Abstract
Lamins are the main components of the nucleoskeleton. They form a protein meshwork that underlies the inner nuclear membrane. Mutations in the LMNA gene coding for A-type lamins (lamins A and C) cause a large panel of human diseases, referred to as laminopathies. These diseases include muscular dystrophies, lipodystrophies and premature aging diseases. Lamin A exhibits a C-terminal region that is different from lamin C and is post-translationally modified. It is produced as prelamin A and it is then farnesylated, cleaved, carboxymethylated and cleaved again in order to become mature lamin A. In patients with the severe Hutchinson–Gilford progeria syndrome, a specific single point mutation in LMNA leads to an aberrant splicing of the LMNA gene preventing the post-translational processing of prelamin A. This leads to the accumulation of a permanently farnesylated lamin A mutant lacking 50 amino acids named progerin. We here report the NMR 1H, 15N, 13CO, 13Cα and 13Cβ chemical shift assignment of the C-terminal region that is specific to prelamin A, from amino acid 567 to amino acid 664. We also report the NMR 1H, 15N, 13CO, 13Cα and 13Cβ chemical shift assignment of the C-terminal region of the progerin variant, from amino acid 567 to amino acid 614. Analysis of these chemical shift data confirms that both prelamin A and progerin C-terminal domains are largely disordered and identifies a common partially populated α-helix from amino acid 576 to amino acid 585. This helix is well conserved from fishes to mammals.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Biological context
The nuclear lamina is a universal feature of metazoan nuclear envelopes (Gerace and Huber 2012; Burke and Stewart 2013). It forms a filamentous layer at the nuclear face of the inner nuclear membrane (Mahamid et al. 2016; Turgay et al. 2017) and is composed primarily of A- and B-type lamins (Xie et al. 2016). Lamins provide structural integrity to the nuclear envelope and contribute to genome organization and function (Gruenbaum and Foisner 2015). Lamin levels correlate with tissue rigidity and with mechanical stability of the nucleus (Swift et al. 2013). The two A-type lamin isoforms lamins A and C encoded by the gene LMNA are mainly expressed in fully differentiated cells. They are largely identical, diverging only in their C-terminal regions. The lamin A C-terminal region undergoes extensive processing. It is produced as prelamin A and is then farnesylated, cleaved, carboxymethylated and cleaved again to lead to mature lamin A. Yet a specific single point mutation in LMNA leads to the internal deletion of 50 amino acids preventing the post-translational processing of prelamin A (Fig. 1). The deletion mutant, called progerin, remains farnesylated and accumulates at the nuclear periphery, causing nuclear architectural defects such as blebs and herniations of the nuclear envelope and thickening of the nuclear lamina (Maraldi and Lattanzi 2007). This mutant causes an extremely rare genetic disorder characterized by premature, rapid aging shortly after birth called Hutchinson–Gilford progeria syndrome (HGPS (De Sandre-Giovannoli et al. 2003; Eriksson et al. 2003)). Progerin is also present to a lower extend in non-pathological cells and is a biomarker of cellular aging (McClintock et al. 2007). The prelamin A-specific C-terminal region is highly conserved from fishes to mammals (Fig. 1). It is essential for binding to other nuclear envelope proteins such as SUN1 (Haque et al. 2010), to histone modifying enzymes such as SIRT6, which is known to deacetylate histone H3 at lysine 9 and 56 (Ghosh et al. 2015), and to the shelterin complex at telomeres (Wood et al. 2014). To further investigate the binding properties of prelamin A and progerin C-terminal regions, before and after post-translational modifications, we have produced and purified these regions and we have described their conformations based on the analysis of their NMR 1H, 15N, 13CO, 13Cα and 13Cβ chemical shifts.
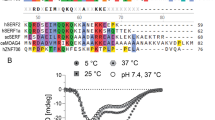
Sequence alignment of human C-terminal region specific to prelamin A, including a representative set of 14 homologous lamins from mammals, frogs to fishes. The upper sequence is that of human prelamin A. The other sequences are marked using their GenBank Identifier number: 109256198, Panthera pardus; 16905, Mus musculus; 101704567, Heterocephalus glaber; 100923083, Sarcophilus harrisii; 104468957, Pterocles guttularis; 102948145, Chelonia mydas; 373673, Xenopus laevis; 733973, Xenopus tropicalis; 102359604, Latimeria chalumnae; 109072855, Cyprinus carpio; 109902034, Oncorhynchus kisutch; 101072260, Takifugu rubripes; 104964806, Notothenia coriiceps; 106528028, Austrofundulus limnaeus. The conserved cysteines are indicated by red arrows. The fragment deleted in progerin is highlighted by a red box. Under the sequence alignment, a graph highlighting the amino acid conservation as calculated by Jalview is displayed (Waterhouse et al. 2009)
Methods and experiments
Protein expression and purification
The human prelamin A peptide PreLamC, from amino acid 567 to amino acid 664, and the human progerin peptide ProgC, from amino acid 567 to amino acid 614, were expressed in Escherichia coli BL21 DE3 Star(Novagen), using a pETM13 vector coding for a GST-tagged protein with a TEV protease site and an additional tryptophan between the cleavage site and the protein. Their cDNAs were optimized for expression in Escherichia coli (Genscript). Moreover, mutated versions of both peptides PreLamC and ProgC in which cysteines are replaced by alanines were also expressed as GST fusion proteins, with a TEV site followed by an additional methionine and an alanine instead of Gly567. These will be further called PreLamCCtoA and ProgCCtoA, respectively. All these peptides were produced using the following protocol. Bacteria were grown in 15N and 13C labelled M9 minimum medium at 37 °C and induced at an optical density of 0.8 with 1 mM Isopropyl β-d-1-thiogalactopyranoside at 37 °C during 3 h. Cells were lysed in 50 mM Tris–HCl, pH 8.0, 300 mM NaCl, 40 mM imidazole, 5% glycerol, 1% Triton X-100 and 1 mM phenylmethanesulfonyl fluoride. Then bacteria were centrifuged at 20,000×g for 20 min. Proteins were purified using Glutathione-Sepharose 4B beads (GE Healthcare) equilibrated with a buffer containing 50 mM Tris–HCl pH 8.0, 150 mM NaCl (and 10 mM dithiothreitol in the case of cysteine-containing peptides). The tag cleavage was performed by adding TEV protease on the beads before washing with the buffer. Peptides collected during washing were concentrated and aliquoted in 1 ml tubes to be heated 10 min at 95 °C. After centrifugation at 14,000×g for 15 min, the supernatants were loaded on a gel filtration column Superdex 75 10/300GL (GE Healthcare) equilibrated in 20 mM Phosphate pH 6.5, 150 mM NaCl, 2 mM dithiothreitol. Finally, all the samples were concentrated up to 600 µM for NMR experiments.
NMR spectroscopy
Most NMR experiments were performed on uniformly 15N and 13C labelled ProgCCtoA, PreLamCCtoA and ProgC peptides in 20 mM phosphate buffer pH 6.5, 150 mM NaCl, 2 mM dithiothreitol, 90%: 10% H20:D20. Only the 1H → 15N nOe experiments were recorded on a 15N labelled sample of ProgC. In the case of PreLamC, a 15N labelled sample was produced and an 1H → 15N HSQC spectrum was recorded. All the NMR experiments were recorded at 283 K on a 600 MHz Bruker Advance II spectrometer equipped with a triple resonance cryogenic probe. 1H, 13C and 15N resonance frequencies were assigned using 3D HNCACB, CBCA(CO)NH, HNCO, HN(CA)CO and HN(CO)(CA)NH experiments. The data were processed using Topspin3.1 (Bruker) and analyzed with CCPNMR (Vranken et al. 2005).
Extent of assignments and data deposition
First, NMR assignment was carried out by analyzing spectra recorded on ProgCCtoA and PreLamCCtoA, which did not oxidize with time. Superimposition of the 1H–15N HSQC NMR spectra of ProgCCtoA (red) and PreLamCCtoA (black) is displayed in Fig. 2a. It revealed that the peaks corresponding to the common lamin fragment, i.e. aa 568 to aa 606 and aa 657 to aa 664, nicely overlap. Only peaks corresponding to residues close to the deletion (Ala605, Gln606 and Ser 657) have different positions in the two spectra. In the case of peptides ProgCCtoA and PreLamCtoA, 100 and 91% (84 out of 92) of 1H–15N pairs, 100 and 93% (90 out of 97) of 13Cα, 100 and 93% (76 out of 82) of 13Cβ and 100 and 91% (88 of out 97) of 13CO resonances were assigned, respectively. The only unassigned PreLamCtoA fragment of more than 3 residues corresponds to Ser645-Tyr646-Leu647-Leu648, i.e. the cleavage site of the last maturation step. Further superimposition of the 1H–15N HSQC NMR spectra of ProgCCtoA (red) and ProgC (blue) is displayed in Fig. 2b. Assignment of most ProgCCtoA NMR signals could be easily transferred to ProgC signals. Analysis of the 1H, 13C and 15N NMR 3D experiments recorded on ProgC was performed in order to confirm the assignment. Thus, the chemical shifts of all 1H–15N pairs, 13Cα, 3Cβ and 13CO resonances of ProgC could be unambiguously assigned. Finally, superimposition of the 1H–15N HSQC NMR spectra of PreLamCCtoA (black) and PreLamC (green) is displayed in Fig. 2c. Here again most 1H–15N HSQC peaks (82%, 75 out of 92) of PreLamC could be assigned from the comparison with PreLamCCtoA. Judging by the narrow range of backbone amide 1H chemical shifts (between 7.8 and 8.6 ppm) and the distribution of the ΔδCα–ΔδCβ values in ProgCCtoA, ProgC and PreLamCCtoA (Fig. 3), all the peptides are intrinsically disordered. However, a partially populated α-helix is present going from aa 576 to aa 585 of ProgCCtoA, ProgC and PreLamCCtoA. Consistently, positive nOe values were measured at 600 MHz for aa 575 to aa 586 of ProgC (Fig. 3, boxed panel). Interestingly, this partially populated α-helix is located in the most conserved region of ProgC and PreLamC (Fig. 1). We propose that lamin A region from aa 576 to aa 585 is a conserved binding site for a yet unidentified partner. This region contains Arg582, a residue mutated into His in patients with a Dunnigan type familial partial lipodystrophy (Speckman et al. 2000). Mutation could impair binding to a lamin partner. More work is now needed in order to identify how the C-terminal region specific to lamin A recognizes its partners at the nuclear envelope. The chemical shift data have been deposited in the BioMagResBank (http://www.bmrb.wisc.edu/) under accession numbers 27375 for ProgCCtoA, 27374 for ProgC and 27376 for PreLamCCtoA.
Superimposition of the 1H–15N HSQC spectra of the different lamin fragments analyzed in this study. All these spectra were recorded at 283 K and 600 MHz. a The ProgCCtoA spectrum (in red) is displayed onto the PreLamCCtoA spectrum (in black). The peaks from ProgCCtoA that do not overlap with peaks from PreLamCCtoA are labeled in purple. b The ProgC spectrum (in blue) is displayed onto the ProgCCtoA spectrum (in red). b The PreLamC spectrum (in green) is displayed onto the PreLamCCtoA spectrum (in black). Peaks corresponding to PreLamC and PreLamCCtoA minor conformations due to proline isomerization are labeled in purple. Indeed, based on their Cβ chemical shifts, proline residues exhibit a trans conformation in the most populated PreLamCCtoA conformation. However, proline residues P576 and P594 clearly show a minor cis conformation that give rise to peaks corresponding to their neighboring residues
Analysis of the 13C chemical shifts of ProgCCtoA, ProgC and PreLamCCtoA. a Superimposition of the differences between the Cα and Cβ secondary chemical shifts of both ProgCCtoA (in red) and ProgC (in blue), and in the boxed panel 1H → 15N heteronuclear nOes recorded on a 15N labelled ProgC sample. b Differences between the Cα and Cβ secondary chemical shifts of PreLamCCtoA
References
Burke B, Stewart CL (2013) The nuclear lamins: flexibility in function. Nat Rev Mol Cell Biol 14:13–24. https://doi.org/10.1038/nrm3488
De Sandre-Giovannoli A, Bernard R, Cau P, Navarro C, Amiel J, Boccaccio I, Lyonnet S, Stewart CL, Munnich A, Le Merrer M, Lévy N, 2003. Lamin a truncation in Hutchinson-Gilford progeria. Science 300:2055. https://doi.org/10.1126/science.1084125
Eriksson M, Brown WT, Gordon LB, Glynn MW, Singer J, Scott L, Erdos MR, Robbins CM, Moses TY, Berglund P, Dutra A, Pak E, Durkin S, Csoka AB, Boehnke M, Glover TW, Collins FS (2003) Recurrent de novo point mutations in lamin A cause Hutchinson-Gilford progeria syndrome. Nature 423:293–298. https://doi.org/10.1038/nature01629
Gerace L, Huber MD (2012) Nuclear lamina at the crossroads of the cytoplasm and nucleus. J Struct Biol 177:24–31. https://doi.org/10.1016/j.jsb.2011.11.007
Ghosh S, Liu B, Wang Y, Hao Q, Zhou Z (2015) Lamin A is an endogenous SIRT6 activator and promotes SIRT6-mediated DNA repair. Cell Rep 13:1396–1406. https://doi.org/10.1016/j.celrep.2015.10.006
Gruenbaum Y, Foisner R (2015) Lamins: nuclear intermediate filament proteins with fundamental functions in nuclear mechanics and genome regulation. Annu Rev Biochem 84:131–164. https://doi.org/10.1146/annurev-biochem-060614-034115
Haque F, Mazzeo D, Patel JT, Smallwood DT, Ellis JA, Shanahan CM, Shackleton S (2010) Mammalian SUN protein interaction networks at the inner nuclear membrane and their role in laminopathy disease processes. J Biol Chem 285:3487–3498. https://doi.org/10.1074/jbc.M109.071910
Mahamid J, Pfeffer S, Schaffer M, Villa E, Danev R, Cuellar LK, Förster F, Hyman AA, Plitzko JM, Baumeister W (2016) Visualizing the molecular sociology at the HeLa cell nuclear periphery. Science 351:969–972. https://doi.org/10.1126/science.aad8857
Maraldi NM, Lattanzi G (2007) Involvement of prelamin A in laminopathies. Crit Rev Eukaryot Gene Expr 17:317–334
McClintock D, Ratner D, Lokuge M, Owens DM, Gordon LB, Collins FS, Djabali K (2007) The mutant form of lamin A that causes Hutchinson-Gilford progeria is a biomarker of cellular aging in human skin. PLoS ONE 2:e1269. https://doi.org/10.1371/journal.pone.0001269
Speckman RA, Garg A, Du F, Bennett L, Veile R, Arioglu E, Taylor SI, Lovett M, Bowcock AM (2000) Mutational and haplotype analyses of families with familial partial lipodystrophy (Dunnigan variety) reveal recurrent missense mutations in the globular C-terminal domain of lamin A/C. Am J Hum Genet 66:1192–1198. https://doi.org/10.1086/302836
Swift J, Ivanovska IL, Buxboim A, Harada T, Dingal PCDP., Pinter J, Pajerowski JD, Spinler KR, Shin J-W, Tewari M, Rehfeldt F, Speicher DW, Discher DE (2013) Nuclear lamin-A scales with tissue stiffness and enhances matrix-directed differentiation. Science 341:1240104. https://doi.org/10.1126/science.1240104
Turgay Y, Eibauer M, Goldman AE, Shimi T, Khayat M, Ben-Harush K, Dubrovsky-Gaupp A, Sapra KT, Goldman RD, Medalia O (2017) The molecular architecture of lamins in somatic cells. Nature 543:261–264. https://doi.org/10.1038/nature21382
Vranken WF, Boucher W, Stevens TJ, Fogh RH, Pajon A, Llinas M, Ulrich EL, Markley JL, Ionides J, Laue ED (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59:687–696. https://doi.org/10.1002/prot.20449
Waterhouse AM, Procter JB, Martin DMA, Clamp M, Barton GJ (2009) Jalview Version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics 25:1189–1191. https://doi.org/10.1093/bioinformatics/btp033
Wood AM, Rendtlew Danielsen JM, Lucas CA, Rice EL, Scalzo D, Shimi T, Goldman RD, Smith ED, Le Beau MM, Kosak ST (2014) TRF2 and lamin A/C interact to facilitate the functional organization of chromosome ends. Nat Commun 5:5467. https://doi.org/10.1038/ncomms6467
Xie W, Chojnowski A, Boudier T, Lim JSY, Ahmed S, Ser Z, Stewart C, Burke B (2016) A-type lamins form distinct filamentous networks with differential nuclear pore complex associations. Curr Biol 26:2651–2658. https://doi.org/10.1016/j.cub.2016.07.049
Acknowledgements
This work was supported by CEA, CNRS and University Paris South, by the French Infrastructure for Integrated Structural Biology (https://www.structuralbiology.eu/ networks/frisbi, Grant Number ANR-10-INSB-05-01, Acronym FRISBI) and by the French Association against Myopathies (AFM; research Grants Nos. 17243 and 20018 to S.Z.-J. and PhD fellowship No. 18159 to C.S.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Celli, F., Petitalot, A., Samson, C. et al. 1H, 13C and 15N backbone resonance assignment of the lamin C-terminal region specific to prelamin A. Biomol NMR Assign 12, 225–229 (2018). https://doi.org/10.1007/s12104-018-9813-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-018-9813-8