Abstract
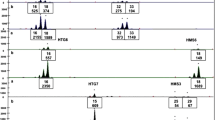
This study investigated the gastrointestinal microbiota in three genetically identical cloned dogs (A, B and C) by somatic cell nuclear transfer. We collected feces from three cloned dogs and their feed to investigate gastrointestinal microbiota using both culture-dependent and culture-independent methods. A total of 962 strains from the feces of cloned dogs were isolated using aerobic and anaerobic culture methods. The dominant microorganisms were Enterococcus faecalis and Enterococcus faecium in all fecal samples. In particular, the fecal sample from cloned dog C had similar proportions of three species (E. faecalis, E. faecium and Lactobacillus murinus). In all, 29 DNA fragments were identified by PCR-denaturing gradient gel electrophoresis (DGGE) analysis. The highest DGGE band intensities were for E. faecalis from cloned dogs A and C and for Clostridium sordellii from cloned dog B, with relative intensities of 15.2, 17.7 and 14.4%, respectively. The other strains identified from the cloned dogs were Chryseobacterium soldanellicola, Escherichia coli, L. murinus, Streptococcus alactolyticus, Weissella confusa and uncultured bacterium. Some microbes isolated from the fecal samples, including C. soldanellicola and W. confuse, were derived from the feed. Overall, gastrointestinal microbiota of all genetically identical cloned dogs, maintained under the same environmental and feeding conditions, showed similar profiles in terms of species diversity analyzed by PCR-DGGE, although there were proportional differences in the amounts of bacterial species. To our knowledge, this is the first report to investigate and compare gastrointestinal microbiota of three genetically identical dogs.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Hong, S. G., M. K. Kim, G. Jang, H. J. Oh, J. E. Park, J. T. Kang, J. O. Koo, T. Kim, M. S. Kwon, B. C. Koo, J. C. Ra, D. Y. Kim, C. Ko, and B. C. Lee (2009) Generation of red fluorescent protein transgenic dogs. Genesis 47: 314–322.
Kim, M. J., H. J. Oh, J. E. Park, G. A. Kim, S. G. Hong, G. Jang, M. S. Kwon, B. C. Koo, T. Kim, S. K. Kang, J. C. Ra, C. Ko, and B. C. Lee (2011) Generation of transgenic dogs that conditionally express green fluorescent protein. Genesis 49: 472–478.
Oh, H. J., M. K. Kim, G. Jang, H. J. Kim, S. G. Hong, J. E. Park, K. Park, C. Park, S. H. Sohn, D. Y. Kim, N. S. Shin, and B. C. Lee (2008) Cloning endangered gray wolves (Canis lupus) from somatic cells collected postmortem. Theriogenol. 70: 638–647.
Christensen, K. L., M. S. Hedemann, H. Jorgensen, J. Stagsted, and K. E. Knudsen (2012) Liquid chromatography-mass spectrometry based metabolomics study of cloned versus normal pigs fed either restricted or ad libitum high-energy diets. J. Proteome Res. 11: 3573–3580.
Clausen, M. R., K. L. Christensen, M. S. Hedemann, Y. Liu, S. Purup, M. Schmidt, H. Callesen, J. Stagsted, and H. C. Bertram (2011) Metabolomic phenotyping of a cloned pig model. BMC Physiol. 11: 14.
Pedersen, R., A. D. Andersen, L. Molbak, J. Stagsted, and M. Boye (2013) Changes in the gut microbiota of cloned and noncloned control pigs during development of obesity: Gut microbiota during development of obesity in cloned pigs. BMC Microbial. 13: 30.
Sutter, N. B. and E. A. Ostrander (2004) Dog star rising: the canine genetic system. Nature reviews. Genetics 5: 900–910.
O’Mahony, D., K. B. Murphy, J. MacSharry, T. Boileau, G. Sunvold, G. Reinhart, B. Kiely, F. Shanahan, and L. O’Mahony (2009) Portrait of a canine probiotic Bifidobacterium—from gut to gut. Vet. Microbial. 139: 106–112.
Neish, A. S. (2009) Microbes in gastrointestinal health and disease. Gastroenterol. 136: 65–80.
Batt, R. M., J. R. Needham, and M. W. Carter (1983) Bacterial overgrowth associated with a naturally occurring enteropathy in the German shepherd dog. Res. Vet. Sci. 35: 42–46.
Biagi, G., I. Cipollini, A. Pompei, G. Zaghini, and D. Matteuzzi (2007) Effect of a Lactobacillus animalis strain on composition and metabolism of the intestinal microflora in adult dogs. Vet. Microbiol. 124: 160–165.
Rondon, M. R., R. M. Goodman, and J. Handelsman (1999) The Earth’s bounty: Assessing and accessing soil microbial diversity. Trends Biotechnol. 17: 403–409.
Schabereiter-Gurtner, C., G. Pinar, W. Lubitz, and S. Rolleke (2001) An advanced molecular strategy to identify bacterial communities on art objects. J. Microbiol. Meth. 45: 77–87.
Gurtner, C., J. Heyrman, G. Pinar, W. Lubitz, J. Swings, and S. Rolleke (2000) Comparative analyses of the bacterial diversity on two different biodeteriorated wall paintings by DGGE and 16S rDNA sequence analysis. Int. Biodeterior. Biodegrad. 229–239.
Moles, L., M. Gómez, H. Heilig, G. Bustos, S. Fuentes, W. de Vos, L. Fernández, J. M. Rodríguez, and E. Jiménez (2013) Bacterial diversity in meconium of preterm neonates and evolution of their fecal microbiota during the first month of life. PloS ONE 8: e66986.
Simoes, C. D., J. Maukonen, J. Kaprio, A. Rissanen, K. H. Pietilainen, and M. Saarela (2013) Habitual dietary intake is associated with stool microbiota composition in monozygotic twins. J. Nutr. 143: 417–423.
Nistal, E., A. Caminero, S. Vivas, J. M. Ruiz de Morales, L. E. Saenz de Miera, L. B. Rodriguez-Aparicio, and J. Casqueiro (2012) Differences in faecal bacteria populations and faecal bacteria metabolism in healthy adults and celiac disease patients. Biochimie 94: 1724–1729.
Jia, J., N. Frantz, C. Khoo, G. R. Gibson, R. A. Rastall, and L. McCartney (2011) Investigation of the faecal microbiota of geriatric cats. Lett. Appl. Microbiol. 53: 288–293.
Cressman, M. D., Z. Yu, M. C. Nelson, S. J. Moeller, M. S. Lilburn, and H. N. Zerby (2010) Interrelations between the microbiotas in the litter and in the intestines of commercial broiler chickens. Appl. Environ. Microbiol. 76: 6572–6582.
Hong, S. W., I. S. Kim, J. S. Lee, and K. S. Chung (2011) Culture-based and denaturing gradient gel electrophoresis analysis of the bacterial community structure from the intestinal tracts of earthworms (Eisenia fetida). J. Microbiol. Biotechnol. 21: 885–892.
Endo, A., Y. Futagawa-Endo, and L. M. Dicks (2009) Lactobacillus and Bifidobacterium diversity in horse feces, revealed by PCR-DGGE. Curr. Microbiol. 59: 651–655.
Jansman, A. J. M. Z., S. J. Koopmans, R. A. Dekker, and H. Smidt (2012) Effects of a simple or a complex starter microbiota on intestinal microbiota composition in caesarean derived piglets. J. Anim. Sci. 90: 433–435.
Rodriguez-Romero, N., L. Abecia, and M. Fondevila (2012) Bacterial profile from caecal contents and soft faeces in growing rabbits given diets differing in soluble and insoluble fibre levels. Anaerobe 18: 602–607.
Kim, M. K., J. G. No, J. J. Park, D. H. Yeom, H. M. Kim, B. H. Choi, D. K. Kim, J. K. Park, and J. G. Yoo (2013) Analysis of growth and hematologic characteristics of cloned puppies. J. Embryo Transf. 28: 229–235.
Jang, G., M. K. Kim, H. J. Oh, M. S. Hossein, Y. H. Fibrianto, S. G. Hong, J. E. Park, J. J. Kim, H. J. Kim, S. K. Kang, D. Y. Kim, and B. C. Lee (2007) Birth of viable female dogs produced by somatic cell nuclear transfer. Theriogenol. 67: 941–947.
Park, J. S., C. J. Sim, and K. D. An (2009) Community structure of bacteria associated with two marine sponges from Jeju island based on 16S rDNA-DGGE profiles. Kor. J. Microbiol. 170–176.
Walter, J., G. W. Tannock, A. Tilsala-Timisjarvi, S. Rodtong, D. M. Loach, K. Munro, and T. Alatossava (2000) Detection and identification of gastrointestinal Lactobacillus species by using denaturing gradient gel electrophoresis and species-specific PCR primers. Appl. Environ. Microbiol. 66: 297–303.
Cetinkaya, Y., P. Falk, and C. G. Mayhall (2000) Vancomycinresistant enterococci. Clin. Microbiol. Rev. 13: 686–707.
Ryan, K. J. and C. G. Ray (2004) Sherris Medical Microbiology. 4th edition ed. McGraw Hill.
Greetham, H. L., C. Giffard, R. A. Hutson, M. D. Collins, and G. R. Gibson (2002) Bacteriology of the Labrador dog gut: A cultural and genotypic approach. J. Appl. Microbiol. 93: 640–646.
Makarova, K., A. Slesarev, Y. Wolf, A. Sorokin, B. Mirkin, E. Koonin, A. Pavlov, and N. Pavlova (2006) Comparative genomics of the lactic acid bacteria. Proc. Natl. Acad. Sci. 15611–15616.
Handl, S., S. E. Dowd, and J. F. Garcia-Mazcorro (2011) Massive parallel 16S rRNA gene pyrosequencing reveals highly diverse fecal bacterial and fungal communities in healthy dogs and cats. FEMS Microbial. Ecol. 76: 301–310.
Chaban, B., M. G. Links, and J. E. Hill (2012) A molecular enrichment strategy based on cpn60 for detection of epsilon-proteobacteria in the dog fecal microbiome. Microb. Ecol. 63: 348–357.
Mentula, S., J. Harmoinen, M. Heikkila, E. Westermarck, M. Rautio, P. Huovinen, and E. Kononen (2005) Comparison between cultured small-intestinal and fecal microbiotas in beagle dogs. Appl. Environ. Microbiol. 71: 4169–4175.
Vanhoutte, T., G. Huys, E. De Brandt, G. C. Fahey, and J. Swings (2005) Molecular monitoring and characterization of the faecal microbiota of healthy dogs during fructan supplementation. FEMS Microbiol. Lett. 249: 65–71.
Konneffer, J. B., Y. Minamoto, and J. S. Suchodolski (2014) Microbiota alterations in acute and chronic gastrointestinal inflammation of cats and dogs. World J Gastroenterol. 20: 16489–16497.
Davis, C. P., D. Cleven, E. Balish, and C. E. Yale (1977) Bacterial association in gastrointestinal tracts of beagle dogs. Appl. Environ. Microbiol. 34: 194–206.
Suchodolski, J. S., C. G. Ruaux, J. M. Steiner, K. Fetz, and D. A. Williams (2005) Assessment of the qualitative variation in bacterial microflora among compartments of the intestinal tract of dogs by use of a molecular fingerprinting technique. Am. J. Vet. Res. 66: 1556–1562.
Stewart, J. A., V. S. Chadwick, and A. Murray (2005) Investigations into the influence of host genetics on the predominant eubacteria in the faecal microflora of children. J. Med. Microbiol. 54: 1239–1242.
Turnbaugh, P. J., M. Hamady, T. Yatsunenko, B. L. Cantarel, A. Duncan, R. E. Ley, M. L. Sogin, W. J. Jones, B. A. Roe, J. P. Affourtit, M. Egholm, B. Henrissat, A. C. Heath, R. Knight, and J. I. Gordon (2009) A core gut microbiome in obese and lean twins. Nature 457: 480–484.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yoo, J.G., Hong, S.W., Kim, JH. et al. Potential application of genetically identical somatic cell nuclear transfer-cloned dogs for gastrointestinal microbiota analysis. Biotechnol Bioproc E 22, 30–36 (2017). https://doi.org/10.1007/s12257-016-0630-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12257-016-0630-6