Abstract
Disclosing the composition of meat products in detail is essential to consumers’ rights. In this study, we developed a rapid method based on loop-mediated isothermal amplification to visibly identify pork DNA in meat products. Four pig specific primers were designed according to the mitochondrial DN1 gene sequence. The analytical sensitivity of the LAMP assay for pork DNA detection is 0.5 pg in our experiment, and the results were not affected by processing temperature. We used the LAMP assay and the industry standard for Chinese entry-exit inspections and quarantines to analyze commercial halal products, and the results were comparable. In conclusion, the LAMP assay has excellent sensitivity and specificity and is a convenient method for pork DNA detection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Meat and meat products play important roles as sources of protein in our daily life (Font and Guerrero 2014). Many commercial food products such as minced meats, meatballs, sausage, and similar items are convenient components of the human diet. Disclosing the constituents of meat products to consumers is necessary. Although there are many countries, such as Malaysia, Indonesia, and Canada, which have clear and definite regulations stating that halal food composition must be labeled in detail, food adulteration is still an international problem. In Turkish, 42 processed meat products were surveyed, such as meatball, salami, sausage, and so on. And found that one sausage sample was labeled as containing 5 % beef, but beef DNA cannot be detected, and a meatball sample labeled as 100 % beef was found to contain chicken (Ulca et al. 2013). In South Africa, 139 processed meat products were detected. The results revealed that 95 of 139 (68 %) samples contained species which were not declared on the product labeling, with the incidence being highest in sausages, burger patties, and deli meats (Cawthorn et al. 2013). In Iran, 224 meat products were collected from different companies and food markets, 6 of 68 fermented sausages (8.82 %), 4 of 48 frankfurters (8.33 %), 4 of 55 hamburgers (7.27 %), 2 of 33 hams (6.6 %), and 1 of 20 cold cutmeat (5 %) were found to contain Haram (unlawful or prohibited) meat (Doosti et al. 2014). Therefore, it is necessary to develop a reliable, sensitive, and rapid method to identify undeclared components in raw, adulterated, and processed meat products.
Several analytical methodologies have been developed, including Fourier transform infrared spectroscopy (Rohman et al. 2011), an electronic nose (Man 2005), gas chromatography/mass spectrometry (Nurjuliana et al. 2011), liquid chromatography with electrochemical detection (Chou et al. 2007), an enzyme-linked immunosorbent assay (Asensio et al. 2008), nanoparticle sensors coupled with optical or fluorescence spectroscopy (Ali et al. 2011a), the polymerase chain reaction (PCR) (Aida et al. 2005), species-specific PCR (Karabasanavar et al. 2014), PCR-restriction fragment length polymorphism (PCR-RFLP) (Ali et al. 2011b), SYBR green real-time PCR (Farrokhi and Jafari Joozani 2011), and TaqMan probe real-time PCR (Ali et al. 2012). Some of these analytical approaches have been used in various countries and organizations as detection method for commercial meat products. However, the methods currently used rely on laboratory and operating of special equipment.
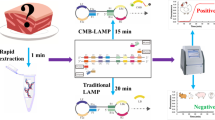
Loop-mediated isothermal amplification (LAMP) is a new nucleic acid amplification method. This method relies on four specific primers, inner primers (FIP and BIP) and outer primers (F3 and B3), which recognize six distinct regions of the target DNA and have high sensitivity and specificity (Notomi et al. 2000). In addition, the LAMP method has the significant advantages of rapidity and ease of operating. Upon incubation in a simple incubator, DNA is amplified under isothermal conditions (60–65 °C) in approximately 60 min by the Bst DNA polymerase. Foremost, during the amplification of the DNA, a large number of pyrophosphate ions is produced, and the mixture thus exhibits visible color in the presence of calcein. Therefore, due to the color-based character of the LAMP assay, this method can be utilized in on-the-spot detection (Tomita et al. 2008). Recently, LAMP detection methods have been widely used for transgenic plants (Chen et al. 2012; Zhang et al. 2013), microorganisms (Denschlag et al. 2012; Niessen et al. 2013; Wang et al. 2014), and viruses (Almasi et al. 2013; Luo et al. 2013). However, there is only one report of the application to meat product identification (Abdulmawjood et al. 2014).
In this study, we designed specific primers and developed a LAMP assay method for the detection of pork DNA in various halal samples.
Materials and Methods
Collection and Preparation of Meat Samples
The fresh meat samples used in this research include pig (Sus scrofa), cattle (Bos taurus), buffalo (Bubalus bubalis), sheep (Ovis aries), goat (Capra hircus), fox (Vulpes vulpes), rabbit (Oryctolagus cuniculus), rat (Rattus norvegicus), dog (Canis familiaris), chicken (Gallus gallus), duck (Anas platyrhynchos), and fish (Cyprinus carpio). The pig, cattle, buffalo, sheep, goat, chicken, duck, and fish meat samples were purchased at Wal-Mart (Changchun, China). The rabbit, rat, and dog meat were provided by the College of Animal Science, Jilin University (Changchun, China). In addition, the fox meat sample was kindly supplied by Dr. Yan Souqing (Jilin University). Other samples were purchased in triplicate from various supermarkets and bazaars located in Changchun, China.
To investigate the efficacy of the method for heat-processed meats, eight pork samples (50–100 g) were either fried with corn oil for 20 min, heated in water at 80 and 100 °C for 30 and 60 min, autoclaved at 121 °C for 30 min, or microwaved for 20 min.
The meat mixtures for mimicking adulteration were prepared by mixing pork into beef or mutton meat at ratios of 50:50, 10:90, 1:99, 0.1:99.9, and 0.01:99.99 to obtain 50, 10, 1, 0.1, and 0.01 % pork meat mixtures, respectively. All the samples were stored at −20 °C prior to DNA extraction.
DNA Isolation
DNA from all the samples was extracted with the TIANamp Genomic DNA kit (Tiangen, China). The isolated DNA was checked for quality, purity, and concentration using the Nanodrop 2000 (Thermo, China), and the DNA samples with OD 260/OD 280 ratios between 1.7 and 1.9 were used for the LAMP amplification.
LAMP Primer Design
The LAMP primer sets were designed using Primer explorer V4 software (http://primerexplorer.jp/), based on the published Sus scrofa mitochondrial ND1 gene (accession number KF888634.1) in the National Center for Biotechnology (NCBI) database. The LAMP primers sequences are shown in Table 1. The oligonucleotide primers were synthesized by GenScript (Nanjing, China).
LAMP Reaction
We optimized the reaction conditions based on Tomita’s report (Tomita et al. 2008). The reaction volume was 25 μL, which included 1.6 μM each of FIP and BIP, 0.2 μM each of F3 and B3, 1.8 mM dNTPs (Tiangen, China), ×1 ThermoPol reaction buffer (20 mM Tris-HCl, 10 mM KCl, 2 mM MgSO4, 10 mM (NH4)2SO4, 0.1 % Triton X-100) (New England Biolabs, USA), 6 mM MgSO4, 0.6 M betaine (Sigma-Aldrich, Canada), 0.5 mM MnCl2, 25 μM calcein (Dojindo, Japan), 8 U of Bst DNA polymerase (New England Biolabs, USA), and 1 μL of template DNA. The reaction mixture was incubated at 64 °C for 1 h and then heated to 80 °C for 5 min to terminate the reaction. The color of the reaction mixture changed from orange to green if the targeted DNA was amplified. Otherwise, the reaction mixture remained orange. Each LAMP assay was repeated three times.
Real-Time PCR
The total DNA of all the commercial samples was used to detect pork DNA using a Real-time PCR Porcine DNA Detection Kit (Takara, China), which is used as the industry standard in China for entry-exit inspections and quarantines (SN/T 2051-2008). The real-time PCR for all the commercial samples was performed according to the manufacturer’s instructions.
Results and Discussion
Designing Specific Primers for Pork DNA
A specific nucleotide sequence screen was conducted via BLASTEN in the NCBI database, and the mitochondrial DN1 gene was selected for the primer design. The primer set consisted of two outer primers (F3 and B3) and two inner primers (FIP and BIP). The inner primers cover two distinct sequences of the target gene (F1c/B1c and F2c/B2c) and are connected with a link of -TTTT-. The sequences of the LAMP primers are given in Table 1, and the detailed locations of the LAMP primers in the target DNA sequences are shown in Fig. 1.
Determining the Specificity of the Primers
The specificity of the LAMP assay primers was determined using 12 meat DNA samples. Each reaction contained 50 ng of genomic DNA as a template. Only the tube containing pig genomic DNA changed color from orange to green and was identified as positive. The other templates (cattle, buffalo, sheep, goat, fox, dog, rabbit, rat, chicken, duck, and fish) did not exhibit nonspecific amplification (Fig. 2). These results indicate that the LAMP assay primers can be used for pork DNA detection.
Most of species identification in meat products is based on PCR-RFLP analysis (Aida et al. 2007; Murugaiah et al. 2009) or real-time PCR (Kesmen et al. 2009; Martin et al. 2009). These methods utilize two primers that bind to the targeted genes and require rigorous species-specific nucleotide sequence in the primer designs. The LAMP assay method employs four primers that bind to specific nucleotide sequences, which ensures specificity and decreases false positives of the detection results.
Determining the Sensitivity of the LAMP Assay
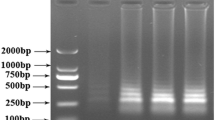
The sensitivity of the LAMP assay was evaluated on the basis of pork genomic DNA that was serially diluted to final concentrations ranging from 50 to 5 × 10−7 ng/μL. Each reaction contained 1 μL of diluted DNA sample as a template. The detection limit of the LAMP assay for pork DNA was found to be 0.5 pg/reaction (Fig. 3).
We then mixed pork DNA into beef or mutton DNA to produce nominal concentrations of 50, 10, 1, 0.1, 0.01, and 0 % to mimic meat adulteration. A total of 50 ng of genomic DNA was used in each reaction, and a mixture of 0.01 % pork in beef could be detected (Fig. 4). The same result was found for pork adulteration in mutton (data not shown).
Compared with other previous studies, this limit of detection represents greater sensitivity than Karabasanavar’s 10 pg (Karabasanavar et al. 2014) detected via species-specific PCR and is similar to Martin’s (Martin et al. 2009) real-time PCR method, which had a detection limit of 0.1 pg. Real-time PCR is now recognized as the most sensitive method to detect nucleic acid molecules, and our LAMP assay has a similar sensitivity. Furthermore, we can detect 0.01 % pork in adulterated sample mixtures, equivalent to 0.1 g of pork adulteration in 1 kg of meat. The sensitivity was thereby satisfactory in detecting adulterated meat.
Effect of Meat Processing on the LAMP Assay
To determine if temperature affects the results of the LAMP assay for pork, we processed pork samples by frying for 20 min, heated in water at 80 and 100 °C for 30 and 60 min, autoclaving at 121 °C for 30 min, and microwaving for 20 min. Although Arslan (Arslan et al. 2006) reported that heat treatment can cause genomic DNA to disintegrate into small fragments, sufficient DNA was obtained for the LAMP assay detection (Fig. 5).
We used frying, heat treatment, and microwave ovens to simulate processing procedures, for comparison with the corresponding fresh specimens. The processed sample volume was smaller but was still adequate for DNA extraction and LAMP assay detection. These results are consistent with previous reports (Sakalar et al. 2012; Karabasanavar et al. 2014). However, Buntjer and Brodmann failed to detect target DNA with a PCR assay due to DNA degradation at high temperatures (Buntjer et al. 1999, Brodmann and Moor 2003).
Commercial Sample Detection Using the LAMP Assay
Commercial meat products are the focus of adulteration reports, especially minced meat (Arslan et al. 2006). Therefore, we selected 42 commercial samples on the market for LAMP assay detection and analysis. Those samples included meatballs, minced meat, sausage, cut meat, and raw braciola, all of which were declared to not contain any pork ingredient.
The results revealed that 5, 5, 2, 2, and 1 samples changed color among the meatballs, minced meat, sausage, braciola, and cut meat samples, respectively. These results are listed in Table 2. In our LAMP test, pork DNA was observed in 15 samples, and 36 % of the selected samples did not conform to the labeling.
To compare the LAMP assay with the current detection method, we employed the Real-time PCR Porcine DNA Detection Kit (Takara, Dalian, China), which is widely used in China as the industry standard for entry-exit inspections and quarantines (SN/T 2051-2008), to analyze the 42 commercial samples. The results are listed in Table 2. Compared with the LAMP test results, there were two discrepant results. In one sample of sausage, our LAMP assay indicated a positive result while the real-time PCR test result was negative. Additionally, a sample of braciola was positive according to the real-time PCR assay but negative according to the LAMP assay. Except for these two results, all of the remaining 40 LAMP assay results corresponded with the official testing standard.
Conclusion
We successfully designed specific pig primers and developed a rapid LAMP assay for the visible detection of pork DNA. Compared with traditional PCR, the LAMP assay has greater analytical specificity. The LAMP assay can be applied directly to processed and commercial samples and is suitable for on-the-spot detection.
References
Abdulmawjood A et al (2014) Development of loop-mediated isothermal amplification (LAMP) assay for rapid and sensitive identification of ostrich meat. PLoS One 9(6), e100717
Aida AA et al (2005) Analysis of raw meats and fats of pigs using polymerase chain reaction for Halal authentication. Meat Sci 69(1):47–52
Aida AA et al (2007) Detection of pig derivatives in food products for halal authentication by polymerase chain reaction–restriction fragment length polymorphism. J Sci Food Agric 87(4):569–572
Ali ME et al (2011a) Swine-specific PCR-RFLP assay targeting mitochondrial cytochrome B gene for semiquantitative detection of pork in commercial meat products. Food Anal Methods 5(3):613–623
Ali ME et al (2011b) Nanobiosensor for detection and quantification of DNA sequences in degraded mixed meats. J Nanomater 2011:32
Ali ME et al (2012) Analysis of pork adulteration in commercial meatballs targeting porcine-specific mitochondrial cytochrome b gene by TaqMan probe real-time polymerase chain reaction. Meat Sci 91(4):454–459
Almasi MA et al (2013) Visual detection of Potato Leafroll virus by loop-mediated isothermal amplification of DNA with the GeneFinder dye. J Virol Methods 192(1-2):51–54
Arslan A et al (2006) Effect of method of cooking on identification of heat processed beef using polymerase chain reaction (PCR) technique. Meat Sci 72(2):326–330
Asensio L et al (2008) Determination of food authenticity by enzyme-linked immunosorbent assay (ELISA). Food Control 19(1):1–8
Brodmann PD, Moor D (2003) Sensitive and semi-quantitative TaqMan™ real-time polymerase chain reaction systems for the detection of beef (Bos taurus) and the detection of the family Mammalia in food and feed. Meat Sci 65(1):599–607
Buntjer J, Lamine A, Haagsma N, Lenstra JA (1999) Species identification by oligonucleotide hybridisation:the influence of processing of meat products.pdf. Sci Food Agric 79(1):53–57
Cawthorn D-M et al (2013) A high incidence of species substitution and mislabelling detected in meat products sold in South Africa. Food Control 32(2):440–449
Chen X et al (2012) Endpoint visual detection of three genetically modified rice events by loop-mediated isothermal amplification. Int J Mol Sci 13(11):14421–14433
Chou CC et al (2007) Fast differentiation of meats from fifteen animal species by liquid chromatography with electrochemical detection using copper nanoparticle plated electrodes. J Chromatogr B Anal Technol Biomed Life Sci 846(1-2):230–239
Denschlag C et al (2012) Hyd5 gene-based detection of the major gushing-inducing Fusarium spp. in a loop-mediated isothermal amplification (LAMP) assay. Int J Food Microbiol 156(3):189–196
Doosti A et al (2014) Molecular assay to fraud identification of meat products. J Food Sci Technol 51(1):148–152
Farrokhi R, Jafari Joozani R (2011) Identification of pork genome in commercial meat extracts for Halal authentication by SYBR green I real-time PCR. Int J Food Sci Technol 46(5):951–955
Font IFM, Guerrero L (2014) Consumer preference, behavior and perception about meat and meat products: an overview. Meat Sci 98(3):361–371
Karabasanavar NS et al (2014) Detection of pork adulteration by highly-specific PCR assay of mitochondrial D-loop. Food Chem 145:530–534
Kesmen Z et al (2009) Identification of meat species by TaqMan-based real-time PCR assay. Meat Sci 82(4):444–449
Luo JG et al (2013) Development of a loop-mediated isothermal amplification assay for rapid detection of bovine parvovirus. J Virol Methods 191(2):155–161
Man Y (2005) Detection of lard adulteration in RBD palm olein using an electronic nose. Food Chem 90(4):829–835
Martin I et al (2009) SYBR-Green real-time PCR approach for the detection and quantification of pig DNA in feedstuffs. Meat Sci 82(2):252–259
Murugaiah C et al (2009) Meat species identification and Halal authentication analysis using mitochondrial DNA. Meat Sci 83(1):57–61
Niessen L et al (2013) The application of loop-mediated isothermal amplification (LAMP) in food testing for bacterial pathogens and fungal contaminants. Food Microbiol 36(2):191–206
Notomi T, Okayama H, Masubuchi H, Yonekawa T, Watanabe K, Amino N, Hase T (2000) Loop-mediated isothermal amplification of DNA. Nucleic Acids Res 28(12), e63
Nurjuliana M et al (2011) Rapid identification of pork for halal authentication using the electronic nose and gas chromatography mass spectrometer with headspace analyzer. Meat Sci 88(4):638–644
Rohman A et al (2011) Analysis of pork adulteration in beef meatball using Fourier transform infrared (FTIR) spectroscopy. Meat Sci 88(1):91–95
Sakalar E et al (2012) Effect of heat processing on DNA quantification of meat species. J Food Sci 77(9):N40–44
Tomita N et al (2008) Loop-mediated isothermal amplification (LAMP) of gene sequences and simple visual detection of products. Nat Protoc 3(5):877–882
Ulca P et al (2013) Meat species identification and Halal authentication using PCR analysis of raw and cooked traditional Turkish foods. Meat Sci 94(3):280–284
Wang X et al (2014) Comparison of conventional PCR, multiplex PCR, and loop-mediated isothermal amplification assays for rapid detection of Arcobacter species. J Clin Microbiol 52(2):557–563
Zhang M et al (2013) One simple DNA extraction device and its combination with modified visual loop-mediated isothermal amplification for rapid on-field detection of genetically modified organisms. Anal Chem 85(1):75–82
Acknowledgments
This work was supported by the Program for Changjiang Scholars and Innovative Research Team in University (No. IRT1248).
Conflict of Interest
The authors declare that they have no competing interests.
Compliance with Ethics Requirements
We must include the following sentence to make sure that readers are aware that there are no ethical issues with human or animal subjects:
This article does not contain any studies with human or animal subjects.
Author information
Authors and Affiliations
Corresponding author
Additional information
Guangyao Ran and Linzhu Ren contributed equally to this work.
Rights and permissions
About this article
Cite this article
Ran, G., Ren, L., Han, X. et al. Development of a Rapid Method for the Visible Detection of Pork DNA in Halal Products by Loop-Mediated Isothermal Amplification. Food Anal. Methods 9, 565–570 (2016). https://doi.org/10.1007/s12161-015-0246-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12161-015-0246-z