Abstract
In this paper, the efficient magnetic dummy molecular imprinted polymers were firstly synthesized by dummy template (raffinose). The dummy molecularly imprinted layer was polymerized on the surface of Fe3O4 nanoparticles modified with NH2 group. The micromorphology and chemical composites of prepared polymers were detailed characterized. Their adsorption and desorption conditions including extraction time, adsorbent dosage, solution pH, washing solvent, and elution solvent were carefully evaluated. The results showed that the obtained magnetic dummy template molecularly imprinted polymers showed uniform sizes, high magnetic responsivity, core–shell structure, and selectively recognition capacity towards five aminoglycoside antibiotics. Under optimum conditions, magnetic solid-phase extraction coupled high performance liquid chromatography-tandem mass spectrometry for enriching and determining aminoglycoside antibiotics in milk samples was developed. This method showed good linearity (50–1000 μg/kg) with r2 larger than 0.9923. The limits of detection (S/N = 3) were from 3.6 to 9.6 μg/kg. With spiked at three different concentration levels, the recoveries fell into 82.6–114.1% with relative standard deviation from 2.8 to 13.2%. The established method based on magnetic dummy molecular imprinted polymers proved to be highly sensitive and repeatable, providing the promising adsorbent for aminoglycoside antibiotics analysis in the complex samples.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Aminoglycoside antibiotics (AAs) are an important class of antibiotics composed of amino sugars and aminocyclic alcohols via oxygen bridges (Wang et al. 2016). AAs have been widely used in animal husbandry and veterinary medicine for inhibiting the growth and reproduction of bacteria and promoting the growth of animals (Yue et al. 2021; Zhang et al. 2020). In spite of their effectively antimicrobial effect, AAs have also been banned for feed addition agent and many countries have established the maximum residue limits of AAs in food owing to the high toxic effects towards the human beings such as nephrotoxicity, ototoxicity, and neuromuscular blockade (Prayle et al. 2010; Jospe-Kaufman et al. 2020). The residual AAs could be easily absorbed by human body through food chain. Therefore, a highly sensitive, rapid, and efficient method to determine AAs in foods is mandatory.
The employed approaches for the determination of AAs involve spectrophotometric, immunochemical, microbiological, and chromatographic techniques (Ghodake et al. 2020; Mabrouk et al. 2020; Xu et al. 2011; Glinka et al. 2020). Among them, solid-phase extraction (SPE) coupled with high performance liquid chromatography-tandem mass spectrometry (HPLC–MS/MS) was the commonly used method in different food matrix (Tarannum et al. 2020).
Although HPLC–MS/MS method could achieve satisfactory limits of detection, the conventional SPE process often lacked flexibility and consumed time. Furthermore, the used enriching or purifying materials (e.g., exchange resin, C18) all lack selectivity towards AAs in both the reverse-phase and the ion-exchange modes in SPE process. These shortcomings could not satisfy high-throughput screening of AAs in complex samples and resulted in significant matrix effect from food constituents (Farouk et al. 2015). In recent years, magnetic solid-phase extraction (MSPE) is a new mode of SPE based on magnetic inorganic material and non-magnetic adsorbent, eliciting growing interest for sample pretreatment. MSPE could overcome the deficiencies of SPE and perform superiorities including easy operations, little or no solvent consumption, and rapid extraction (Yu et al. 2019; Li et al. 2018). The adsorbent plays a key role for enriching targets and decides the performance of MSPE. Thus, in order to develop a highly efficient MSPE process for AAs in food samples, the new selective adsorbent should be designed.
Molecularly imprinted polymers (MIPs), as artificial-synthetic biomimetic recognition materials, have been widely utilized as promising adsorbent for a huge range of target molecules in MSPE process, showing high selectivity, excellent mechanical and chemical stability, and allowing to work in harsh conditions (Moura et al. 2018; Hassan et al. 2018; Hu et al. 2018). In view of applications of MIPs for AAs detection, only streptomycin as template was used to form polymers (Ji et al. 2013) and recent several studies directly bought expensive SPE-aminoglycoside cartridge from Sigma-Aldrich (Moreno-González et al. 2015, 2017; Savoy et al. 2018). However, the further applications of traditional MIPs not only consumed large amounts of standard products, but also were greatly limited owing to the unavoidable template leakage, influencing the accurate quantization of the analytes. To avoid those deficiencies, the synthetic strategy of MIPs with the structural analogs of analytes, named dummy templates, has been developed and the obtained magnetic dummy template-MIPs (MDMIPs) were successfully used for enriching different trace analytes in MSPE process (Tan et al. 2016; Zhao et al. 2018; Guo et al. 2019; Gao et al. 2018; Dinc et al. 2019; Yuan et al. 2020; Marć and Wieczorek 2020; Fu et al. 2019). Recently, the dummy molecularly imprinted polymers based on template-raffinose by precipitation polymerization were synthesized by our team and dummy molecularly imprinted polymers (DMIPs) cartridge was applied to enrich AAs in environmental water samples (Zhang et al. 2020). Considering the mentioned superiority of MSPE, it is necessary to develop magnetic enriching process. To the best of our knowledge, there have been no articles reporting the use of magnetic dummy molecular imprinted polymers for determining AAs so far.
In this paper, based on the previous study (Zhang et al. 2020), the cheap, nontoxic compound-raffinose was used as dummy template to furtherly synthesize magnetic polymers by surface imprinting techniques. DMIPs shell was polymerized on the surface of Fe3O4 and lessened the molecule diffusion barrier, resulting in easily removing template. The synthesis of MDMIPs for extracting and enriching AAs was firstly described. The adsorption characteristics and the selectivity of MDMIPs were investigated by binding experiments. The MDMIPs as the adsorbent coupled with hydrophilic interaction-high performance liquid chromatography-tandem mass spectrometry (HILIC-HPLC–MS/MS) were successfully applied to analyze AAs in milk.
Experimental
Materials
Methacrylic acid (MAA), trimethylolpropane triacrylate (TRIM), tetraethyl orthosilicate (TEOS, 98%), (3-aminopropyl) triethoxysilane (APTES, 98%), formic acid (FA, 98%), and ammonium acetate were bought from Sigma-Aldrich (St. Louis, MO, USA). The initiator 2,2-azobisisobutyronitrile (AIBN, > 98%), FeCl3·6H2O (99%), and FeCl2·4H2O (98%) were supplied by the Reagent & H. V. Chemical Co. Ltd. (Shanghai, China). Deionized water obtained from a Milli-Q system (France) was used. Hexane (Hex), methanol (MeOH), and acetonitrile (ACN) were supplied by Tedia (Fairfield, USA). Raffinose (RAF) and six AAs including kanamycin sulfate (KAM), apramycin sulfate (APM), gentamycin sulfate (GNT), tobramycin (TOM), paromomycin sulfate (PRM), and spectinomycin pentahydrate dihydrochloride (SPC) were purchased from Dr. Ehrenstorfer Gmbh (Augsburg, Germany). These standards were stored under refrigeration at 4 °C and used to prepare working standard solutions.
Apparatus
FT-IR spectra were collected using a spectrum 100 instrument (PerkinElmer, USA). Morphological characterization was carried out on a scanning electron microscopy (S-8200 SEM, Japan) and a transmission electron microscope instrument (TEM LIBRA-200 FE, Germany), respectively. The magnetization curves were obtained by a LakeShore7407 vibrating sample magnetometer (VSM, USA). KQ-500DB ultrasonic cleaner (Kunshan, China) was also used. An HPLC system (ultraLC-100, AB Sciex, USA) coupled with a triple quadrupole mass spectrometer (AB-4000, USA) with an electrospray ionization source was used to analyze AAs. The detailed parameters of mass spectrometer and chromatographic conditions were shown in supplementary materials (Table S1 and Section S1.1).
Synthesis of NH2-functionalized Magnetic Nanoparticles
The Fe3O4 nanoparticles were prepared with chemical co-precipitation method according to reported literature (Mashhadizadeh et al. 2013) and the detailed procedures were listed in supplementary documents (Section S1.2).
The synthesized Fe3O4 (0.5 g) were dispersed in 120 mL ethanol–water (5:1, v/v) in 250 mL three-neck flask, followed by the addition of 1.0 mL of ammonium hydroxide (25 wt%) and 2.0 mL of TEOS sequentially. After the mixture reacted for 24 h at room temperature under continuous stirring, 1.5 mL APTES was added into the mixture and vigorously stirred for another 24 h at 40 °C. The obtained Fe3O4@SiO2-NH2 microspheres were collected by magnetic separation and alternatively washed three times with deionized water and ethanol. Finally, the Fe3O4@SiO2-NH2 microspheres were dried under vacuum at 80 °C for 24 h.
Synthesis of Magnetic Imprinted Polymers
0.l mmol template-RAF and 1.4 mmol functional monomer-MAA were firstly dissolved in 10 mL methanol in a 150 mL round-bottom flask, and the flask was shaken for 30 min to pre-polymerize. After Fe3O4@SiO2-NH2 microspheres (100 mg) were dispersed in the pre-polymerization solution, 1.0 mmol TRIM and 30 mg initiator-AIBN were added in the flask flowing with ultrasonication for 5 min. The mixtures were deoxygenated with a stream of nitrogen. The flask was sealed and placed in water bath at 60 °C for 24 h. Then, MDMIPs were separated by magnet and eluted by methanol-acetic acid solution (9:1, v/v) until RAF could not be detected. Finally, the polymers were washed by methanol and dried in accordance with the above method. The magnetic dummy non-imprinted polymers (MDNIPs) were prepared following the same procedure without RAF molecule.
Magnetic Solid-phase Extraction
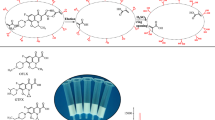
A schematic representation of proposed MSPE process based on MDMIPs was shown in Fig. 1. The detailed procedure included the following steps: 30 mg MDMIPs adsorbents were sequentially conditioned with 2 mL methanol and 2 mL ultra-pure water with vortex for 1 min in 10 mL tube, and rapidly separated by the magnet. Then, 5 mL milk extracts were added into this tube and vortexed vigorously for 10 min in an oscillator to completely adsorb analytes. The adsorbents were separated again from the mixture with external magnet. After washing with 2 mL ACN, 2 mL 0.5% FA-H2O:ACN (50:50 v/v, 1 mL for 3 min ultrasonication, repeated 2 times) was used to elute AAs. After magnetic separation, the desorption solution was transferred to a new 5 mL tube and filtered through a 0.22 microfiltration membrane for HPLC–MS/MS analysis.
Binding Experiments
To study the adsorbing performance of MDMIPs, dynamic absorption and static adsorption experiments were carried out, respectively. Firstly, the sized and washed magnetic adsorbents (10.0 mg) were added into 2 mL of 20 μg/g of GNT water solution with different periods of shaking time (5–180 min) to set the saturation adsorption time. Then, the magnetic adsorbents (10 mg) were mixed into 2.0 mL of GNT solution ranging from 10 to 1000 μg/g at saturation adsorption time. After centrifugation, the supernatant was diluted and analyzed by HPLC–MS/MS to quantify the concentration of free GNT. The adsorption capacity was calculated according to Scatchard equation (Eq. 1).
where Q and Qmax were the amount of GNT adsorbed to polymers and the maximum number of binding sites, respectively; Ce and KD were the free GNT concentration at equilibrium and the equilibrium dissociation constant, respectively.
Milk Samples
The whole cow milk and the skimmed cow milk samples were purchased from the local market in the city of Yantai. Three milliliters of trichloroacetic acid (5% containing 0.5% EDTA-2Na m/v) was added to 2 g of milk samples and the mixture was vortexed for 3 min, then subjected to an ultrasonic bath for 5 min. After that, the samples were centrifuged for 10 min at 6000 g and the supernatant was collected into a tube. The above procedures were repeated with two times for extraction and the supernatant was merged together.
Results and Discussion
Characterization of Polymers
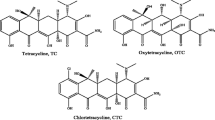
Selection of appropriate dummy template molecule offers an effective way to avoid the shortcoming of template leakage. In this study, due to the similar spatial structure of three or four six-membered ring with AAs (Fig. S1) and the abundant OH groups providing hydrogen bonding interaction with COOH in MAA, the RAF compound was selected as dummy template to synthesize MDMIPs.
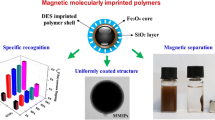
To understand morphology and chemical composites of the synthesized polymers, SEM, TEM, and FT-IR were used to characterize Fe3O4@SiO2-NH2, Fe3O4@DMIPs, and Fe3O4@DNIPs (Fig. 2). SEM graphs revealed the mean diameter of Fe3O4@SiO2-NH2 was about 20 nm. After polymerizing the imprinted layers, the diameter of Fe3O4@DMIPs increased to approximately 500 nm with a relatively narrow particle size distribution. This indicated that the dummy imprinted layers were successfully polymerized on the surface of Fe3O4 by the hydrogen bond interaction between the functional groups of template-RAF, monomer-MAA, crosslinking agent-TRIM, and NH2 group of APTES modified on the surface of Fe3O4. Compared to Fe3O4@SiO2-NH2, the TEM images of Fe3O4@DMIPs were found to be composed of the Fe3O4 core (dark) and the polymer shell (gray) with the thickness of approximately 60 nm, furtherly illustrated the uniform polymerization of DMIP shell on Fe3O4 and there were hardly any free Fe3O4 NPs.
Figure 3A showed the adsorption peaks of polymers measured by Fourier transform infrared. The spectra of Fe3O4@SiO2-NH2 displayed the specific peak at 1603 cm−1 assigned to N–H stretching vibration, illustrating the NH2 group of APTES successfully modified on Fe3O4@SiO2 (Liu et al. 2020). The characteristic peak at 966 cm−1, assigned to glycosidic bond existed in RAF (curve a) and MDMIPs (curve c), while MNIPs synthesized using the same procedure without the template molecule RAF did not show the peak (curve d) (Zhang et al. 2020). After the MDMIPs were washed with HAc-MeOH, the intensity of 966 cm−1 was disappeared, indicating the template-RAF was removed. Also, absorptions of 1732 cm−1 and 1259 cm−1 resulting from carboxyl groups in the functional monomer-MAA existed in all the three polymers (curves c, d, and e) (Ji et al. 2013). Those spectra of FT-IR demonstrated the coatings of DMIPs on the surface of magnetic nanoparticles.
Furthermore, in order to easily separate MDMIPs from matrix solution in MSPE process, the magnetic property of MDMIPs was evaluated by the vibrating sample magnetometer at room temperature. As showed in Fig. 3B, both remanence and coercivity were zero, confirming that the polymers were paramagnetic. The magnetic saturation value of MDMIPs was 12.4 emu·g−1 without magnetic hysteresis. Additionally, the MDMIP microspheres could be rapidly collected from the sample solution with external magnet (Fig. S2). The sample became clear and transparent after magnetic separation, indicating that the MSPE process could simplify the pretreatment steps for complex matrix.
Binding Study and Scatchard Analysis
To evaluate the binding properties of MDMIPs and MNIPs, equilibrium absorption and Scatchard analysis were employed. The saturation adsorption time (120 min) was obtained by adding 10 mg of magnetic adsorbents into 2 mL of 20 μg/g GNT water solutions (Fig. S3). In static adsorption experiment, the same dosages of magnetic adsorbents were mixed with GNT water solution (2.0 mL) from 10 to 1000 μg/g for 120 min, respectively (Fig. 4a). In spite of both the increasing adsorption amounts of MDMIPs and MNIPs for GNT with the increasing initial concentration, the higher binding capacity (16.0 mg/g) of MDMIPs than MNIPs (9.0 mg/g) at the initial concentration of 500 mg/L was performed owing to the specific binding sites of GNT molecules on MDMIPs. Furthermore, the apparent maximum binding amount (Qmax) and the equilibrium dissociation constant (KD) were calculated by Scatchard equation (Eq. 1). As shown in Fig. 4b, two different linear regression equations were achieved for MDMIPs. According to the equilibrium theory, it was inferred that the binding sites in MDMIPs were heterogeneous. The theoretical maximum adsorption amounts were calculated to be 8.7 mg/g for the low affinity sites and 20.5 mg/g for the high affinity sites, and the calculated values for KD were 17.39 μg mL−1 and 129.81 μg mL−1, respectively. All of above results indicated the favorable adsorbing capacity of MDMIPs for GNT.
Optimization of Magnetic Solid-phase Extraction Conditions
To ensure the analytes completely being adsorbed and desorbed from MDMIPs adsorbent, the adsorption and the desorption parameters including extraction time, adsorbent dosage, solution pH, washing solvents, elution solvent, and elution volume were optimized, respectively. Five-milliliter milk extracts with spiked concentration of 1 mg/kg of each AAs were used in the following optimizing experiments.
Extraction Time
During the equilibrium-based MSPE process, extraction time is one of the primary factors which influence the extraction recovery. The extraction time from 1 to 20 min was investigated (Fig. 5a). The results indicated that AAs recovery both improved (e.g., KAM from 42.6 to 91.0%) with the increasing extraction time from 1 to 10 min, and then did not obviously change from 10 to 15 min. When the extraction time was more than 15 min, the recovery of AAs tended to lightly decrease. This might be the occurrence of desorption. Thus, the extraction time of 10 min was chosen for the complete extraction time.
Adsorbent Dosage
In order to obtain the highest recovery for enriching AAs, the adequate amount of MDMIPs is a critical factor. In this experiment, 10 to 40 mg of adsorbents were evaluated for extracting AAs in 5 mL milk extracts. Then, oscillation was performed for 10 min, and 2 mL H2O was used for washing. From Fig. 5b, it was showed that the recovery of five AAs reached a plateau while the added amount was 30 mg. Therefore, the optimum amount of adsorbents was 30 mg in the following experiments.
Solution pH
The recognition sites in the MDMIPs mainly depend on the hydrogen bonding interaction between OH or NH2 groups in AAs and COOH in MAA. The pH value of the sample extracts is an important factor affecting not only the behavior of the liquid charge present on the surface of MDMIPs, but also the physical–chemical properties of analytes (Fang et al. 2021; Dinali et al. 2021). In this study, milk extracts with different pH in the range of 3.0–11.0 adjusted by ammonia and formic acid were studied. As shown in Fig. 5c, the recoveries of AAs increased from 3 to 7 and then decreased substantially. This might be that the strongest recognition capacity of imprinted polymers towards targets could be performed while the solution pH was close to their pKa. Therefore, the optimized pH value was at 7.
Washing Solvents
In the adsorbing process for AAs based on MDMIPs, the washing step could minimize unspecific interactions between the un-targeted molecules (e.g., organic acid, amino acid, carbohydrate) and the binding sites, effectively reducing the matrix effect. So, different washing solvents including 2 mL of Hex, ACN, MeOH, and H2O were investigated, respectively. In this experiment, after MDMIPs adsorbent adsorbed AAs, different solvents were added and vortexed for 2 min, respectively. Figure 5d showed that the higher recoveries of most AAs were achieved when 2 mL ACN used as washing solvent. So, 2 mL ACN was applied.
Desorption Conditions
After extracting AAs under optimum conditions (5 mL milk extracts, 30 mg adsorbents, 10 min of extraction time, 2 mL ACN for washing), the analytes should be desorbed from MDMIPs before detected by instruments. Different elution conditions have great influence on the desorbing efficiency. In order to obtain maximum efficiency, optimum elution condition should be established. Firstly, different elution solvents including 3 mL 1% FA-H2O and 1% FA-ACN:H2O (20:80, 50:50, 80:20 v/v) were investigated, respectively. After ultrasonic elution for 3 min, the elution ability of 1% FA-H2O:ACN (50:50 v/v) showed the highest recovery than other solvents (Fig. 5e). Then, the concentration of FA in elution solvent was also optimized. One milliliter of H2O:ACN (50:50 v/v) including 0.1%, 0.5%, and 1% FA was investigated. The recoveries improved with the increasing concentration of FA, a satisfactory recovery could be obtained when the concentration of FA reaches 0.5%, while the recoveries almost unchanged with 1% FA. Furthermore, the elution volumes were compared (Fig. S4) and it was found that 1 mL 0.5% FA-H2O:ACN (50:50 v/v) with ultrasonic elution for 3 min by two repeats was the optimal volume and desorption time. Therefore, 2 mL 0.5%FA-H2O:ACN (50:50 v/v) was finally selected as elution solvent.
Specific Recognition
As shown from Fig. 6, the molecular structure of SPC has relative structure with RAF. In order to evaluate the specific recognition ability of MSPE based on MDMIPs, the milk samples spiked at 1 μg/mL of each AAs were treated by MDMIPs. The recovery of SPC (17.7%) was obviously much lower than KAM, APM, GNT, TOM, and PRM (88.3%, 92.0%, 90.0%, 97.3%, 95.2%), indicating that MDMIPs possessed excellent class-specific recognition capacity towards those five targeted AAs.
Recyclability of MDMIPs
Because MDMIPs adsorbent is the key factor deciding the final results and the detecting cost, it is necessary to verify the recyclability of MDMIPs. After MDMIPs were completely eluted by 0.5% FA-50% ACN and furtherly rinsed by 3 mL MeOH and 3 mL deionized water, they were reused to extract AAs from milk samples. From Fig. 7, it could be seen that even after 6 recycling uses of MDMIPs, the recoveries of AAs were more than 80%.
Method Validation
The blank milk extracts without AAs were obtained by treating liquid milk with the MSPE process described in “Synthesis of Magnetic Imprinted Polymers” and “Magnetic Solid-phase Extraction” sections. The standard calibration curves were established using the blank milk extracts at six different concentration levels (50, 100, 200, 500, 800, and 1000 µg/kg for each AAs) and analysis were performed with five replicates for 50 µg kg−1. Linearity, correlation coefficient (r2), and limits of detection (LODs) were carefully studied. The results showed that all the analytes exhibited good linearity with r2 larger than 0.9923 (Table 1). The LODs (S/N = 3) were from 3.6 to 9.6 μg/kg. Those data illustrated the developed method was sensitive and applicable.
Analysis of Real Milk Samples
In order to further verify the accuracy of the method, the whole cow milk and the skimmed cow milk samples were obtained in local market in the city of Yantai and analyzed by MSPE-HPLC–MS/MS based on MDMIPs under optimized conditions. None of AAs was detected in actual milk samples. Both the two types of liquid milk were also spiked at three different concentration levels of AAs (50, 100, 200 µg/kg), and the typical chromatogram has been listed in Fig. S5. Table 2 showed that the recoveries fell into 82.6–114.1% and relative standard deviations (RSDs) were from 2.8 to 13.2%. Those results furtherly proved that the developed method possessed high accuracy and good repeatability.
Conclusions
This study showed that the efficient magnetic dummy molecular imprinted polymers with core–shell structure were successfully synthesized. The obtained MDMIPs showed uniform sizes, high magnetic responsivity, and avoided template leakage in the sample pretreatment, making simple operation and good selective enriching capacity towards AAs. Under optimum conditions, MSPE-HPLC–MS/MS method was successfully applied to determine AAs in milk samples. The established method proved to be highly sensitive and repeatable, providing the promising potential adsorbent and method for AAs analysis in the complex samples.
References
Dinali LAF, Oliveira HL, Teixeira LS, Borges WS, Borges KB (2021) Mesoporous molecularly imprinted polymer core@shell hybrid silica nanoparticles as adsorbent in microextraction by packed sorbent for multiresidue determination of pesticides in apple juice. Food Chem 345:128745. https://doi.org/10.1016/j.foodchem.2020.128745
Dinc M, Esen C, Mizaikoff B (2019) Recent advances on core-shell magnetic molecularly imprinted polymers for biomacromolecules. Trac-Trend Anal Chem 144:202–217. https://doi.org/10.1016/j.trac.2019.03.008
Fang L, Miao YX, Wei D, Zhang Y, Zhou YC (2021) Efficient removal of norfloxacin in water using magnetic molecularly imprinted polymer. Chemosphere 262:128032. https://doi.org/10.1016/j.chemosphere.2020.128032
Farouk F, Azzazy HME, Niessen WMA (2015) Challenges in the determination of aminoglycoside antibiotics, a review. Anal Chim Acta 890:21–43. https://doi.org/10.1016/j.aca.2015.06.038
Fu X, Zhu DL, Huang L, Yan XY, Liu SM, Wang CH (2019) Superparamagnetic core-shell dummy template molecularly imprinted polymer for magnetic solid-phase extraction of food additives prior to the determination by HPLC. Microchem J 150:104169. https://doi.org/10.1016/j.microc.2019.104169
Gao D, Wang DD, Fu QF, Wang LJ, Zhang KL, Yang FQ, Xia ZN (2018) Preparation and evaluation of magnetic molecularly imprinted polymers for the specific enrichment of phloridzin. Talanta 178:299–307. https://doi.org/10.1002/jssc.201601116
Ghodake G, Shinde S, Saratale RG, Kadam A, Saratale GD, Syed A, Marraiki N, Elgorban AM, Kim DY (2020) Silver nanoparticle probe for colorimetric detection of aminoglycoside antibiotics: picomolar-level sensitivity toward streptomycin in water, serum, and milk samples. J Sci Food Agric 100:874–884. https://doi.org/10.1021/acsami.9b00277
Glinka M, Wojnowski W, Wasik A (2020) Determination of aminoglycoside antibiotics: current status and future trends. Trac-Trend Anal Chem 131:116034. https://doi.org/10.1016/j.trac.2020.116034
Guo LH, Ma XG, Xie XW, Huang RF, Zhang MY, Li J, Zeng GL, Fan YM (2019) Preparation of dual-dummy-template molecularly imprinted polymers coated magnetic graphene oxide for separation and enrichment of phthalate esters in water. Chem Eng J 361:245–255. https://doi.org/10.1016/j.cej.2018.12.076
Hassan AHA, Moura SL, Ali FHM, Moselhy WA, Sotomayors MDPT, Pividori MI (2018) Electrochemical sensing of methyl parathion on magnetic molecularly imprinted polymer. Biosens Bioelectron 118:181–187. https://doi.org/10.1016/j.bios.2018.06.052
Hu M, Huang P, Suo L, Wu FY (2018) Polydopamine-based molecularly imprinting polymers on magnetic nanoparticles for recognition and enrichment of ochratoxins prior to their determination by HPLC. Microchim Acta 185:300. https://doi.org/10.1002/jssc.201401142
Ji SL, Zhang FF, Luo X, Yang BC, Jin GW, Yan JY, Liang XM (2013) Synthesis of molecularly imprinted polymer sorbents and application for the determination of aminoglycosides antibiotics in honey. J Chromatogr A 1313:113–118. https://doi.org/10.1016/j.chroma.2013.08.072
Jospe-Kaufman M, Siomin L, Fridman M (2020) The relationship between the structure and toxicity of aminoglycoside antibiotics. Bioorg Med Chem Lett 30(13):127218. https://doi.org/10.1016/j.bmcl.2020.127218
Li N, Jiang HL, Wang XL, Wang X, Xu GJ, Zhang BB, Wang LJ, Zhao RS, Lin JM (2018) Recent advances in graphene-based magnetic composites for magnetic solid-phase extraction. Trac-Trend Anal Chem 102:60–74. https://doi.org/10.1016/j.trac.2018.01.009
Liu Y, Li ZQ, Jia L (2020) Synthesis of molecularly imprinted polymer modified magnetic particles for chiral separation of tryptophan enantiomers in aqueous medium. J Chromatogr A 1622:461147. https://doi.org/10.1016/j.chroma.2020.461147
Mabrouk MM, Noureldin HAM, Badr IHA, Saad AHK (2020) Simple spectrofluorimetric methods for determination of veterinary antibiotic drug (apramycin sulfate) in pharmaceutical preparations and milk samples. Spectrochim Acta A Mol Biomol Spectrosc 224:117395. https://doi.org/10.1016/j.saa.2019.117395
Marć M, Wieczorek PP (2020) The preparation and evaluation of core-shell magnetic dummy-template molecularly imprinted polymers for preliminary recognition of the low-mass polybrominated diphenyl ethers from aqueous solutions. Sci Total Environ 724:138151. https://doi.org/10.1016/j.scitotenv.2020.138151
Mashhadizadeh MH, Amoli-Diva M, Pourghazi K (2013) Magnetic nanoparticles solid phase extraction for determination of ochratoxin A in cereals using high-performance liquid chromatography with fluorescence detection. J Chromatogr A 1320:17–26. https://doi.org/10.1016/j.chroma.2013.10.062
Moreno-González D, Lara FJ, Jurgovská N, Gámiz-Gracia L, García-Campaña AM (2015) Determination of aminoglycosides in honey by capillary electrophoresis tandem mass spectrometry and extraction with molecularly imprinted polymers. Anal Chim Acta 891:321–328. https://doi.org/10.1016/j.aca.2015.08.003
Moreno-González D, Hamed AM, García-Campaña AM, Gámiz-Gracia L (2017) Evaluation of hydrophilic interaction liquid chromatography-tandem mass spectrometry and extraction with molecularly imprinted polymers for determination of aminoglycosides in milk and milk-based functional foods. Talanta 171:74–80. https://doi.org/10.1021/ac9006667
Moura SL, Fajardo LM, Cunha LDA, Sotomayor DPT, Machado FBC, Ferrao LFA, Pividori MI (2018) Theoretical and experimental study for the biomimetic recognition of levothyroxine hormone on magnetic molecularly imprinted polymer. Biosens Bioelectron 107:203–210. https://doi.org/10.1016/j.bios.2018.01.028
Prayle A, Watson A, Fortnum H, Smyth A (2010) Side effects of aminoglycosides on the kidney, ear and balance in cystic fibrosis. Thorax 65:654–658. https://doi.org/10.1080/00032719.2020.1734606
Savoy MC, Woo PM, Ulrich P, Tarres A, Mottier P, Desmarchelier A (2018) Determination of 14 aminoglycosides by LC-MS/MS using molecularly imprinted polymer solid phase extraction for clean-up. Food Addit Contam 35:674–685. https://doi.org/10.1080/19440049.2018.1433332
Tan L, He R, Chen KC, Peng RF, Huang C, Yang R, Tang YW (2016) Ultra-high performance liquid chromatography combined with mass spectrometry for determination of aflatoxins using dummy molecularly imprinted polymers deposited on silica-coated magnetic nanoparticles. Microchim Acta 183:1469–1477. https://doi.org/10.1002/jssc.201601445
Tarannum N, Khatoon S, Dzantiev BB (2020) Perspective and application of molecular imprinting approach for antibiotic detection in food and environmental samples: a critical review. Food Control 118:107381. https://doi.org/10.1016/j.foodcont.2020.107381
Wang Y, Li S, Zhang F, Lu YF, Yang BC, Zhang F, Liang XM (2016) Study of matrix effects for liquid chromatography-electrospray ionization tandem mass spectrometric analysis of 4 aminoglycosides residues in milk. J Chromatogr A 1437:8–14. https://doi.org/10.1002/jssc.201200344
Xu N, Qu C, Ma W, Xu L, Liu L, Kuang H, Xu C (2011) Development and application of one-step ELISA for the detection of neomycin in milk. Food Agric Immunol 22:259–269. https://doi.org/10.1080/09540105.2011.569882
Yu X, Li MZ, Zhao MZ, Lau SHS, Tan HR, The WJ, Yang HS, Zheng C, Zhang YQ (2019) Quantification of aflatoxin B1 in vegetable oils using low temperature clean-up followed by immuno-magnetic solid phase extraction. Food Chem 275:390–396. https://doi.org/10.1016/j.foodchem.2018.09.132
Yuan XC, Yuan YX, Gao X, Xiong ZL, Zhao LS (2020) Magnetic dummy-template molecularly imprinted polymers based on multi-walled carbon nanotubes for simultaneous selective extraction and analysis of phenoxy carboxylic acid herbicides in cereals. Food Chem 333:127540. https://doi.org/10.1016/j.foodchem.2020.127540
Yue FL, Li FL, Kong QQ, Guo YM, Sun X (2021) Recent advances in aptamer-based sensors for aminoglycoside antibiotics detection and their applications. Sci Total Environ 762:143129. https://doi.org/10.1016/j.scitotenv.2020.143129
Zhang Z, Cao XL, Zhang ZP, Yin JG, Wang DN, Xu YN, Zheng W, Li XY, Zhang QS, Liu LW (2020) Synthesis of dummy-template molecularly imprinted polymer adsorbents for solid phase extraction of aminoglycosides antibiotics from environmental water samples. Talanta 208:120–385. https://doi.org/10.1016/j.talanta.2019.120385
Zhao QY, Zhao HT, Yang X, Zhang H, Dong AJ, Wang J, Li B (2018) Selective recognition and fast enrichment of anthocyanins by dummy molecularly imprinted magnetic nanoparticles. J Chromatogr A 1572:9–19. https://doi.org/10.1016/j.chroma.2018.08.029
Acknowledgements
This work was supported by the Natural Science Foundation of Shandong Province (ZR2019BC091; ZR2018MB027) and Graduate Innovation Foundation of Yantai University, GIFYTU.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Ethics Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed Consent
Informed consent is not applicable in this study.
Conflict of Interest
Xiaolin Cao declares that he has no conflict of interest. Zheng Zhang declares that he has no conflict of interest. Guangyang Liu declares that he has no conflict of interest. Ziping Zhang declares that he has no conflict of interest. Jungang Yin declares that he has no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 284 kb)
Rights and permissions
About this article
Cite this article
Cao, X., Zhang, Z., Liu, G. et al. Preparation of Magnetic Dummy Template Molecularly Imprinted Polymers for the Determination of Aminoglycosides Antibiotics in Milk. Food Anal. Methods 14, 2111–2120 (2021). https://doi.org/10.1007/s12161-021-02042-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12161-021-02042-z