transporter (KUP and KEA) in transgenic lines was significantly lower than the WT, while the transcript abundance of DERB, NRT1/PTR, ERDL6, LEA14 and SPSA was significantly up-regulated (Fig. 7a, Table 1). qRT-PCR analysis verified the down-regulation of KUP2, KUP6, KEA4 (Fig. 7b) and the up-regulation of ERDL6 and SPSA (Fig. 7c) in transgenic lines. No significant difference was observed for NPF8.1 and LEA14 (Fig S4).
The expression of stress-responsive genes. a The heatmap illustrating the expression of stress-responsive genes of RNA-seq. Log10FPKM is used to quantify the expression level of relative genes. WT: wild type. T-1, T-2, T-3: transgenic tobacco lines. b qPCR assays illustrating the expression of potassium regulation genes. c qPCR assays illustrating the expression of ERDL6 and SPSA
Discussion
As a component of the plant signal transduction network, phospholipid signal requires the involvement of various phospholipid hydrolases such as PLC and PLD. PLC, a member of the phospholipase family, has been shown to play an important role in plant growth and development and response to environmental stress. Previous studies have shown that PLC2 is involved in auxin biosynthesis and signaling, thereby modulating the fertility of Arabidopsis (Li et al. 2015). Single and double mutant of Arabidopsis atplc3/atplc9 exhibited a heat-sensitive phenotype, while the double mutant was more severely affected by heat shock. Heterologously expressed AtPLC9 has considerable potential to enhance heat tolerance in cereal crops (Hong et al. 2008; Zheng et al. 2012; Liu et al. 2020). AtPLC4 positively regulates the increase of intracellular Ca2+ in response to salt signals, thereby inducing plant responses to salt stress (Xia et al. 2017). ZmPLC1 may enhance plant drought tolerance by restoring the composition of unsaturated membrane lipid levels under drought stress (Zhai et al. 2013). Overexpression of TaPI-PLC1-2B significantly enhanced salt and drought tolerance in transgenic Arabidopsis (Wang et al. 2020). GmPI-PLC7 may enhance drought and salt tolerance in soybean through the ABA signaling pathway and the SOS-related calcium-signaling pathway (Chen et al. 2021). The involvement of rice PLC in salt stress has been well investigated: OsPLC3 plays a positive role in rice salt stress response by regulating the aboveground/underground Na+ content and the sodium/potassium ion ratio. Similarly, OsPLC1-mediated calcium signaling is involved in Na+ accumulation (Li et al. 2017). OsPLC4 modulates the osmotic stress response in rice seedlings through lipid- and Ca2+-mediated signaling (Deng et al. 2019).
To identify key genes for salt resistance in poplar, we have obtained the cDNA fragments in response to high salt stress using the cDNA-AFLP method. One of the cDNA fragments contains an AP2/ERF domain (GenBank accession number: GW672629) and shares high homology with the poplar ERF76 gene (Wang et al. 2011). The interaction between the PsnPLC and ERF was further analyzed; meanwhile, this pair of genes shares similar expression patterns under salt treatment (Wang et al. 2018). Although ERF has been demonstrated to improve salinity tolerance (Yao et al. 2016a, b), the functions of PLC in wooden plant remains unclear. To evaluate the role of poplar PLC, we cloned the full-length cDNA of PsnPLC from P. simonii x P. nigra. The phylogenetic analysis indicated that PsnPLC shares high sequence similarity with poplar PLC (Fig. 1). The conserved motifs can be found in most of the submitted PLCs, except for AtPLC8 and AtPLC9 which do not contain the EF-hand domain in the N-terminus (Fig. 2). The chromosome distribution results show that PLC genes are distributed on chromosome 1, 8, 9, and 10 in poplar (Fig. 3a). To delineate the evolutionary relationship of poplar PLC, we identified four tandem duplication events (Fig. 3a) and two segmental duplication events (Fig. 3b). Notably, both segmental duplication events associate with Chromosome 8, implying that Chromosome 8 may have played a central role in the evolution of poplar PLC genes. Interestingly, the segmental duplication event also occurred in Potri.018g109200, a gene that does not belong to the PLC family but encodes a PLC-X catalytic domain. Its encoding protein contains a transcriptional activation domain (AP2/B3 domain) and a PLC catalytic domain (PLC-X domain), and the structure of the “recombinant” protein may explain the previously identified protein interactions between the AP2/ERF transcription factor and PsnPLC (Wang et al. 2018). Based on these findings, we may further investigate the phospholipase C modulation network and the relations between PsnPLC and transcription factors.
The expression of plant PLCs can be induced by multiple environmental factors, including dehydration and salt stress, cold stress, and ABA (Hirayama et al. 1995; Kim et al. 2004; Zhai et al. 2005; Zhang et al. 2014; Deng et al. 2019; Iqbal et al. 2020). In this study, transient accumulation of PsnPLC transcripts in poplar leaves under salt stress was observed, with peak expression appearing at 6 h (Fig. 4). Under NaCl treatment, PtoPLC1 from Populus tomentosa (Zhang et al. 2015), and the homologous genes from soybean (Glycine max), GmPLC3, GmPLC6, GmPLC7, and GmPLC10 (Wang et al. 2015), showed similar expression patterns to PsnPLC. Therefore, it can be predicted that PsnPLC may be transiently induced by salt stress. It should be noted that the expression level of PsnPLC was down-regulated at 12–24 h, but up-regulated again at 48 h. The fluctuating expression may be driven by the crosslinks of multiple phospholipid signals, maintaining the balance of phospholipase activity. The recovery at 48 h and other time points might be due to the reactivated PsnPLC-mediated response signals induced by the continuous cellular damage.
To further understand the role of PsnPLC in salt stress tolerance, we analyzed the physiological traits of the transgenic tobacco lines overexpressing PsnPLC. Under normal conditions, there were no phenotypic differences between the transgenic lines and WT (Fig. 5a). However, under salt stress, we observed better growth and increased plant height in the transgenic tobacco (Fig. 5b). The SOD and POD activities and proline content of the transgenic strains were measured and compared with WT, aiming to investigate if overexpression of PsnPLC could maintain the cellular ROS homeostasis and stable osmotic pressure. SOD and POD assays suggested that PsnPLC could mitigate the adverse effect of salt stress by regulating ROS scavenging enzymes (Fig. 5c, d). The proline content of the transgenic lines was also increased significantly under salt stress (Fig. 5e), suggesting that PsnPLC could maintain a balanced osmotic pressure by mediating the synthesis of osmo-regulators (Natarajan et al. 2012). Histochemical staining assay of ROS accumulation with NBT revealed that transgenic lines suffered less cellular damage under NaCl treatment (Fig. 5f). MDA content, an indicator of cell membrane oxidative damage (Tsikas 2017), was significantly lower in transgenic tobacco than in WT (Fig. 5g). Overall, the physiological trait analysis suggested the salt-dependent tolerance of PsnPLC except for SOD. The SOD activity in T-1 and T-2 was higher than WT under both control and salt conditions (Fig. 5d), demonstrating that the SOD activity can be influenced by PsnPLC. However, its mechanism differs from other salt response signals.
To gain insight into the salt response mechanism of PsnPLC, we further analyzed the RNA-seq data using transgenic tobacco against WT. The DEGs were mapped to the corresponding GO terms and KEGG pathways. Gene ontology analysis showed that the principle enriched GO terms include “integral component of membrane” (CC), “metal ion binding” (MF), “carbohydrate metabolic process” (BP), and “metabolic process” (BP) (Fig. 6a), which are all closely associated with the functions of the membrane. PsnPLC could hydrolyze its substrates on the cell membrane, leading to a dramatic change in the lipid composition and fluidity. As a result, overexpression of PsnPLC might directly interfere with the above membrane-related GO functions. The top enriched KEGG pathways include “plant hormone signal transduction”, “toll-like receptor signaling pathway”, “starch and sucrose metabolism”, “glutathione metabolism”, “alanine, aspartate, and glutamate metabolism” and “butanoate metabolism”, and the rich factor of these pathways varies from 0.047 to 0.1 (Fig. 6b). Previous research has demonstrated that PLC is closely related to phytohormones in the cell guarding signaling, reproductive development, and stomatal closure (Mills et al. 2004; Singh et al. 2015; Li et al. 2015). Promoter analysis also showed that the promoter region of Potri.010G188800.1 (PsnPLC homologous gene) contains the cis-elements involved in ABA, auxin, and MeJA responsiveness (Table 2). PsnPLC-mediated control of endogenous hormone content may lead to the environmental stress adaption of transgenic lines. Belonging to the TLR signaling pathway, most RLKs receive signals through their extracellular ligand recognition domain and transduce the message to effector molecules (Vaid et al. 2013). PsnPLC may alleviate salt stress in collaboration with RLKs, which can act as a switch to activate downstream signals. The DEGs related to “starch and sucrose metabolism” as well as “carbohydrate metabolic process” could directly regulate sugar biosynthesis, transportation, and degradation (Wang et al. 2019). Thus, the soluble sugars derived from this pathway can function as osmolytes, energy sources, and signal molecules in combating abiotic stress (Boriboonkaset et al. 2013; Li et al. 2014). Overexpression of PsnPLC not only elevated the activity of SOD and POD but regulated the metabolism of GSTs. 13 out of 14 DEGs clustered into the “glutathione metabolism” pathway were up-regulated. These enzymes play an important role in maintaining the redox hemostasis in transgenic lines (Gill and Tuteja 2010; Csiszár et al. 2014). “Alanine, aspartate and glutamate metabolism” and “butanoate metabolism”, the KEGG pathways with the highest rich factor, are closely related to the metabolism of gamma-aminobutyrate (GABA). Endogenous GABA enhances salt tolerance by maintaining photosynthetic capacity and modulating absorption of Na+ (Wu et al. 2020; Khanna et al. 2021).
Previous study has shown that overexpression of GmPI-PLC7 may activate the expression of drought- or salt-responsive genes to meditate stress responses (Chen et al. 2021). In our study, overexpression of PsnPLC might also alter the expression of salt-responsive genes (Fig. 7 and Table. 1). Among them, KUP and KEA may positively regulate osmotic resistances by influencing the K+-mediated ABA response, stomatal behavior, and osmotic homeostasis (Osakabe et al. 2013; Li et al.2018; Ou et al. 2018; Cai et al. 2021). Since ABA-related activities may be directly influenced by the overexpression of PLC, the involvement of the K+ transporter may not be required in the ABA signal pathways, leading to the decreased expression of KUP and KEA in transgenic tobacco. Overexpression of NRT1/PTR can promote nitrate absorption, which alleviates the chloride toxicity and maintains the photosynthesis under salt stress (Teakle and Tyerman 2010; Zhao et al. 2021). LEA14 has been demonstrated to enhance Arabidopsis salt stress tolerance by maintaining the stability and cellular protein and membrane (Jia et al. 2014), and ERDL6 and SPSA-catalyzed subcellular sugars can act as osmoprotectants (Poschet et al. 2011; Volkert et al. 2014). As a phosphatidylinositol hydrolase, PsnPLC cannot directly regulate the transcript abundance of stress-responsive genes, whereas it may affect the expression of related genes indirectly. PLC can catalyze the hydrolysis of substrates into secondary messenger IP3 and DAG. IP3 and DAG play a key role in the signal cascade, activating a series of cellular activities. These cellular activities may regulate the expression of stress-responsive genes. The differential expression of these genes may explain the putative functions of PsnPLC in mediating salt and phospholipid signals; however, the underlying molecular mechanism needs further exploration.
In sum, a phosphatidylinositol-specific phospholipase C gene from Populus simonii x P. nigra, PsnPLC, which could be transiently induced by salt stress, might significantly enhance salt tolerance in transgenic tobacco by scavenging ROS and maintaining cellular osmotic pressure in a salt-dependent manner. RNA-seq data analysis suggested that the functions of PsnPLC might be closely related to the activities of the cell membrane and the expression level of salt-responsive genes involving potassium transportation and sugar metabolism. Therefore, PsnPLC could play an essential role in mediating salt and phospholipid signals, indicating its potential application in molecular breeding.
Data Availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
Abbreviations
- PI-PLC:
-
Phosphatidylinositol-specific phospholipase C
- PIP2 :
-
Phosphoinositide (4,5) bisphosphate
- DGK:
-
Diacylglycerol kinase
- PA:
-
Phosphatidic acid
- TBA:
-
Thiobarbituric acid
- POD:
-
Peroxidase
- SOD:
-
Superoxide dismutase
- NBT:
-
Nitro blue tetrazolium
- TL:
-
Transgenic line
- WT:
-
Wild type
- DEG:
-
Differentially expressed genes
- URG:
-
Up-regulated genes
- DRG:
-
Down-regulated genes
- MDA:
-
Malondialdehyde
- GABA:
-
Gamma-aminobutyrate
References
Abd-El-Haliem AM, Joosten MH (2017) Plant phosphatidylinositol-specific phospholipase C at the center of plant innate immunity. J Integr Plant Biol 59:164–179
Bailey TL, Mikael B, Buske FA et al (2009) MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res 37:W202–W208
Boriboonkaset T, Theerawitaya C, Yamada N et al (2013) Regulation of some carbohydrate metabolism-related genes, starch and soluble sugar contents, photosynthetic activities and yield attributes of two contrasting rice genotypes subjected to salt stress. Protoplasma 250:1157–1167
Cai K, Zeng F, Wang J et al (2021) Identification and characterization of HAK/KUP/KT potassium transporter gene family in barley and their expression under abiotic stress. BMC Genomics 22:317
Chen C, Chen H, Zhang Y et al (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13:1194–1202
Chen ZF, Ru JN, Sun GZ et al (2021) Genomic-wide analysis of the PLC family and detection of GmPI-PLC7 responses to drought and salt stresses in soybean. Front Plant Sci 12:631470. https://doi.org/10.3389/fpls.2021.631470
Chinnusamy V, Zhu J, Zhu JK (2006) Salt stress signaling and mechanisms of plant salt tolerance. Genet Eng 27:141
Csiszár J, Horváth E, Váry Z et al (2014) Glutathione transferase supergene family in tomato: Salt stress-regulated expression of representative genes from distinct GST classes in plants primed with salicylic acid. Plant Physiol Bioch 78:15–26
Deng XJ, Yuan S, Cao HS et al (2019) Phosphatidylinositol-hydrolyzing phospholipase C4 modulates rice response to salt and drought. Plant Cell Environ 42(2):536–548
Dickson EJ, Jensen JB, Hille B (2014) Golgi and plasma membrane pools of PI(4)P contribute to plasma membrane PI(4,5)P2 and maintenance of KCNQ2/3 ion channel current. P Natl Acad Sci USA 111:E2281–E2290
Gallois P, Marinho P (1995) Leaf Disk transformation using Agrobacterium tumefaciens-expression of heterologous genes in tobacco. Plant Gene Transfer and Expression Protocols, Springer, New York, pp 39–48
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48:909–930
Hirayama T, Ohto C, Mizoguchi T et al (1995) A gene encoding a phosphatidylinositol-specific phospholipase C is induced by dehydration and salt stress in Arabidopsis thaliana. Proc Natl Acad Sci USA 92:3903–3907
Hong TL, Gao F, Guo LL et al (2008) The calmodulin-binding protein kinase 3 is part of heat-shock signal transduction in Arabidopsis thaliana. Plant J 55(5):760–773
Iqbal S, Ali U, Fadlalla T et al (2020) Genome wide characterization of phospholipase A & C families and pattern of lysolipids and diacylglycerol changes under abiotic stresses in Brassica napus L. Plant Physiol Biochem 147:101–112
Jia F, Qi S, Li H et al (2014) Overexpression of late embryogenesis abundant 14 enhances Arabidopsis salt stress tolerance. Biochem Biophys Res Commun 454(4):505–511
Kadamur G, Ross EM (2013) Mammalian phospholipase C. Annu Rev Physiol 75:127–154
Khanna RR, Jahan B, Iqbal N et al (2021) GABA reverses salt-inhibited photosynthetic and growth responses through its influence on NO-mediated nitrogen-sulfur assimilation and antioxidant system in wheat. J Biotechnol 325:73–82
Kim YJ, Kim JE, Lee JH et al (2004) The Vr-PLC3 gene encodes a putative plasma membrane-localized phosphoinositide-specific phospholipase C whose expression is induced by abiotic stress in mung bean (Vigna radiata L). Febs Lett 556(1–3):127–136
Letunic I, Bork P (2021) Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res 23:127–128
Li C, Jia H, Chai Y et al (2014) Abscisic acid perception and signaling transduction in strawberry: A model for non-climacteric fruit ripening. Plant Signal Behav 6(12):1950–1953
Li L, He Y, Wang Y et al (2015) Arabidopsis PLC2 is involved in auxin-modulated reproductive development. Plant J 84(3):504–515
Li L, Wang F, Yan P et al (2017) A phosphoinositide-specific phospholipase C pathway elicits stress-induced Ca2+ signals and confers salt tolerance to rice. New Phytol 214(3):1172–1187
Li W, Xu G, Alli A et al (2018) Plant HAK/KUP/KT K+ transporters: function and regulation. Semin Cell Dev Biol 74:133–141
Liu Y, Liu X, Wang X et al (2020) Heterologous expression of heat stress-responsive AtPLC9 confers heat tolerance in transgenic rice. BMC Plant Biol 20:514. https://doi.org/10.1186/s12870-020-02709-5
Meijer H, Munnik T (2003) Phospholipid-based signaling in plants. Annu Rev Plant Biol 54(1):265–306
Mills LN, Hunt L, Leckie CP et al (2004) (2004) The effects of manipulating phospholipase C on guard cell ABA-signalling. J Exp Bot 55(395):199–204
Musille PM, Kohn JA, Ortlund EA (2013) Phospholipid-driven gene regulation. FEBS Lett 587:1238–1246
Natarajan SK, Zhu W, Liang X et al (2012) Proline dehydrogenase is essential for proline protection against hydrogen peroxide induced cell death. Free Radic Biol Med 53:1181–1191
Osakabe Y, Arinaga N, Umezawa T et al (2013) Osmotic stress responses and plant growth controlled by potassium transporters in Arabidopsis. Plant Cell 25:609–624
Ou W, Mao X, Huang C et al (2018) Genome-wide identification and expression analysis of the KUP family under abiotic stress in cassava (Manihot esculenta Crantz). Front Physiol 9:17
Pokotylo I, Kolesnikov Y, Kravets V et al (2014) Plant phosphoinositide-dependent phospholipases C: variations around a canonical theme. Biochimie 96:144–157
Poschet G, Hannich B, Raab S et al (2011) A novel Arabidopsis vacuolar glucose exporter is involved in cellular sugar homeostasis and affects the composition of seed storage compounds. Plant Physiol 157(4):1664–1676
Singh A, Bhatnagar N, Pandey A et al (2015) Plant phospholipase C family: regulation and functional role in lipid signaling. Cell Calcium 58(2):139–146
Stevenson-Paulik J, Phillippy BQ (2010) Inositol polyphosphates and kinases. In Lipid Signaling in Plants, Book Series: Plant Cell Monographs; Minnik T, Ed. Springer-Verlag, Berlin: Berlin, Germany 16:161–174. https://doi.org/10.1007/978-3-642-03873-0_11
Tamura K, Peterson D, Peterson N et al (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Teakle NL, Tyerman SD (2010) Mechanisms of Cl− transport contributing to salt tolerance. Plant Cell Environ 33(4):566–589
Tsikas D (2017) Assessment of lipid peroxidation by measuring malondialdehyde (MDA) and relatives in biological samples: analytical and biological challenges. Anal Biochem 524:13–30
Vaid N, Macovei A, Tuteja N (2013) Knights in action: lectin receptor-like kinases in plant development and stress responses. Mol Plant 6:1405–1418
Volkert K, Stefan D, Lars MV et al (2014) Loss of the two major leaf isoforms of sucrose-phosphate synthase in Arabidopsis thaliana limits sucrose synthesis and nocturnal starch degradation but does not alter carbon partitioning during photosynthesis. J Exp Bot 65(18):5217–5522
Wang X (2004) Lipid signaling. Curr Opin Plant Biol 7(3):329–336
Wang L, Zhou BR, Wu LL et al (2011) Differentially expressed genes in Populus simonii x Populus nigra in response to NaCl stress using cDNA-AFLP. Plant Sci 180(6):796–801
Wang F, Deng Y, Zhou Y et al (2015) Genome-wide analysis and expression profiling of the phospholipase C gene family in soybean (Glycine max). PLoS ONE 10(9):e0138467
Wang L, Sun Y, Xia XL et al (2018) Screening of proteins interacting with ERF transcriptional factor from Populus simonii x P.nigra by yeast two-hybrid method. Biotechnol Biotec Eq. 32:543–549
Wang M, Dai W, Du J et al (2019) ERF109 of trifoliate orange (Poncirus trifoliata (L.) Raf.) contributes to cold tolerance by directly regulating expression of Prx1 involved in antioxidative process. Plant Biotechnol J 17:1316–1332
Wang X, Liu Y, Li Z et al (2020) Genome-wide identification and expression profile analysis of the phospholipase C gene family in wheat (Triticum aestivum L). Plants 9(7):885
Wu X, Jia Q, Ji S et al (2020) Gamma-aminobutyric acid (GABA) alleviates salt damage in tomato by modulating Na+ uptake, the GAD gene, amino acid synthesis and reactive oxygen species metabolism. BMC Plant Biol 20(1):465
Xia K, Wang B, Zhang J et al (2017) Arabidopsis phosphoinositide-specific phospholipase C4 negatively regulates seedling salt tolerance. Plant Cell Environ 40:1317–1331
Xue H, Chen X, Li G (2007) Involvement of phospholipid signaling in plant growth and hormone effects. Curr Opin Plant Biol 10:483–489
Yao W, Wang L, Zhou B et al (2016a) Over-expression of poplar transcription factor ERF76 gene confers salt tolerance in transgenic tobacco. J Plant Physiol 198:23–31
Yao W, Wang S, Zhou B et al (2016b) Transgenic poplar overexpressing the endogenous transcription factor ERF76 gene improves salinity tolerance. Tree Physiol 7:896–908
Zhai S, Sui Z, Yang A et al (2005) Characterization of a novel phosphoinositide-specific phospholipase C from Zea mays and its expression in Escherichia coli. Biotechnol Lett 27(11):799–804
Zhai S, Gao Q, Liu X et al (2013) (2013) Overexpression of a Zea mays, phospholipase C1 gene enhances drought tolerance in tobacco in part by maintaining stability in the membrane lipid composition. Plant Cell Tissue Organ Cult 115(2):253–262
Zhang K, Jin CC, Wu LZ et al (2014) Expression analysis of a stress-related phosphoinositide-specific phospholipase C gene in wheat (Triticum aestivum L.). PLoS ONE. https://doi.org/10.1371/journal.pone.0105061
Zhang J, Zhang Z, Zhu D et al (2015) Expression and initial characterization of a Phosphoinositide-specific phospholipase C from Populus tomentosa. J Plant Biochem Biotechnol 24:338–346
Zhao L, Chen P, Liu P et al (2021) Genetic effects and expression patterns of the nitrate transporter (NRT) gene family in Populus tomentosa. Front Plant Sci 12:661635
Zheng SZ, Liu YL, Li B et al (2012) Phosphoinositide-specific phospholipase C9 is involved in the thermotolerance of Arabidopsis. Plant J 69(4):689–700
Funding
This work was supported by the Heilongjiang Provincial Natural Science Foundation of China (LH2019C059) and Excellent Youth Foundation of Heilongjiang Academy of Sciences (CXJQ2021GJS02).
Author information
Authors and Affiliations
Contributions
Yao Sun and Lei Wang conceived and designed the research. All the authors performed the experiments. Yao Sun, Xin Sun and Lei Wang analyzed the data. Yao Sun prepared figures and tables. Yao Sun and Lei Wang authored or reviewed drafts of the paper. All the authors read and approved the final draft.
Corresponding author
Ethics declarations
Conflict of Interest
The authors have no conflicts of interest to declare that are relevant to the content of this article.
Ethics Approval
Not applicable.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sun, Y., Li, Y., Sun, X. et al. Overexpression of a Phosphatidylinositol-Specific Phospholipase C Gene from Populus simonii × P. nigra Improves Salt Tolerance in Transgenic Tobacco. J. Plant Biol. 65, 365–376 (2022). https://doi.org/10.1007/s12374-022-09359-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12374-022-09359-0