Abstract
Synapse organizers are essential for the development, transmission, and plasticity of synapses. Acting as rare synapse suppressors, the MAM domain containing glycosylphosphatidylinositol anchor (MDGA) proteins contributes to synapse organization by inhibiting the formation of the synaptogenic neuroligin-neurexin complex. A previous analysis of MDGA2 mice lacking a single copy of Mdga2 revealed upregulated glutamatergic synapses and behaviors consistent with autism. However, MDGA2 is expressed in diverse cell types and is localized to both excitatory and inhibitory synapses. Differentiating the network versus cell-specific effects of MDGA2 loss-of-function requires a cell-type and brain region-selective strategy. To address this, we generated mice harboring a conditional knockout of Mdga2 restricted to CA1 pyramidal neurons. Here we report that MDGA2 suppresses the density and function of excitatory synapses selectively on pyramidal neurons in the mature hippocampus. Conditional deletion of Mdga2 in CA1 pyramidal neurons of adult mice upregulated miniature and spontaneous excitatory postsynaptic potentials, vesicular glutamate transporter 1 intensity, and neuronal excitability. These effects were limited to glutamatergic synapses as no changes were detected in miniature and spontaneous inhibitory postsynaptic potential properties or vesicular GABA transporter intensity. Functionally, evoked basal synaptic transmission and AMPAR receptor currents were enhanced at glutamatergic inputs. At a behavioral level, memory appeared to be compromised in Mdga2 cKO mice as both novel object recognition and contextual fear conditioning performance were impaired, consistent with deficits in long-term potentiation in the CA3-CA1 pathway. Social affiliation, a behavioral analog of social deficits in autism, was similarly compromised. These results demonstrate that MDGA2 confines the properties of excitatory synapses to CA1 neurons in mature hippocampal circuits, thereby optimizing this network for plasticity, cognition, and social behaviors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Synapse organizers act as molecular codes that help determine synapse assembly, maturation, and function during brain development [1,2,3,4]. Gene linkage studies have implicated these organizers in the pathogenesis of most neurodevelopmental and neuropsychiatric disorders [5, 6]. Canonical synaptic complexes consisting of presynaptic neurexins (Nrxs) and postsynaptic neuroligins (NLs) bind across the synaptic cleft to coordinate the molecular organization of synapses [2, 4, 7]. They accomplish this by initiating the intracellular recruitment of machinery required for synapse validation including scaffolds, ion channels, and components of vesicular release [4, 7]. Presynaptic Nrxs and postsynaptic NLs are complex gene families, with different isoforms that selectively function at excitatory or inhibitory synapses [8], and mutations in both families of synapse organizers are contributing factors in autism and other brain disorders [6].
Nrxs are located at the presynapse where they organize the molecular networks required for vesicular release, but can also influence postsynaptic properties through multiple postsynaptic binding partners including leucine-rich-repeat transmembrane neuronal proteins, calsyntenins, and neuroligins [8,9,10]. Neuroligins are localized to the postsynaptic membrane of glutamatergic (NL1 and NL3), GABAergic (NL2, NL3, and NL4), and glycinergic (NL4) synapses [11,12,13,14]. NL1 is expressed at glutamatergic synapses where it directly interacts with membrane-associated guanylate kinase scaffolding proteins including PSD-95 [15, 16].
The NL–Nrx complex is regulated by meprin, A-5 protein, and receptor protein-tyrosine phosphatase mu [MAM] domain-containing glycosylphosphatidylinositol anchor proteins (MDGAs), which are established as repressors of synapse development [17,18,19,20,21,22,23]. MDGAs contain six extracellular immunoglobulin-like (Ig) domains, a fibronectin type III-like domain, a MAM domain, and a glycosylphosphatidylinositol anchor [24]. Recent crystal structure analysis has elucidated the binding interactions through which MDGAs suppress NL–Nrx-mediated synapse development. Structurally, two MDGA2 molecules form a triangular structure that encompasses NL dimers, simultaneously occluding Site 1 on both NLs which constitute the Nrx binding regions [25,26,27]. Ig domains 1–3 mediate this direct steric hindrance [18, 19]. In humans, Mdga1 mutations are associated with schizophrenia [28] and Mdga2 copy number variants have been reported in 10 unrelated cases of autism [29]. As such, MDGA2 is a brain-specific cell surface protein that falls within a major autism spectrum disorder (ASD) pathway through the prevention of NL–Nrx interactions [30].
MDGAs are broadly expressed throughout the brain including superficial cortical layers, the hippocampus, the amygdala, the cerebellum, the thalamus, and the olfactory bulb [20]. The expression of MDGAs is elevated during late embryonic and early postnatal periods but also persists into adulthood, particularly in the cortex and hippocampus [19, 21, 22, 24]. Within the hippocampus, MDGA2 is localized to multiple cell types, including interneurons [22], whereas MDGA1 appears to be restricted to pyramidal neurons [21]. At the subcellular level, we previously reported diffuse localization of recombinant MDGA2 throughout neurons [22]. However, the use of a potentially more sensitive tag revealed a selective concentration of MDGA2 at inhibitory synapses [31].
The fundamental importance of MDGAs for synapse development is supported by the perinatal lethal phenotype of mice lacking MDGA2, which necessitates the use of heterozygotes (Mdga2+/−) for in vivo analysis [22]. Initial findings in vitro suggested that MDGAs negatively regulate GABAergic synapses, as MDGA1 overexpression reduces, and knockdown increases, GABAergic synapse density in cultured neurons [18, 19]. In addition, co-expression of MDGA1 with NL2, the GABAergic-selective neuroligin, inhibits the ability of NL2 to induce presynaptic differentiation in contacting axons. Based on these results, MDGA2 was similarly hypothesized to inhibit NL2-mediated GABAergic synapse development. However, MDGA2 appears to selectively repress excitatory synapses onto pyramidal neurons in vivo. Mdga2+/− mice exhibit upregulation of glutamatergic synapses, AMPAR-mediated transmission, and GluA1 surface expression [20, 22]. CRISPR/Cas9-mediated knockout of Mdga2 in CA1 pyramidal neurons of organotypic slices also enhances AMPAR currents, consistent with the reports in Mdga2+/− mice [32].
Levels of both NL1 and PSD-95 family proteins are likewise increased in multiple brain regions lacking MDGA2. These synaptic deficits occur in parallel with altered long-term potentiation (LTP) and impaired memory. Mdga2+/− mice further demonstrate behaviors consistent with autism, including reduced social interactions and repetitive jumping [22], although these phenotypes appear to be influenced by the genetic background [17]. Importantly, the lack of effect on inhibitory synapses in vivo may be due to compensation by MDGA1 at inhibitory synapses within this neuronal population [22].
Reconciling these findings requires consideration of alternative models proposing how MDGA2 determines neural circuit properties. One such model is the “cell-autonomous” model, which predicts that a reduction of MDGA2 will increase excitatory and inhibitory synapses exclusively in the cells in which it is expressed [20]. Importantly, MDGA2 function was initially characterized in vivo using heterozygotes derived from a germline knockout of Mdga2, limiting our understanding of how MDGA2 regulates synapse development and function within distinct cell populations. In addition, a major knowledge gap exists regarding how MDGA2 uniquely regulates synapse properties in development relative to mature neural circuits.
Here, we sought to overcome these limitations by selectively deleting Mdga2 from CA1 pyramidal neurons in adult mice. The CA1 region was targeted as this allows direct comparison with previous studies [26], in a well-characterized brain region that is functionally linked to the cognitive deficits in autism. In addition, using a “late” knockout strategy provides a more precise method for exploring how MDGA2 within CA1 pyramidal neurons uniquely impacts neuronal, synaptic, and behavioral outputs, while differentiating constitutive versus ongoing requirements for MDGA2 during development.
Materials and Methods
Animals
Mdga2 floxed mice were generated by Biocytogen Pharmaceuticals Co., Ltd (Beijing, China) by injecting Cas9/sgRNA into fertilized eggs (Fig. S1); they were maintained on the C57BL/6J background. The generation of Mdga2 floxed mice was as previously described [33]. The single guide RNAs (sgRNAs) were designed specifically for the introns upstream and downstream of Exon2 using online CRISPR design tools (https://wge.stemcell.sanger.ac.uk//, https://cctop.cos.uni-heidelberg.de:8043/, https://crisprgold.mdc-berlin.de/index.php). The designed sgRNAs were then screened for on-target activity using UCATM (Universal CRISPR Activity Assay), a sgRNA activity detection system developed by Biocytogen that is simpler and more sensitive than MSDase assay. The sequences of screened sgRNAs were as follows: 5′Guide, 5′-GGGGGGGGGGTTCATGTACA-GGG-3′; 3′Guide, 5′-TGACATAATTGTACCTGAAGAGG-3′.
The mice were housed 4–6 per cage with food and water available ad libitum and were maintained in a room at 21 ± 2 °C under a 12 h/12 h dark/light cycle (light on at 07:00 and off at 19:00). All experiments were approved by the Zhejiang University Animal Use and Care Committee and performed following the National Institutes of Health (NIH) Guide for the Care and Use of Laboratory Animals. All the mice used were male unless otherwise specified. All mice were 5–6 weeks old at the time of injection and allowed to recover for 6–7 weeks prior to experiments. Genomic DNA was extracted from the tail of 4–6-week-old males for genetic identification of Mdga2 floxed mice. The sequences of PCR primers were as follows: Forward 5′-TTGTGCGCTGATGCGAGATAGAGAG-3′; Reverse 5′-GGGGATTTGAACTAAAGGCACAGG-3′ (Sangon Biotech, China).
Stereotaxic Surgery and Viral Injection
For stereotaxic surgery, the mice were deeply anesthetized with 2% pentobarbital sodium and fixed in a stereotaxic frame (RWD, China). An incision was made in the midline of the scalp, and a craniotomy was performed above the hippocampus region. The virus AAV2/9-CaMKIIα-CRE-P2A-mCherry-WPRE-hGH-pA (1.26 × 1011 vg/mL, 1:20, 200 nL, PT-0739, BrainVTA, China) or the control virus: AAV2/9-CaMKIIα-mCherry-WPRE-hGH-pA (2.57 × 1011 vg/mL, 1:20, 200 nL, PT-0108, BrainVTA, China) was injected bilaterally into the hippocampus, area CA1 (ML, ± 1.7 mm; AP, − 2.15 mm; DV, − 1.23 mm) through a pulled-glass micropipette with a 10–20 μm tip diameter connected to a 10 μL microliter syringe (Hamilton Co., USA). The AAV was delivered using a stereotactic injector (Pump 11 Elite, Harvard Apparatus, USA) at a rate of 30 nL/min. After injection, the micropipettes were left in place for an additional 10 min to allow for diffusion before the pipette was slowly withdrawn.
Immunohistochemistry
Mice were deeply anesthetized with isoflurane and transcardially perfused with cold 0.1 mol/L PBS followed by chilled 4% paraformaldehyde (PFA, P0099, Beyotime Biotechnology, China) in PBS (C0221A, Beyotime Biotechnology, China). The brains were removed and then post-fixed overnight at 4 °C in PFA. Coronal brain sections (30 μm thick) were cut on a VT1200S vibratome (Leica Microsystems, Germany) for immunofluorescence. The sections were incubated in blocking buffer (QuickBlock™, P0260, Beyotime Biotechnology, China) for 1 h at 37 °C, and incubated with the primary antibodies anti-CaMKII alpha (1:300, rabbit, ab5683, Abcam, USA), anti-VGAT (1:200, mouse, #131011, Synaptic Systems, Germany), anti-GAD65 (1:200, mouse, AB_528264, Developmental Studies Hybridoma Bank, USA), and anti-VGluT1 (1:800, guinea pig, AB5905, Millipore, USA) at 4 °C for 24 h, followed by the corresponding secondary antibodies conjugated to Alexa 488 or 647 (Thermo Fisher, USA) for 2 h at 37 °C. After washing, the sections were coverslipped using Dapi Fluoromount-G for nuclear labeling. Fluorescence signals were visualized under a confocal microscope (SP8, Leica Microsystems, Germany) using the same parameters for each marker among sections.
For quantification of excitatory (VGluT1) and inhibitory (GAD65) synaptic markers in brain sections, images were manually thresholded to define puncta, and the total area of vGluT1 and GAD65 per tissue area was measured. Measurements were made on a minimum of three sites per animal for each region and a total of six animals for each group, measuring the same regions for both VGluT1 and GAD65. Analysis was applied using ImageJ software (NIH, USA) and GraphPad Prism (GraphPad Software, USA).
Quantitative Reverse Transcription PCR (qRT-PCR)
Total RNA was extracted from the hippocampus using TRIzol reagent (Invitrogen) according to the manufacturer’s instructions. Total RNA was quantified using a spectrophotometer (Nanodrop 2000, Thermo Fisher). All the samples presented 260/280 nm ratios between 1.8 and 2.0. Next, cDNA was synthesized using the PrimeScript™ RT reagent kit (Takara). Real-time quantitative PCR was then performed using TB Green® Premix Ex Taq™ (Tli RNase H Plus, Takara) on the Real-Time PCR Detection System (CFX Connect™, BIO-RAD, USA). The primer sequences of Mdga2 and GAPDH were as follows: Mdga2 (Forward 5′-TGGTTACTGG-ACATCCACGC-3′, Reverse 5′-GAGAGGGTGTAGGCTCTCCT-3′), GAPDH (Forward 5′-GGTGAAGGTCGGTGTGAACG-3′, Reverse 5′-CTCGCTCCTGGAA-GATGGTG-3′). The results were analyzed using the 2(−ΔΔCt) method.
In Situ Hybridization (ISH)
In situ hybridization was carried out using the BaseScope Reagent Kit v2–RED (323900, Advanced Cell Diagnostics, USA). Assays were performed following guidelines provided by the supplier (Advanced Cell Diagnostics, USA). Briefly, brain sections were incubated in BaseScope hydrogen peroxide, followed by target retrieval, and protease III solution. The sections were then hybridized with the probe (1259031-C1, BA-Mm-Mdga2-3zz-st, Advanced Cell Diagnostics, USA) and incubated in amplification solutions. Signals were detected using BaseScope™ Fast RED.
Western Blot
Tissue samples were lysed with Syn-PER Reagent (#87793, Thermo Scientific, USA) and PMSF (ST506, Beyotime Biotechnology, China) at a ratio of Syn-PER Reagent: PMSF = 100:1 and were centrifuged at 1200 g for 10 min at 4 °C to obtain supernatant which was transferred to new tubes. Then the samples were centrifuged at 15000 g for 20 min at 4 °C to remove the supernatant and obtain the synaptosome pellet. Finally, Syn-PER Reagent was added to suspend the synaptosome pellet. The BCA protein assay kit (P0010S, Beyotime Biotechnology, China) was used to quantify the protein concentration. Protein (20 μg) was loaded onto 8% SDS-PAGE gels for electrophoretic separation and transferred onto PVDF membranes (IPVH00010, Millipore, USA). The PVDF membranes were then blocked with 5% BSA in TBST buffer for 1 h at RT and subsequently incubated with primary antibodies at 4 °C. The primary antibodies were as follows: anti-β-actin (1:5000, mouse, #3700, Cell Signaling Technology, USA), anti-β-actin (1:5000, rabbit, #4970, Cell Signaling Technology, USA), anti-MDGA1 (1:2,000, rabbit, #421002, Synaptic Systems, Germany), anti-Mdga2 (1:1000, rabbit, orb2939, Biorbyt, UK), anti-NL1 (1:2000, mouse, #129111, Synaptic Systems, Germany), anti-NL2 (1:2000, mouse, #129511, Synaptic Systems, Germany), anti-NL3 (1:2000, mouse, #129311, Synaptic Systems, Germany), anti-VGAT (1:1000, mouse, #131011, Synaptic Systems, Germany), anti-VGluT1 (1:2000, rabbit, #135303, Synaptic Systems, Germany), anti-Synapsin1/2 (1:4000, guinea pig, #106004, Synaptic Systems, Germany), anti-Gephyrin (1:1000, mouse, #75-443, NeuroMab, USA), anti-PSD95 (1:2000, mouse, #75-348, NeuroMab, USA), anti-GluA1 (1:2000, rabbit, AB1504, Sigma-Aldrich, USA) and anti-GluN1 (1:2000, mouse, MAB363, Sigma-Aldrich, USA). Appropriate secondary antibodies were added for 1 h at RT at the following dilutions: anti-mouse 680 conjugate (1:5000, goat, #5257, Cell Signaling Technology, USA), anti-mouse 800 conjugate (1:5000, goat, #5470, Cell Signaling Technology, USA), anti-rabbit 800 conjugate (1:5000, goat, #5151, Cell Signaling Technology, USA) and anti-guinea pig 800 conjugate (1:5000, goat, #925-32411, LI-COR, USA). Immunoreactive bands were visualized using Odyssey CLx (LI-COR, USA), and data were quantified using Fiji/ImageJ (NIH, USA).
Ex Vivo Slice Electrophysiology Recording
Mice were anesthetized with isoflurane and then perfused with 20 mL ice-cold cutting solution (oxygenated with 95% O2 + 5% CO2) containing (in mmol/L): 120 choline chloride, 2.5 KCl, 7 MgSO4, 0.5 CaCl2, 1.25 NaH2PO4, 26 NaHCO3, and 25 glucose, with 3 mmol/L sodium pyruvate and 5 mmol/L sodium ascorbate added. The brain was rapidly removed from the skull and quickly chilled in an oxygenated cutting solution. Coronal slices (300 μm) containing the hippocampus were cut in a cold-cutting solution using a VT1200S vibratome (Leica, Germany). The slices were recovered in cutting solution for 15 min at 34 °C, followed by oxygenated artificial cerebrospinal fluid (ACSF) containing (in mmol/L): 124 NaCl, 2.5 KCl, 2 MgSO4, 2.5 CaCl2, 1.25 NaH2PO4, 22 NaHCO3, and 10 glucose, for at least 1 h at 25 °C before recording. ACSF was continuously gassed with 95% O2 and 5% CO2.
Whole-cell patch-clamp recordings were made using an infrared differential interference contrast microscope (BX51WI, Olympus, Japan) equipped with two automatic manipulators (MP285, Sutter Instrument Co, USA) and a highly sensitive CCD camera (IR-1000E, DAGE-MTI, USA). A single slice was transferred to the recording chamber and continuously perfused with oxygenated ACSF at a rate of ~ 2 mL/min. The temperature of the ASCF was maintained at 32 ± 1 °C with an automatic temperature controller (TC-324C, Warner Instrument Co. USA). Recording electrodes were made from filamented borosilicate glass capillary tubes (inner diameter, 0.84 μm) using a vertical pipette puller (PC-100, Narishige, Japan). The pipette resistance was 3–5 MΩ. Series resistance (Rs) was in the range of 10–20 MΩ and monitored throughout the experiments. If Rs changed > 20% during recording, the data were excluded. Data were sampled at 10 kHz and filtered at 3 kHz using a patch-clamp amplifier (MultiClamp 700B, Axon, USA) and collected with pCLAMP 10.6 software (Molecular Devices, USA).
To evoke action potentials (APs), cells were recorded in current-clamp mode, and increasing in steps of 50 pA (0–250 pA, 1 s) or depolarizing current pulse ramps (250 pA over 1 s) were injected [34]. The patch electrodes were filled with K-gluconate pipette solution containing (in mmol/L), 130 K-gluconate, 5 NaCl, 1 MgCl2, 10 HEPES, 0.2 EGTA, 2 MgATP, and 0.1 NaGTP. The pH was adjusted to 7.3–7.4 with KOH and osmolarity was adjusted to 285 mOsm with sucrose. The AP threshold was identified at the point where the AP was initiated and showed a > 10-fold change in the rate of rise. The fast afterhyperpolarization potential (fAHP) was measured as the difference between the peak following the spike and the threshold potential. AP amplitude was calculated as the maximum voltage during the spike minus the spike threshold voltage. AP half-width was measured at half-height between the threshold and the peak of the AP. The rheobase was measured as the current at the first AP threshold.
To record miniature excitatory postsynaptic currents (mEPSCs), the membrane potential was held at − 70 mV with 1 μmol/L tetrodotoxin (TTX) and 100 μmol/L picrotoxin (PTX) added into the bath solution. To record spontaneous EPSC (sEPSCs), 100 μmol/L PTX was added to the bath solution. For recordings of mEPSCs and sEPSCs, the patch electrodes were filled with a Cs-methanesulfonate-based pipette solution containing (in mmol/L) 130 Cs-methanesulfonate, 5 NaCl, 1 MgCl2, 10 HEPES, 0.2 EGTA, 2ATP-Mg, 0.1 GTP-Na, and 5 QX314. The pH was adjusted to 7.3–7.4 with CsOH and osmolarity was adjusted to 285 mOsm with sucrose.
To record mIPSCs, the membrane potential was held at − 70 mV with 20 μmol/L cyanquixaline (CNQX), 20 μmol/L APV (DL-2-amino-5-phosphonovaleric acid), and 1 μmol/L TTX added to the bath solution. To record sIPSCs, 20 μmol/L CNQX and 20 μmol/L APV were added to the bath solution. For recordings of mIPSC and sIPSC, the patch electrodes were filled with a CsCl-based pipette solution containing (in mmol/L) 100 CsCl, 30 Csmethanesulfonate, 5 NaCl, 1 MgCl2, 10 HEPES, 0.2 EGTA, 2 ATP-Mg, and 0.1 GTP-Na. The pH was adjusted to 7.3–7.4 with CsOH and osmolarity was adjusted to 285 mOsm with sucrose.
For paired-pulse ratio recording, pyramidal neurons were clamped at − 70 mV in the presence of 100 μmol/L PTX, and the patch electrodes were filled with Cs-methanesulfonate pipette solution. A concentric bipolar stimulating electrode was placed in the stratum radiatum to evoke EPSCs in CA1 pyramidal cells. Paired stimuli were delivered with different inter-stimulus intervals (25, 50, 100, and 200 ms). The ratios were calculated as 2nd EPSC/1st EPSC.
To measure LTP at the Schaffer collateral-CA1 synapses in hippocampal slices, whole-cell recording was made in virus-infected CA1 neurons with ACSF that contained 100 μmol/L PTX. The patch electrodes were filled with a 130 mmol/L Cs-methanesulfonate pipette solution. Evoked EPSPs (eEPSPs) were recorded in current clamp mode and current injection was manually applied to maintain the cell near a stable resting potential of − 70 mV. Stimulus intensity was adjusted to produce eEPSPs of 2–7 mV. then LTP was induced by a compressed theta-burst stimulation (cTBS) protocol (Fig. 5B) applied in voltage clamp mode (− 10 mV) after recording a 15 min baseline. LTP was quantified by normalizing the data collected in the last 15 min to the mean value of the slope of baseline eEPSPs, which were recorded for 15 min before LTP induction. Input-output curves of AMPAR and NMDAR were generated by increasing stimulation in a multiplicative manner in the presence of PTX (100 μmol/L), in whole-cell mode. The evoked AMPAR-mediated EPSCs were recorded at − 70 mV and NMDAR-mediated EPSCs were recorded at + 40 mV. NMDA receptor-mediated EPSCs were determined 50 ms after the peak of the current response. The patch electrodes were filled with a 130 mmol/L Cs-methanesulfonate pipette solution. “Threshold” stimulation was determined by using a stimulation intensity that evoked a response of ~ 15 pA (1×). To measure the AMPA/NMDA current ratio, cells were held at − 70 mV to record AMPAR-mediated EPSCs and at + 40 mV to record NMDAR-mediated EPSCs, in the presence of PTX. NMDA receptor-mediated EPSCs were determined 50 ms after the peak of the current response when the contribution of the AMPAR current was minimal. The current–voltage (I/V) curves of AMPA and NMDA receptor currents were derived at holding potentials ranging from − 80 mV to + 40 mV. Currents were normalized to peak responses at − 80 mV and + 40 mV, respectively.
TTX, PTX, APV, and CNQX were from Tocris unless otherwise specified. Stock solutions of PTX and CNQX were made by dissolution in dimethyl sulfoxide. Stock solutions of TTX and APV were made in sterilized ddH2O.
Behavioral Tests
The open field test, novel object recognition, three-chamber social interaction test, and contextual fear conditioning were carried out 6 weeks after the expression of the virus. Mice were handled for 20 min per day over 5–7 days before the behavioral tests to minimize the impact of handling stress.
Open Field Test
Mice were brought to the room and acclimated for at least 30 min before the test. The open field chamber was made of transparent plastic (40 cm × 40 cm) and a 20 cm × 20 cm center square was color marked. Individual mice were placed in the center of the chamber and their behavior was monitored for 10 min with an overhead video-tracking system (Ethovision XT, Noldus, Netherlands). The time spent in the center area and the total distance traveled were monitored throughout the experiment.
Novel Object Recognition Test
The experiment was divided into three stages. In the first stage, during the adaptation period, the mice explored freely for 10 min in an open field chamber (40 cm × 40 cm and 40 cm in height). In the second stage, during the familiarity period, two identical objects were placed in the chamber, 10 cm away from each side of the wall. Each mouse was placed into the chamber with its back to the object at an equal distance from each object. The exploration time (touching the object with the mouth or nose and approaching to within ~ 2–3 cm of the object) spent on each object and the frequency of exploration for each object within 5 min epochs were recorded with the overhead video-tracking system. 1 h after exposure to the now familiar objects, mice were reintroduced into the open field for 10 min with two objects, a copy of a familiar object and a new object, and the time spent exploring either one of the objects was recorded. The recognition index (RI) was calculated as follows: RI = novel object exploration time/total exploration time.
Three-Chamber Social Test
The three-chamber social test apparatus consisted of three rectangular boxes (each 20 cm in length, 40 cm in width, and 20 cm in height) and two identical cylindrical cages (15 cm in diameter and 20 cm in height). The cages were located at either end of the apparatus. Each box was separated by a transparent plexiglass plate with 10 cm × 10 cm openings that allowed the mice to move between chambers. Mice acclimated to the room for at least 1 h before the behavioral test and acclimated to the three-chamber apparatus for 10 min immediately before testing. After the adaptation period, the mouse was put back into the middle chamber. In the first stage of the experiment, a novel mouse (Stranger 1) was randomly put into the cylindrical cage on one side, and the other side was empty. The exploration time and the contact time between Stranger 1 and the vacant cylindrical cage were recorded for 10 min. In the second stage of the experiment, a second novel mouse (Stranger 2) was put into the empty cylindrical cage on the other side. The time spent sniffing each container was recorded and analyzed between Stranger 1 (the now familiar mouse) and Stranger 2 (the novel mouse) within 10 min epochs. At the end of the 10 min testing period, the mice were removed, and the apparatus and cages were washed with 75% ethanol, followed by drying with a paper towel to remove odors.
Contextual Fear Conditioning
Mice were placed into the conditioning chamber (60 cm × 60 cm × 60 cm) for 2 min to explore the testing chamber and develop a representation of the context. This was followed by three unconditioned stimuli (footshock; 2 s, 0.5 mA) at 1 min intervals. The chamber was cleaned with 75% ethanol after testing. To evaluate the memory retention of contextual fear conditioning, the mice were placed back in the conditioning chamber 24 h later. The freezing behavior (absence of all movement except respiration) was videotaped and analyzed using the CleverSys FreezeScan system (CleverSys Inc.) and expressed as the percentage of time freezing.
Statistical Analysis
All data are presented as the mean ± SEM. An unpaired Student’s t-test was used to compare the two groups. For comparison among three or more groups, one-way ANOVA with post-hoc Bonferroni’s t-test was used. For data with more than one independent variable, two-way ANOVA with post-hoc Bonferroni’s t-test was used. In all cases, P < 0.05 was considered to be statistically significant.
Results
Generation and Characterization of Mdga2 Floxed Mice
To understand how MDGA2 regulates synaptic properties at single-cell resolution, we generated Mdga2 floxed mice by injecting Cas9/sgRNA into fertilized eggs (Fig. S1 and Fig. 1A). The Mdga2 floxed alleles were validated by genotyping (Fig. 1B). As expected from the conditional nature of the floxed alleles, the mice developed and bred normally.
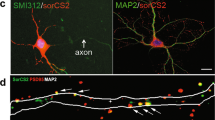
Generation of Mdga2 conditional knockout transgenic mice. A Diagram of the Mdga2 conditional allele targeting strategy. Cre-mediated recombination excises exon 2 and results in a knockout allele. Arrows indicate the positions of PCR primers used for verifying the presence of the wild-type (WT) and the floxed alleles. B PCR genotyping of genomic DNA from Mdga2f/f, Mdga2f/+, and WT mice using forward (F) and reverse (R) primers which flank the 5′-loxp site and show a WT band at 282 bp and a flox-inserted band at 386 bp. C Schematic of the experimental procedure. Verification of Mdga2 knockout efficiency was assessed by qPCR and western blot (WB) 5–6 weeks after virus injection in CA1. D Representative images showing the AAVs injection site in the hippocampal CA1. Scale bar, 500 μm. E Quantification of MDGA2 mRNA levels in the hippocampus with virus-infected CA1 PNs. n = 4 mice for Con, n = 6 mice for Cre; P = 0.0052, unpaired t test. F and G Immunoblot and quantification of MDGA2 and MDGA1 proteins in AAV-infected CA1 6–7 weeks after viral injections. Protein levels were normalized to the loading control (β-actin) and then normalized to the mean of the Con value; n = 3 mice per group; MDGA2, P = 0.0318; MDGA1, P = 0.3676; unpaired t test. β-actin is used as a loading control. P2, synaptosome pellet; DAPI, labels nuclei. All data are presented as the mean ± SEM. n.s., no significant difference; *P < 0.05, **P < 0.01.
To delete Mdga2 selectively in CA1 pyramidal neurons (PNs), we stereotactically delivered AAV2/9-CaMKII-Cre-mCherry (CA1-Mdga2-cKO) or AAV2/9-CaMKII-mCherry (control) bilaterally into the CA1 of Mdga2 floxed/floxed (Mdga2f/f) mice at the age of 5–6 weeks (Fig. 1C, D). Staining with an anti-CaMKII antibody revealed that the AAV targeting, assessed by mCherrry expression, was exclusively in CaMKII+ excitatory neurons, and the efficiency of viral infection in CaMKII+ neurons was ~ 83% (Fig. S2A, B). The spread of the virus (mCherry) was largely restricted to CA1 (Fig. S2C). To assess whether Mdga2 was successfully deleted in CA1 after Cre-recombinase expression, real-time quantitative PCR (qPCR), in situ hybridization, and western blotting (WB) was performed 6–7 weeks after the viral injections (Fig. 1C). We found that the mRNA expression of MDGA2 was partially reduced in the hippocampus or CA1 of Mdga2f/f mice injected with AAV2/9-CaMKII-Cre-mCherry compared to Mdga2f/f mice injected with AAV2/9-CaMKII-mCherry (Figs 1E and S2D). Immunoblots also demonstrated that MDGA2 protein levels were significantly decreased in the dorsal hippocampus after stereotactic Cre-recombinase administration, while the level of another homologous MDGA protein, MDGA1, remained unchanged (Fig. 1F, G). In addition, other associated synaptic organizing proteins, such as NL1, NL2, and NL3, also remained unchanged after stereotactic Cre-recombinase administration (Fig. S3A–D), suggesting the expression of these synapse organizers was not affected. Thus, these results indicated that Mdga2 was successfully deleted in CA1 PNs following Cre-recombinase expression.
Adult Knockout of Mdga2 Increases the Excitability of CA1 Pyramidal Neurons
To determine whether Mdga2 deficiency alters neuronal properties, including intrinsic excitability and resting membrane potential (RMP), AAV2/9-CaMKII-Cre-mCherry or AAV2/9-CaMKII-mCherry was administered to 5–6 week-old Mdga2f/f mice. Whole-cell patch-clamp recordings were made to measure the excitability of virus-infected CA1 PNs using ex vivo slice electrophysiology 6–7 weeks later (Fig. 2A, B). APs in virus-infected CA1 PNs were induced by injecting depolarizing currents in incremental steps (Δ = 50 pA, 1 s duration). As shown in Fig. 2C and D, the AP firing rate was significantly greater in the Mdga2-cKO (Cre) mice than in the control (Con) mice, while the RMP (Fig. 2E) remained unaltered. The input resistance (Rin) was significantly increased (Fig. 2F), suggesting that the CA1 PNs are more excitable in Mdga2 cKO mice. Further analysis of the active membrane properties showed that the fAHP (Fig. 2G), AP threshold (Fig. 2H), AP amplitude, and AP half width (Fig. 2I, J) were also unaltered after loss of MDGA2. To confirm this result, another protocol was applied to measure neuronal firing upon the injection of ramped depolarizing currents (Fig. 2K). Consistent with the previous result, the loss of MDGA2 increased AP numbers recorded in parallel with a decrease in the rheobase (Fig. 2L, M). These results suggest that CA1 PNs are hyperactive after Mdga2 deficiency.
Adult knockout of Mdga2 in hippocampal CA1 pyramidal neurons elevates neuronal excitability. A Schematic of the experimental procedure. B Patching of virus-infected CA1 PNs under transmitted (left) and fluorescent light microscopy (right). Scale bar, 10 μm. C Representative traces of action potential (AP) firing in response to injection of current in neurons from control (Con) and CA1-Mdga2-cKO (Cre) mice. D Mdga2-cKO in CA1 PNs increases the number of APs against the injected current strength. n = 29 neurons, 3 mice for control (Con) group; n = 22 neurons, 3 mice for Mdga2-cKO (Cre) group; P < 0.0001; F(1, 350) = 40.48; two-way ANOVA. E Mdga2-deficiency in CA1 PNs had no effect on resting membrane potential (RMP). n = 29 neurons, 3 mice for control (Con) group; n = 22 neurons, 3 mice for Mdga2-cKO (Cre) group; P = 0.1183, unpaired t test. F Mdga2-deficiency in CA1 PNs increases the input resistance (Rin). n = 29 neurons, 3 mice for control (Con) group; n = 22 neurons, 3 mice for Mdga2-cKO (Cre) group; P = 0.0483, unpaired t test. G Mdga2 deficiency in CA1 PNs does not affect fAHP. n = 29 neurons, 3 mice for control (Con) group; n = 22 neurons, 3 mice for Mdga2-cKO (Cre) group; P = 0.5599, unpaired t test. H Mdga2 deficiency in CA1 PNs has no effect on the AP threshold. n = 29 neurons, 3 mice for control (Con) group; n = 22 neurons, 3 mice for Mdga2-cKO (Cre) group; P = 0.1591, unpaired t test. I Mdga2-deficiency in CA1 PNs does not affect AP amplitude. n = 29 neurons, 3 mice for control (Con) group; n = 22 neurons, 3 mice for Mdga2-cKO (Cre) group; P = 0.3953, unpaired t test. J Mdga2-deficiency in CA1 PNs has no effect on AP half-width. n = 29 neurons, 3 mice for control (Con) group; n = 22 neurons, 3 mice for Mdga2-cKO (Cre) group; P = 0.3730, unpaired t test. K Representative spiking traces showing the firing of CA1 PNs upon the injection of a depolarizing current ramp (0–250 pA, 1 s). L Summary plots of AP numbers with ramped depolarizing current injected into CA1 pyramidal neurons, as in K. n = 27 neurons, 3 mice for control (Con) group; n = 22 neurons, 3 mice for Mdga2-cKO (Cre) group; unpaired t test, P = 0.0107. M Mdga2-deficiency in CA1 PNs decreases the rheobase. n = 27 neurons, 3 mice for control (Con) group; n = 22 neurons, 3 mice for Mdga2-cKO (Cre) group; P = 0.0130, unpaired t test. All data are presented as the mean ± SEM. n.s., no significant difference; *P < 0.05, **P < 0.01.
Adult Knockout of Mdga2 in CA1 Pyramidal Neurons Enhances Excitatory but not Inhibitory Synaptic Transmission
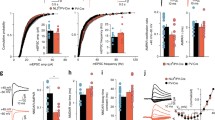
To begin to probe the contribution of MDGA2 to synaptic function within a distinct cell population, whole-cell voltage-clamp recordings were made in CA1 PNs in acute brain slices from Mdga2-cKO and control mice. As shown in Fig. 3A, compared with controls, MDGA2 reduction significantly increased the frequency and amplitude of mEPSCs in the Mdga2-cKO mice (Fig. 3B, C). Consistent with these effects, a significant leftward shift in the cumulative probability curve of the mEPSC frequency (Fig. 3B) and a rightward shift in amplitude curves (Fig. 3C) were found. Additional exploration of effects on inhibitory transmission onto CA1 PNs was assayed by recording mIPSCs. As shown in Fig. 3D, Mdga2-cKO did not detectably change the mean and cumulative probability curve of the frequency (Fig. 3E) or amplitude (Fig. 3F) of mIPSCs. Further analysis of mEPSC and mIPSC kinetics (rise time, decay, and half-width; Fig. S4A–F) showed that the kinetics of mEPSCs and mIPSCs in CA1 PNs were not altered by late Mdga2 knockout. Moreover, the frequency and the amplitude of sEPSCs were also increased in Mdga2-cKO mice (Fig. 3G–I), while there were no detectable changes in the frequency and the amplitude of sIPSCs (Fig. 3J–L). Thus, the reduction of MDGA2 in CA1 PNs selectively enhances excitatory but not inhibitory synaptic transmission.
Adult Mdga2 knockout in hippocampal CA1 pyramidal neurons elevates excitatory but not inhibitory synaptic strength. A Representative mEPSC traces in CA1 pyramidal neurons. B Mdga2-cKO (Cre) neurons exhibit a significant increase in mEPSC frequency compared with control (Con) neurons. n = 14 neurons, from 3 mice for Con, n = 21 neurons, from 4 mice for Cre; P = 0.0044, unpaired t test. This increase results in a leftward shift in the cumulative probability curve corresponding to reduced inter-event intervals. C Mdga2-cKO (Cre) neurons exhibit a significant increase in mEPSC amplitude compared with control (Con) neurons; P = 0.0374, unpaired t test. D Representative mIPSC traces. E mIPSC frequency does not differ between the Cre and Con groups. n = 27 neurons, from 3 mice for Con, n = 20 neurons, from 3 mice for Cre; P = 0.4632, unpaired t test. F mIPSC amplitude does not differ between the Cre and Con groups; P = 0.2137, unpaired t test. G Representative sEPSC traces in CA1 pyramidal neurons. H Mdga2-cKO (Cre) neurons exhibit a significant increase in sEPSC frequency compared with control (Con) neurons. n = 13 neurons, from 4 mice for Con, n = 18 neurons, from 4 mice for Cre; P = 0.0039, unpaired t test. This increase results in a leftward shift in the cumulative probability curve corresponding to reduced inter-event intervals. I Mdga2-cKO (Cre) neurons exhibit a significant increase in sEPSC amplitude compared with control (Con) neurons; P = 0.0464, unpaired t test. J Representative sIPSC traces. K sIPSC frequency does not differ between the Cre and Con groups. n = 12 neurons, from 3 mice for Con, n = 11 neurons, from 3 mice for Cre; P = 0.8096, unpaired t test. L sIPSC amplitude does not differ between the Cre and Con groups; P = 0.3698, unpaired t test. All data are presented as the mean ± SEM. n.s., no significant difference; *P < 0.05, **P < 0.01.
Adult Knockout of Mdga2 in CA1 Pyramidal Neurons Increases Excitatory but not Inhibitory Synapse Density in CA1
The increased frequency of mEPSCs and sEPSCs suggests that adult KO of Mdga2 may selectively enhance the function or density of glutamatergic pre-synaptic terminals in CA1. However, no differences were found in paired-pulse facilitation (Fig. 4A, B) or the expression of synapsin1/2 (Fig. S5A, B), suggesting that the increased frequency of mEPSCs and sEPSCs was not due to changes in neurotransmitter release probability. As MDGA2 has been reported to selectively regulate the formation of synapses [18, 19, 22], we next assessed whether the mature PNs lacking MDGA2 demonstrate changes in synaptic density.
Adult knockout of Mdga2 in mature CA1 neurons increases excitatory but not inhibitory synapse density in CA1. A Representative traces of paired-pulse stimulation at a 50 ms. B Paired-pulse facilitation (PPF) is normal in Mdga2-cKO (Cre) mice. n = 9 neurons, from 3 mice for control (Con) group; n = 7 neurons from 3 mice for Mdga2-cKO (Cre) group; P = 0.7652; F(1, 56) = 0.0901; two-way ANOVA. C Immunoblot and quantification of VGluT1 protein in AAV-infected CA1 6–7 weeks after viral injections. P2, synaptosome pellet. Values for control mice were taken as 1; n = 3 mice per group; P = 0.0295, unpaired t test. β-actin was used as a loading control. D Immunoblot and quantification of VGAT protein in AAV-infected CA1 6–7 weeks after viral injections. Values for control mice were taken as 1; n = 3 mice per group; P = 0.4538, unpaired t test. β-actin was used as a loading control. E Representative confocal images from CA1 layers (SO, SR, SLM) of control (Con) and Mdga2-cKO (Cre) brain sections immunolabeled with the excitatory presynaptic marker VGluT1. SO, stratum oriens; SP, stratum pyramidale; SR, stratum radiatum; SLM, stratum lacunosum moleculare. Scale bar, 10 μm. F Quantification of the total area of vGluT1 shows a significant increase in the CA1 layers (SO, SR, SLM) of Mdga2-cKO (Cre) mice. n = 6 mice per group; SO, P < 0.01; SR, P < 0.001; SLM, P < 0.01; unpaired t test. G Representative confocal images from CA1 layers (SO, SP, SR, SLM) of control (Con) and Mdga2-cKO (Cre) brain sections immunolabeled with the inhibitory presynaptic marker VGAT. Scale bar, 10 μm. H Quantification of the total area of VGAT shows no change in the CA1 layers (SO, SP, SR, SLM) of Mdga2-cKO (Cre) mice. n = 6 mice per group; SO, P = 0.2736; SP, P = 0.7757; SR, P = 0.7190; SLM, P = 0.7935; unpaired t test. All data are presented as the mean ± SEM. n.s., no significant difference; *P < 0.05, **P < 0.01, ***P < 0.001.
To achieve this, the protein expression of VGlut1 and VGAT, representing glutamatergic and GABAergic presynaptic terminals respectively were analyzed and an immunohistochemical analysis of VGlut1 and VGAT was performed in coronal brain sections. Immunoblots demonstrated that VGluT1 levels increased but VGAT protein expression remained unchanged (Fig. 4C, D). In addition, the punctate immunofluorescence of VGlut1 was significantly increased in the stratum oriens (SO), stratum radiatum (SR), and stratum lacunosum-moleculare (SLM) in sections from Mdga2-cKO mice, relative to control mice (Fig. 4E, F). However, quantitation of VGAT revealed no changes in the VGAT+ area in any CA1 layers (SO, SP, SR, and SLM), compared with the control mice (Fig. 4G, H). Similarly, quantitation of GAD65, another marker for GABAergic terminals, also revealed no changes in the Mdga2-cKO (Cre) mice (Fig. S6A, B). These results were consistent with the findings from western blot experiments, demonstrating an increase of glutamatergic but not GABAergic synapses in CA1 following Mdga2 knockout.
Furthermore, we also assessed the expression of the postsynaptic scaffolding proteins gephyrin and PSD95, in addition to the GluA1 AMPAR and GluN1 NMDAR subunits in the dorsal hippocampus, after stereotactic Cre-recombinase administration (Fig S5A, C–F). However, no differences were found between the two groups. Collectively, these results suggest that Mdga2 selectively limits glutamatergic inputs on CA1 PNs, at least in part by controlling the density of excitatory but not inhibitory synapses in this region.
Adult Knockout of Mdga2 Selectively Alters AMPAR-Mediated Synaptic Transmission and Impairs Long-Term Potentiation in CA1 Pyramidal Neurons
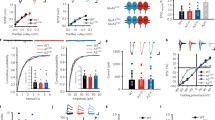
To determine whether synaptic plasticity was altered following adult knockout of Mdga2 in CA1 PNs, LTP was assessed in virus-infected neurons from the Mdga2-cKO and control mice. Stimulation of CA3 PN axon collaterals in CA1 evoked EPSPs subthreshold for evoking APs (Fig. 5A). After recording a baseline (15 min), we delivered three episodes of theta burst stimulation (TBS), each stimulus train consisting of 5 shocks at 100 Hz delivered 5 times at 5 Hz (75 stimuli in total; see Fig. 5B schematic). Comparisons of the LTP magnitude during the last 15 min of recording revealed that the maintenance of LTP was impaired in MDAG2-cKO mouse slices relative to controls (Fig. 5C–E).
Adult knockout of Mdga2 in hippocampal CA1 pyramidal neurons impairs long-term potentiation (LTP) and increases basal synaptic transmission. A Schematic of a hippocampal brain slice for LTP experiments, showing the positioning of recording and stimulating electrodes. SC, Schaffer collaterals. B Summary of the induction protocol for compressed theta burst stimulation (cTBS). C Averaged EPSPs recorded before (1, dashed line) and after (2, solid line) LTP induction, showing the interval (3 ms) used to measure the initial slope. D LTP is impaired in Mdga2-cKO (Cre) neurons. n = 15 neurons, from 5 mice for the control (Con) group; n = 9 neurons, from 5 mice for Mdga2-cKO (Cre) group. P < 0.0001; F(1, 880) = 115.6; two-way ANOVA. E Summary of the mean LTP magnitude at 30–45 min (2) after LTP induction for panel B. n = 15 neurons, from 5 mice for the control (Con) group; n = 9 neurons, from 5 mice for Mdga2-cKO (Cre) group; P = 0.0030, unpaired t test. F Input/output curves to measure relative basal synaptic strength in control (Con) and Mdga2-cKO (Cre) neurons. Example traces while increasing stimulation as a multiple of threshold (~ 15 pA) stimulation. n = 10 neurons, from 3 mice for control (Con) group; n = 10 neurons, from 4 mice for Mdga2-cKO (Cre) group; the data were analyzed in separate t-tests at each stimulus intensity; *P < 0.05. G The AMPA to NMDA ratio is elevated in Mdga2-cKO (Cre) neurons. Responses were recorded at − 70 mV for AMPA and + 40 mV for NMDA receptor-mediated currents. n = 14 neurons, from 3 mice for the control (Con) group; n = 18 neurons, from 4 mice for Mdga2-cKO (Cre) group. P = 0.0039, unpaired t test. H Representative traces of evoked EPSCs in CA1 PNs when the holding potential was increased from − 80 to + 40 mV in 20-mV steps. I and J Mdga2-cKO has no effect on the current-voltage relationships of AMPAR-mediated EPSCs and NMDAR-mediated EPSCs. Current amplitudes were normalized to that recorded at − 80 (I) or + 40 mV (J). n = 11 neurons, from 3 mice for the control (Con) group; n = 11 neurons, from 4 mice for Mdga2-cKO (Cre) group. All data are presented as the mean ± SEM, *P < 0.05, **P < 0.01.
Previous reports have suggested that Mdga2 deletion may selectively alter different components of synaptic transmission [32], which may reflect developmentally restricted roles for NL1 in modulating synaptic currents. To establish the cell-selective impact of MDGA2 loss on glutamatergic transmission, we tested the input/output properties of Mdga2-deficient CA1 neurons by applying graded increases in stimulus intensity while recording isolated AMPAR currents. The responses of AMPAR-dependent EPSCs were significantly elevated in Mdga2-cKO neurons relative to control neurons (Fig. 5F). To further test if the AMPAR component was uniquely affected, we assessed the AMPA/NMDA ratio, which was also significantly elevated in Mdga2-cKO neurons (Fig. 5G). Next, we also measured the input-output curves of NMDAR but failed to detect any difference between Mdga2-cKO neurons and control neurons (Fig S7A, B). Analysis of I/V curves for AMPA and NMDA receptor currents demonstrated that the basal function of these receptors appeared to be intact, and the shift in the AMPA/NMDA ratio did not reflect compromised NMDAR function (Fig. 5H–J). Thus, adult knockout of Mdga2 in CA1 PNs impaired LTP, while upregulating the AMPAR component of evoked synaptic transmission.
Adult Knockout of Mdga2 in CA1 Pyramidal Neurons Impairs Recognition and Contextual Fear Memory, and Reduces Social Interactions
To assess the behavioral impact of the loss of Mdga2 from CA1 PNs, we applied a series of behavioral analyses 6–8 weeks after viral infection (Fig. 6A). Mice were first subjected to the open field test. The results showed both the Mdga2-cKO and control mice had the normal motor ability and spent similar amounts of time occupying the center of the open field (Fig. 6Bi–Biii), indicative of preserved locomotor function and normal anxiety levels. Recognition memory was assessed using novel object recognition, a simple, low-stress task (Fig. 6Ci). Whereas control mice spent considerably more time with the novel object than the familiar object (Fig. 6Cii), Mdga2-cKO mice failed to distinguish between the familiar and novel objects (Fig. 6Ciii). Accordingly, Mdga2-cKO mice showed a lower recognition index, suggesting an impaired ability to distinguish between familiar and novel objects (Fig. 6Civ).
Adult knockout of Mdga2 in CA1 pyramidal neurons impairs recognition and contextual fear memory and reduces social interactions but not social novelty preference. A Experimental paradigm for behavioral testing. Bi Representative traces of 10 min in open field tests. Bii No difference in time spent in the center between the two groups. n = 8 mice for control (Con), n = 9 mice for Mdga2-cKO (Cre); P = 0.6358, unpaired t test. Biii No difference in total distance traveled between the two groups. n = 8 mice for control (Con), n = 9 mice for Mdga2-ccKO (Cre); P = 0.6365, unpaired t test. Ci Schematic of the novel object recognition test. Cii Control mice spend significantly more time with the novel object. n = 8 mice for Con; P = 0.0064, unpaired t test. Ciii No difference in time spent with the novel and familiar object in Mdga2-cKO (Cre) mice; n = 9 mice for Cre; P = 0.4247, unpaired t test. Civ Reduced recognition index in Mdga2-cKO (Cre) mice; n = 8 mice for Con, n = 9 mice for Cre; P = 0.0300, unpaired t test. Di Schematic of the contextual fear conditioning test. Dii The overall ANOVA was significant but post hoc analysis demonstrated that individual shock trial performance was not different between groups across trials in the curve of fear acquisition. n = 8 mice for Con, n = 9 mice for Cre; F(1, 60) = 2.633, P = 0.1099; two-way ANOVA. Diii Mdga2-cKO (Cre) mice show a significant reduction in freezing time when placed back in the test chamber 24 h after training. n = 8 mice for Con, n = 9 mice for Cre; P = 0.0186, unpaired t test. Ei Schematic of the three-chamber social interaction test. Eii Average heat map of time spent in the social interaction test and social novelty preference test for control and Mdga2-cKO (Cre) mice. Eiii Control mice prefer Stranger 1 (S1) mice over the Middle (M) and Empty (E) chambers, whereas Mdga2-cKO (Cre) mice spend equal time in either the S1 or empty chambers. n = 8 mice for control (Con), n = 9 mice for Mdga2-cKO (Cre); F(2, 45) = 29.66, two-way ANOVA. Eiv Both control and Mdga2-cKO (Cre) mice prefer Stranger 2 (S2) mice over the Middle (M) and Stranger 1 (S1) mouse chambers. n = 8 mice for control (Con), n = 9 mice for Mdga2-cKO (Cre); F(2, 45) = 49.27, two-way ANOVA. Ev The total distance traveled by both groups is comparable during the social interaction test. n = 8 mice for control (Con), n = 9 mice for Mdga2-cKO (Cre); P = 0.7561, unpaired t test. All data are presented as the mean ± SEM. n.s., no significant difference; *P < 0.05, **P < 0.01, ****P < 0.0001.
Hippocampus-dependent memory was also measured using contextual fear conditioning (Fig. 6Di). Mdga2-cKO mice learned to associate the conditioning chamber with a footshock at rates similar to control mice, showing increased freezing at early time points (Fig. 6Dii). However, Mdga2-cKO mice demonstrated impaired contextual memory performance at 24 h following training compared with control mice (Fig. 6Diii). Collectively, these results indicate that the adult knockout Mdga2 in CA1 PNs is sufficient to induce deficits in a cognitive function requiring the hippocampus.
Given that social aversion is a hallmark of autism, and gene linkage analysis has linked Mdga2 to neurodevelopmental disorders, we next assessed the impact of the adult KO of Mdga2 in CA1 PNs on social behavior. Virus-infected mice were examined in the three-chamber test, which measures the preference for interacting with a novel conspecific relative to an empty chamber (Fig. 6Ei). Control mice spent significantly more time with social targets (a stranger mouse, S1), compared with middle and empty chambers (E). Conversely, Mdga2-cKO mice failed to show a preference for social interactions, consistent with a general social aversion (Fig. 6Eii, Eiii). Interestingly, this deficit did not extend to social novelty as both Mdga2-cKO and control mice spent more time with another stranger mouse (Stranger 2; novel mouse) compared to the middle chamber or Stranger 1 (familiar mouse) in a social novelty test (Fig. 6Eiv). The total distance traveled for both groups was comparable, suggesting preference was not influenced by locomotor function (Fig. 6Ev). Thus, although Mdga2-cKO mice demonstrated impaired social interactions, social novelty preference and locomotor function were intact. Collectively, these data suggest that the adult knockout of Mdga2 in CA1 PNs impairs engagement in certain types of social interactions, which may be offset by social novelty.
Discussion
MDGA2 appears to be essential for proper brain development, as full knockout significantly reduces pup survival rates [22]. This necessitates using heterozygotes for exploring the functions of MDGA2 in young adult mice, subject to compensatory mechanisms, further complicated by residual MDGA2 that would be present in heterozygotic neurons. Moreover, diffuse expression across cell types further limits the interpretation of results, as this approach fails to differentiate cell-type selective versus network and developmental effects. Here, we addressed these deficiencies using a “late” conditional knockout strategy to delete Mdga2 from CA1 pyramidal neurons postnatally.
Consistent with previous reports using global knockout mice [22], mEPSC frequency was increased at glutamatergic inputs converging on neurons lacking MDGA2, with no detectable change in mIPSC properties. Moreover, increased sEPSC frequency and amplitude with unchanged sIPSC properties were also recorded. In addition, we also found increased excitability in CA1 PNs lacking MDGA2, which may be partially due to the increased input resistance and decreased rheobase. No changes were found in NL2 protein levels nor the inhibitory scaffold protein, gephyrin. Similarly, LTP at Schaffer collateral inputs onto this region were impaired, likely contributing to the recognition and contextual memory deficits in Mdga2-cKO mice. Our data are consistent with an ongoing and selective requirement for MDGA2 in maintaining suppression of glutamatergic synapses onto CA1 pyramidal neurons.
As a first step in determining why LTP is impaired, measures of the AMPA/NMDA ratio demonstrated a marked upregulation of the AMPAR component. However, WB analysis failed to detect changes in the canonical glutamatergic receptor subunits crucial for LTP (GluA1, GluN1) and an I/V analysis suggested that NMDAR function is preserved. Furthermore, no change was found in the input-output relationship of NMDAR-mediated EPSCs. This finding is consistent with recent studies showing that Mdga2 knock-out increases AMPAR-mediated EPSCs without affecting NMDAR-mediated EPSCs [32]. Accordingly, impaired synaptic plasticity may not be the result of impaired NMDARs, which are required for the induction of LTP, but rather may reflect a synaptic saturation resulting from exaggerated AMPAR currents.
Notably, others have demonstrated that Mdga2 KD decreases NMDAR EPSCs while leaving AMPAR currents unaltered [35]. However, these results may reflect the use of different species (mice versus rats), with different knockout strategies in different growth stages, which may lead to divergent outcomes. Interestingly, initial analysis in Mdga2+/− mice showed a nuanced plasticity phenotype, with single-train LTP enhanced and multiple-train LTP impaired. Theta-burst LTP was not tested in this model, however, the trend towards altered plasticity is consistent with a central role for MDGA2 within CA1 PNs for optimizing the conditions for LTP. Moreover, the levels of GluA1 were increased in Mdga2+/− mice which was not found in the current cKO model. This suggests that during critical periods of early development, MDGA2 may regulate the establishment of the synaptic proteome, but is not required for ongoing maintenance of the complement of at least some plasticity-related proteins. Overall, these findings provide evidence that MDGA2 helps sustain network properties within a critical operating range for normal plasticity and associated cognitive performance. Given the prominent role of LTP-like mechanisms in memory formation [36,37,38], the LTP deficit within CA1 is likely a major contributing factor to the recognition and contextual memory deficits found here.
A hallmark feature of autism is aversion to social engagement. Late Mdga2-cKO revealed a functional divergence in the requirement for MDGA2 in social interactions relative to social novelty preference, with Mdga2-cKOs avoiding social interactions but still preferring a novel conspecific in the social preference test. A maintained preference for novelty coupled with a lack of differences between controls and Mdga2-cKOs in the open field test supports that anxiety (either novelty-induced or an intrinsic predisposition) did not contribute to the social deficits, indicating that the behavioral change was restricted to the social domain. In addition, the partial rescue of social preference in the social novelty task provides evidence that these types of experiences (meeting novel conspecifics) may counter tendencies towards social aversion in an autism model.
Several points of divergence between Mdga2 germline KO and cKO mice advance our understanding of the constitutive versus ongoing requirements for MDGA2 in neural circuits. VGlut1 is upregulated in both Mdga2+/− mice and the cKO mouse SR, whereas this effect extends to the SO and SLM in cKOs. In addition, NL1 and PSD-95 family MAGUKs were elevated in Mdga2+/− mice, whereas expression of these proteins showed an increasing trend with no significant differences in cKO mice. These differences suggest that germline knockout of Mdga2 may exacerbate changes in synaptic protein complement that are determined during early periods of brain development. The conserved increase in mEPSC frequency and amplitude suggests that MDGA2 plays an ongoing and prominent role in establishing basal excitatory synaptic strength early in development that extends into adulthood, with a parallel role in limiting glutamatergic synapse number across developmental stages [32].
Interestingly, MDGA1 and MDGA2 suppress inhibitory synapse development in cultured neurons [18, 19, 22]. However, mIPSC properties and GABAergic presynaptic development appeared normal in mature neurons lacking MDGA2. This is consistent with the constitutive loss of MDGA2 in heterozygotic neurons which resulted in a selective de-repression of excitatory synapse development. These discrepancies suggest that developmental stage, network-driven functional divergence, or synaptogenic co-factors help tune the regulatory roles for MDGA2 under different experimental conditions. Recent evidence suggests that MDGA1 suppresses APP-induced GABAergic synapse regulation through direct interactions with presynaptic APP [33]. Furthermore, the catecholaminergic metabolite homovanillic acid is upregulated in the Mdga1−/− mouse hippocampus and cortex, indicating increased dopamine turnover [39]. These findings suggest that the presence of co-factors or neuromodulatory inputs in slice conditions could bias MDGA2 towards glutamatergic inputs on pyramidal neurons.
Recent evidence has also shown that MDGAs are diffusely expressed in dendrites, are highly mobile, and reside in extra/perisynaptic spaces where they may sequester NLs in an activity-dependent fashion [32]. Along with the current study, further investigation is required to identify the molecular correlates of these constraints, how neuronal activity modulates MDGA activity patterns, and how these organizers operate in different cell populations (interneurons, astrocytes) to set the parameters for synapse formation and function.
The synaptic plasticity and cognitive impairments reported here are consistent with an ongoing requirement for MDGA2 within CA1 PNs for optimizing network dynamics for encoding new experiences in mature brain circuits. Both recognition and contextual memory require processing within CA1 for encoding [40,41,42,43,44], and the LTP deficits indicate that activity-dependent synapse changes are insufficient for supporting memory. Interestingly, NL1 overexpression results in a similar deficit in LTP [45], which was interpreted as the result of exaggerated spine maturation which would render spines, the primary sites of glutamatergic synaptic contact, less capable of adaptation in response to synaptic activity. In addition, the TBS-induced LTP is predicated, in part, on a disinhibition mechanism that facilitates postsynaptic depolarization, leaving open the untested question as to whether the shift in E/I balance found here changes the kinetics of synaptic responses such that the TBS-driven disinhibition is muted, thereby limiting the depolarization necessary for LTP. Collectively, these results suggest that selective removal of MDGA2 in the adult brain recapitulates several of the key cellular and behavioral phenotypes found in constitutive knockouts while identifying clear distinctions across conditions that advance our understanding of MDGA2 function. Accordingly, our data provide primary evidence that MDGA2 regulates glutamatergic inputs converging on CA1 PNs, and synaptic plasticity in fully developed networks, helping maintain circuit properties within an optimal range for normal cognitive function and social behavior.
Although MDGA2 has been localized to both inhibitory and excitatory synapses, functional changes of GABAergic synapses within area CA1 following MDGA2 reduction have not been detected. Interestingly, we previously found that knocking down MDGA1 in neurons derived from Mdga2+/− mice results in an increase in mIPSCs [26], suggesting that MDGA1 may be compensating under conditions in which MDGA2 is reduced. Alternatively, recent findings indicate that MDGAs play developmentally restricted roles that switch during maturation, with MDGA1 partially redistributing from glutamatergic to GABAergic synapses over time in rat hippocampal cell cultures [36]. It will be interesting to determine if MDGA2 likewise demonstrates maturational redistribution or developmentally restricted roles at select synapse types. Finally, it remains to be determined whether analyses focused on interneurons or astrocytes will identify a more central role for MDGA2 in mediating GABAergic synapses that are formed on or supported by these alternative cell types.
Important questions remain surrounding the connection between MDGA2-mediated cellular alterations and the behavioral readouts of impaired memory and social deficits. At a qualitative level, increased overall excitability and increased synaptic noise would reduce the fidelity of information processed within these circuits, perhaps disrupting the high-resolution information processing necessary for discriminating objects, forming memories, or engaging in social interactions. Future studies are needed to determine if reducing neuronal excitability or normalizing synaptic properties (density, function, or plasticity) results in a corresponding rescue of behavioral outputs.
MDGAs play roles in maintaining proper synaptic patterning in the developing brain. Our data extends the known range of MDGA influence to mature neural circuits. Along with previous studies, our findings are consistent with developmentally restricted roles for MDGA2, suggesting it limits excitation and maintains synaptic balance in adult pyramidal neurons, whereas functions early in development appear to involve sustaining the density, strength, and synaptic protein composition of newly forming synapses. Given that loss of function mutations in MDGAs increase the risk for ASD, schizophrenia, and other neuropsychiatric disorders [23, 28, 29], our findings have important implications for understanding the synaptic basis of these disorders and future treatments. Our findings potentially narrow the focus of therapeutic targeting strategies going forward, suggesting that MDGA2 reconstitution selectively within CA1 PNs is a viable target for restoring network and cognitive function in carriers of MDGA2 mutations.
References
Südhof TC. Synaptic neurexin complexes: A molecular code for the logic of neural circuits. Cell 2017, 171: 745–769.
Siddiqui TJ, Craig AM. Synaptic organizing complexes. Curr Opin Neurobiol 2011, 21: 132–143.
Siddiqui TJ, Pancaroglu R, Kang Y, Rooyakkers A, Craig AM. LRRTMs and neuroligins bind neurexins with a differential code to cooperate in glutamate synapse development. J Neurosci 2010, 30: 7495–7506.
Craig AM, Kang Y. Neurexin-neuroligin signaling in synapse development. Curr Opin Neurobiol 2007, 17: 43–52.
Cao X, Tabuchi K. Functions of synapse adhesion molecules neurexin/neuroligins and neurodevelopmental disorders. Neurosci Res 2017, 116: 3–9.
Südhof TC. Neuroligins and neurexins link synaptic function to cognitive disease. Nature 2008, 455: 903–911.
Varoqueaux F, Aramuni G, Rawson RL, Mohrmann R, Missler M, Gottmann K. Neuroligins determine synapse maturation and function. Neuron 2006, 51: 741–754.
Graf ER, Zhang X, Jin SX, Linhoff MW, Craig AM. Neurexins induce differentiation of GABA and glutamate postsynaptic specializations via neuroligins. Cell 2004, 119: 1013–1026.
Pettem KL, Yokomaku D, Luo L, Linhoff MW, Prasad T, Connor SA, et al. The specific α-neurexin interactor calsyntenin-3 promotes excitatory and inhibitory synapse development. Neuron 2013, 80: 113–128.
Siddiqui TJ, Tari PK, Connor SA, Zhang P, Dobie FA, She K, et al. An LRRTM4-HSPG complex mediates excitatory synapse development on dentate gyrus granule cells. Neuron 2013, 79: 680–695.
Budreck EC, Scheiffele P. Neuroligin-3 is a neuronal adhesion protein at GABAergic and glutamatergic synapses. Eur J Neurosci 2007, 26: 1738–1748.
Chubykin AA, Atasoy D, Etherton MR, Brose N, Kavalali ET, Gibson JR, et al. Activity-dependent validation of excitatory versus inhibitory synapses by neuroligin-1 versus neuroligin-2. Neuron 2007, 54: 919–931.
Song JY, Ichtchenko K, Südhof TC, Brose N. Neuroligin 1 is a postsynaptic cell-adhesion molecule of excitatory synapses. Proc Natl Acad Sci U S A 1999, 96: 1100–1105.
Varoqueaux F, Jamain S, Brose N. Neuroligin 2 is exclusively localized to inhibitory synapses. Eur J Cell Biol 2004, 83: 449–456.
Irie M, Hata Y, Takeuchi M, Ichtchenko K, Toyoda A, Hirao K, et al. Binding of neuroligins to PSD-95. Science 1997, 277: 1511–1515.
Levinson JN, Chéry N, Huang K, Wong TP, Gerrow K, Kang R, et al. Neuroligins mediate excitatory and inhibitory synapse formation: Involvement of PSD-95 and neurexin-1beta in neuroligin-induced synaptic specificity. J Biol Chem 2005, 280: 17312–17319.
Fertan E, Wong AA, Purdon MK, Weaver ICG, Brown RE. The effect of background strain on the behavioral phenotypes of the Mdga2+ /- mouse model of autism spectrum disorder. Genes Brain Behav 2021, 20: e12696.
Pettem KL, Yokomaku D, Takahashi H, Ge Y, Craig AM. Interaction between autism-linked MDGAs and neuroligins suppresses inhibitory synapse development. J Cell Biol 2013, 200: 321–336.
Lee K, Kim Y, Lee SJ, Qiang Y, Lee D, Lee HW, et al. MDGAs interact selectively with neuroligin-2 but not other neuroligins to regulate inhibitory synapse development. Proc Natl Acad Sci USA 2013, 110: 336–341.
Connor SA, Elegheert J, Xie Y, Craig AM. Pumping the brakes: Suppression of synapse development by MDGA-neuroligin interactions. Curr Opin Neurobiol 2019, 57: 71–80.
Connor SA, Ammendrup-Johnsen I, Kishimoto Y, Karimi Tari P, Cvetkovska V, Harada T, et al. Loss of synapse repressor MDGA1 enhances perisomatic inhibition, confers resistance to network excitation, and impairs cognitive function. Cell Rep 2017, 21: 3637–3645.
Connor SA, Ammendrup-Johnsen I, Chan AW, Kishimoto Y, Murayama C, Kurihara N, et al. Altered cortical dynamics and cognitive function upon haploinsufficiency of the autism-linked excitatory synaptic suppressor Mdga2. Neuron 2016, 91: 1052–1068.
Wang R, Dong JX, Wang L, Dong XY, Anenberg E, Jiang PF, et al. A negative regulator of synaptic development: MDGA and its links to neurodevelopmental disorders. World J Pediatr 2019, 15: 415–421.
Litwack ED, Babey R, Buser R, Gesemann M, O’Leary DD. Identification and characterization of two novel brain-derived immunoglobulin superfamily members with a unique structural organization. Mol Cell Neurosci 2004, 25: 263–274.
Elegheert J, Cvetkovska V, Clayton AJ, Heroven C, Vennekens KM, Smukowski SN, et al. Structural mechanism for modulation of synaptic neuroligin-neurexin signaling by MDGA proteins. Neuron 2017, 96: 242–244.
Gangwar SP, Zhong X, Seshadrinathan S, Chen H, Machius M, Rudenko G. Molecular mechanism of MDGA1: Regulation of neuroligin 2: Neurexin trans-synaptic bridges. Neuron 2017, 94: 1132–1141.e4.
Kim JA, Kim D, Won SY, Han KA, Park D, Cho E, et al. Structural insights into modulation of neurexin-neuroligin trans-synaptic adhesion by MDGA1/neuroligin-2 complex. Neuron 2017, 94: 1121–1131.e6.
Kähler AK, Djurovic S, Kulle B, Jönsson EG, Agartz I, Hall H, et al. Association analysis of schizophrenia on 18 genes involved in neuronal migration: MDGA1 as a new susceptibility gene. Am J Med Genet B Neuropsychiatr Genet 2008, 147B: 1089–1100.
Bucan M, Abrahams BS, Wang K, Glessner JT, Herman EI, Sonnenblick LI, et al. Genome-wide analyses of exonic copy number variants in a family-based study point to novel autism susceptibility genes. PLoS Genet 2009, 5: e1000536.
Bourgeron T. A synaptic trek to autism. Curr Opin Neurobiol 2009, 19: 231–234.
Loh KH, Stawski PS, Draycott AS, Udeshi ND, Lehrman EK, Wilton DK, et al. Proteomic analysis of unbounded cellular compartments: Synaptic clefts. Cell 2016, 166: 1295–1307.e21.
Toledo A, Letellier M, Bimbi G, Tessier B, Daburon S, Favereaux A, et al. MDGAs are fast-diffusing molecules that delay excitatory synapse development by altering neuroligin behavior. Elife 2022, 11: e75233.
Kim J, Kim S, Kim H, Hwang IW, Bae S, Karki S, et al. MDGA1 negatively regulates amyloid precursor protein-mediated synapse inhibition in the hippocampus. Proc Natl Acad Sci USA 2022, 119: e2115326119.
Huang SH, Liu WZ, Qin X, Guo CY, Xiong QC, Wang Y, et al. Association of increased amygdala activity with stress-induced anxiety but not social avoidance behavior in mice. Neurosci Bull 2022, 38: 16–28.
Bemben MA, Sandoval M, Le AA, Won S, Chau VN, Lauterborn JC, et al. Contrasting synaptic roles of MDGA1 and Mdga2. bioRxiv 2023. https://doi.org/10.1101/2023.05.25.542333.
Nabavi S, Fox R, Proulx CD, Lin JY, Tsien RY, Malinow R. Engineering a memory with LTD and LTP. Nature 2014, 511: 348–352.
Whitlock JR, Heynen AJ, Shuler MG, Bear MF. Learning induces long-term potentiation in the hippocampus. Science 2006, 313: 1093–1097.
Bliss TV, Collingridge GL. A synaptic model of memory: Long-term potentiation in the hippocampus. Nature 1993, 361: 31–39.
Hossain MR, Jamal M, Tanoue Y, Ojima D, Takahashi H, Kubota T, et al. MDGA1-deficiency attenuates prepulse inhibition with alterations of dopamine and serotonin metabolism: An ex vivo HPLC-ECD analysis. Neurosci Lett 2020, 716: 134677.
Broadbent NJ, Gaskin S, Squire LR, Clark RE. Object recognition memory and the rodent hippocampus. Learn Mem 2010, 17: 5–11.
Wan H, Aggleton JP, Brown MW. Different contributions of the hippocampus and perirhinal cortex to recognition memory. J Neurosci 1999, 19: 1142–1148.
Parkinson JK, Murray EA, Mishkin M. A selective mnemonic role for the hippocampus in monkeys: Memory for the location of objects. J Neurosci 1988, 8: 4159–4167.
Winocur G, Gilbert M. The hippocampus, context, and information processing. Behav Neural Biol 1984, 40: 27–43.
Abel T, Nguyen PV, Barad M, Deuel TA, Kandel ER, Bourtchouladze R. Genetic demonstration of a role for PKA in the late phase of LTP and in hippocampus-based long-term memory. Cell 1997, 88: 615–626.
Dahlhaus R, Hines RM, Eadie BD, Kannangara TS, Hines DJ, Brown CE, et al. Overexpression of the cell adhesion protein neuroligin-1 induces learning deficits and impairs synaptic plasticity by altering the ratio of excitation to inhibition in the hippocampus. Hippocampus 2010, 20: 305–322.
Acknowledgments
We thank Dr. Ann Marie Craig for her early guidance and vision. This work was supported by the National Natural Science Foundation of China (82001203, 82173819, 81871012, and 81571263), the Scientific Research Fund of Zhejiang Provincial Education Department (Y201839276), the Scientific Research Foundation of Zhejiang University City College (X-202103), the R&D Project of Zhejiang (2022C03034), the Natural Science Foundation of Zhejiang Province (LQ23C090001), and a Canada Research Chair Award (P2018-0246).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors claim that there are no conflicts of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, X., Lin, D., Jiang, J. et al. MDGA2 Constrains Glutamatergic Inputs Selectively onto CA1 Pyramidal Neurons to Optimize Neural Circuits for Plasticity, Memory, and Social Behavior. Neurosci. Bull. 40, 887–904 (2024). https://doi.org/10.1007/s12264-023-01171-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12264-023-01171-1