Abstract
MiRNAs are non-coding RNAs that play important roles in the pathogenesis of human diseases by regulating target gene expression in specific cells or tissues. Previously we identified colorectal cancer (CRC) associated MIR196B, which was specifically up-regulated in CRC cells and tissue. We also identified 18 putative MIR196B target genes by comparing the mRNAs down-regulated in MIR196B-overexpressed cells with MIR196B target genes predicted by public bioinformatics tools. In this study, we verified the association between MIR196B and three genes, HOXA5, HOXB6 and GLTP. HOXA5, HOXB6 and GLTP transcripts were directly down-regulated by MIR196B. The mRNA and proteins levels of HOXA5, HOXB6 and GLTP were also down-regulated in CRC cells by the up-regulated MIR196B. GLTP protein expression was decreased in CRC tissues compared to adjacent non-tumor tissues. These results suggest that HOXA5, HOXB6, and GLTP were direct target genes of MIR196B in CRC cells, and that the up-regulated MIR196B in CRC tissue regulates the expression levels of HOXA5, HOXB6, and GLTP during colorectal carcinogenesis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
MicroRNAs (miRNAs) are endogenously synthesized small noncoding RNA molecules of approximately 19–24 nucleotides. MiRNAs contribute to the regulation of crucial biological processes, such as cell proliferation, apoptosis, differentiation, and angiogenesis by controlling the stability and translation of target mRNAs [1, 2]. They are also implicated in the pathogenesis of various diseases as tumor suppressor genes or oncogenes [3, 4]. Accumulating evidence suggests that miRNAs act as tumor suppressors or oncogenes by targeting genes involved in cell proliferation, survival, apoptosis, and metastasis [5,6,7].
In our previous study, MIR196B was identified as a colorectal cancer (CRC) associated miRNA by miRNA expression profiling of CRC tissues versus healthy colorectal tissues, which showed that its expression was significantly up-regulated in CRC tissues [8]. MIR196B is a member of the MIR196 gene family (MIR196A1, MIR196A2, and MIR196B), which are transcribed from three different genes and are located in the homeobox (HOX) gene cluster regions in humans [9, 10]. The MIR196A1 gene is located on Chr. 17q21.32 between the HOXB9 and HOXB10 genes. The MIR196A2 gene is located between HOXC10 and HOXC9 on Chr. 12q13.13. The gene for MIR196B is located in an evolutionarily conserved region between HOXA9 and HOXA10 on Chr. 7p15.2. The mature nucleotide sequences of MIR196A1 and MIR196A2 are identical, whereas mature MIR196B differs from MIR196A by one nucleotide [10].
We previously detected 18 target genes of MIR196B by comparing the mRNA microarray analysis of MIR196B-overexpressed human colon cells with the bioinformatics-predicted candidate target genes, and suggested that MIR196B regulates FAS cell surface death receptor (FAS)-mediated apoptosis in colorectal cancer cells [8].
In this study, we validated the expression levels of HOXA5, HOXB6, and glycolipid transfer protein (GLTP) in MIR196B-transfected CRC cells, and showed that HOXA5, HOXB6 and GLTP are the direct target genes of MIR196B.
Material and Methods
Patients and Tissue Samples
The tissue samples used in this study were provided by the Biobank of Wonkwang University Hospital, a member of the National Biobank of Korea. With approval from the institutional review board and informed consent from the patients, we obtained colon cancer tissue from 2 colon cancer patients (1 male and 1 female) and rectal cancer tissue from 2 rectal cancer patients (2 males). The mean ages of the colon cancer and rectal cancer patients were 64.8 and 72 years, respectively. The colon cancer and matched normal colon tissue samples were used to analyze GLTP protein expression.
Cell Culture
The human CRC cell lines SW480 and Caco2 were obtained from the Korea Cell Line Bank (KCLB, Seoul, Korea) and American Type Culture Collection (ATCC, Manassas, VA, USA), respectively. SW480 cells were cultured in RPMI 1640 (HyClone, Logan, UT, USA) supplemented with 10% FBS in 5% CO2 at 37 °C in a humidified atmosphere. Caco2 cells were cultured in Alpha-MEM (HyClone) supplemented with 20% FBS in 5% CO2 at 37 °C in a humidified atmosphere.
RNA Extraction and Quantitative RT-PCR (qRT-PCR)
Total RNA was isolated with TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to our previous reports [11, 12]. After digestion with DNase and cleanup, RNA samples were quantified, aliquoted, and stored at −80 °C. Total RNA isolated from tissue samples and/or cultured cells was used as a template to synthesize cDNA for qRT-PCR analysis in a StepOne Real-time PCR system (Applied Biosystems, Foster City, CA, USA).
The differential miRNA expression patterns were validated with the TaqMan qRT-PCR assay (Applied Biosystems) or the NCode VILO miRNA cDNA Synthesis kit for qRT-PCR and EXPRESS SYBR GreenER miRNA qRT-PCR kit (Invitrogen). Furthermore, qRT-PCR with SYBR Green dye (Applied Biosystems) was used to assess mRNA expression. RNU48 (for TaqMan qRT-PCR) or 5.8S (for SYBR qRT-PCR) and GAPDH were used as endogenous controls for the qRT-PCR of miRNA and mRNA, respectively. Each sample was run in triplicate.
MIR196B Transfection
SW480 cells (3 × 105) or Caco2 cells (1.5 × 105) were plated onto 6-well or 10-cm culture plates and cultured as described above. MIR196B (hsa-miR-196b, Pre-miR miRNA Precursor AM17100, Product ID: PM12946) was commercially synthesized (Ambion, Austin, TX, USA) and transfected at a concentration of 50 nmol/L using Lipofectamine RNAiMAX (Invitrogen) or siPORT™ NeoFX™ transfection agent (Ambion) according to the manufacturers’ recommendations. Cells were harvested 24–48 h (for miRNA and mRNA) or 48–72 h (for protein) after transfection for RNA or protein extraction.
Plasmid Construct and Luciferase Assays
Wild-type or mutant fragments of the 3′ untranslated region (UTR) of HOXA5, HOXB6 and GLTP containing the predicted binding site for MIR196B, were amplified by PCR using the primer set shown in Table 1. The PCR product was cloned into the pmirGLO Dual-Luciferase miRNA Target Expression Vector (Promega, Madison, WI, USA). Analysis of the luciferase assay results was carried out according to our previously described method (Mo et al., 2015) [8]. Briefly, cells (5 × 104/well) were seeded onto 24-well plates and co-transfected with wild-type or mutant HOXA5, HOXB6 and GLTP constructs (500 ng/well) or with 50 nM MIR196B or MIR-1 (negative control) using Lipofectamine 2000 (Invitrogen Life Technologies) or siPORT™ NeoFX™ Transfection Agent (Ambion). Firefly and Renilla luminescence was measured 24 h after transfection using the Dual-Glo Luciferase Assay System (Promega) according to the manufacturer’s instructions. Non-transfected cells were used for background subtraction, and the ratio of firefly reporter luminescence to Renilla reporter luminescence (control) was calculated. All experiments were performed in triplicate and repeated at least three times.
Antibodies and Western Blot Analysis
SW480 or Caco2 cells (2 × 105 cells/well) were seeded onto 6-well plates and incubated for 72 h. Whole cell lysates were prepared by incubation in RIPA buffer supplemented with a protease inhibitor mixture for 30 min at 4 °C. Protein was collected by centrifugation at 12,000 rpm for 30 min at 4 °C. Equal amounts of protein (50 μg; determined by the Bradford assay) were boiled in Laemmli buffer, subjected to 12.0% or 15.0% SDS-PAGE, and then transferred onto PVDF membranes. The membranes were incubated with blocking buffer (5% BSA in TBS containing 0.1% Tween-20 [TBS-T]) for 2 h at room temperature. Membranes were then incubated overnight at 4 °C with primary antibodies to HOXA5 (Pierce Biotechnology) and HOXB6 and GLTP (Santa Cruz Biotechnology), and washed three times for 10 min per wash with TBS-T. Membranes were incubated with HRP-conjugated rabbit or mouse IgG secondary antibodies for 1 h at room temperature. After washing three times for 10 min in TBS-T, protein was detected with ECL solution (Millipore Corporation, Billerica, MA, USA) and the FluorChem E System (Cell Biosciences, Santa Clara, CA, USA). After protein detection, some membranes were stripped with stripping buffer for 1 h at room temperature and re-probed with an antibody to GAPDH (0411; Santa Cruz Biotechnology), which was used as a loading control. Protein expression was evaluated using the ImageJ software (version 1.44; http://rsbweb.nih.gov/ij/index.html).
Statistical Analysis
Each experiment was repeated at least three times with consistent results. All of the data were represented as mean ± standard deviation(SD). Statistical differences were analyzed by the Student′s t-test, and p-values of less than 0.05 (p < 0.05) were regarded as statistically significant.
Results
Validation and Expression of HOXA5 and HOXB6 Genes
The expression levels of HOXA5 and HOXB6 transcripts in un-transfected and transfected cells were compared by qRT-PCR. As shown in Fig. 1a, HOXA5 and HOXB6 mRNA levels were downregulated in SW480 and Caco2 cells transfected with pre-MIR196B, when compared to levels in the un-transfected control cells (Fig. 1a).
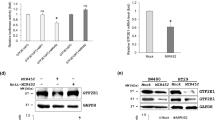
MIR196B directly down-regulates HOXA5 and HOXB6 expression. (a) Validation of MIR196B target genes. The expression of HOXA5 and HOXB6 was downregulated by MIR196B overexpression in SW480 or Caco2 cells. CRC cells were transfected with precursor MIR196B. Total RNA extracts were prepared 48 h after transfection, and HOXA5 and HOXB6 mRNA levels were examined by qRT-PCR. Values were normalized to levels in un-transfected Caco2 cells in three independent experiments. (b) Sequence alignment of the MIR196B seed region and the putative binding sites in the HOXA5 and HOXB6 3′ UTRs. The wild-type (WT) or mutant (MT) 3′ UTR of the HOXA5 or HOXB6 gene, containing the MIR196B target sequence, was cloned into a pmirGLO Dual-Luciferase miRNA Target Expression Vector. Each luciferase reporter construct was co-transfected into SW480 cells with either pre-MIR1 as a negative control or pre-MIR196B. After 24 h, luciferase activity was measured in the cell extracts. In the presence of the WT HOXA5 (c) or HOXB6 (d) 3′ UTR, transfection with pre-MIR196B inhibited luciferase activity (P < 0.01; two-sided t-test). Inhibition was not observed with the reporter constructs containing the MT 3′ UTR target sequences or with MIR1 transfection. (e) Western blot analysis in MIR196B- or mock-transfected SW480 cells. The HOXA5 and HOXB6 bands were quantified relative to the appropriate loading controls using the ImageJ software and are shown relative to the protein level in mock-transfected cells. The data are representative of three independent experiments
HOXA5 and HOXB6 Are Direct Targets of MIR196B
To determine whether MIR196B directly regulates HOXA5 and HOXB6 mRNA, we used a luciferase reporter system. The predicted MIR196B recognition sites in the HOXA5 and HOXB6 3′ UTRs were cloned into a pmirGLO Dual-Luciferase miRNA Target Expression Vector (pmirGLO-HOXA5 WT and pmirGLO-HOXB6 WT). We also generated mutated (MT) MIR196B binding sites by site-directed mutagenesis (pmirGLO-HOXA5 MT and pmirGLO-HOXB6 MT) (Fig. 1b). The base pairing between the MIR196B seed sites and the HOXA5 and HOXB6 mRNA targets is depicted in Fig. 1b. Subsequently, the vectors were co-transfected into SW480 cells with pre-MIR196B or pre-MIR1. MIR196B inhibited luciferase activity in cells transfected with pmirGLO-HOXA5 WT and pmirGLO-HOXB6 WT (~27% and ~30%, respectively, vs. controls) (Fig. 1c and d). In contrast, transfection of MIR196B did not reduce luciferase activity in cells transfected with pmirGLO-HOXA5 MT or pmirGLO-HOXB6 MT vectors, and transfection of pre-MIR1 did not reduce the luciferase activity in cells transfected with the WT or MT constructs (Fig. 1c and d).
MIR196B Regulates HOXA5 and HOXB6 Expression in CRC Cells
We investigated whether MIR196B regulated HOXA5 and HOXB6 protein expression. MIR196B was transfected into SW480 cells, and western blot analyses were performed using cells isolated 48 h after transfection. HOXA5 and HOXB6 expression levels were significantly downregulated in cells transfected with MIR196B (Fig. 1e). These results suggest that the expression of HOXA5 and HOXB6 is directly regulated by MIR196B in CRC cells.
GLTP Is a Direct Target of MIR196B
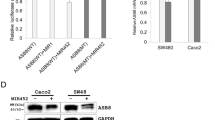
To demonstrate a direct interaction between the GLTP 3′ UTR region and MIR196B, we cloned the WT GLTP 3′ UTR region, which is predicted to interact with MIR196B into a luciferase vector (Fig. 2a). Luciferase activity was reduced by approximately 25%, when cells were co-transfected with pre-MIR196B (P < 0.01, Fig. 2b). As a control experiment, we cloned a mutated GLTP 3′ UTR sequence lacking ten of the complementary bases. As expected, repression of luciferase activity was abolished when the interaction between MIR196B and its target 3′ UTR was disrupted (Fig. 2b). As additional control experiments, MIR1 instead of MIR196B was co-transfected with the WT and MT GLTP 3′ UTR constructs. Transfection of pre-MIR1 did not affect the luciferase activity of either construct (Fig. 2b). These results suggest that MIR196B directly regulates GLTP expression in CRC cells.
GLTP is a direct target gene of MIR196B. (a) Sequence alignment of the MIR196B seed region and the putative binding sites in the GLTP 3′ UTRs. The wild-type (WT) or mutant (MT) 3′ UTR of the GLTP gene containing the MIR196B target sequence was cloned and transfected in CRC cells. (b) After 24 h, luciferase activity was measured in the cell extracts. In the presence of the WT GLTP 3′ UTR, transfection with pre-MIR196B inhibited luciferase activity (P < 0.01; two-sided t-test). Inhibition was not observed with the reporter constructs containing the MT 3′ UTR target sequences or with MIR1 transfection. Luciferase activity was determined using the dual luciferase assay. Results are shown as the relative firefly luciferase activity normalized to Renilla luciferase activity. (c) GLTP mRNA and protein levels in MIR196B- or mock-transfected Caco2 or SW480 cells. Total RNA or protein extracts were prepared 48 h or 72 h after transfection, and GLTP mRNA or protein levels were examined by qRT-PCR or western blot analysis, respectively (P < 0.01; two-sided t-test). (d) GLTP expression in 4 pairs of human CRC tissue samples and adjacent normal colon tissue samples. The data are representative of three independent experiments
We also investigated whether MIR196B regulated GLTP mRNA and protein levels in SW480 cells. The GLTP mRNA level was lower in SW480 cells transfected with pre-MIR196B than in un-transfected control cells (P < 0.01, Fig. 2c). GLTP protein expression was also downregulated in MIR196B-overexpressing cells (P < 0.01, Fig. 2c).
GLTP Expression Levels in Human CRC Tissues
On the basis of the findings described above, we evaluated GLTP expression in 4 human CRC tissues and the matching normal CRC tissues by western blotting. GLTP protein expression was decreased (3 of the 4 pairs) in CRC tissues compared to adjacent non-tumorous tissues (Fig. 2d).
Discussion
MiRNAs have been implicated as important regulators of gene expression in a variety of biological processes, as well as in various diseases as tumor suppressor genes or oncogenes [2,3,4]. In our previous study, we showed that MIR196B expression was up-regulated in human CRC tissues compared to matching healthy CRC tissues, and also suggested that MIR196B regulates FAS expression levels in CRC cells [8]. We used mRNA microarray analysis and bioinformatics tools to identify MIR196B target genes in CRC. Of the 18 genes identified in our previous study, five genes (HOXA5, HOXA9, HOXB6, HOXB7, and HOXC8) were members of the HOX gene family. This is not surprising because MIR196B is located between HOXA9 and HOXA10 on chromosome 7. Other studies have also previously identified HOXA9 [9, 13, 14], HOXB7 [15], and HOXC8 [16] as MIR196B target genes.
In this study, we showed that HOXA5 and HOXB6 are also MIR196B target genes in CRC cells (Fig. 1). HOXA5 and HOXB6 (Fig. 1a) expression was down-regulated by MIR196B overexpression in SW480 cells. Furthermore, HOXA5 and HOXB6 protein levels were also downregulated (Fig. 1e). These results suggest that the expression of HOXA5 and HOXB6 was directly regulated by MIR196B in CRC cells.
HOXA5 binds directly to a putative HOX-binding motif in the p53 promoter region. A reduction in the HOXA5 expression level correlates with the loss of p53 expression in breast cancer (Raman et al., 2000) [17]. The loss of HOXA5 gene expression in human breast cancer also correlates with the progression to higher-grade lesions [18], mammary gland homeostasis, and carcinogenesis [19]. HOXA5 participates in the developmental regulation of the lung. HOXA5−/− mice exhibited impaired postnatal lung development, indicating that HOXA5 has a critical role in lung ontogeny [20]. HOXA5 was also involved in the development and patterning of the mouse lung [21]. Furthermore, abnormal expression of HOXA5 was associated with lung tumorigenesis [22]. Recently, MIR196A has been shown to be significantly upregulated in non-small cell lung cancer (NSCLC) tissues, and that MIR196A regulates NSCLC cell proliferation, migration, and invasion, partially via the downregulation of HOXA5 [23]. HOXA5 and HOXB6 were verified as MIR196B targets in this study. Thus, MIR196B may correlate with the downregulation of HOXA5 expression in early colorectal carcinogenesis.
GLTP is a small cytosolic single-polypeptide protein that is able to catalyze the intermembrane transfer of glycosphingolipids between two membranes [24, 25]. The overexpression of human GLTP was found to dramatically alter the cell phenotype [26]. Human GLTP is located on chromosome 12 (12q24.11) and its expression is regulated by specific protein-1 (Sp1) and Sp3 [27, 28]. These results suggest that GLTP could be a significant player in cellular sphingolipid and overall lipid metabolism [29, 30]. The accumulating results suggested that glycolipids play an important role in biological functions such as recognition and cell signaling events as well as oncogenesis [31,32,33]. Although there have been many reports on glycolipid function, the exact functions and regulation of GLTP still remain unclear.
In this study, we showed that GLTP is a direct target of MIR196B in CRC cells (Fig. 2b). Our result also showed that the up-regulated MIR196B modulates the mRNA and protein levels of GLTP in CRC cells (Fig. 2c). As expected, the expression of GLTP protein is down-regulated in CRC tissues compared to adjacent non-tumorous tissues (Fig. 2d). Thus, these results suggest that the up-regulated MIR196B levels in CRC cells may regulates GLTP expression levels during colorectal carcinogenesis and therefore affect cell–cell communication or functions in CRC tissues.
Although we did not study in more detail the signaling mechanisms of HOXA5, HOXB6 and GLTP by the regulation of MIR196B, our results suggest that HOXA5, HOXB6 and GLTP were direct target genes of MIR196B in CRC cells. As the result, the up-regulated MIR196B in the CRC tissue regulate the expression levels of HOXA5, HOXB6 and GLTP during colorectal carcinogenesis. Our results could provide a valuable resource for further functional studies of HOXA5, HOXB6 and GLTP genes in the oncogenesis of various organs.
References
Ambros V (2004) The functions of animal microRNAs. Nature 431:350–355. https://doi.org/10.1038/nature02871
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297. https://doi.org/10.1016/S0092-8674(04)00045-5
Garzon R, Fabbri M, Cimmino A, Calin GA, Croce CM (2005) MicroRNA expression and function in cancer. Trends Mol Med 12:580–587. https://doi.org/10.1016/j.molmed.2006.10.006
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, Sweet-Cordero A, Ebert BL, Mak RH, Ferrando AA, Downing JR, Jacks T, Horvitz HR, Golub TR (2005) MicroRNA expression profiles classify human cancers. Nature 435:834–838. https://doi.org/10.1038/nature03702
Brennecke J, Hipfner DR, Stark A, Russell RB, Cohen SM (2003) Bantam encodes a developmentally regulated microRNA that controls cell proliferation and regulates the proapoptotic gene hid in drosophila. Cell 113:25–36. https://doi.org/10.1016/S0092-8674(03)00231-9
Ma L, Teruya-Feldstein J, Weinberg RA (2007) Tumour invasion and metastasis initiated by microRNA-10b in breast cancer. Nature 449:682–688. https://doi.org/10.1038/nature06174
Su H, Yang JR, Xu T, Huang J, Xu L, Yuan Y, Zhuang SM (2009) MicroRNA-101, down-regulated in hepatocellular carcinoma, promotes apoptosis and suppresses tumorigenicity. Cancer Res 69:1135–1142. https://doi.org/10.1158/0008-5472.CAN-08-2886
Mo JS, Alam KJ, Kang IH, Park WC, Seo GS, Choi SC, Kim HS, Moon HB, Yun KJ, Chae SC (2015) MicroRNA 196B regulates FAS-mediated apoptosis in colorectal cancer cells. Oncotarget 6:2843–2855. https://doi.org/10.18632/oncotarget.3066
Yekta S, Shih IH, Bartel DP (2004) MicroRNA-directed cleavage of HOXB8 mRNA. Science 304:594–596. https://doi.org/10.1126/science.1097434
Tanzer A, Amemiya CT, Kim CB, Stadler PF (2005) Evolution of microRNAs located within Hox gene clusters. J Exp Zool B Mol Dev Evol 304:75–85. https://doi.org/10.1002/jez.b.21021
Mo JS, Alam KJ, Kim HS, Lee YM, Yun KJ, Chae SC (2016) MicroRNA 429 regulates mucin gene expression and secretion in murine model of colitis. J Crohns Colitis 10:850–859. https://doi.org/10.1093/ecco-jcc/jjw033
Alam KJ, Mo JS, Han SH, Park WC, Kim HS, Yun KJ, Chae SC (2017) MicroRNA 375 regulates proliferation and migration of colon cancer cells by suppressing the CTGF-EGFR signaling pathway. Int J Cancer 141:1614–1629. https://doi.org/10.1002/ijc.30861
Li Z, Huang H, Chen P, He M, Li Y, Arnovitz S, Jiang X, He C, Hyjek E, Zhang J, Zhang Z, Elkahloun A, Cao D, Shen C, Wunderlich M, Wang Y, Neilly MB, Jin J, Wei M, Lu J, Valk PJ, Delwel R, Lowenberg B, Le Beau MM, Vardiman J, Mulloy JC, Zeleznik-Le NJ, Liu PP, Zhang J, Chen J (2012) miR-196b directly targets both HOXA9/MEIS1 oncogenes and FAS tumour suppressor in MLL-rearranged leukaemia. Nat Commun 3:688. https://doi.org/10.1038/ncomms1681
Liu Y, Zheng W, Song Y, Ma W, Yin H (2013) Low expression of miR-196b enhances the expression of BCR-ABL1 and HOXA9 oncogenes in chronic myeloid leukemogenesis. PLoS One 8:e68442. https://doi.org/10.1371/journal.pone.0068442
How C, Hui AB, Alajez NM, Shi W, Boutros PC, Clarke BA, Yan R, Pintilie M, Fyles A, Hedley DW, Hill RP, Milosevic M, Liu FF (2013) MicroRNA-196b regulates the Homeobox B7-vascular endothelial growth factor Axis in cervical cancer. PLoS One 8:e67846. https://doi.org/10.1371/journal.pone.0067846
Li Y, Zhang M, Chen H, Dong Z, Ganapathy V, Thangaraju M, Huang S (2010) Ratio of miR-196s to HOXC8 mRNA correlates with breast cancer cell migration and metastasis. Cancer Res 70:7894–7904. https://doi.org/10.1158/0008-5472.CAN-10-1675
Raman V, Martensen SA, Reisman D, Evron E, Odenwald WF, Jaffee E, Marks J, Sukumar S (2000) Compromised HOXA5 function can limit p53 expression in human breast tumours. Nature 405:974–978. https://doi.org/10.1038/35016125
Henderson GS, van Diest PJ, Burger H, Russo J, Raman V (2006) Expression pattern of a homeotic gene:HOXA5, in normal breast and in breast tumors. Cell Oncol 28:305–313. https://doi.org/10.1155/2006/974810
Gendronneau G, Lemieux M, Morneau M, Paradis J, Têtu B, Frenette N, Aubin J, Jeannotte L (2010) Influence of Hoxa5 on p53 TumorigenicOutcome in mice. Am J Pathol 176:995–1005. https://doi.org/10.2353/ajpath.2010.090499
Mandeville I, Aubin J, LeBlanc M, Lalancette-Hébert M, Janelle MF, Tremblay GM, Jeannotte L (2006) Impact of the loss of Hoxa5 function on lung alveogenesis. Am J Pathol 169:1312–1327. https://doi.org/10.2353/ajpath.2006.051333
Packer AI, Mailutha KG, Ambrozewicz LA, Wolgemuth DJ (2000) Regulation of the Hoxa4 and Hoxa5 genes in the embryonic mouse lung by retinoic acid and TGFbeta1: implications for lung development and patterning. Dev Dyn 217:62–74. https://doi.org/10.1002/(SICI)1097-0177(200001)217:1<62::AID-DVDY6>3.0.CO;2-U
Golpon HA, Geraci MW, Moore MD, Miller HL, Miller GJ, Tuder RM, Voelkel NF (2001) HOX genes in human lung: altered expressionin primary pulmonary hypertension and emphysema. Am J Pathol 158(3):955–966. https://doi.org/10.1016/S0002-9440(10)64042-4
Liu XH, Lu KH, Wang KM, Sun M, Zhang EB, Yang JS, Yin DD, Liu ZL, Zhou J, Liu ZJ, De W, Wang ZX (2012) MicroRNA-196a promotes non-small cell lung cancer cell proliferation and invasion through targeting HOXA5. BMC Cancer 12:348. https://doi.org/10.1186/1471-2407-12-348
Mattjus P, Pike HM, Molotkovsky JG, Brown RE (2000) Charged membrane surfaces impede the protein mediated transfer of glycosphingolipids between phospholipid bilayers. Biochemistry 39:1067–1075. https://doi.org/10.1021/bi991810u
Tuuf J, Mattjus P (2007) Human glycolipid transfer protein–intracellular localization and effects on the sphingolipid synthesis. Biochim Biophys Acta 1771:1353–1363. https://doi.org/10.1016/j.bbalip.2007.09.001
Gao Y, Chung T, Zou X, Pike HM, Brown RE (2011) Human glycolipid transfer protein (GLTP) expression modulates cell shape. PLoS One 6:e19990. https://doi.org/10.1371/journal.pone.0019990
Zou X, Chung T, Lin X, Malakhova ML, Pike HM, Brown RE (2008) Human glycolipid transfer protein (GLTP) genes: organization, transcriptional status and evolution. BMC Genomics 9:72. https://doi.org/10.1186/1471-2164-9-72
Zou X, Gao Y, Ruvolo VR, Gardner TL, Ruvolo PP, Brown RE (2011) Human glycolipid transfer protein gene (GLTP) expression is regulated by Sp1 and Sp3. J Biol Chem 286:1301–1311. https://doi.org/10.1074/jbc.M110.127837
Kjellberg MA, Backman AP, Ohvo-Rekila H, Mattjus P (2014) Alternation in the glycolipid transfer protein expression causes changes in the cellular Lipidome. PLoS One 9:e97263. https://doi.org/10.1371/journal.pone.0097263
Mattjus P (2009) Glycolipid transfer proteins and membrane interaction. Biochim Biophys Acta 1788:267–272. https://doi.org/10.1016/j.bbamem.2008.10.003
Kojima N, Hakomori S (1991) Cell adhesion, spreading, and motility of GM3- expressing cells based on glycolipid-glycolipid interaction. J Biol Chem 266:17552–17558
Boggs JM, Menikh A, Rangaraj G (2000) Trans interaction between galactosyl ceramide and cerebroside sulphate across opposed bilayers. Biophys J 78:874–885. https://doi.org/10.1016/S0006-3495(00)76645-8
Schnaar RL (2004) Glycolipid-mediated cell-cell recognition in inflammation and nerve regeneration. Arch Biochem Biophys 426:163–172. https://doi.org/10.1016/j.abb.2004.02.019
Acknowledgements
The biospecimens for this study were provided by the Biobank of Wonkwang University Hospital, a member of the National Biobank of Korea, which is supported by the Ministry of Health and Welfare. This research was supported by Wonkwang University in 2017.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest Statement
The authors declare that they have no conflicts of interests.
Informed Consent
All persons gave their informed consent prior to their inclusion in the study.
Human and Animal Studies
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
This article does not contain any studies with animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Mo, JS., Park, YR. & Chae, SC. MicroRNA 196B Regulates HOXA5, HOXB6 and GLTP Expression Levels in Colorectal Cancer Cells. Pathol. Oncol. Res. 25, 953–959 (2019). https://doi.org/10.1007/s12253-018-0399-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12253-018-0399-3