Abstract
Meiosis is a distinctive type of cell division that reorganizes genetic material between generations. The initial stages of meiosis consist of several crucial steps which include double strand break, homologous chromosome pairing, break repair and crossover. Crossover frequency varies depending on the position on the chromosome, higher at euchromatin region and rare at heterochromatin, centromeres, telomeres and ribosomal DNA. Crossover positioning is dependent on various factors, especially epigenetic modifications. DNA methylation, histone post-translational modifications, histone variants and non-coding RNAs are most probably playing an important role in positioning of crossovers on a chromosomal level as well as hotspot level. DNA methylation negatively regulates crossover frequency and its effect is visible in centromeres, pericentromeres and heterochromatin regions. Pericentromeric chromatin and heterochromatin mark studies have been a centre of attraction in meiosis. Crossover hotspots are associated with euchromatin regions having specific chromatin modifications such as H3K4me3, H2A.Z. and H3 acetylation. This review will provide the current understanding of the epigenetic role in plants during meiotic recombination, chromosome synapsis, double strand break and hotspots with special attention to euchromatin and heterochromatin marks. Further, the role of epigenetic modifications in regulating meiosis and crossover in other organisms is also discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Meiotic division is essential in all the sexually propagating eukaryotes. Meiosis maintains stable chromosome numbers across generations, as it ensures that every gamete receives only half the number of chromosomes of a parent. Meiotic cells undergo single round of DNA replication and condensation before two rounds of chromosomal segregation (Yelina et al. 2015a, b; Dawe 1998; Mercier et al. 2015). The major differences between mitosis and meiosis are the ploidy of progeny cells and additional steps undergoing in Prophase I (Mercier et al. 2015). In Arabidopsis thaliana, meiosis starts with double strand breaks (DSBs) at the leptotene stage and recombination process commences. Pairing between homologous chromosomes occurs at zygotene via the polymerization of the synaptonemal complex (SC) and recombination proceeds. Synapsis completes by pachytene, SC disassembles in the diplotene stage and the chiasmata structure displays the crossover sites between homologous chromosomes. Chromosomal condensation occurs during diakinesis and bivalents are visible (Sims et al. 2021; Ross et al. 1996; Mercier et al. 2015). Segregation of homologs takes place during meiosis I and segregation of sister chromatids occur during meiosis II to generate 4 gametes. DSB repair happens through crossovers (COs) and non-crossovers (NCOs) (shown in Fig. 1). It’s common in all the plants to have more DSBs than the resulting crossovers. Most of these breaks end up as Non-crossovers (NCOs) and sister chromatids are used as a template in this repair pathway. Not just crossing overs, non-crossovers (gene conversions) are also important for maintaining diversity (Mercier et al. 2015).
Steps occurring in the Arabidopsis recombination process is shown in brief. A Homologous chromosomes are shown in two contrasting colours (blue and pink). Estimated frequency of each crossover pathway is represented in brackets. The prominent proteins involved in each step are mentioned on the left side. (MRNS complex- MRE11-RAD50-NBS1/XRS2-SAE2/COM1, MUS81-MMS AND UV SENSITIVE81, FANCM-FANCONO ANEMIA COMPLEMENTATION GROUP M, DHJ-Double Holliday Junction). B Some of the important meiotic stages are represented at the lower panel of the figure
Meiotic crossovers use homologous chromosomes as a template and shuffle the genetic information. It is a necessary for each pair of homologs to undergo a minimum of one crossover for proper segregation i.e., obligatory crossing over (Darlington et al. 1932). Recombination has been one of the major factors in driving speciation and variation. However, its effect on the genome varies as the recombination sites are unevenly distributed. Certain regions of the genome are more prone to crossover events than others and these regions are known as recombination hotspots (Marand et al 2019). In most flowering plants, crossing overs are abundant in sub-telomeric and chromosomal arms whereas crossovers are scarce in ribosomal DNA, pericentromeric and centromeric regions (i.e., U-shaped trend). The exceptions are plants like Nelumbo nucifera and Camellia sinensis which have higher recombination rates in the middle of the chromosome (Brazier and Glemin et al. 2022). The longer the chromosomal length, the higher the chance of distal bias (Haenel et al. 2018).
Epigenetic modifications are known to have been involved in meiotic regulation and crossover (Yelina et al. 2015a, b). It refers to a layer of information that exists beyond what is encoded in the DNA sequence. Epigenetic modifications can be divided into four categories: DNA methylation, histone post-translational modifications, chromatin remodelling and non-coding RNAs (ncRNAs) (Portela and Esteller 2010; Dziegielewski and Ziolkowski 2021). All these modifications are very dynamic in nature which means they could be stage-specific, tissue-specific, loci-specific. These dynamic structural changes can affect processes like transcription, DNA repair, chromosome segregation and transposon movements. Chromatin states in a chromosome are differentiated into euchromatin (enriched with active genes), facultative heterochromatin (enriched with repressed genes) and constitutive heterochromatin (enriched with transposons and repeats). Epigenetic modifications regulate directly (in cis) or indirectly through regulating meiotic gene expression (in trans) (Mirouze et al. 2012). Most of the studies on plant meiosis are available in male meiosis. It is technically difficult to isolate intact embryo sac mother cell with analysable chromosomes (Koul et al. 2023). The recombination steps take most of the time of meiotic phase indicating that it’s a time taking and highly regulated process (Prusicki et al. 2019; Osman et al. 2021).
All the epigenetic classes interplay with each other and reinforce through many positive and negative feedback mechanisms (Portela and Esteller 2010). The epigenetic factors initiating and determining the crossover sites in flowering plants still remain a puzzle.
Meiosis in plants: an overview
In plants, meiotic studies with recombination patterns are available in Arabidopsis, barley, wheat, maize, rice, potato and tomato (Anderson et al. 2014; other references are mentioned in Table 1). Meiotic events and genetic regulation of meiotic division in plants are more or less the same as observed in Arabidopsis. The meiotic processes of Arabidopsis are briefly discussed here. Arabidopsis thaliana stands out as a model plant for studying chromatin anomalies and meiotic recombination because of the already available resources with chromosome spreading protocols (Ross et.al. 1996), crossover numbers (Francis et al. 2007) and lesser chromosomal number (2n = 10). The process of meiotic recombination involves the exchange of genetic material between homologous chromosomes that occurs in several steps, which can be summarized as follows: (1) Chromosome condensation (2) DNA double-strand break formation (3) Resection (4) Strand invasion and double holiday junction formation (5) Resolution (Fig. 1).
During early prophase I of meiosis, the replicated homologous chromosomes condense. Double strand breaks (DSBs) are the starting point for meiotic recombination. They are catalysed by SPORULATION 11 (SPO11) (Grelon et al. 2001). DSBs are then processed by MRE11-RAD50-NBS1/XRS2-SAE2/COM1 (MRNS) complex and EXO I generates 3’ overhang single-stranded DNA (ssDNA). The 3’ overhangs are bounded by heterotrimeric replication protein A (RPA) complex and loading of recombinases like RADIATION SENSITIVE51 (RAD51) and DISRUPTED MEIOTIC cDNA1 (DMC1) takes place during leptotene stage (Li et al. 2004; Couteau et al. 1999). In pachytene, the synaptonemal complex (SC) is completely formed between aligned maternal and paternal chromosomes forming synapsis between the bivalents. SC is the zipper like tripartite proteinaceous structure formed between chromatin loops made up of central elements and lateral axes. One of the single-stranded DNA ends invades the homologous chromosome and pairs up with a complementary strand on the homologous chromosome. DNA synthesis follows, extending the invading strand and displacing the original strand of the homologous chromosome. This leads to the creation of the Holliday junction, a structure where the two DNA duplexes are intertwined. In diplotene, the Holliday junction is resolved in two ways: either by cutting the junction horizontally to produce two non-crossover products or by cutting it vertically to produce two crossover products. As the homologous chromosomes separate, X-shaped structures called chiasmata are formed. Chromosomes further separate and condense by diakinesis stage. The resulting chromosomes are different from the original chromosomes as they contain a mixture of genetic information from each parent (for a detailed recombination process, see the review; Mercier et al. 2015).
Homologous recombination (HR) happens in both somatic cells and meiotic cells. In somatic cells, it is predominantly repaired through Non-Homologous End Joining pathway (NHEJ) and rarely through HR. Sister chromatids act as a template in somatic cells, while homologous chromosome act as a template in meiotic cells. It has been known that DMC1 favours inter-homolog recombination (IH) and crossing over in meiotic cells while RAD51 favours sister chromatid recombination in both mitotic and meiotic cells. RAD51 and DMC1 function independently and are spatially separated during meiosis (Kurzbauer et al. 2012). Their foci do not colocalize and the DMC1 triggers down-regulation of RAD51 exchange activity through an unknown signalling pathway however the presence of RAD51 is essential during chromosomal strand exchange (Da Ines et al. 2022).
In Arabidopsis, on an average of 250 DSBs, 12 recombination events takes place per meiosis. Out of these 12, 10 are class I COs and the other two are class II COs (Mercier et al. 2015). Homolog-dependent repair of a DSB can occur through Class I, Class II and NCO pathways. Class I is interference dependent and relies on a group of proteins called ZMM and two other conserved proteins called MutL HOMOLOGS (MLH1 and MLH3) (Jean et al. 1999; Mercier et al. 2015). That is why class I is also called as ZMM pathway. Class II is interference independent (little or no interference) and MMS AND UV SENSITIVE81 (MUS81) contribute to it (Higgins et al. 2008; Mercier et al. 2015). Sturtevant (1915) described interference as follows: “The occurrence of one crossing-over in a given chromosome pair prevents another nearby” . Class I contributes to above 70–80 per cent of all the crossovers and class II plays just a minor role. Although Class II is interference independent, class I and class II COs can detect and interfere with each other (Anderson et al. 2014). FANCONO ANEMIA COMPLEMENTATION GROUP M (FANCM) and RECQ4 helicases direct recombination intermediates from class II CO pathway to synthesis-dependent-strand-annealing NCO pathway (Crismani et al. 2012; Lorenz et al. 2012; Mercier et al. 2015). Apart from these three, there is a non-interference pathway described in Solanum lycopersicum called FANCD2 (FANCONO ANEMIA D2) (Kurzbauer et al. 2018). There is limited information available about this pathway. It is unclear why many pathways have to be evolved and interplay with each other. According to population genetics perspective, the substrate of each pathway could have the deciding factor in evolving different pathways (Ziolkowski et al. 2023). Ribosomal DNA (rDNA) arrays regulate their crossover numbers through NHEJ pathway (Sims et al. 2019).
In Arabidopsis, crossovers can be estimated through classical genetic analysis using segregation of markers in the progeny, chiasmata counting, visual assay of fluorescent pollen tetrads (FTLs) and immunostaining of MLH1 foci (Berchowitz and Copenhaver 2008; De Muyt et al. 2009; Mercier et al. 2015; Francis et al. 2007). FTLs are as best as Chlamydomonas reinhardtii zoospores and Saccharomyces tetrads systems to determine the crossover rates from a region of interests (Francis et al. 2007). Deep tetrad software ease the process through it’s high-throughput analysis of tetrads and yield very quick results (Lim et al. 2020). Flow cytometry based protocols can achieve high-throughput analysis but they miss out on calculating double crossovers and gene conversion measurements (as pollen tetrads -qrt1 mutants are not used in the protocol) (Yelina et al. 2013). Fluorescent antibodies usage against MLH1 and MUS81 foci gives the number of class I COs and class II COs, respectively (Mercier et al. 2015).
Factors affecting crossover positioning
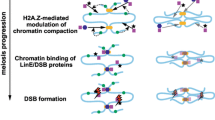
Epigenetic modifications, such as histone modifications, DNA methylation and DNA remodelers can influence crossover positioning by altering the accessibility of DNA to the recombination machinery. The process of homologous chromosome pairing during meiosis can influence the positioning of crossovers. Telomeres attach to the nuclear membrane in the early prophase I stage and form bouquet-shaped chromatin organisation. In grasses, there is asynchronous timing of axis maturation and SC formation leading to early maturation and localisation of ZMM proteins in distal regions. It is believed early interaction can lead to early designation of crossover sites (Higgins et al. 2012, 2022). Pro-Crossover ZMM class of proteins like HUMAN ENHANCER OF INVASION 10 (HEI10) diffuse and coarsen across the chromosome. Only these foci turn into crossovers (Morgan et al. 2021). Crossing over in plants is tightly regulated by the ZMM pathway and synaptonemal complex (Durand et al. 2022). The DNA sequences called DNA motifs or sequence motifs, are found in the vicinity of crossover hotspots. They vary from plant to plant. Surprisingly, transposons also overlap with crossover locations. (shown in Table 1). Structural variants like inversions, transpositions and indels can also show local CO suppression and determine CO positioning (Rowan et al. 2019).
Epigenetic factors involved in the regulation of meiosis
Histone modifications in meiotic regulation
In eukaryotes, DNA wraps around the positively charged proteins called histones and this DNA–protein complex is called chromatin. About 146 bp of DNA make a tight 1.65 negative super helical turns around an octamer of core histone proteins. The histone octamer consists of a central heterotetramer of histones H3 and H4, flanked by two heterodimers of histones H2A and H2B (i.e., two copies of H3, H4, H2A and H2B each) (Fischle et al. 2003).
A variety of posttranslational modifications have been discovered on tails such as acetylation, methylation, ubiquitination, phosphorylation, sumoylation and ribosylation (some of the marks are shown in Fig. 2). The histone modifications are seen in both amino-terminal (N-terminal) and carboxy-terminal tails. Modifications are abundant in the H3 histone N-terminal tail. The modifications acting as hallmark of actively transcribed regions are called active chromatin marks and the marks abundant in non-transcribed regions are called repressive chromatin marks. These modifications are written by writer proteins, read by reader proteins and removed by eraser proteins. Some of the proteins can do multiple roles. All these effector proteins help to precisely balance the location of histone marks. For example, SET DOMAIN GROUP8 (SDG8) in Arabidopsis encodes for a histone methyl transferase that have both a writer domain called SET domain and a reader domain called CW domain. JUMONJI 30 (JMJ30), a histone demethylase acts as an eraser for the same histone mark H3K36me3 (Cheng et al. 2020).
Acetylation of histone proteins involves the addition of an acetyl group to lysine residues, which neutralizes the positive charge of the lysine (+ 1 to 0) and weakens the interaction between the histone and the negatively charged DNA. This leads to a more relaxed chromatin structure and increased accessibility of the DNA to transcription factors and other regulatory proteins. Methylation of histone proteins occurs on lysine or arginine residues and can have different effects depending on the specific site and degree of methylation. Phosphorylation of histone proteins occurs on serine, threonine, and tyrosine residues and can affect chromatin structure and gene expression in a variety of ways (Peterson et al. 2004). The role of histone marks can change from one organism to the another. H3K4me2 is an active chromatin mark in animals and Arabidopsis but it acts as a repressive mark in rice (Liu et al. 2019; Cheng et al. 2020; Tock et al. 2021). Histone euchromatin (active) and heterochromatin (repressive) marks are represented in Table 2.
Cohesin subunit AtSYN4 recruits Histone H2A monoubiquitination (H2Aub1) mark specifically into the genomic loci showing that there could be prominent interplay between cohesin subunits and histone modification enzymes (Zhang et al. 2023). Transcription factors also recruit the chromatin modifying enzymes leading to the accumulation of specific histone marks (Zhang et al. 2015) (Both cohesin and transcription factor recruitment are shown in Fig. 3).
Diagram showing the DSBs generally localise with euchromatin features like low nucleosomal occupancy, DNA methylation (Cm) and sequence motifs. Most of the DSBs in Arabidopsis overlap with promoter regions. Cohesins and Transcriptions factors (TF) recruit various kinds of histone modification enzymes. Histone methyl transferases-(HMTs) responsible for euchromatin is shown in green colour and for heterochromatin is shown in brown colour
In Arabidopsis, recombination hotspots tend to be located in gene-rich regions of the genome and these are highly correlated with active chromatin modifications, including H2A.Z, histone H3 Lys4 trimethylation (H3K4me3) (Choi et al. 2013), low nucleosomal density, low DNA methylation (Choi et al. 2018; Underwood et al. 2018).
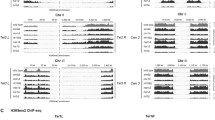
In Arabidopsis, Histone H3 Lys 4 trimethylation (H3K4me3), an active euchromatin mark, is highly enriched near SPO11-1 oligos but not highly correlated with double strand breaks at fine level. In the DSB site, exonucleases release SPO11-oligonucleotide complexes. These oligos provide a high-resolution profile of meiotic DSB patterns genome-wide (Choi et al. 2018). SET DOMAIN GROUP 2 (SDG2) encodes for the H3K4me3 mark and the loss of function mutant sdg2 has not shown any defect in meiosis. Its role is limited to post meiotic microspore development, especially chromatin decondensation in the pollen vegetative nucleus (Pinon et al. 2017). In Saccharomyces cerevisiae (budding yeast), H3K4me3 is enriched in the vicinity of DSB sites and the mutant showed a drastic reduction in DSBs (Borde et al. 2009). Spp1 subunit of COMPASS complex recognises H3K4me3 rich hotspots and Mer2 axis and recruits the DSB site to the chromosomal axis for the action of Spo11 (Acquaviva et al. 2013; Lambing et al. 2020; Yelina et al. 2015a, b). Surprisingly in Schizosaccharomyces pombe (fission yeast) H3K9ac is more significantly enriched than H3K4me3 in hotspots (Yamada et al. 2013). PR DOMAIN CONTAINING 9 (PRDM9) dependent H3K4me3 majorly determines recombination hotspots in humans and mice (Baudat et al. 2009). PRDM9 (meiosis-specific histone lysine methyltransferase with SET domain) binds to DNA at the hotspot and trimethylates H3K4 as well as H3K36, which causes nucleosomes to move apart creating favourable site for SPO11 to produce double strand break (Parvanov et al. 2010; Diagourage et al. 2018; Bhattacharya et al. 2019; Powers et al. 2016) (Fig. 4). DSBs are formed in the prdm9 mutant, but are formed at different locations. It means PRDM9 does not initiate the meiotic recombination but is required only for determination of hotspot position and DSBs (Brick et al. 2012). CXXC finger-domain protein, a component of H3K4 methyltransferase in mice is involved in regulating meiotic gene expression (Ki et al. 2022). However, plants lack PRDM9 homologs and CXXC kind of finger-domain proteins. Almost in all the organisms H3K4me3 is seen in the vicinity of recombination hotspots but the mark is probably controlled through different molecular mechanisms.
In Arabidopsis, Histone 3 lysine 9 dimethylation (H3K9me2), a repressive mark, is abundant in transposable elements and repetitive sequences. H3K9me2 mark is controlled by methyl transferases called SUPPRESSOR OF VARIEGATION HOMOLOG 4/KRYPTONITE (KYP/SUVH4/5/6) and histone demethylases called IBM1 (INCREASE IN BONSAI METHYLATION). Mutants of KYP/SUVH4/5/6 have shown an increase in recombination without any chromosomal defects (Underwood et al. 2018) whereas demethylase mutant ibm1 has displayed incomplete synapsis, chromosome entanglement, reduction of recombination during meiosis and fertility. The number of chiasmata per cell in ibm mutants is less compared to the WT (He et al. 2022). These defects are due to IBM1-mediated alterations in gene expression and interaction with cohesin cofactor PDS5s in downstream (Cheng et al. 2022; He et al. 2022). It means that timely maintenance of H3K9 demethylation is more important than H3K9 methylation in plant meiosis. Whereas in mammalian meiosis H3K9 methylation is important for prophase progression. Loss of heterochromatic H3K9 methylation leads to improper synapsis, leading to abnormal chromosome segregation and infertility in mice (Tachibana et al. 2007; Takada et al. 2021). Similar to Arabidopsis, drosophila and fission yeast H3K9me2 mutants are reported to have more crossovers (Ellermeier et al. 2010; Peng and Karpen 2009). In fission yeast, H3K9 methylation marks attract mitotic cohesion complex and limit DSBs around centromeres. It’s unknown in plants how H3K9me2 is limiting DSBs (Nambiar and Smith 2018). In higher organisms, pericentromeres are evolved to have 100 times lower crossovers than chromosomal arms. Recombination in pericentromeres is known to have mis-segregation and aneuploidy (for example: Down syndrome) (Nambiar and Smith 2016). Whereas in Arabidopsis, cytologically there are no chromosomal defects found despite increasing pericentromeric crossovers (Underwood et al. 2018).
REC8, a sister chromatid cohesin is associated with suppression of DSBs and meiotic crossovers (Lambing et al. 2020). In fission yeast, REC8 is enriched in the centromeres. In H3K9me2 mutants of fission yeast, REC8 loading is greatly reduced and leading to chromosomal segregation defects (Ellermeier et al. 2010). Despite the fact that the REC8 and H3K9me2 peaks overlap in Arabidopsis, REC8 loading remained normal in H3K9me2 and non-CG DNA methylation mutants. It’s also possible that other heterochromatin marks are driving the REC8 localisation in Arabidopsis (Lambing et al. 2020). In yeast, H3 histone modifications are involved in meiotic recombination and highly acetylated in recombination hotspots (Borde et al. 2009; Merker et al. 2008; Yamada et al. 2004). However, in Arabidopsis, H3 Histone hyperacetylation with overexpression line gcn5-related histone N-acetyltransferase (mcc1), displayed difficulty in segregation, altered chiasmata distribution, leading to the abortion of half of the gametes (Perrella et al. 2010).
In maize, DSBs are distributed uniformly in a chromosome but crossovers follow the usual U-shaped trend (higher CO rate in chromosomal ends and lower CO rate in centromeres). Not even 20 percent of hotspots are associated with H3K4me3 mark. As the majority of Maize genome is filled with transposons, DSBs are abundant in transposons but only the genic DSBs turned into crossovers (He et al. 2017). So, with all these unique features maize has to be considered as an outlier. In allopolyploids like wheat, spatiotemporal asymmetry of meiotic programs is possibly favouring distal pattern of crossover localisation (Higgins et al. 2022; Osman et al. 2021). Along with the distinct pattern, H3K4me3, H3K9me3, H3K27me3 and H3K27me2 marks are also preferentially enriched in distal parts of wheat chromosomes (Osman et al. 2021; Tock et al. 2021). Facultative heterochromatin mark H3K27me3 is positively correlated along with (ASYNAPTIC 1) ASY1 and DMC1 peaks with crossover regions. This polycomb repressive mark is enriched in the crossover active distal regions of chromosomes and is highly associated with disease responsive genes (nucleotide binding leucine rich-repeat proteins) (Tock et al. 2021; Liu et al. 2021). H3K27me2 is also enriched in the distal area of chromosomes, but it’s associated with crossover suppression. Both the marks are mutually exclusive and the H3K27me2 role in supressing crossovers is also observed in Arabidopsis and maize (Liu et al. 2021).
In rice and soybean, Histone 3 lysine 36 trimethylation (H3K36me3), an active transcription mark is negatively correlated with crossovers (Ma et al. 2023; Marand et al. 2017). Even in yeast, H3K36me3 negatively correlates with DSBs and meiotic crossovers (Hansen et al. 2011; Merker et al. 2008). Further, H3K36me3 mark decreases resection and promoting non-homologous end joining pathway (NHEJ) (Pai et al. 2014). The immunolocalization analysis of various H3 methylation and acetylation modifications throughout all phases of meiosis in Aegilops sp., Secale cereale and Arabidopsis thaliana showed that their distribution and dynamics of histone marks are species specific and stage specific. This implies that there is an evolutionary divergence in histone language (Oiliver et al. 2013).
DNA methylation in meiotic regulation
DNA methylation mainly occurs at the fifth position of cytosine, resulting in 5′-methylcytosine (5-mC). In general, hypermethylation and hypomethylation are found in heterochromatin and euchromatin regions, respectively (Zhang et al. 2018). In plants, DNA methylation occurs at cytosine residue in all three sequence contexts such as symmetric CG, CHG and asymmetric CHH (where H is any nucleotide except G) while CG methylation is predominant in animals. As of now, only 5 methyltransferases are known in Arabidopsis. They are METHYLTRANSFERASE 1 (MET1), CHROMOMETHYLASE 2 (CMT2), CHROMOMETHYLASE 3 (CMT3), DOMAINS REARRANGED METHYLTRANSFERASE 1 (DRM1) and DOMAINS REARRANGED METHYLTRANSFERASE 2 (DRM2). DRM2 is responsible for DNA methylation in all three sequence contexts (CG, CHG and CHH), whereas MET1 and CMT3 maintain CG and CHG methylation, respectively. CMT2 and DRM2 are jointly responsible for maintaining CHH methylation in long heterochromatic transposable elements (TEs) and short euchromatic TEs, respectively (Fang et al. 2021).
In Arabidopsis, the centromeres are highly heterochromatic regions that are epigenetically silenced by H3K9me2 and DNA methylation in CG and non-CG sequence contexts. MET1 maintains CG methylation all over the chromosome (Mirouze et al. 2012). Loss of CG methylation in met1 is shown to have more SPO-11-1-oligos, reduction of nucleosome density, slight gain of H3K4me3 and transcriptional reactivation of repetitive sequences in centromeres. The met1 mutant got triggered into crossover remodelling and resulted in having complex phenotype of increased chromosomal arms, centromere recombination and decreased pericentromeric recombination (Choi et al. 2018; Yelina et al. 2012). In total, DSB foci and crossover number remain the same but the crossovers are redistributed to chromosomal arms from pericentromeres. The redistribution is limited to only interfering crossovers (class I COs) (Choi et al. 2018; Yelina et al. 2015a, b). In somatic cells of met1 mutant, there is a redistribution of epigenetic marks among H3K9me2, DNA methylation and H3K27me3 in PcG target genes (Deleris et al. 2012). Ectopic H3K9me2 marks noticed in the proximal region of chromosomes could be due to the redistribution of chromatin marks. Along with the alteration in chromatin modifications, crossover interference mechanisms might be responsible for this complex phenotype (Yelina et al. 2015a, b).
The chromatin remodeler DDM1 (DECREASED DNA METHYLATION 1) gene is responsible for DNA methylation and heterochromatin maintenance. In the ddm1 mutant, CG methylation is reduced throughout the genome especially in heterochromatin regions. FTL markers between euchromatin regions showed more recombination rate but the pericentric heterochromatin FTL markers showed similar recombination rate as of control (Bessudo and Levy 2012). DDM1 is also responsible for maintaining classical Mendelian trait segregation. This is possibly through excessive gene conversion happening in the heterochromatin (Ali et al. 2021). A similar study in mouse, DNA methylation mutant dnmt3, has shown increased DSBs formation and gain of H3K4me3 marks in retrotransposon regions (Zamudio et al. 2015).
The mutation in the H3K9 methyltransferase (KYP/SUVH4/5/6) or the CHG methylation maintenance gene CMT3 (CG methylation remains same), showed a significant increase SPO11-1-oligos and crossovers in pericentromeric regions. Both class I and class II crossovers are elevated in the mutant (Underwood et al. 2018). Overall, both CG and non-CG methylation are able to inhibit DSBs, but only CHG methylation has the ability to inhibit crossovers at pericentromeric region (Choi et al. 2018). In somatic cells, CHG methylation and H3K9me2 are mutually linked and dependent on each other. Proteins encoding these epigenetic modifications are able to recognise each other (Du et al. 2012). Additional binding proteins like AGENET DOMAIN CONTAINING P1 (AGDP1) also link H3K9me2 and non-CG DNA methylation (Zhang et al. 2018). A positive reinforcing loop is noticed in meiotic chromosomes also i.e., promoting each other’s presence (Underwood et al. 2018). In all the plants studied, methylation has been negatively impacting crossing over. An unusual occurrence in the form of CHH and CHG methylation peaks is observed in rice hotspot regions (Marand et al. 2019). Rate of recombination in epigenetic mutants of Arabidopsis, categorized by region of chromosome, are represented in Table 3.
Histone variants in meiotic regulation
In addition to histone covalent modifications, there is an additional tier of variation called histone variants. Histone variants are versions of proteins that differ from the canonical histones in their amino acid sequence. Dozens of histone variants like H2A, H2B and H3 families have been discovered in eukaryotes (for example, H2A.Z) (Jamge et al. 2023). There is a great correlation and interplay between Histone modifications and these variants, which means that a variant is strongly associated with a specific mark (Loppin et al. 2020). For example, H3.3 is strongly associated with H3 acetylation and H3K36 methylation, whereas H3.1 is associated with H3K27me1. Histone variants can be incorporated into chromatin in a replication-independent manner, meaning that they are added to pre-existing chromatin rather than during DNA replication. This allows cells to modify chromatin structure in response to different signals and cellular states (Jamge et al. 2023).
Histone variant H2A.Z foci colocalize with crossover regions, DMC1 and RAD51. The knockdown mutant (arp6) has shown low DSB which led to low crossover numbers (Choi et al. 2013). ACTIN RELATED PROTEIN 6 (ARP6), a subunit of SWR1 ATP dependent remodelling complex is necessary for H2A.Z deposition and regulates meiotic gene expression during megasporogenesis (Qin et al. 2014). In fission yeast, the variant promotes chromatin compaction through cohesins (Yamada et al. 2018). Although the variant promotes DSBs, it is not enriched around the hotspots (Yamada et al. 2018). Phosphorylated H2A.X signals DNA damage response and invites chromatin remodelling complexes (Rogakou et al. 1998; Kuo 2021). This mark is generally used in immunolocalization studies to determine DSBs (Higgins et al. 2012). H2A.W, a heterochromatin mark is independent of other heterochromatin features and causes heterochromatin condensation (Yelagandula et al. 2014). HTA6 and HTA7 are the two major genes responsible for H2A.W variant deposition and proteins are enriched on the pericentric heterochromatin (Yelagandula et al. 2014; Kuo 2021). The recombination rate is increased only in the heterochromatin regions of hta6-1 hta7 mutants although the mutants showed normal cytology. Double mutants of the hta7 cmt3 did not show to any significant rise in crossovers compared to the single mutant (hta7). The overall study has shown that the H2A.W variant is a repressor of meiotic recombination (Kuo 2021).
Noncoding RNAs in meiotic regulation
Noncoding RNAs are classified as small RNAs (sRNAs), medium size ncRNAs (mncRNAs) and long-size ncRNAs (lncRNAs). A large number of non-coding RNAs are upregulated in the events of meiosis. They have a prominent role in controlling meiotic gene expression, chromosome condensation and centromere organisation. Transcriptome studies in various plants have shown the significance of noncoding RNAs in meiosis (Jiang et al. 2023; Dziegielewski and Ziolkowski 2021). Small interfering RNAs (siRNAs) are important for transposon silencing and DNA methylation. Mutants of miRNA biogenesis and function, dcl, hyl1, hen1, hst and ago1, have shown higher expression of SPO11, DMC1, RAD51, MSH4, and MUS81 (Dziegielewski and Ziolkowski 2021).
Other factors affecting crossover rate
Chromatin loops
Meiotic chromosomes are organized in loops and attached to the chromosome axis which interacts with chromatin to regulate meiotic recombination. At the macro level, chromosomes appear as X-shaped structures. But the chromatin is packed into various kinds of loops to serve different functions. Larger loops form shorter axes, thus leading to fewer COs and vice-versa (Wang et al. 2019) (Fig. 5). Anchor regions of long-range cis chromatin loops are associated with H3K27me3 and maintain chromatin organisation in Arabidopsis (Sun et al. 2023). H3K27me3 deposition also influences chromatin compartmentalisation and gene co-regulation (Huang et al. 2021). There is a need to verify the role of the dynamic chromatin loops in choosing and initiating crossovers.
Abiotic stress
Meiotic crossover frequency is plastic and modulated by different factors like age, temperature, nutrient availability, developmental stage, and behaviour (Modliszewski and Copenhaver 2017; see the review). Crossover rate in Arabidopsis male meiosis follows a U-shaped trend in response to temperature range between 5 and 28 °C. CO rate is highest at 5 °C and 28 °C but the lowest at its optimal growth temperature 20 °C (Lloyd et al. 2018). The observed high meiotic crossover frequency at elevated temperature (28 °C) is derived from only the type I pathway. There is no increase in ds breaks, just the non-crossovers are converted into crossovers in this scenario (Modliszewski et al. 2018). Heat stress can also induce altered transposon activation, leading to the rearrangement of chromatin organisation (Sun et al. 2020). This can lead to chromatin decondensation and the activation of heterochromatin transcription (Tittel-Elmer et al. 2010).
Sudden exposure to temperature stress severely decreases the duration of early prophase stages, but the crossover maturation phases got prolonged. Mutants related to the recombination pathway like spo11-1, dmc1, msh4 did not show this pattern, implying that prolong maturation phase is recombination-dependent (Braet et al. 2022). Heavy stress inhibits meiotic recombination via reduced ds breaks and homolog asynapsis. In barley, the distribution of recombination events is shifted to centromere under high temperature (Philips et al. 2015). The same shifting pattern is noticed in wheat under temperature stress, but it’s not as significant as in the case of Barley (Coulton et al. 2020). In crop plants, many of the essential genes lie in recombination cold spots. Heat treatment can give different combinations of progeny due to the change in crossover distribution. Although the mechanisms are unknown, the mild heat stress can be tested in heterosis programmes.
Conclusion and future perspective
Meiotic recombination in plants is tightly regulated at different levels. It’s maintained by various genetic, epigenetic, genome structure and environmental factors. Epigenetic factors influence the recombination landscape directly as well as indirectly through meiotic gene expression (overall model is shown in Fig. 6). On a local scale, crossover positioning is majorly done by histone marks, DNA methylation and structural variants. On a broad scale, the spatiotemporal asymmetry model and diffusion mediated ZMM proteins coarsening model are perfectly explaining the distal region preference in crossovers and crossover interference. Only grasses have shown additional factors like time of meiotic events, pre-meiotic DNA replication along with usual chromatin features and pro CO genes (Higgins et al. 2022). Pro crossover enzymes like HEI10 and anti-crossover factors like RECQ4 control the crossover positioning at the genome level (Morgan et al. 2021). Positioning is initially determined by DSB sites and pro-crossover enzymes like HEI10 in a suitable chromatin environment do the next layer of selection. The accumulation of HEI10 determines the location of crossovers. The factors determining the location of HEI10 accumulation and coarsening are unknown.
Chromatin features can influence DSB proteins, recombination machinery and meiotic genes. The number of DSBs influences the total number of crossovers (Xue et al. 2018). Through this web of interactions, chromatin features control crossing over in one way or the other. Deciphering the chromatin features and mechanisms involved in crossover mechanisms will ease breeding programmes and help us to make smart crops. Increasing crossovers in pericentromeric regions has been achieved through various heterochromatin mutants in Arabidopsis. The heterochromatin marks (like H3K9me2) are also responsible for heterochromatin stability, so transferring these results into food crops has to be done in a careful manner (Peng and Karpen 2009). There is also a great variation in epigenetic control of meiosis among the model organisms. Only yeast has shown great similarity with recombination control as in plant systems. Undoubtedly, open chromatin is required for cross overs but the present data shows that all the euchromatin features are not positively correlated with crossovers and vice-versa. Most of our food crops are polyploids and the data availability is very limited (Bomblies 2023). So, it’s a field waiting to be explored. Research on holocentric plants can give insights into chromatin features and their relation to recombination (Hofstatter et al. 2021). Studies with heterochromatin marks have been extensive and there are many euchromatin marks waiting for us to study their role in meiosis.
References
Acquaviva L, Drogat J, Dehé PM, de la Saint-André CR, Géli V (2013) Spp1 at the crossroads of H3K4me3 regulation and meiotic recombination. Epigenetics 8:355–360. https://doi.org/10.4161/epi.24295
Ali S, Zhang T, Lambing C, Wang W, Zhang P, Xie L, Wang J, Khan N, Zhang Q (2021) Loss of chromatin remodeler DDM1 causes segregation distortion in Arabidopsis thaliana. Planta. https://doi.org/10.1007/s00425-021-03763-5
Anderson LK, Lohmiller LD, Tang X, Hammond DB, Javernick L, Shearer L, Basu-Roy S, Martin OC, Falque M (2014) Combined fluorescent and electron microscopic imaging unveils the specific properties of two classes of meiotic crossovers. Proc Natl Acad Sci USA 111:13415–13420. https://doi.org/10.1073/pnas.1406846111
Baudat F, Buard J, Grey C, Fledel-Alon A, Ober C, Przeworski M, Coop G, De Massy B (2009) PRDM9 is a major determinant of meiotic recombination hotspots in humans and mice. Science 327:836–840. https://doi.org/10.1126/science.118343
Berchowitz LE, Copenhaver GP (2008) Fluorescent Arabidopsis tetrads: a visual assay for quickly developing large crossover and crossover interference data sets. Nat Protoc 3:41–50. https://doi.org/10.1038/nprot.2007.491
Bhattacharyya T, Walker M, Powers NR, Brunton C, Fine AD, Petkov PM, Handel MA (2019) Prdm9 and meiotic cohesin proteins cooperatively promote DNA double-strand break formation in mammalian spermatocytes. Curr Biol 29:1002–1018. https://doi.org/10.1016/j.cub.2019.02.007
Bomblies K (2023) Learning to tango with four (or more): the molecular basis of adaptation to polyploid meiosis. Plant Reprod 36:107–124. https://doi.org/10.1007/s00497-022-00448-1
Borde V, Robine N, Lin W, Bonfils S, Géli V, Nicolas A (2009) Histone H3 lysine 4 trimethylation marks meiotic recombination initiation sites. EMBO J 28:99–111. https://doi.org/10.1038/emboj.2008.257
Brazier T, Glémin S (2022) Diversity and determinants of recombination landscapes in flowering plants. PLoS Genet. https://doi.org/10.1371/journal.pgen.1010141
Brick K, Smagulova F, Khil P et al (2012) Genetic recombination is directed away from functional genomic elements in mice. Nature 485:642–645. https://doi.org/10.1038/nature11089
Cheng K, Xu Y, Yang C, Ouellette L, Niu L, Zhou X, Chu L, Zhuang F, Liu J, Wu H, Charron JB, Luo M (2020) Histone tales: lysine methylation, a protagonist in Arabidopsis development. J Exp Bot 71:793–807. https://doi.org/10.1093/jxb/erz435
Cheng J, Xu L, Bergér V, Bruckmann A, Yang C, Schubert V, Grasser KD, Schnittger A, Zheng B, Jiang H (2022) H3K9 demethylases IBM1 and JMJ27 are required for male meiosis in Arabidopsis thaliana. New Phytol 235(6):2252–2269. https://doi.org/10.1111/nph.18286
Choi K, Zhao X, Kelly KA, Venn O, Higgins JD, Yelina NE, Hardcastle TJ, Ziolkowski PA, Copenhaver GP, Franklin FCH, Mcvean G, Henderson IR (2013) Arabidopsis meiotic crossover hot spots overlap with H2A.Z nucleosomes at gene promoters. Nat Genet 45:1327–1338. https://doi.org/10.1038/ng.2766
Choi K, Zhao X, Tock AJ, Lambing C, Underwood CJ, Hardcastle TJ, Serra H, Kim J, Cho HS, Kim J, Ziolkowski PA (2018) Nucleosomes and DNA methylation shape meiotic DSB frequency in Arabidopsis thaliana transposons and gene regulatory regions. Genome Res 28:532–546
Coulton A, Burridge AJ, Edwards KJ (2020) Examining the effects of temperature on recombination in wheat. Front Plant Sci. https://doi.org/10.3389/fpls.2020.00230
Couteau F, Belzile F, Horlow C, Grandjean O, Vezon D, Doutriaux M-P (1999) Random chromosome segregation without meiotic arrest in both male and female meiocytes of a dmc1 mutant of Arabidopsis. Plant Cell 11:1623–1634. https://doi.org/10.1105/tpc.11.9.1623
Crismani W, Girard C, Froger N, Pradillo M, Santos JL, Chelysheva L, Copenhaver GP, Horlow C, Mercier R (2012) FANCM limits meiotic crossovers. Science 336:1588–1590. https://doi.org/10.1126/science.1220381
Da Ines O, Bazile J, Gallego ME, White CI (2022) DMC1 attenuates RAD51-mediated recombination in Arabidopsis. PLoS Genet. https://doi.org/10.1371/journal.pgen.1010322
Darlington CD, Dark SOS (1932) The origin and behaviour of chiasmata, II Stenobothrus parallelus. Cytologia 3:169–185. https://doi.org/10.1508/cytologia.3.169
Darrier B, Rimbert H, Balfourier F, Pingault L, Josselin AA, Servin B, Navarro J, Choulet F, Paux E, Sourdille P (2017) High-resolution mapping of crossover events in the hexaploid wheat genome suggests a universal recombination mechanism. Genetics 206:1373–1388. https://doi.org/10.1534/genetics.116.196014
Dawe RK (1998) Meiotic chromosome organization and segregation in plants. Annu Rev Plant Physiol Plant Mol Biol 49:371–395. https://doi.org/10.1146/annurev.arplant.49.1.371
De Jaeger-Braet J, Krause L, Buchholz A, Schnittger A (2022) Heat stress reveals a specialized variant of the pachytene checkpoint in meiosis of Arabidopsis thaliana. Plant Cell 34:433–454. https://doi.org/10.1093/plcell/koab257
Deleris A, Stroud H, Bernatavichute Y, Johnson E, Klein G, Schubert D, Jacobsen SE (2012) Loss of the DNA methyltransferase MET1 induces H3K9 hypermethylation at PcG target genes and redistribution of H3K27 trimethylation to transposons in Arabidopsis thaliana. PLoS Genet. https://doi.org/10.1371/journal.pgen.1003062
Demirci S, van Dijk ADJ, Sanchez Perez G, Aflitos SA, de Ridder D, Peters SA (2017) Distribution, position and genomic characteristics of crossovers in tomato recombinant inbred lines derived from an interspecific cross between Solanum lycopersicum and Solanum pimpinellifolium. Plant J 89:554–564. https://doi.org/10.1111/tpj.13406
Diagouraga B, Clément JA, Duret L, Kadlec J, de Massy B, Baudat F (2018) PRDM9 methyltransferase activity is essential for meiotic DNA double-strand break formation at its binding sites. Mol Cell 69:853–865. https://doi.org/10.1016/j.molcel.2018.01.033
Du J, Zhong X, Bernatavichute YV, Stroud H, Feng S, Caro E, Vashisht AA, Terragni J, Chin HG, Tu A, Hetzel J (2012) Dual binding of chromomethylase domains to H3K9me2-containing nucleosomes directs DNA methylation in plants. Cell 151(1):167–180. https://doi.org/10.1016/j.cell.2012.07.034
Durand S, Lian Q, Jing J, Ernst M, Grelon M, Zwicker D, Mercier R (2022) Joint control of meiotic crossover patterning by the synaptonemal complex and HEI10 dosage. Nat Commun. https://doi.org/10.1038/s41467-022-33472-w
Dziegielewski W, Ziolkowski PA (2021) License to regulate: noncoding RNA special agents in plant meiosis and reproduction. Front Plant Sci 12:662185. https://doi.org/10.3389/fpls.2021.662185
Ellermeier C, Higuchi EC, Phadnis N, Holm L, Geelhood JL, Thon G, Smith GR (2010) RNAi and heterochromatin repress centromeric meiotic recombination. Proc Natl Acad Sci U S A 107:8701–8705. https://doi.org/10.1073/pnas.0914160107
Fang J, Leichter SM, Jiang J, Biswal M, Lu J, Zhang Z-M, Ren W, Zhai J, Cui Q, Zhong X, Song J (2021) Substrate deformation regulates DRM2-mediated DNA methylation in plants. Sci Adv. https://doi.org/10.1126/sciadv.abd922
Fischle W, Wang Y, Allis CD (2003) Histone and chromatin cross-talk. Curr Opin Cell Biol 15:172–183. https://doi.org/10.1016/S0955-0674(03)00013-9
Francis KE, Lam SY, Harrison BD, Bey AL, Berchowitz LE, Copenhaver GP (2007) Pollen tetrad-based visual assay for meiotic recombination in Arabidopsis. Proc Natl Acad Sci U S A 104:3913–3918. https://doi.org/10.1073/pnas.0608936104
Grelon M, Vezon D, Gendrot G, Pelletier G (2001) AtSPO11-1 is necessary for efficient meiotic recombination in plants. EMBO J 20:589–600. https://doi.org/10.1093/emboj/20.3.589
Haenel Q, Laurentino TG, Roesti M, Berner D (2018) Meta-analysis of chromosome-scale crossover rate variation in eukaryotes and its significance to evolutionary genomics. Mol Ecol 27:2477–2497. https://doi.org/10.1111/mec.14699
Hansen L, Kim NK, Mariño-Ramírez L, Landsman D (2011) Analysis of biological features associated with meiotic recombination hot and cold spots in Saccharomyces cerevisiae. PLoS One. https://doi.org/10.1371/journal.pone.0029711
He Y, Wang M, Dukowic-Schulze S, Zhou A, Tiang CL, Shilo S, Sidhu GK, Eichten S, Bradbury P, Springer NM, Buckler ES, Levy AA, Sun Q, Pillardy J, Kianian PMA, Kianian SF, Chen C, Pawlowski WP (2017) Genomic features shaping the landscape of meiotic double-strand-break hotspots in maize. Proc Natl Acad Sci U S A 114:12231–12236. https://doi.org/10.1073/pnas.1713225114
He L, Huang H, Bradai M, Zhao C, You Y, Ma J, Zhao L, Lozano-Durán R, Zhu JK (2022) DNA methylation-free Arabidopsis reveals crucial roles of DNA methylation in regulating gene expression and development. Nat Commun. https://doi.org/10.1038/s41467-022-28940-2
Higgins JD, Buckling EF, Franklin FCH, Jones GH (2008) Expression and functional analysis of AtMUS81 in Arabidopsis meiosis reveals a role in the second pathway of crossing-over. Plant J 54:152–162. https://doi.org/10.1111/j.1365-313X.2008.03403.x
Higgins JD, Perry RM, Barakate A, Ramsay L, Waugh R, Halpin C, Armstrong SJ, Franklin FCH (2012) Spatiotemporal asymmetry of the meiotic program underlies the predominantly distal distribution of meiotic crossovers in barley. Plant Cell 24:4096–4109. https://doi.org/10.1111/j.1365-313X.2008.03403.x
Higgins JD, Osman K, Desjardins SD, Henderson IR, Edwards KJ, Franklin FHC (2022) Unravelling mechanisms that govern meiotic crossover formation in wheat. Biochem Soc Trans 50:1179–1186. https://doi.org/10.1042/BST20220405
Hofstatter PG, Thangavel G, Castellani M, Marques A (2021) Meiosis Progression and recombination in holocentric plants: what is known? Front Plant Sci 12:658296. https://doi.org/10.3389/fpls.2021.658296
Huang Y, Sicar S, Ramirez-Prado JS, Manza-Mianza D, Antunez-Sanchez J, Brik-Chaouche R, Rodriguez-Granados NY, An J, Bergounioux C, Mahfouz MM, Hirt H, Crespi M, Concia L, Barneche F, Amiard S, Probst AV, Gutierrez-Marcos J, Ariel F, Raynaud C, Latrasse D, Benhamed M (2021) Polycomb-dependent differential chromatin compartmentalization determines gene coregulation in Arabidopsis. Genome Res 31:1230–1244. https://doi.org/10.1101/gr.273771.120
Jamge B, Lorković ZJ, Axelsson E, Osakabe A, Shukla V, Yelagandula R, Akimcheva S, Kuehn AL, Berger F (2023) Histone variants shape chromatin states in Arabidopsis. Elife. https://doi.org/10.7554/eLife.87714
Jean M, Pelletier J, Hilpert M, Belzile F, Kunze R (1999) Isolation and characterization of AtMLH1, a MutL homologue from Arabidopsis thaliana. Mol Gen Genet 262:633–642. https://doi.org/10.1007/s004380051126
Jiang Y, N’Diaye A, Koh CS, Quilichini TD, Shunmugam ASK, Kirzinger MW, Konkin D, Bekkaoui Y, Sari E, Pasha A et al (2023) The coordinated regulation of early meiotic stages is dominated by non-coding RNAs and stage-specific transcription in wheat. Plant J 114:209–224
Ki BS, Shim SH, Park C et al (2022) Epigenetic regulator Cfp1 safeguards male meiotic progression by regulating meiotic gene expression. Exp Mol Med 54:1098–1108. https://doi.org/10.1038/s12276-022-00813-0
Kumar Koul K, Nagpal R (2023) Female meiosis in plants, and differential recombination in the two sexes: a perspective. Nucleus 66:195–223. https://doi.org/10.1007/s13237-023-00417-7
Kuo (2021) The role of Arabidopsis thaliana heterochromatin component H2A.W in meiotic recombination. Dissertation, University of Cambridge
Kurzbauer MT, Pradillo M, Kerzendorfer C, Sims J, Ladurner R, Oliver C, Janisiw MP, Mosiolek M, Schweizer D, Copenhaver GP, Schlögelhofer P (2018) Arabidopsis thaliana FANCD2 promotes meiotic crossover formation. Plant Cell 30:415–428. https://doi.org/10.1105/tpc.17.00745
Kurzbauer MT, Uanschou C, Chen D, Schlögelhofer P (2012) The recombinases DMC1 and RAD51 are functionally and spatially separated during meiosis in Arabidopsis. Plant Cell 24(5):2058–2070. https://doi.org/10.1105/tpc.112.098459
Lambing C, Tock AJ, Topp SD, Choi K, Kuo PC, Zhao X, Osman K, Higgins JD, Franklin FHC, Henderson IR (2020) Interacting genomic landscapes of REC8-cohesin, chromatin, and meiotic recombination in Arabidopsis. Plant Cell 32:1218–1239. https://doi.org/10.1105/TPC.19.00866
Li W, Chen C, Markmann-Mulisch U, Timofejeva L, Schmelzer E, Ma H, Reiss B (2004) The Arabidopsis AtRAD51 gene is dispensable for vegetative development but required for meiosis. Proc Natl Acad Sci U S A 101:10596–601
Lim EC, Kim J, Park J, Kim EJ, Kim J, Park YM, Cho HS, Byun D, Henderson IR, Copenhaver GP, Hwang I, Choi K (2020) DeepTetrad: high-throughput image analysis of meiotic tetrads by deep learning in Arabidopsis thaliana. Plant J 101:473–483. https://doi.org/10.1111/tpj.14543
Liu X, Yang S, Yu CW, Chen CY, Wu K (2016) Histone acetylation and plant development. Enzymes 40:173–199
Liu Y, Liu K, Yin L, Yu Y, Qi J, Shen WH, Zhu J, Zhang Y, Dong A (2019) H3K4me2 functions as a repressive epigenetic mark in plants. Epigenet Chromatin. https://doi.org/10.1186/s13072-019-0285-6
Liu Y, Yuan J, Jia G, Ye W, Jeffrey Chen Z, Song Q (2021) Histone H3K27 dimethylation landscapes contribute to genome stability and genetic recombination during wheat polyploidization. Plant J 105:678–690. https://doi.org/10.1111/tpj.15063
Lloyd A, Morgan C, Franklin FCH, Bomblies K (2018) Plasticity of meiotic recombination rates in response to temperature in Arabidopsis. Genetics 208:1409–1420. https://doi.org/10.1534/genetics.117.300588
Loppin B, Berger F (2020) Histone variants: the nexus of developmental decisions and epigenetic memory. Annu Rev Genet 54:121–149. https://doi.org/10.1146/annurev-genet-022620-100039
Lorenz A, Osman F, Sun W, Nandi S, Steinacher R, Whitby MC (2012) The fission yeast FANCM ortholog directs non-crossover recombination during meiosis. Science 336:1585–1588. https://doi.org/10.1126/science.1220111
Ma X, Fan L, Zhang Z, Yang X, Liu Y, Ma Y, Pan Y, Zhou G, Zhang M, Ning H, Kong F, Ma J, Liu S, Tian Z (2023) Global dissection of the recombination landscape in soybean using a high-density 600K SoySNP array. Plant Biotechnol J 21:606–620. https://doi.org/10.1111/pbi.13975
Marand AP, Jansky SH, Zhao H, Leisner CP, Zhu X, Zeng Z, Crisovan E, Newton L, Hamernik AJ, Veilleux RE, Buell CR, Jiang J (2017) Meiotic crossovers are associated with open chromatin and enriched with Stowaway transposons in potato. Genome Biol. https://doi.org/10.1186/s13059-017-1326-8
Marand AP, Zhao H, Zhang W, Zeng Z, Fang C, Jiang J (2019) Historical meiotic crossover hotspots fueled patterns of evolutionary divergence in rice. Plant Cell 31:645–662
Melamed-Bessudo C, Levy AA (2012) Deficiency in DNA methylation increases meiotic crossover rates in euchromatic but not in heterochromatic regions in Arabidopsis. Proc Natl Acad Sci U S A. https://doi.org/10.1073/pnas.1120742109
Mercier R, Mézard C, Jenczewski E, Macaisne N, Grelon M (2015) The molecular biology of meiosis in plants. Annu Rev Plant Biol 66:297–327. https://doi.org/10.1146/annurev-arplant-050213-035923
Merker JD, Dominska M, Greenwell PW, Rinella E, Bouck DC, Shibata Y, Strahl BD, Mieczkowski P, Petes TD (2008) The histone methylase Set2p and the histone deacetylase Rpd3p repress meiotic recombination at the HIS4 meiotic recombination hotspot in Saccharomyces cerevisiae. DNA Repair 7:1298–1308. https://doi.org/10.1016/j.dnarep.2008.04.009
Mirouze M, Lieberman-Lazarovich M, Aversano R, Bucher E, Nicolet J, Reinders J, Paszkowski J (2012) Loss of DNA methylation affects the recombination landscape in Arabidopsis. Proc Natl Acad Sci U S A 109:5880–5885. https://doi.org/10.1073/pnas.1120841109
Modliszewski JL, Copenhaver GP (2017) Meiotic recombination gets stressed out: CO frequency is plastic under pressure. Curr Opin Plant Biol 36:95–102. https://doi.org/10.1016/j.pbi.2016.11.019
Modliszewski JL, Wang H, Albright AR, Lewis SM, Bennett AR, Huang J, Ma H, Wang Y, Copenhaver GP (2018) Elevated temperature increases meiotic crossover frequency via the interfering (Type I) pathway in Arabidopsis thaliana. PLoS Genet. https://doi.org/10.1371/journal.pgen.1007384
Morgan C, Fozard JA, Hartley M, Henderson IR, Bomblies K, Howard M (2021) Diffusion-mediated HEI10 coarsening can explain meiotic crossover positioning in Arabidopsis. Nat Commun. https://doi.org/10.1038/s41467-021-24827-w
De Muyt A, Mercier R, Mezard C, Grelon M. (2009) Meiotic recombination and crossovers in plants. Meiosis. Kargel, Basel, 5:pp14–25.
Naish M, Alonge M, Wlodzimierz P, Tock AJ, Abramson BW, Schmücker A, Mandáková T, Jamge B, Lambing C, Kuo P, Yelina N, Hartwick N, Colt K, Smith LM, Ton J, Kakutani T, Martienssen RA, Schneeberger K, Lysak MA, Berger F, Bousios A, Michael TP, Schatz MC, Henderson IR (2021) The genetic and epigenetic landscape of the Arabidopsis centromeres. Science 1979:374. https://doi.org/10.1126/science.abi7489
Nambiar M, Smith GR (2016) Repression of harmful meiotic recombination in centromeric regions. Semin Cell Dev Biol 54:188–197. https://doi.org/10.1016/j.semcdb.2016.01.042
Nambiar M, Smith GR (2018) Pericentromere-specific cohesin complex prevents meiotic pericentric DNA double-strand breaks and lethal crossovers. Mol Cell 71:540–553. https://doi.org/10.1016/j.molcel.2018.06.035
Oliver C, Pradillo M, Corredor E, Cuñado N (2013) The dynamics of histone H3 modifications is species-specific in plant meiosis. Planta 238:23–33. https://doi.org/10.1007/s00425-013-1885-1
Osman K, Algopishi U, Higgins JD, Henderson IR, Edwards KJ, Franklin FCH, Sanchez-Moran E (2021) Distal bias of meiotic crossovers in hexaploid bread wheat reflects spatio-temporal asymmetry of the meiotic program. Front Plant Sci. https://doi.org/10.3389/fpls.2021.631323
Pai CC, Deegan RS, Subramanian L, Gal C, Sarkar S, Blaikley EJ, Walker C, Hulme L, Bernhard E, Codlin S, Bähler J, Allshire R, Whitehall S, Humphrey TC (2014) A histone H3K36 chromatin switch coordinates DNA double-strand break repair pathway choice. Nat Commun. https://doi.org/10.1038/ncomms5091
Parvanov ED, Petkov PM, Paigen K (2010) Prdm9 controls activation of mammalian recombination hotspots. Science 327:835
Peng JC, Karpen GH (2009) Heterochromatic genome stability requires regulators of histone H3 K9 methylation. PLoS Genet. https://doi.org/10.1371/journal.pgen.1000435
Peterson CL, Laniel M-A (2004) Histones and histone modifications. Curr Biol 14:546–551. https://doi.org/10.1016/j.cub.2004.07.007
Phillips D, Jenkins G, Macaulay M, Nibau C, Wnetrzak J, Fallding D, Colas I, Oakey H, Waugh R, Ramsay L (2015) The effect of temperature on the male and female recombination landscape of barley. New Phytol 208:421–429. https://doi.org/10.1111/nph.13548
Pinon V, Yao X, Dong A, Shen WH (2017) SDG2-mediated H3K4me3 is crucial for chromatin condensation and mitotic division during male gametogenesis in Arabidopsis. Plant Physiol 174:1205–1215. https://doi.org/10.1104/pp.17.00306
Portela A, Esteller M (2010) Epigenetic modifications and human disease. Nat Biotechnol 28:1057–1068. https://doi.org/10.1038/nbt.1685
Powers NR, Parvanov ED, Baker CL, Walker M, Petkov PM, Paigen K (2016) The meiotic recombination activator PRDM9 trimethylates both H3K36 and H3K4 at recombination hotspots in vivo. PLoS Genet. https://doi.org/10.1371/journal.pgen.1006146
Prusicki MA, Keizer EM, Van-Rosmalen RP, Komaki S, Seifert F, Müller K, Wijnker E, Fleck C, Schnittger A (2019) Live cell imaging of meiosis in Arabidopsis thaliana. eLife. https://doi.org/10.7554/eLife.42834.001
Qin Y, Zhao L, Skaggs MI, Andreuzza S, Tsukamoto T, Panoli A, Wallace KN, Smith S, Siddiqi I, Yang Z, Yadegari R, Palanivelu R (2014) ACTIN-RELATED PROTEIN6 regulates female meiosis by modulating meiotic gene expression in Arabidopsis. Plant Cell 26:1612–1628. https://doi.org/10.1105/tpc.113.120576
Rogakou EP, Pilch DR, Orr AH, Ivanova VS, Bonner WM (1998) DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. J Biol Chem 273:5858–5868. https://doi.org/10.1074/jbc.273.10.5858
Ross KJ, Fransz P, Jones GH (1996) A light microscopic atlas of meiosis in Arabidopsis thaliana. Chromosome Res 4:507–516. https://doi.org/10.1007/BF02261778
Rowan BA, Heavens D, Feuerborn TR, Tock AJ, Henderson IR, Weigel D (2019) An ultra high-density Arabidopsis thaliana crossover map that refines the influences of structural variation and epigenetic features. Genetics 213:771–787. https://doi.org/10.1534/genetics.119.302406
Shilo S, Melamed-Bessudo C, Dorone Y, Barkai N, Levy AA (2015) DNA crossover motifs associated with epigenetic modifications delineate open chromatin regions in Arabidopsis. Plant Cell 27:2427–2436. https://doi.org/10.1105/tpc.15.00391
Si W, Yuan Y, Huang J, Zhang X, Zhang Y, Zhang Y, Tian D, Wang C, Yang Y, Yang S (2015) Widely distributed hot and cold spots in meiotic recombination as shown by the sequencing of rice F2 plants. New Phytol 206:1491–1502. https://doi.org/10.1111/nph.13319
Sims J, Copenhaver GP, Schlögelhofer P (2019) Meiotic DNA repair in the nucleolus employs a nonhomologous end-joining mechanism. Plant Cell 31:2259–2275. https://doi.org/10.1105/TPC.19.00367
Sims J, Schlögelhofer P, Kurzbauer MT (2021) From microscopy to nanoscopy: defining an Arabidopsis thaliana meiotic atlas at the nanometer scale. Front Plant Sci 12:672914. https://doi.org/10.3389/fpls.2021.672914
Sturtevant AH (1915) The behavior of the chromosomes as studied through linkage. Z.Ver-erbungslehre 13:234–287. https://doi.org/10.1007/BF01792906
Sun L, Cao Y, Li Z, Liu Y, Yin X, Deng XW, He H, Qian W (2023) Conserved H3K27me3-associated chromatin looping mediates physical interactions of gene clusters in plants. J Integr Plant Biol 00:1–17. https://doi.org/10.1111/jipb.13502
Sun L, Jing Y, Liu X, Li Q, Xue Z, Cheng Z, Wang D, He H, Qian W (2020) Heat stress-induced transposon activation correlates with 3D chromatin organization rearrangement in Arabidopsis. Nat Commun 11(1):1886. https://doi.org/10.1038/s41467-020-15809-5
Tachibana M, Nozaki M, Takeda N, Shinkai Y (2007) Functional dynamics of H3K9 methylation during meiotic prophase progression. EMBO J 26:3346–3359. https://doi.org/10.1038/sj.emboj.7601767
Takada Y, Yaman-Deveci R, Shirakawa T, Sharif J, Tomizawa S, Miura F, Ito T, Ono M, Nakajima K, Koseki Y, Shiotani F, Ishiguro K, Ohbo K, Koseki H (2021) Maintenance DNA methylation in pre-meiotic germ cells regulates meiotic prophase by facilitating homologous chromosome pairing. Development. https://doi.org/10.1242/dev.194605
Tittel-Elmer M, Bucher E, Broger L, Mathieu O, Paszkowski J et al (2010) Stress-induced activation of heterochromatic transcription. PLoS Genet. https://doi.org/10.1371/journal.pgen.1001175
Tock AJ, Holland DM, Jiang W, Osman K, Sanchez-Moran E, Higgins JD, Edwards KJ, Uauy C, Franklin FCH, Henderson IR (2021) Crossover-active regions of the wheat genome are distinguished by DMC1, the chromosome axis, H3K27me3, and signatures of adaptation. Genome Res 31:1614–1628. https://doi.org/10.1101/gr.273672.120
Underwood CJ, Choi K, Lambing C, Zhao X, Serra H, Borges F, Simorowski J, Ernst E, Jacob Y, Henderson IR, Martienssen RA (2018) Epigenetic activation of meiotic recombination near Arabidopsis thaliana centromeres via loss of H3K9me2 and non-CG DNA methylation. Genome Res 28:519–531. https://doi.org/10.1101/gr.227116.117
Wang S, Veller C, Sun F, Ruiz-Herrera A, Shang Y, Liu H, Zickler D, Chen Z, Kleckner N, Zhang L (2019) Per-nucleus crossover covariation and implications for evolution. Cell 177:326–338. https://doi.org/10.1016/j.cell.2019.02.021
Xue M, Wang J, Jiang L, Wang M, Wolfe S, Pawlowski WP, Wang Y, He Y (2018) The number of meiotic double-strand breaks influence crossover distribution in Arabidopsis. Plant Cell 30:2628–2638. https://doi.org/10.1105/tpc.18.00531
Yamada T, Ki M, Hirota K, Kon N, Wahls WP, Hartsuiker E, Murofushi H, Shibata T, Ohta K (2004) Roles of histone acetylation and chromatin remodeling factor in a meiotic recombination hotspot. EMBO J 23:1792–1803. https://doi.org/10.1038/sj.emboj.7600138
Yamada S, Ohta K, Yamada T (2013) Acetylated histone H3K9 is associated with meiotic recombination hotspots, and plays a role in recombination redundantly with other factors including the H3K4 methylase Set1 in fission yeast. Nucleic Acids Res 41:3504–3517. https://doi.org/10.1093/nar/gkt049
Yamada S, Kugou K, Ding DQ, Fujita Y, Hiraoka Y, Murakami H, Ohta K, Yamada T (2018) The conserved histone variant H2A.Z illuminates meiotic recombination initiation. Curr Genet 64:1015–1019. https://doi.org/10.1007/s00294-018-0825-9
Yelagandula R, Stroud H, Holec S, Zhou K, Feng S, Zhong X, Muthurajan UM, Nie X, Kawashima T, Groth M, Luger K, Jacobsen SE, Berger F (2014) The histone variant H2A.W defines heterochromatin and promotes chromatin condensation in Arabidopsis. Cell 158:98–109. https://doi.org/10.1016/j.cell.2014.06.006
Yelina NE, Choi K, Chelysheva L, Macaulay M, de Snoo B, Wijnker E, Miller N, Drouaud J, Grelon M, Copenhaver GP, Mezard C, Kelly KA, Henderson IR (2012) Epigenetic remodeling of meiotic crossover frequency in Arabidopsis thaliana DNA methyltransferase mutants. PLoS Genet. https://doi.org/10.1371/journal.pgen.1002844
Yelina NE, Ziolkowski PA, Miller N, Zhao X, Kelly KA, Muñoz DF, Mann DJ, Copenhaver GP, Henderson IR (2013) High-throughput analysis of meiotic crossover frequency and interference via flow cytometry of fluorescent pollen in Arabidopsis thaliana. Nat Protoc 8:2119–2134. https://doi.org/10.1038/nprot.2013.13
Yelina N, Diaz P, Lambing C, Henderson IR (2015a) Epigenetic control of meiotic recombination in plants. Sci China Life Sci 58:223–231. https://doi.org/10.1007/s11427-015-4811-x
Yelina NE, Lambing C, Hardcastle TJ, Zhao X, Santos B, Henderson IR (2015b) DNA methylation epigenetically silences crossover hot spots and controls chromosomal domains of meiotic recombination in Arabidopsis. Genes Develop 29:2183–2202. https://doi.org/10.1101/gad.270876
Zamudio N, Barau J, Teissandier A, Walter M, Borsos M, Servant N, Bourc’his D (2015) DNA methylation restrains transposons from adopting a chromatin signature permissive for meiotic recombination. Genes Dev 29:1256–1270. https://doi.org/10.1101/gad.257840
Zhang T, Cooper S, Brockdorff N (2015) The interplay of histone modifications—writers that read. EMBO Rep 16:1467–1481. https://doi.org/10.15252/embr.201540945
Zhang C, Du X, Tang K, Yang Z, Pan L, Zhu P, Luo J, Jiang Y, Zhang H, Wan H, Wang X, Wu F, Tao WA, He XJ, Zhang H, Bressan RA, Du J, Zhu JK (2018) Arabidopsis AGDP1 links H3K9me2 to DNA methylation in heterochromatin. Nat Commun. https://doi.org/10.1038/s41467-018-06965-w
Zhang Y, Ma M, Liu M, Sun A, Zheng X, Liu K, Yin C, Li C, Jiang C, Tu X, Fang Y (2023) Histone H2A monoubiquitination marks are targeted to specific sites by cohesin subunits in Arabidopsis. Nat Commun. https://doi.org/10.1038/s41467-023-36788-3
Ziolkowski PA (2023) Why do plants need the ZMM crossover pathway? A snapshot of meiotic recombination from the perspective of interhomolog polymorphism. Plant Reprod 36:43–54. https://doi.org/10.1007/s00497-022-00446-3
Acknowledgements
We thank Sheena Shah for proofreading the manuscript. This work was supported by Faculty Research Programme Grant-Institution of Eminence (IoE) (IoE Grant No. /IoE/2021/12/FRP).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
We declare that we have no competing interest.
Consent for publication
All authors consented for this publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chowdary, K.V.S.K.A., Saini, R. & Singh, A.K. Epigenetic regulation during meiosis and crossover. Physiol Mol Biol Plants 29, 1945–1958 (2023). https://doi.org/10.1007/s12298-023-01390-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-023-01390-w