Abstract
Gut microbiota have been linked to immune thrombocytopenia (ITP) and Henoch–Schönlein purpura (HSP) in recent studies, but a cause-and-effect relationship is unclear. We used Mendelian randomization (MR) to assess causal relationships between gut microbiota and HSP/ITP using summary statistics from the GWAS dataset of the international MiBioGen and FinnGen consortium. The IVW method was used as the main evaluation indicator. MR analysis of 196 intestinal flora and HSP/ITP/sTP phenotypes showed that 12 flora were potentially causally associated with ITP, 6 with HSP, and 9 with sTP. The genes predicted that genus Coprococcus3 (p = 0.0264, OR = 2.05, 95% CI 1.09–3.88)and genus Gordonibacter (p = 0.0073, OR = 1.38; 95% CI 1.09–1.75) were linked to a higher likelihood of developing ITP. Additionally, family Actinomycetaceae (p = 0.02, OR = 0.51, 95% CI 0.28–0.90) and order Actinomycetales (p = 0.0199, OR = 0.50, 95% CI 0.28–0.90) linked to reduced HSP risk. Genus Ruminococcaceae UCG013 (p = 0.0426, OR = 0.44, 95% CI 0.20–0.97) negatively correlated with sTP risk. Our MR analyses offer evidence of a possible cause-and-effect connection between certain gut microbiota species and the likelihood of HSP/ITP.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
HSP and ITP are both prevalent immune-mediated bleeding disorders characterized by distinct symptoms of purpura [1, 2]. ITP, also known as idiopathic thrombocytopenic purpura or immune thrombocytopenic purpura, is a condition characterized by a low platelet count caused by autoantibodies targeting platelet antigens [3]. ITP is a hematologic disorder that is relatively common, with the highest incidence observed in the pediatric population [4]. Approximately 40% of cases occur in children under 10 years of age [5]. The estimated overall incidence of ITP in the United States is approximately 3.3 cases per 100,000 persons per year [3]. The incidence of ITP in pregnant women ranges from 1 to 10 per 10,000 cases. ITP accounts for approximately 1–4% of all cases of thrombocytopenia during pregnancy and is the primary cause of thrombocytopenia in the early stages of pregnancy [6]. The primary clinical symptom of ITP is hemorrhage in the skin and mucous membranes [7]. The primary clinical symptom of ITP is hemorrhage in the skin and mucous membranes [8]. It is generally believed that the abnormal immune response, which mediates platelet destruction, and the disorder of platelet production caused by the impairment of bone marrow megakaryocyte maturation and poor platelet production, play a key role in the pathogenesis [9]. Early treatment does not reduce the incidence of chronic ITP, but it can help restore platelet levels quickly, improve quality of life, and reduce the occurrence of severe bleeding or complications [10]. In Henoch–Schönlein purpura (HSP) or immunoglobulin A vasculitis (IgA), small blood vessels in the skin, kidneys, and gastrointestinal system are affected [1].This condition is classified as a small-vessel leukocytoclastic vasculitis resulting from the deposition of immune complexes. It can present as either a systemic or localized disease, with commonly affected organs including the skin, kidneys, gastrointestinal system, and joints [1]. With a prevalence of 20.4 per 100,000, HSP is the most common form of systemic vasculitis in children [11].
There is mounting evidence that the gut microbiota plays a crucial role in maintaining metabolic and immune system balance [12,13,14]. Dysregulation of the gut microbiota can contribute to the development of autoimmune and metabolic diseases, as well as hematologic disorders such as rheumatoid arthritis (RA) [15]. In recent years, numerous studies have explored the connection between gut microbiota and various diseases, including rheumatoid arthritis (RA) [15], ankylosing spondylitis (AS) [16], inflammatory bowel disease (IBD) [17], immune disorders like systemic lupus erythematosus (SLE) [18], and metabolic disorders such as type II diabetes mellitus [19] and atherosclerotic cardiovascular disease (ACVD) [20]. Changes in the abundance of gut microbiota, such as Methylobacterium, Sphingomonas, and Staphylococcus, can impact serum salicylate levels. The latter has been found to inhibit platelet production in megakaryocytes and induce the development of ITP. The activation status of platelets in patients with ITP has been associated with gut flora [21].
Xiaomin Yu et al. discovered that specific lipid metabolites, including RvD2, eicosatetraenoic acid, monooleic acid, and phosphatidylcholine, were up-regulated in the gut microbiota of patients with ITP. They also found that these metabolites could regulate platelet homeostasis, reduce platelet activation, and consequently impact platelet function [22]. Furthermore, the microbiota can affect the effectiveness of ITP treatment. Research has shown that the microbiota found in the airways of asthma patients can cause resistance to corticosteroids [23]. The study revealed significant differences in the gut microbiota of patients with corticosteroid-resistant ITP compared to both healthy individuals (3 genera and 13 species) and patients with corticosteroid-sensitive ITP (21 genera and 59 species). This indicates that the imbalance of microbiota differs between corticosteroid-sensitive and drug-resistant ITP patients [24]. Disturbed intestinal flora is found in children with HSP. This could play a role in the onset of HSP by impacting the production and breakdown of unsaturated fatty acids [25,26,27].
While there is a connection between gut flora and ITP, there is a lack of comprehensive data regarding the link between gut microbiota and ITP. It is crucial to rule out any secondary causes of thrombocytopenia for an accurate clinical diagnosis of ITP. Therefore, exploring the association between gut microbiota and secondary thrombocytopenia (sTP) could provide valuable understanding. To identify specific pathogenic bacterial taxa and screen for genetic variants, we utilized genome-wide association study (GWAS) data and employed MR analysis [28]. Furthermore, we examined the possible cause-and-effect connection between the gut microbiota and disease susceptibility genes like HSP, ITP, and sTP.
Study design and methods
Study design
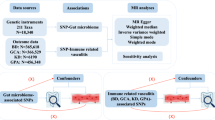
A summary of the study's design is depicted in Fig. 1. This study strictly adheres to the three hypotheses of MR analyses: the association hypothesis, which posits that instrumental variables (IVs) are strongly correlated with exposure. The independent hypothesis states that IVs are not influenced by confounding variables between the exposure and the outcome. IVs only have an effect on the outcome through exposure and do not directly affect the exclusive hypothesis of the outcome.
Data sources of gut microbiota
The information on the microbiome of the human gut was collected from the GWAS dataset of the MiBioGen International Consortium [28]. The research combined extensive, multiethnic 16S ribosomal RNA gene sequencing and genotyping data from 24 different groups, including 18,340 participants in the United States, United Kingdom, Germany, Finland, Denmark, the Netherlands, Canada, and South Korea. The data were adjusted for major components such as age, sex, technical covariates, and study population. Upon analysis of the data, 15 taxa of bacteria with unknown or unspecified names were excluded, resulting in the inclusion of 196 taxa, comprising 119 genera, 32 families, 20 orders, 16 classes, and 9 phyla. The data for this MR analysis were derived from publicly available GWAS genetic aggregate data, thus obviating the need for additional ethical approval or consent for participation.
Data sources for HSP/ITP/sTP
The aggregated outcomes data are sourced from data R9 published by the FinnGen Consortium. HSP included 856 cases and 391,567 controls, ITP included 810 cases and 391,613 controls, and sTP included 298 cases and 392,125 controls. Detailed information about the outcome data from the included studies is summarized in Table 1. In 810 cases of ITP patients, females accounted for 52.7% (427 cases) and males accounted for 47.3% (383 cases).The mean age at first event for females was 43.7 years, and for males it was 54.73 years. The death rate for the entire study period, spanning from 1998 to 2019, was 0.08, with a Hazard Ratio (HR) and a 95% Confidence Interval (CI) of 5.61 (3.19, 9.86). Furthermore, the death rates at both the 15-year and 5-year marks were both 0.01, with HR (95% CI) values of 2.08 (1.20, 3.62) and 5.43 (3.55, 8.30) respectively. In 856 cases of HSP patients, females accounted for 59.1% (506 cases) and males accounted for 40.9% (350 cases). The mean age at first event for females was 36.25 years, and for males it was 43.10 years. The death rate for the entire study period from 1998 to 2019 was calculated to be 0.13,with HR (95% CI) of 9.04 (5.81, 14.06). Additionally,the death rates at both the 15-year and 5-year marks were determined to be 0.02 and 0.01 respectively,with HR (95% CI) values of 3.23 (1.95, 5.36) and 6.71 (3.91, 11.51). In 298 cases of sTP patients, it was found that females accounted for 52.7% (157 cases) and males accounted for 47.3% (141 cases). The mean age at the first event for females was approximately 53.73 years, while for males it was approximately 59.08 years old.
Instruments selection
The selection criteria and steps for IVs were as follows: to obtain a satisfactory number of IVs, we selected p < 1.0 × 10–5 as the threshold to screen for significantly correlated single nucleotide polymorphisms (SNPs). 2. We used the "TwoSampleMR" package to remove linkage disequilibrium by setting the linkage disequilibrium (LD) threshold to R2 < 0.001 and the aggregation distance to 10,000 kb. 3. The distance is 10,000 kb. 3. When quantifying using the F-statistic, we set F > 10 to exclude weak instrumental variables. To calculate the F-value, we extracted the beta, standard error (SE), and p value (p) for each SNP. The F-value was calculated as F = Beta2/SE2.
Statistical analysis
In this study, four methods, namely IVW, MR-Egger regression, weighted median, and weighted mode, were utilized to investigate the potential causal relationship between the gut microbiota and HSP, ITP, and sTP. The IVW method served as the primary analytical approach. The weighted median approach operated under the assumption that more than 50% of the IVs were plausible. To address potential horizontal pleiotropy, concerns were evaluated through intercept tests for MR-Egger regression, Mendelian Randomization Pleiotropy Residual Sum and Outlier (MR-PRESSO) analysis, and sensitivity analyses. The reliability of MR results may be compromised if the p-value of the intercept test in MR-Egger regression is less than 0.05. If the p-value of MR-PRESSO is less than 0.05, it is necessary to identify outliers and remove them from the IVs before conducting MR analysis. In addition, we tested for heterogeneity among the included data SNPs using Cochran's Q value. If a p-value is less than 0.05, it indicates the presence of heterogeneity. In this case, we opt for the random-effects IVW approach. Conversely, if the p-value is greater than or equal to 0.05, we use a fixed-effects model analysis. Finally, the relationships between human gut microbiota and diseases were quantified as odds ratios (OR) and their corresponding 95% confidence intervals (CI).
To assess causality more rigorously, we adjusted for the number of bacterial taxa associated with each attribute using the Bonferroni method, which required setting more stringent significance p-values. For example, we used the following adjusted p-values: Door: 0.05/9 (5.6 × 10–3), Class: 0.05/16 (3.1 × 10–3), Family: 0.05/32 (1.56 × 10–3), and Genus: 0.05/119 (4.20 × 10–4). If the p-value of the results of this MR analysis falls between the corrected threshold and 0.05, we would consider the bacterial taxon as providing suggestive evidence of a potential causal association with the disease and performed a reverse MR analysis. The MR analyses were conducted using R 3.6.3 software and the "TwoSampleMR" and "MR-PRESSO" packages. The code for the study is provided in Supplementary Table 1.
Results
In this MR analysis of gut flora with HSP/ITP/sTP, F values were calculated for 196 gut microbial SNPs, with the F-statistics for SNPs ranging from 12.6 to 32.5, all of which were above 10, indicating a lower likelihood of experiencing weak instrument bias (Supplementary Table 2).
Causal effects of gut microbiota on ITP
This research uncovered 12 causal connections between the gut microbiota and ITP (Supplementary Table 3). The genetically forecasted genus Coprococcus3 (p = 0.0264, OR = 2.05, 95% CI 1.09–3.88) and genus Gordonibacter (p = 0.0073, OR = 1.38, 95% CI 1.09–1.75) exhibited positive associations with the likelihood of ITP as determined by the IVW method (Fig. 2). The connection between genus Coprococcus3 (p = 0.0360, OR = 2.45, 95% CI 1.06–5.66) and genus Gordonibacter (p = 0.025, OR = 1.41, 95% CI 1.04–1.89) and ITP remained steady in the weighted-median method. Furthermore, we observed that five bacterial characteristics were tentatively linked to a reduced risk of ITP in the IVW method (p = 0.0172, OR = 0.71, 95% CI 0.53–0.94 for class Methanobacteria; p = 0.0177, OR = 0.61, 95% CI 0.41–0.92 for family BacteroidalesS24.7 group; p = 0.0314, OR = 0.55, 95% CI 0.32–0.95 for family Lachnospiraceae; p = 0.0172, OR = 0.71, 95% CI 0.53–0.94 for family Methanobacteriaceae; p = 0.0429, OR = 0.62, 95% CI 0.39–0.98 for genus Eubacteriumhalliigroup; p = 0.0377, OR = 0.74, 95% CI 0.56–0.98 for genus Eubacteriumruminantiumgroup; p = 0.0103, OR = 0.70, 95% CI 0.53–0.92 for genus Allisonella; p = 0.0432, OR = 0.60, 95% CI 0.37–0.98 for genus Coprococcus2; p = 0.0192, OR = 0.69, 95% CI 0.50–0.94 for order Bacillales; p = 0.0172, OR = 0.71, 95% CI 0.53–0.94 for order Methanobacteriales). However, the results from the weighted median method did not support such a causal effect. The MR-Egger and MR-PRESSO tests indicated no evidence of horizontal pleiotropy or outliers (p > 0.05). Furthermore, the results of Cochrane's Q-test revealed no significant heterogeneity (p > 0.05) (Table 2). Additionally, leave-one-out analyses demonstrated that none of the identified causal associations were affected by any individual independent variable (Supplementary Fig. 1). None of the outcomes from Bonferroni's corrected test indicated the presence of a causal effect.
Causal effects of gut microbiota on HSP
This research uncovered six causal connections between the gut microbiota and HSP (Supplementary Table 4). The genetically forecasted family Actinomycetaceae (p = 0.0200, OR = 0.51, 95% CI 0.28–0.90) and order Actinomycetales (p = 0.0199, OR = 0.50, 95% CI 0.28–0.90;) exhibited negative associations with the likelihood of HSP as determined by the IVW method (Fig. 2). The connection between family Actinomycetaceae (p = 0.0236, OR = 0.45, 95% CI 0.23–0.90), order Actinomycetales (p = 0.0341, OR = 0.45, 95% CI 0.21–0.94), and HSP remained steady in the weighted-median method. In addition, the increased presence of the genetically predictive genus Bilophila (p = 0.0178, OR = 1.65, 95% CI 1.09–2.51), genus FamilyXIIIAD3011group (p = 0.0215, OR = 1.87, 95% CI 1.10–3.17) and genus Roseburia (p = 0.0411, OR = 1.61, 95% CI 1.02–2.54) were found to be linked to the development of HSP using the IVW method. Conversely, the presence of genus Marvinbryantia (p = 0.0363, OR = 0.59, 95% CI 0.37–0.97) was associated with a decreased likelihood of HSP occurrence. Nevertheless, the findings obtained from the weighted median method did not provide evidence for a causal effect. The MR-Egger and MR-PRESSO tests indicated no evidence of horizontal pleiotropy or outliers (p > 0.05). Furthermore, the results of Cochrane's Q-test revealed no significant heterogeneity (p > 0.05) (Table 2). Additionally, leave-one-out analyses demonstrated that none of the identified causal associations were affected by any individual independent variable (Supplementary Fig. 2). None of the outcomes from Bonferroni's corrected test indicated the presence of a causal effect.
Causal effects of gut microbiota on sTP
This research uncovered 9 causal connections between the gut microbiota and sTP (Supplementary Table 5). The genetically forecasted genus RuminococcaceaeUCG013 (p = 0.0426, OR = 0.44, 95% CI 0.20–0.97), family Methanobacteriaceae (p = 0.0016, OR = 0.47, 95% CI 0.29–0.75), order Methanobacteriales (p = 0.0016, OR = 0.47, 95% CI 0.29–0.75), class Methanobacteria (p = 0.0016, OR = 0.47, 95% CI 0.29–0.75) exhibited negative associations with the likelihood of sTP as determined by the IVW method (Fig. 2). The association between genus RuminococcaceaeUCG013 (p = 0.0337, OR = 0.32, 95% CI 0.11–0.92), family Methanobacteriaceae (p = 0.0325, OR = 0.50, 95% CI 0.27–0.94), order Methanobacteriales (p = 0.0359, OR = 0.50, 95% CI 0.27–0.65), class Methanobacteria (p = 0.0328, OR = 0.50, 95% CI 0.27–0.95), and sTP remained steady in the weighted-median method.
Furthermore, we noted that four bacterial characteristics were tentatively linked to an increased likelihood of sTP using the IVW method ( p = 0.0045, OR = 3.71, 95% CI 1.50–9.18 for genus Collinsella; p = 0.0341, OR = 2.40, 95% CI 1.07–5.39 for genus Phascolarctobacterium; p = 0.0107, OR = 2.06, 95% CI 1.18–3.60 for genus Prevotella9; p = 0.0269, OR = 2.08, 95% CI 1.09–3.98 for genus Adlercreutzia) and one bacterial trait was suggestively associated with a lower risk of sTP (p = 0.0235, OR = 0.46, 95% CI 0.23–0.90 for genus Slackia). Nevertheless, the findings obtained from the weighted median method did not provide evidence for a causal effect. The MR-Egger and MR-PRESSO tests indicated no evidence of horizontal pleiotropy or outliers (p > 0.05). Furthermore, the results of Cochrane's Q-test revealed no significant heterogeneity (p > 0.05) (Table 2). Additionally, leave-one-out analyses demonstrated that none of the identified causal associations were affected by any individual independent variable (Supplementary Fig. 2). None of the outcomes from Bonferroni's corrected test indicated the presence of a causal effect.
Finally, our inverse MR analysis using the IVW method did not reveal any potential inverse association of gut flora with these three diseases.
Discussion
Our research employed magnetic resonance analysis and comprehensive genome-wide association study (GWAS) data to explore potential causal connections between gut microbiota and HSP, ITP, and sTP at the genetic prediction level, yielding findings with strong causal explanatory capability. This two-sample MR investigation identified a combined total of 12 bacterial taxa potentially linked to the susceptibility of ITP, 6 bacterial taxa associated with the susceptibility of HSP, and 9 bacterial taxa associated with the susceptibility of sTP. Given the potential for bias in IVW analyses, additional MR methods may help to identify causal relationships. Our sensitivity analyses, using various MR methods and restricted IV groups, found that two bacterial taxa, genus Coprococcus3 and genus Gordonibacter, were associated with the risk of ITP. Additionally, two bacterial taxa, such as order Methanobacteriales and class Methanobacteria, were associated with the risk of HSP. Four bacterial taxa, including genus RuminococcaceaeUCG013, family Methanobacteriaceae, order Methanobacteriales, and class Methanobacteria, were found to be correlated with sTP. In addition, MR analyses between gut flora and sTP and ITP identified the presence of the same instrumental variables and bacterial taxa with potential causal associations, such as family Methanobacteriaceae order, order Methanobacteriales, and class Methanobacteria, have all been shown to have a negative correlation with sTP and ITP. This indicates that there is a common mechanism or potential interference in the correlation between Methanobacteria and ITP and sTP.
Our study revealed that genus Coprococcus3 (OR: 2.05; 95% CI 1.09, 3.88; p = 0.0264) and genus Gordonibacter (OR: 1.38; 95% CI 1.09, 1.75; p = 0.0073) were positively associated with the risk of ITP. Coprococcus is a significant member of the family Lachnospiraceae (phylum Firmicutes), an important genus of intestinal bacteria, with most of the strains being isolated from feces. It is also a significant producer of butyric acid. In a research conducted on a cohort of middle-aged individuals from Southwest China, it was observed that there was an inverse relationship between the abundance of Firmicutes and the levels of IgM. Additionally, the ratio of Firmicutes/Bacteroidetes was found to have a positive correlation with IgG and IgM levels [29]. It has been reported that the abundance of the phylum. Firmicutes and the ratio of Firmicutes/Bacteroidetes are correlated with Th17-Tregs anti-differentiation, which is regulated by IL-6 [30]. These findings suggest that they play a crucial role in regulating the immune homeostasis of systemic inflammation [31]. Upregulation of IL-6 has been associated with autoimmune diseases such as rheumatoid arthritis, multiple sclerosis, and asthma [22, 32].Additionally, a negative correlation was found between Th17 cells and elevated levels of Roseburia and Coprococcus in patients with primary Sjögren's syndrome [33]. An elevated presence of Coprococcus has been noted in female HLA-B27 positive patients compared to healthy controls [34]. Additionally, Coprococcus was found to be lower in the IgA nephropathy group but higher in the IgA vasculitis group [35]. The mechanisms through which Coprococcus3 may impact ITP could involve several factors. It has been demonstrated that C. faecalis regulates the production of the cytokines IL-1β and IL-6, thereby coordinating the inflammatory response during infection [36]. Stimulation with Candida albicans mycelia resulted in increased levels of IL-1β and IL-6 in circulating peripheral blood mononuclear cells, and this reaction was specifically linked to Coprococcus. Additionally, this finding showed a negative correlation with the production of IL-22 induced by S. aureus [37]. In contrast, IL-6 levels were significantly higher in patients with ITP [22]. Curiously, bacteria of the genus Coprococcus have also been researched for their anti-inflammatory effects, as they help suppress immune responses and lessen the severity of allergic reactions. The unique role of Coprococcus3 in ITP warrants further investigation.
Coprococcus is strongly associated with lipid metabolism. A study of 534 healthy adult Dutch volunteers found that more than 20% of metabolites were strongly associated with platelet activation markers, with the majority being lipid-related. This suggests that the gut microbiota plays a critical role in regulating platelet function [38]. In patients with ITP, the gut microbiota regulates elevated lipid metabolites such as RvD2, eicosatetraenoic acid, monooleic acid, and phosphatidylcholine. RvD2 has been shown to reduce platelet activation and maintain platelet function. Eicosatetraenoic acid is believed to regulate platelet homeostasis. Monooleic acid and phosphatidylcholine are both considered key compounds that promote platelet activation [22]. Metabolic profiling revealed a negative correlation between steatosis and aromatic and branched-chain amino acids, as well as glycoprotein acetyl groups, along with alpha diversity and fecal cocci 3 [39]. Anaerobic Coprococcus3 was associated with a lower incidence of steatosis. In contrast, Hoyles et al. found through liver biopsy that the genus Coprococcus was associated with a lower presence of hepatocyte balloons [40]. We hypothesize that Coprococcus3 may be involved in regulating platelet activation and homeostasis by influencing lipid metabolism, but further comprehensive studies are required to confirm this.
Gordonibacter, a group of Gram-positive bacteria belonging to the family Eggerthellaceae, is capable of metabolizing dietary ellagitannins, which are hydrolyzed to ellagic acid and then into urolithin [41]. Urolithin is a dibenzopyrone metabolite that has high bioavailability and shows anti-inflammatory activity in vivo [42]. It also promotes the browning of adipocytes, enhances cholesterol metabolism, inhibits the growth of transplanted tumors, mitigates inflammation, and down-regulates neuronal amyloid formation through the β3-AR/PKA/p38MAPK, ERK/AMPKα/SREBP1, PI3K/AKT pathways [43]. However, the mechanism of Gordonibacter in ITP has not been studied. Furthermore, Lachnospiraceae, Methanobacteria, and Bacteroidales S24, which are associated with ITP, exhibit a negative correlation with ITP. This suggests that these species may offer protection against the disease. Xiaomin Yu and his team found that the Lachnospiraceae NC2004 was depleted in ITP patients, consistent with the results of this analysis. However, the genus. Bacteroides in ITP patients showed enrichment [22].
Our study revealed a potential negative correlation between family Actinomycetaceae, order Actinomycetales and the risk of developing HSP. HSP is a systemic vasculitis primarily induced by IgA-mediated immune complex deposition in the vessel wall, often involving IgG class as well. In recent research, the activation of immune cells, particularly granulocytes, results in the release of inflammatory factors [44]. At present, there was currently limited evidence to suggest a direct correlation between Actinomycetaceae and HSP. However, Actinomycetaceae may be associated with certain infectious diseases that could potentially impact platelet function. Further research was needed to explore any potential relationship between Actinomycetaceae and HSP. However, Our study revealed a potential positive correlation between family Defluviitaleaceae, genus DefluviitaleaceaeUCG011 and the risk of developing HSP. In contrast, the genus Sutterella showed a negative correlation with HSP, as determined by MR analysis using Finnish data available on the OPEN GWAS website (GWAS ID: Finn-b-d3_allergpura). The family Defluviitaleaceae and its genus Defluviitaleaceae UCG011 may have a modulating effect on granulocytes through CD11c. Neutrophil infiltration plays an important role as a mediator during vascular injury in HSP [45]. CD11c is an integrin typically expressed on dendritic cells and is involved in multiple immune responses [46, 47]. The Defluviitaleaceae UCG011 has been found to be positively associated with urticaria and granulomatous polyangiitis [46, 48]. Sutterella is the type genus of the family Sutterellaceae within the order Burkholderiales of the class β-Proteobacteria. The genus Sutterella does not directly induce significant inflammation, but rather acts by degrading IgA, affecting local or widespread immune barriers. This mechanism is implicated in the pathogenesis of ulcerative colitis [49]. A study published in Nature revealed that elevated levels of Sutterella in the gut microbiota resulted in reduced IgA levels in the feces or intestinal mucosa. The presence of Sutterella resulted in the degradation of IgA, and both live bacteria and bacterial lysates degraded the free or bound secreted components of IgA due to the production and secretion of IgA proteases [50]. We hypothesize that this may be one of the primary reasons for the negative correlation between Sutterella and HSP.
This study has some limitations. Firstly, the data analyzed was sourced from open databases and aggregated. The study population is not homogeneous, consisting mostly of individuals of European origin, but also including a small number of participants from other ethnicities. Therefore, it is not possible to further explore the correlation between intestinal flora and disease through subgroup analysis. It is also not possible to directly generalize the results of the analysis to other racial groups. Secondly, the smallest category of gut flora analyzed in this study was the genus, which limited the exploration of the causal relationship between gut microbiota and disease at the level of specific bacterial species. Thirdly, we chose p < 1.0 × 10–5 as the screening criterion for gut microbiota in order to obtain sufficient IV. Fourth, Data obtained from public databases usually consist of results from genome-wide association studies, including genotype frequencies, genetic variations, and disease associations. However, they generally lack clinical information. As a result, we are unable to conduct subgroup analyses on important clinical characteristics of ITP patients, such as gender, age, new diagnosis or chronic phase, treatment regimens, prognosis, and mortality. These clinical characteristics are crucial for discussing the relationship between the pathogenesis of ITP and gut microbiota. In future clinical studies, we will make efforts to collect and enhance these essential clinical details to facilitate a more comprehensive discussion on the connection between the pathogenesis of ITP and gut microbiota.
Conclusion
In summary, the current study indicates a potential cause-and-effect association between the composition of gut microbiota and the development of ITP and HSP, as inferred from the examination of Mendelian randomization involving 196 gut microbiota. The genes predicted that genera Coprococcus3 and genus Gordonibacter were found to be correlated with a heightened susceptibility to ITP. Family Actinomycetaceae, order Actinomycetales was negatively associated with HSP. This study identified specific microbiota associated with ITP, HSP by MR analysis. These findings may offer insights into the pathogenesis of these conditions and propose new therapeutic approaches.
Data availability
The data for this study were sourced from publicly available databases. The data on HSP/ITP/sTP was obtained from the FinnGen release data R9 (https://www.finngen.fi/en/access_results), while the data on the human gut microbiome was acquired from the GWAS dataset of the MiBioGen International Consortium (https://mibiogen.gcc.rug.nl/).
References
Pillebout E, Sunderkotter C. IgA vasculitis. Semin Immunopathol. 2021;43(5):729–38.
Cooper N, Ghanima W. Immune thrombocytopenia. N Engl J Med. 2019;381(10):945–55.
Ramey C, LePera A. A possible case of nitrofurantoin-associated immune thrombocytopenia in a healthy 45-year-old Caucasian female. Cureus. 2023;15(2): e34654.
Zeller B, Rajantie J, Hedlund-Treutiger I, et al. Childhood idiopathic thrombocytopenic purpura in the Nordic countries: epidemiology and predictors of chronic disease. Acta Paediatr. 2005;94(2):178–84.
Bussel J, Cooper N, Boccia R, Zaja F, Newland A. Immune thrombocytopenia. Expert Rev Hematol. 2021;14(11):1013–25.
Gernsheimer T, James AH, Stasi R. How I treat thrombocytopenia in pregnancy. Blood. 2013;121(1):38–47.
Wang LY, Liu KK, Chu JH, et al. Influencing factors in the chronicity of immune thrombocytopenia in children. Zhongguo Shi Yan Xue Ye Xue Za Zhi. 2021;29(3):881–6.
Zufferey A, Kapur R, Semple JW. Pathogenesis and therapeutic mechanisms in immune thrombocytopenia (ITP). J Clin Med. 2017;6(2):16.
Provan D, Arnold DM, Bussel JB, et al. Updated international consensus report on the investigation and management of primary immune thrombocytopenia. Blood Adv. 2019;3(22):3780–817.
Heitink-Polle K, Uiterwaal C, Porcelijn L, et al. Intravenous immunoglobulin vs observation in childhood immune thrombocytopenia: a randomized controlled trial. Blood. 2018;132(9):883–91.
Nikolaishvili M, Pazhava A, Di Lernia V. Viral infections may be associated with Henoch–Schonlein Purpura. J Clin Med. 2023;12(2):697.
Clemente JC, Ursell LK, Parfrey LW, Knight R. The impact of the gut microbiota on human health: an integrative view. Cell. 2012;148(6):1258–70.
Maynard CL, Elson CO, Hatton RD, Weaver CT. Reciprocal interactions of the intestinal microbiota and immune system. Nature. 2012;489(7415):231–41.
Rooks MG, Garrett WS. Gut microbiota, metabolites and host immunity. Nat Rev Immunol. 2016;16(6):341–52.
Zhang X, Zhang D, Jia H, et al. The oral and gut microbiomes are perturbed in rheumatoid arthritis and partly normalized after treatment. Nat Med. 2015;21(8):895–905.
Wen C, Zheng Z, Shao T, et al. Correction to: Quantitative metagenomics reveals unique gut microbiome biomarkers in ankylosing spondylitis. Genome Biol. 2017;18(1):214.
Franzosa EA, Sirota-Madi A, Avila-Pacheco J, et al. Gut microbiome structure and metabolic activity in inflammatory bowel disease. Nat Microbiol. 2019;4(2):293–305.
Collison J. Gut microbiota linked to kidney disease in SLE. Nat Rev Rheumatol. 2019;15(4):188.
Qin J, Li Y, Cai Z, et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature. 2012;490(7418):55–60.
Jie Z, Xia H, Zhong SL, et al. The gut microbiome in atherosclerotic cardiovascular disease. Nat Commun. 2017;8(1):845.
Liu C, Cheng L, Ji L, et al. Intestinal microbiota dysbiosis play a role in pathogenesis of patients with primary immune thrombocytopenia. Thromb Res. 2020;190:11–9.
Yu X, Zheng Q, He Y, et al. Associations of gut microbiota and fatty metabolism with immune thrombocytopenia. Front Med (Lausanne). 2022;9: 810612.
Goleva E, Harris JK, Robertson CE, Jackson LP, Martin RJ, Leung D. Airway microbiome and responses to corticosteroids in corticosteroid-resistant asthma patients treated with acid suppression medications. J Allergy Clin Immunol. 2017;140(3):860-862.e1.
Wang Y, Liu F, Zhang G, et al. Gut microbiome alterations and its link to corticosteroid resistance in immune thrombocytopenia. Sci China Life Sci. 2021;64(5):766–83.
Wang X, Zhang L, Wang Y, et al. Gut microbiota dysbiosis is associated with Henoch–Schonlein Purpura in children. Int Immunopharmacol. 2018;58:1–8.
Wen M, Dang X, Feng S, et al. Integrated analyses of gut microbiome and host metabolome in children with Henoch–Schonlein purpura. Front Cell Infect Microbiol. 2021;11: 796410.
Wang JK, Yan B, Zhao JM, Yuan LP. Effect of gut microbiota from Henoch–Schonlein purpura patients on acid-sensitive ion channel 3 expression and intestinal motility in germ-free rats. BMC Pediatr. 2021;21(1):536.
Kurilshikov A, Medina-Gomez C, Bacigalupe R, et al. Large-scale association analyses identify host factors influencing human gut microbiome composition. Nat Genet. 2021;53(2):156–65.
Shen X, Miao J, Wan Q, et al. Possible correlation between gut microbiota and immunity among healthy middle-aged and elderly people in southwest China. Gut Pathog. 2018;10:4.
Ruiz L, Lopez P, Suarez A, Sanchez B, Margolles A. The role of gut microbiota in lupus: what we know in 2018? Expert Rev Clin Immunol. 2018;14(10):787–92.
Zheng X, Wang D. The adenosine A2A receptor agonist accelerates bone healing and adjusts Treg/Th17 cell balance through interleukin 6. Biomed Res Int. 2020;2020:2603873.
Zhu XM, Shi YZ, Cheng M, Wang DF, Fan JF. Serum IL-6, IL-23 profile and Treg/Th17 peripheral cell populations in pediatric patients with inflammatory bowel disease. Pharmazie. 2017;72(5):283–7.
Xin X, Wang Q, Qing J, et al. Th17 cells in primary Sjogren’s syndrome negatively correlate with increased roseburia and coprococcus. Front Immunol. 2022;13: 974648.
Stoll ML, Sawhney H, Wells PM, et al. The faecal microbiota is distinct in HLA-B27+ ankylosing spondylitis patients versus HLA-B27+ healthy controls. Clin Exp Rheumatol. 2023;41(5):1096–104.
Hu X, Fan R, Song W, et al. Landscape of intestinal microbiota in patients with IgA nephropathy, IgA vasculitis and Kawasaki disease. Front Cell Infect Microbiol. 2022;12:1061629.
Zhao J, Pan X, Hao D, et al. Causal associations of gut microbiota and metabolites on sepsis: a two-sample Mendelian randomization study. Front Immunol. 2023;14:1190230.
Schirmer M, Smeekens SP, Vlamakis H, et al. Linking the human gut microbiome to inflammatory cytokine production capacity. Cell. 2016;167(4):1125-1136.e8.
Vadaq N, Schirmer M, Tunjungputri RN, et al. Untargeted plasma metabolomics and gut microbiome profiling provide novel insights into the regulation of platelet reactivity in healthy individuals. Thromb Haemost. 2022;122(4):529–39.
Alferink L, Radjabzadeh D, Erler NS, et al. Microbiomics, metabolomics, predicted metagenomics, and hepatic steatosis in a population-based study of 1,355 adults. Hepatology. 2021;73(3):968–82.
Hoyles L, Fernandez-Real JM, Federici M, et al. Molecular phenomics and metagenomics of hepatic steatosis in non-diabetic obese women. Nat Med. 2018;24(7):1070–80.
Ngom II, Hasni I, Lo CI, et al. Taxono-genomics and description of Gordonibacter massiliensis sp. nov., a new bacterium isolated from stool of healthy patient. New Microbes New Infect. 2020;33:100624.
Selma MV, Tomas-Barberan FA, Beltran D, Garcia-Villalba R, Espin JC. Gordonibacter urolithinfaciens sp. nov., a urolithin-producing bacterium isolated from the human gut. Int J Syst Evol Microbiol. 2014;64(Pt 7):2346–52.
Zhang M, Cui S, Mao B, et al. Ellagic acid and intestinal microflora metabolite urolithin A: a review on its sources, metabolic distribution, health benefits, and biotransformation. Crit Rev Food Sci Nutr. 2023;63(24):6900–22.
Cartin-Ceba R, Peikert T, Specks U. Pathogenesis of ANCA-associated vasculitis. Curr Rheumatol Rep. 2012;14(6):481–93.
Fu SQ, Guo YJ, Song CL. Expression and significance of endothelial microparticles in children with Henoch-Schonlein purpura. Zhongguo Dang Dai Er Ke Za Zhi. 2018;20(10):831–4.
Chen Y, Tang S. Gut microbiota and immune mediation: a Mendelian randomization study on granulomatosis with polyangiitis. Front Immunol. 2023;14:1296016.
Wang Q, Su X, He Y, et al. CD11c participates in triggering acute graft-versus-host disease during bone marrow transplantation. Immunology. 2021;164(1):148–60.
Shi YZ, Tao QF, Qin HY, Li Y, Zheng H. Causal relationship between gut microbiota and urticaria: a bidirectional two-sample mendelian randomization study. Front Microbiol. 2023;14:1189484.
Kaakoush NO. Sutterella species, IgA-degrading bacteria in ulcerative colitis. Trends Microbiol. 2020;28(7):519–22.
Moon C, Baldridge MT, Wallace MA, et al. Vertically transmitted faecal IgA levels determine extra-chromosomal phenotypic variation. Nature. 2015;521(7550):90–3.
Funding
This work was supported by grants from Medical Science and Technology Project of Henan (LHGJ20200469).
Author information
Authors and Affiliations
Contributions
Chendong Jiang and Enwu Yuan designed this work and analyzed the data; Chendong Jiang, Shu Deng, Xiaohan Ma and Juan Song collected and analyzed the data. Jinpeng Li helped compile the data; Chendong Jiang and Enwu Yuan wrote the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no competing financial interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Jiang, C., Deng, S., Ma, X. et al. Mendelian randomization reveals association of gut microbiota with Henoch–Schönlein purpura and immune thrombocytopenia. Int J Hematol 120, 50–59 (2024). https://doi.org/10.1007/s12185-024-03777-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12185-024-03777-1