Abstract
Before use in practice, it is necessary to precisely identify and characterize a new probiotic candidate. Eight animal lactobacilli and collection strain Lactobacillus reuteri CCM 3625 were studied from the point of saccharide fermentation profiles, bile salt resistance, antibiogram profiles, and influence of bile on sensitivity to antibiotics. Studied lactobacilli differed in their sugar fermentation ability determined by API 50CHL and their identification based on these profiles did not correspond with molecular-biological one in most cases. Survival of strains Lactobacillus murinus C and L. reuteri KO4b was not affected by presence of bile. The resistance of genus Lactobacillus to vancomycin and quinolones (ofloxacin, ciprofloxacin) was confirmed in all strains tested. This study provides the new information about oxgall (0.5 and 1 %) effect on the lactobacilli antibiotic susceptibility. Antibiotic profiles were not noticeably affected, and both bile concentrations tested had comparable impact on the lactobacilli antibiotic sensitivity. Interesting change was noticed in L. murinus C, where the resistance to cephalosporins was reverted to susceptibility. Similarly, susceptibility of L. reuteri E to ceftazidime arose after incubation in both concentration of bile. After influence of 1 % bile, Lactobacillus mucosae D lost its resistance to gentamicin. On the base of gained outcomes, the best probiotic properties manifested L. reuteri KO4b, Lactobacillus plantarum KG4, and L. reuteri E due to their survival in the presence of bile.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Changeable physiological conditions along human gastrointestinal tract (GIT) determine its microbiota composition. Each microbial inhabitant has to adapt to specific environmental niche and develops appropriate mechanisms for withstanding of diverse stress types (Lebeer et al. 2008). Lactobacillus spp. that play a pivotal role in human gut and represents the most commonly used probiotic genus has to survive GIT passage in vital form. Except acid in the stomach (average pH 2) (Siciliano and Mazzeo 2012) and bile in the intestine and colon (concentration varies between 40 and 1 mmol/L) (Fontana et al. 2013), digestion enzymes also affect microbial composition of GIT. Surrounding commensal and opportune pathogenic microflora represents also a rival competing for nutrients and host cell receptors with probiotic cells (Horošová et al. 2006).
All GIT stress factors influence the ability of probiotic bacteria to maintain their health benefits to host macroorganism. Lactobacilli stress coping is evolved in several mechanisms. Acid stress tolerance is provided by three main manners: (i) maintaining the intracellular pH homeostasis; (ii) reparation of DNA and proteins damaged due to low pH; (iii) modification of cell envelope architecture (Lebeer et al. 2008). The cross-adaptation between acid and bile resistance is present, considering some of the common adaptation mechanisms (Lorca and de Valdez 2009). Bile salt hydrolases (coding by bsh genes) are responsible for specific adaptation mechanism of GIT lactobacilli to bile (Begley et al. 2005a; Patel et al. 2010). The host may also profit from this adaptation mechanism. Cholesterol level decrease, as one of probiotic mechanisms of action, can be partially ascribed to bile salt hydrolase activity (Liong and Shah 2005; Ebringer et al. 2008; Turková et al. 2013).
Probiotic bacteria are potential reservoir of antibiotic resistance genes that can be spread to surrounding microbiota, even pathogenic. In comparison with intrinsic resistance (genes located on bacterial chromosome), the acquired resistance (genes located on plasmids and transposons) represents the real risk in practical use due to possible resistance transfer (Ashraf and Shah 2011; Fukao and Yajima 2012). Postantibiotic diarrhea is one of the health disorders that probiotic bacteria are able to moderate (Sepp et al. 2011). In the case of postantibiotic diarrhea recovery, use of probiotics is very common. Certain strain to be applied should be resistant to antibiotic used in therapy (Clementi and Aquilanti 2011; Dušková and Karpíšková 2013). For these reasons, it is essential to know the antibiogram profile of probiotic bacterium.
The present study is focused on antibiotic susceptibility determination, ability to survive in presence of bile, bile effect on antibiotic susceptibility, and comparison of API 50CHL and molecular-biological identification (previous works: Bilková et al. 2008; Kiňová Sepová and Bilková 2013) of potential probiotic lactobacilli for human or veterinary use.
Material and methods
Bacterial strains and growth conditions
Several bacterial strains were isolated from stomach mucosas of breast-fed lamb and goatling. Lamb isolates were identified as Lactobacillus murinus C, Lactobacillus mucosae D, and Lactobacillus reuteri E (Bilková et al. 2008); goatling as L. reuteri KO4b, L. reuteri KO4m, L. reuteri KO5, Lactobacillus plantarum KG1z, and L. plantarum KG4 (Kiňová Sepová and Bilková 2013). L. reuteri CCM 3625 was purchased from Czech Collection of Microorganisms (Brno, Czech Republic). Lactobacilli were cultivated in MRS broth (Oxoid, Great Britain) at 37 °C in anaerobic conditions for 18 h.
API 50CHL
Carbohydrate fermentation profiles were found out using API 50CHL kit (bioMérieux, France) according to the manufacturer’s recommendations.
Antibiotic susceptibility
Susceptibility to selected antimicrobial substances was determined by disc diffusion method according to Coyle (2005). Discs with antimicrobial substances (Tables 2 and 3) were purchased from Oxoid (Great Britain). Testing of antimicrobial susceptibility of lactobacilli after influence of 0.5 and 1.0 % bile salts was performed according to Elkins and Mullis (2004). Preincubation with bile salts lasted 12 h at 37 °C.
Bile resistance
Experiment was performed by modified method according to De Boever and Verstraete (1999). Briefly, bacterial cultures after 18 h anaerobical incubation were settled by centrifugation, washed twice in physiological saline, and adjusted to ca 4.8 × 107 CFU/mL. Cultures were further diluted in MRS broth (Oxoid, Great Britain) supplemented with 0 (control sample), 0.5 or 1 % oxgall (BiomarkTM Laboratories, India) in ratio 1:9. After 12 h of incubation, the number of survived bacteria was determined by serial dilutions and plating on MRS agar. The results were expressed by the percentage of growth in the presence of bile salts compared to the control.
Statistical analysis
Experiments were repeated in three or six parallels. The statistical comparison between control and tested samples was performed by Student’s t-test. Statistical significant differences between control and sample are expressed as ns non-significant difference; *p < 0.05; **p < 0.01; and ***p < 0.001.
Results and discussion
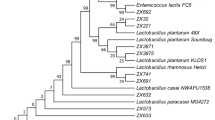
Antibiograms and survival in bile and its effect on antibiotic susceptibility of eight lactobacilli probiotic candidates were studied. As the exact identification of probiotic microorganism is essential (FAO/WHO 2001), molecular-biological identification (partial 16S rDNA sequencing (Bilková et al. 2008; Kiňová Sepová and Bilková 2013)) was compared with biochemical one (API 50CHL; Table 1). The cohesive results were gained only in L. plantarum isolates. In other isolates, some discrepancies using API 50CHL were observed. Number of Lactobacillus species is still increasing (to date 201 species, 29 subspecies) (Euzéby 2014), and therefore not all of them are covered in database. Actually, API 50CHL database includes 16 species and 3 subspecies of lactobacilli and for example L. reuteri, which was firstly described in 1980 (Kandler et al. 1980), is still missing in version V5.1. Heterogeneity of genus Lactobacillus (Collins et al. 1991; Claesson et al. 2007) gives also the evidence of various results.
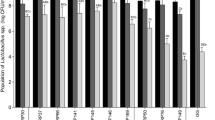
To overcome the stress conditions in GIT, gut-associated lactobacilli are able to metabolize conjugated bile salts by hydrolysis (De Boever and Verstraete 1999). Furthermore, GIT strains adapt more progressively to the bile salts presence (Ruiz et al. 2013) in comparison with bacteria originated from other niches. The probiotic candidates isolated from Slovak bryndza cheese did not show such high rate of survival in the presence of 0.5 and 1 % bile (Belicová et al. 2013), than our strains originating from animals GIT. Survival of L. reuteri E in the presence of 1 % bile was significantly better than in 0.5 % (Fig. 1). Survival of strains L. murinus C and L. reuteri KO4b was not significantly impacted by the presence of bile in both concentrations used. The rest strains were more sensitive to higher bile concentrations.
Survival of tested lactobacilli in the presence of 0.5 and 1 % bile C Lactobacillus murinus C, D Lactobacillus mucosae D, E Lactobacillus reuteri E, KO4b Lactobacillus reuteri KO4b, KO4m Lactobacillus reuteri KO4m, KO5 Lactobacillus reuteri KO5, KG1z Lactobacillus plantarum KG1z, KG4 Lactobacillus plantarum KG4, CCM 3625 Lactobacillus reuteri CCM 3625. Control sample corresponds to 100 %. Values are calculated from six independent experiments as arithmetical means ± SD. Statistical significant differences between control and sample are expressed as ns non-significant difference; *p < 0.05; **p < 0.01; ***p < 0.001
Susceptibility of tested lactobacilli to 11 commonly used antibiotics was determined by disc diffusion assay and interpreted according to Charteris et al. (1998). Since the official breakpoints for lactobacilli are not given, diameters of inhibitory zones are also introduced (Table 2). MRS agar was used due to the poor growth of strains on Müller-Hinton agar, although, components of MRS medium can inactivate some antibiotics, e.g., imipenem (Ammor et al. 2007). The selection of appropriate cultivation medium for lactic acid bacteria was proclaimed also by the other authors (Klare et al. 2005; Dušková and Karpíšková 2013). All tested strains were resistant to vancomycin, ofloxacin, and ciprofloxacin (Table 2) in accordance with the literature findings. Vancomycin resistance was described as intrinsic in this genus (Bernardeau et al. 2008). Resistance to quinolones (ofloxacin and ciprofloxacin) corresponds with findings of other authors (Hummel et al. 2007), and it is also proposed to be intrinsic in lactobacilli. Detected susceptibility in lactobacilli to penicillin, ampicillin, and erythromycin was also described before (Danielsen and Wind 2003; Klare et al. 2005)
Influence of bile on antibiotics susceptibility was tested after preincubation of lactobacilli in the presence of 0.5 and 1 % bile. In comparison to previous experiment, antibiotic susceptibility profiles were not markedly modified and both bile concentrations had comparable effect (Table 3). Interesting result was noticed in L. murinus C which lost its resistance to ceftazidime and cefotaxime. After preincubation with 1 % bile, L. mucosae D lost its gentamicin resistance. Similarly, susceptibility of L. reuteri E to ceftazidime arose after exposition to bile. Increased diameter of inhibition zones of L. mucosae D to quinolones, penicillin, and ampicillin was detected.
Bile itself has surfactant properties and is able to emulsify and solubilize lipids. These detergent properties result also in an antimicrobial effect due to its membrane activity (Begley et al. 2005b). Modification of bacterial antibiotic susceptibility after their exposition to bile could be due the loss of semipermeability, which eases molecules passage. Our observation suggests that this effect could be synergic with activity of beta-lactams. In some cases, the diameters of inhibition zones were larger after action of bile (Elkins and Mullis 2004), and in some cases, the resistance was reverted to susceptibility (ceftazidime and cefotaxime in L. murinus C and ceftazidime in L. reuteri E).
The present study completed previous characterization of potential probiotic lactobacilli strains. Due to declared intrinsic resistance to vancomycin (Bernardeau et al. 2008) and quinolones (Hummel et al. 2007) in genus Lactobacillus, tested strains do not represent a risk of this resistance genes transfer between bacterial species. There is an assumption that antibiotic resistance of majority of tested strains will be not changed excessively after action of bile in host gut. As possible probiotics, the most promising strains seem to be L. reuteri KO4b, L. plantarum KG4, and L. reuteri E for their good survival in bile comparable to control. However, after complex in vitro studies (e.g., determination of transmissible antibiotic resistance), the in vivo evaluation of beneficial attributes on human and/or animal model is necessary before use in practice.
References
Ammor MS, Flórez AB, Mayo B (2007) Antibiotic resistance in non-enterococcal lactic acid bacteria and bifidobacteria. Food Microbiol 24:559–570
Begley M, Hill C, Gahan CGM (2005a) Bile salt hydrolase activity in probiotics. Appl Environ Microbiol 72:1729–1738
Begley M, Gahan CGM, Hill C (2005b) The interaction between bacteria and bile. FEMS Microbiol Res 29:625–651
Belicová A, Mikulášová M, Dušinský R (2013) Probiotic potential and safety properties of Lactobacillus plantarum from Slovak bryndza cheese. Biomed Res Int 2013. doi:10.1155/2013/760298
Bernardeau M, Vernoux JP, Henri-Dubernet S, Guéguen M (2008) Safety assessment of dairy microorganisms: the Lactobacillus genus. Int J Food Microbiol 126:278–285
Bilková A, Kiňová Sepová H, Bilka F, Bukovský M, Balažová A, Bezáková L (2008) Identification of newly isolated lactobacilli from the stomach mucus of lamb. Acta Facult Pharm Univ Comenianae 55:64–72
Charteris WP, Kelly PM, Morelli L, Collins JK (1998) Antibiotic susceptibility of potentially probiotic Lactobacillus species. J Food Prot 61:1636–1643
Claesson MJ, van Sinderen D, O’Toole PW (2007) The genus Lactobacillus – a genomic basis for understanding its diversity. FEMS Microbiol Lett 269:22–28
Clementi F, Aquilanti L (2011) Recent investigations and updated criteria for the assessment of antibiotic resistance in food lactic acid bacteria. Anaerobe 17:394–398
Collins MD, Rodrigues U, Ash C, Aguirre M, Farrow JAE, Martine-Murcia A, Phillips BA, Williams AM, Wallbanks S (1991) Phylogenetic analysis of the genus Lactobacillus and related lactic acid bacteria as determined by reverse transcriptase sequencing of 16S rRNA. FEMS Microbiol Lett 77:5–12
Coyle MB, Ed (2005) Manual of antimicrobial susceptibility testing. American Society for Microbiology
Danielsen M, Wind A (2003) Susceptibility of Lactobacillus spp. to antimicrobial agents. Int J Food Microbiol 82:1–11
De Boever P, Verstraete W (1999) Bile salts deconjugation by Lactobacillus plantarum 80 and its implication for bacterial toxicity. J Appl Microbiol 87:345–352
Dušková M, Karpíšková R (2013) Antimicrobial resistance of lactobacilli isolated from food. Czech J Food Sci 31:27–32
Ebringer L, Ferenčík M, Krajčovič J (2008) Beneficial effects of milk and fermented dairy products – review. Folia Microbiol 53:378–394
Elkins CA, Mullis LB (2004) Bile-mediated aminogycosyte sensitivity in Lactobacillus species likely results from increased membrane permeability attributable to cholic acid. App Environ Microbiol 70:7200–7209
Euzéby JP 2014 List of prokaryotic names with standing in nomenclature. http://www.bacterio.net/lactobacillus.html Accessed 28 March 2014
FAO/WHO (2001) Health and nutritional properties of probiotics in food including powder milk with live lactic acid bacteria. http://www.who.int/foodsafety/publications/fs_management/en/probiotics.pdf Accessed 7 April 2014
Fontana L, Bermudez-Brito M, Plaza-Diaz J, Munoz-Quezada S, Gil A (2013) Sources, isolation, characterization and evaluation of probiotics. Brit J Nutr 109:35–50
Fukao M, Yajima N (2012) Antibiotic resistant bacteria – a continuous challenge in the new millennium. In: Pana M (ed) Assessment of antibiotic resistance in probiotic lactobacilli, 1st edn. InTech, Rijeka, pp 503–512
Horošová K, Bujňáková D, Kmeť V (2006) Effect of lactobacilli on E. coli adhesion to Caco-2 cells in vitro. Folia Microbiol 51:281–282
Hummel AS, Hertel C, Holtzapfel WA, Franz CMAP (2007) Antibiotic resistances of starter and probiotic strains of lactic acid bacteria. Appl Environ Microbiol 73:730–739
Kandler O, Stetter KO, Kohl R (1980) Lactobacillus reuteri sp. nov., a new species of heterofermentative lactobacilli. Zbl Bakt Hyg I Abt Orig C 1:264–269
Kiňová Sepová H, Bilková A (2013) Isolation and identification of new lactobacilli from goatling stomach and investigation of reuterin production in Lactobacillus reuteri strains. Folia Microbiol 58:33–38
Klare I, Konstabel C, Müller-Bertling S, Reissbrodt R, Huys G, Vanncaneyt M, Swings J, Goossens H, Witte W (2005) Evaluation of new broth media for microdilution antibiotic susceptibility testing of lactobacilli, pediococci, lactococci and bifidobacteria. Appl Environ Microbiol 71:8982–8986
Lebeer S, Vanderleyden J, De Keersmaecker SCJ (2008) Genes and molecules of Lactobacilli supporting probiotic action. Microbiol Mol Biol Rev 72:728–764
Liong MT, Shah NP (2005) Bile salt deconjugation ability, bile salt hydrolase activity and cholesterol co-precipitation ability of lactobacilli strains. Int Dairy J 15:391–398
Lorca GL, de Valdez GF (2009) Lactobacillus molecular biology. From genomics to probiotics. In: Ljung Å, Wadström T (eds) Lactobacillus stress responses, 1st edn. Caister Academic Press, Norfolk, pp 115–138
Patel AK, Singhania RR, Pandey A, Chincholkar SB (2010) Probiotic bile salt hydrolase: current developments and perspectives. Appl Biochem Biotechnol 162:166–180
Ruiz L, Margolles A, Sánchez B (2013) Bile resistance mechanisms in Lactobacillus and Bifidobacterium. Front Microbiol. doi:10.3389/fmicb.2013.00396
Sepp E, Štšepetova J, Smidt I, Rätsep M, Kõljalg S, Lõivukene K, Mändar R, Jaanimäe L, Löhr HI, Natås OB, Naaber P (2011) Intestinal lactoflora in Estonian and Norwegian patients with antibiotic associated diarrhea. Anaerobe 17:407–409
Siciliano RA, Mazzeo MF (2012) Molecular mechanisms of probiotic action: a proteomic perspective. Curr Opin Microbiol 15:1–7
Turková K, Mavrič A, Narat M, Rittich B, Španová A, Rogelj I, Matjašić BB (2013) Evaluation of Lactobacillus strains for selected probiotic properties. Folia Microbiol 58:261–267
Acknowledgments
We are thankful to Professor Dušan Mlynarčík, DrSc., for critical reading of manuscript.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hyacinta, M., Hana, K.S., Andrea, B. et al. Bile tolerance and its effect on antibiotic susceptibility of probiotic Lactobacillus candidates. Folia Microbiol 60, 253–257 (2015). https://doi.org/10.1007/s12223-014-0365-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-014-0365-8