Abstract
Background
Aerobic glycolysis has a pivotal role in the carcinogenic process. The current understanding of the functional role and mechanism of UCHL3-related aerobic glycolysis in pancreatic cancer is far from comprehensive, therefore requires an in-depth analysis on this aspect.
Methods
In the present research, the expressions of ubiquitin carboxyl-terminal hydrolase L3 (UCHL3), lactate dehydrogenase A (LDHA) and Forkhead box protein M1 (FOXM1) were detected by qRT-PCR, Western blot and immunohistochemistry. The effects of UCHL3 knockdown or overexpression on pancreatic cancer cells were examined by determining cell viability and colony formation. Aerobic glycolysis was assessed according to glucose uptake, lactic acid production, and lactate dehydrogenase (LDH) activity. Dual-luciferase reporter assay was performed to detect LDHA promoter activity.
Results
The results showed that UCHL3 expression was significantly increased in the pancreatic cancer tissues and cells, and that knocking down UCHL3 noticeably inhibited cell viability and aerobic glycolysis. Further investigations revealed that LDHA expression was promoted by UCHL3 and could be reduced by shFOXM1, and that low-expressed LDHA partly reversed the inhibition of aerobic glycolysis induced by overexpressed UCHL3.
Conclusions
To conclude, this study demonstrates that UCHL3 plays a carcinogenic role by promoting aerobic glycolysis in pancreatic cancer, suggesting that UCHL3 may be a potential diagnostic and therapeutic target for the treatment of cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Pancreatic cancer is one of the most frequent malignant tumors to the digestive tract all over the world [1]. The latest statistics demonstrated that in developed countries, as compared with other malignancies, the incidence of pancreatic cancer is the ninth highest for women and the tenth highest for men, with its mortality ranking the fourth highest [2]. China is also experiencing a steady increase in the incidence and mortality of pancreatic cancer [3]. Majority of patients with pancreatic cancer are often diagnosed at a later stage and, therefore, miss the optimal opportunity of surgical resection. Moreover, the prognosis of pancreatic cancer is largely poor, even the combined therapy of radiotherapy and chemotherapy is implemented, thus, the median survival time for pancreatic cancer patients is mostly 6–9 months, with a 5-year survival rate lower than 5% [4, 5]. Currently, unclear early symptoms, early distant metastasis, late diagnosis and ineffective treatment are seen as main factors contributing to the low survival rate of pancreatic cancer patients [5, 6]. Therefore, accurate and sensitive markers of pancreatic cancer should be developed to improve the early diagnosis of cancer.
In 1924, the German biochemist Otto Warburg first proposed that in addition to peroxidation and phosphorylation, energy metabolism of tumor cells could be achieved through glycolysis even in the case with sufficient oxygen, and such a phenomenon is referred to as Warburg effect or aerobic glycolysis [7]. Tumor aerobic glycolysis participates in the regulation of tumor genesis and development, adapts to the rapid division of tumor cells by regulating the energy metabolism, biosynthesis and REDOX state of cells, thus promoting the proliferation of tumor cells and regulating tumor metastasis [8,9,10]. Therefore, for the development of new targets more effective strategies to the treatment pancreatic cancer, it is highly important to investigating the molecular mechanism of aerobic glycolysis in the regulation of metastasis of the cancer.
Lactate dehydrogenase A (LDHA), which is a key enzyme in the glycolytic pathway, converts pyruvate to lactic acid, and it shows a high expression in many tumors, such as in hepatocellular carcinoma [11], oral squamous cell carcinoma [12]. Moreover, LDHA plays an important role in promoting the proliferation, metastasis and invasion of malignant tumor cells [12, 13].
UCHL3 belongs to UCH family, and current studies demonstrated that UCHL3 can be specifically hydrolyzed to remove the ubiquitin small molecules attached to the carbon terminal of the protein substrate. Researchers indicated that the deubiquitination enzyme UCHL3 has a critical function in DNA repair, biological development and tumorigenesis. When DNA damage occurs, UCHL3 could specifically remove the ubiquitination formed by RAD51 after ATM flow acidification, thus stabilizing the BRCA2-RAD51 complex [14]. UCHL3 is mainly located in the acrosome and flagella of sperm cells, and its protein expression level is significantly lower in asthenospermia and oligoasthenospermia patients than that in the normal group [15, 16]. Knocking down UCHL3 in prostate cancer cell line RWPE1 cells induces the molecular alterations of EMT (reflected by down-regulated expressions of E-cadherin and β-catenin and up-regulated vimentin), thus significantly promotes cell migration and invasion [17]. Although the specific biological functions and regulatory mechanisms of UCHL3 remained to be fully determined, UCHL3 has been widely proven to play an important role in many biological functions. Currently, the role of the deubiquitination enzyme UCHL3 in glycolysis remained as a research gap to be bridged.
Our study aimed to reveal the expression profile of UCHL3 in pancreatic cancer tissues and to examine its biological role in the progression of cancer and the Warburg effect. The current results showed that UCHL3 bound to the LDHA promoter activated LDHA expression and ultimately promoted pancreatic cancer progression. Our findings suggest that UCHL3 could serve as a novel biomarker and a promising therapeutic target for the prognosis and treatment of pancreatic cancer.
Materials and methods
Sample collection
Pancreatic cancer tissue specimens and adjacent normal tissues were collected from 31 pancreatic cancer patients attended our hospital between June 2018 and October 2019, and all the tissues remained untreated by drugs. The participants have signed the informed consent and the research was approved by the Ethics Committee of the Seventh Affiliated Hospital of Sun Yat-sen University. The experiments were conducted in accordance with the Helsinki declaration.
Cell culture
Pancreatic cancer cell lines (BxPC-3, AsPC-1, Capan-2, and PANC-1) and normal pancreatic ductal epithelial cell line HPDE6-C7 were purchased from American Type Culture Collection (Manassas, VA, USA). The cell lines were grown in DMEM medium (Hyclone, Logan, UT, USA) supplemented with 10% FBS (Invitrogen, Grand Island, NY, USA), 1% 100 U/mL penicillin, and 100 μg/mL streptomycin (HyClone), and maintained in a humidified atmosphere with 5% CO2 at 37 °C. The cells in the logarithmic growth phase were used for further experiments.
Cell transfection
The BxPC-3 cells and PANC-1 cells at a cell density of 6 × 105/well were inoculated into 6-well plates and grew to 60% confluence. UCHL3 plasmid, mock plasmid or Forkhead box protein M1c (FOXM1c or FOXM1), NC or LDHA-shRNA, Scramble were transfected into BxPC-3 cells, and UCHL3#1/#2-shRNA, shRNA-NC or FOXM1-shRNA, shRNA-NC were transfected into PANC-1 cells using Lipofectamine2000 (Invitrogen; Thermo Fisher Scientific, Inc., CA, USA). After transfection for 48 h, the cells were harvested for the following experiments.
Cell viability
Cell viability was determined by performing Cell Counting Kit-8 (CCK-8; Wako, Osaka, Japan). Briefly, the cells (1.0 × 105/well) were seeded into 6-well plates for 48 h, then added with CCK-8 solution (10 μL) for a 2-h incubation at 37 °C. The absorbance was measured using a microplate reader at OD 450 nm (Bio-Rad Laboratories, Inc., Hercules, CA, USA).
qRT-PCR
TRIzol (Invitrogen, Carlsbad, CA, USA) was for the extraction of the total RNA from tissues and cells. Following the instructions of the manufacture, PrimeScript RT Reagent Kit (Invitrogen, Shanghai, China) was used to reverse-transcribe RNA into cDNA. RT-PCR was performed using Bio-Rad CFX96 and SYBR Green Premix Ex Taq II (Takara, Dalian, China). GAPDH served as an internal reference gene, and 2(−ΔΔCt) method was used for calculating the data. The primer sequences of the genes are shown in Table 1.
Western Blot
RIPA lysis buffer was used to extract total proteins, and protein concentration was detected by BCA method. Total proteins were separated by SDS-PAGE electrophoresis and then transferred to PVDF membranes. The membranes were blocked with 5% skimmed milk for 2 h and then incubated with the primary antibodies at 4 °C overnight. Horseradish peroxidase-labelled secondary antibody (ab205719, 1:5000, abcam) was added to the membranes for 1-h incubation at room temperature. Finally, the bands were developed with chemiluminescence using hypersensitive ECL (Guangzhou Xiangbo Biotechnology Co., Ltd.).
Co-immunoprecipitation (co-IP)
BxPC-3 cells were washed with pre-chilled phosphate-buffered saline (PBS) and lysed. Cell lysates were boiled for 15 min and diluted with NTEN lysis buffer containing protease inhibitors and deubiquitination inhibitors in a 1:10 ratio. Finally, immunoprecipitation was performed with antibodies and protein A/G beads for 4 h at 4 °C. The immunocomplexes were subjected to western blot assay after washing the beads.
Glucose consumption and lactate production
The glucose consumption and lactate production were, respectively, detected by the Glucose Uptake Assay Kit (Colorimetric, Abcam) and Lactate Assay Kit (Sigma-Aldrich), according to the instructions of manufacturers.
LDH activity assay
The BxPC-3 cells and PANC-1 cells were seeded into 96-well plates and cultured to a density of 80%. The lactate dehydrogenase (LDH) activity was determined using the LDH Assay Kit (C0017, Beyotime, China). A Bio-Rad microplate reader was used to measure the absorbance at the reference wavelengths of 490 nm and 600 nm.
Immunohistochemistry (IHC)
Formalin-fixed, paraffin-embedded pancreatic cancer tissues and paired normal tissue blocks were cut into Sects. (5 µm). After dewaxing the sections in xylene, and rehydrated by distilled water in gradient concentrations (100%, 95%, and 75%) of ethanol, the samples were immersed in 10 mmol/L citrate buffer (pH 6.0) and treated in a thermostatic water bath. Antigen repair was performed at 4 °C for 2 h. The sections were first treated with 0.3% hydrogen peroxide to block endogenous peroxidase activity and then processed by 0.3% hydrogen peroxide to block endogenous peroxidase activity. Subsequently, the sections were incubated with anti-UCHL3 antibody at 4 °C overnight and then with biotinylated goat anti-rabbit IgG (antGene, Wuhan, China). After washing, diaminobenzidine and sumac were added to the sections for detecting the immunoreactive proteins.
Statistical analysis
All the experimental data are presented as the mean ± SD, and p < 0.05 was considered as statistically significant. Statistical significance was determined by Student’s t test or analysis of variance (ANOVA) among groups, followed by Dunnett-t test. All the statistical analyses were performed using GraphPad Prism 7.
Results
The expression of UCHL3 was upregulated in clinical pancreatic cancer specimens and cell lines
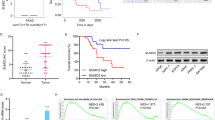
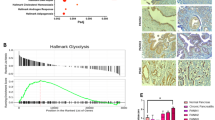
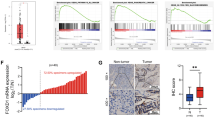
To examine the function of UCHL3 in pancreatic cancer, the expression of UCHL3 in pancreatic cancer tissues and their adjacent normal tissues were detected by qRT-PCR, immunochemistry and Western blot. The data showed that the mRNA expression of UCHL3 was significantly increased in pancreatic cancer tissues in comparison with that in the adjacent normal tissues (Fig. 1a). In immunochemistry, UCHL3 showed an obviously higher expression (Fig. 1b). Consistently, the protein expression of UCHL3 in the four human pancreatic cancer cell lines (BxPC-3, AsPC-1, Capan-2 and PANC-1) were significantly upregulated compared with that in normal epithelial cell line HPDE6-C7 (Fig. 1c, d). For further experiment, BxPC-3 cells with the highest UCHL3 expression and PANC-1 cells with the lowest UCHL3 expression were used for overexpressing and knocking down UCHL3, respectively.
Knocking down UCHL3 inhibited aerobic glycolysis of pancreatic cancer cells
The transfection efficiency of UCHL3 was investigated by conducting a Western blot. The transfection of a plasmid expressing precursor UCHL3 into BxPC-3 cells successfully upregulated miR-936 expression in the cells. UCHL3 was knocked down by transfecting shUCHL3#1 and shUCHL3#2 into the PANC-1 cells, and the inhibition rate of shUCHL3#1 reached 40%. Thus, shUCHL3#1 was selected for further experiments (Fig. 2a). MTT assay results showed that overexpression UCHL3 increased the proliferation of BxPC-3 cells (Fig. 2b). Next, we investigated the effect of UCHL3 overexpression on aerobic glycolysis and observed that glucose uptake, lactic acid production and LDH activity were significantly increased by overexpressed UCHL3 in the BxPC-3 cells (Fig. 2c). These phenomena were also verified by knocking down UCHL3 in the PANC-1 cells (Fig. 2d) because the data showed that knockdown of UCHL3 suppressed the proliferation of PANC-1 cells and significantly inhibited glucose uptake, lactic acid production and LDH activity (Fig. 2e).
Knocking down UCHL3 inhibited aerobic glycolysis of pancreatic cancer cells. a Transfection efficiency of UCHL3 was determined by Western blot and RT-qPCR. b Overexpressed UCHL3 promoted BxPC-3 cell viability. c Overexpressed UCHL3 promoted glucose uptake, lactate production and LDH activity of BxPC-3 cells. e Knocking down UCHL3 inhibited PANC cell viability. f Overexpressed UCHL3 promoted glucose uptake, lactate production and LDH Activity of PANC cells
LDHA expression was promoted by UCHL3
LDHA expression is correlated with aerobic glycolysis in cancer, including in oral squamous cell carcinoma, pancreatic cancer. We also found that lactic acid production and LDH activity of pancreatic cancer cells were affected by LDHA, thus, the relationship between UCHL3 and LDHA were explored. The data revealed that the protein expression of LDHA was upregulated by overexpressed UCHL3 in BxPC-3 cells, and downregulated by UCHL3 knockdown in PANC-1 cells (Fig. 3a–c). qRT-PCR assay on BxPC-3 cells and PANC-1 cells also demonstrated similar results (Fig. 3d, e). To further confirm the regulatory relationship between UCHL3 and LDHA, dual-luciferase assay was performed for detecting LDHA promoter activity, and the result showed that relative luciferase activity of LDHA was positively correlated with UCHL3 expression (Fig. 3f, g).
A positive correlation between UCHL3 and LADH. a, b, c LADH expression was promoted by UCHL3 in BxPC-3 cells and inhibited by shUCHL3 in PANC cells at the protein level. d, e LADH expression was promoted by UCHL3 in BxPC-3 cells and inhibited by shUCHL3 in PANC cells at mRNA level. f, g LDHA promoter activity was promoted by UCHL3 and inhibited by shUCHL3 in HEK-293 cells
LDHA expression was promoted by UCHL3 through upregulating FOXM1 expression
Studies have found that UCHL3/FOXM1 signal and LDHA/FOXM1 signal have regulatory effects on tumors. However, the relationship among the three genes has not been reported. We found that the protein expressions of FOXM1 and UCHL3 were positively correlated (Fig. 4a–c). Next, BxPC-3 cells and PANC-1 cells were, respectively, transfected by FOXM1 plasmid and shFOXM1 to determine the relationship between FOXM1 and LDHA. Western blot results indicated that the expressions of FOXM1 and LDHA were promoted by FOXM1 plasmid in BxPC-3 cells but inhibited by shFOXM1 in PANC-1 cells (Fig. 4d–f). Co-immunoprecipitation experiments showed that UCHL3 directly interact with FOXM1 (Fig. 4g). These results suggested a regulatory relationship among the three genes. Subsequently, BxPC-3 cells were co-transfected with UCHL3 plasmid and shFOXM1 to detect LDHA expression, and we observed that the mRNA and protein expressions of LDHA were increased by overexpressed UCHL3 but reduced by FOXM1 inhibition (Fig. 4h, i). Dual-luciferase assay also revealed that the increased relative luciferase activity of LDHA by overexpression of UCHL3 was reversed by shFOXM1 (Fig. 4j).
LDHA expression was promoted by UCHL3 through up-regulating FOXM1 expression. a, b, c FOXM1 expression was promoted by UCHL3 in BxPC-3 cells and inhibited by shUCHL3 in PANC cells at the protein level. d, e, f LDHA expression was promoted by FOXM1 in BxPC-3 cells and inhibited by shFOXM1 in PANC cells at protein level. g Co-immunoprecipitation experiments showed that UCHL3 directly interact with FOXM1. h, i LDHA expression was promoted by UCHL3, but was reversed by shFOXM1 at mRNA and protein levels. j LDHA promoter activity was promoted by UCHL3, but was reversed by shFOXM1 at mRNA and protein levels
UCHL3 promoted aerobic glycolysis of BxPC-3 cells through regulating LDHA
Here, whether the role of UCHL3 in aerobic glycolysis of BxPC-3 cells was correlated with LDHA was investigated. BxPC-3 cells were co-transfected with UCHL3 and shLDHA, and the transfection efficiency was determined by Western blot (Fig. 5a, b). MTT assay data revealed that UCHL3 significantly increased the cell proliferation compared with a negative control group, but was partially reversed by shLDHA (Fig. 5c). Glucose uptake, lactic acid production and LDH activity were greatly increased by overexpression of UCHL3, and these phenomena were rescued by shLDHA (Fig. 5d–f).
UCHL3 promoted aerobic glycolysis of BxPC-3 cells through regulating LDHA expression. a, b There was a positive correlation between UCHL3 and LADH at protein level. c UCHL3 promoted viability of BxPC-3 cells through upregulating LDHA expression. d, e, f UCHL3 promoted glucose uptake, lactate production and LDH activity in BxPC-3 cells via regulating LDHA
Discussion
This study found that the expression of UCHL3 was obviously up-regulated in pancreatic cancer tissues and cell lines, and that knocking down UCHL3 inhibited the proliferation and aerobic glycolysis of pancreatic cancer cells. Furthermore, LDHA expression was regulated by UCHL3 through FOXM1. The proliferation and aerobic glycolysis of pancreatic cancer cells were accelerated by UCHL3, while shLDHA was found to reverse those phenomena.
Ubiquitination plays an important role in many cellular biological processes through a variety of protein regulatory mechanisms, such as the regulation of protein activity and degradation, modifying protein function, and altering protein–protein interactions [18, 19]. Siah E3 ubiquitin ligase 1 delays the development of pancreatic carcinogenesis through inhibiting the ubiquitination and degradation of Wnt1-inducible signaling pathway protein-1 (WISP1) [20]. Enforced expression of USP21 accelerates murine PDAC tumor growth and USP21 deubiquitination and promotes cancer cell stemness [21]. Ubiquitin-like protein 4A (UBL4A) exerts an antitumor effect on autophagy-related proliferation and metastasis of pancreatic ductal adenocarcinoma [22]. Ubiquitin exists particularly in ancestral eukaryote UCHL3, which therefore retains ubiquitin throughout evolution [23]. UCHL3, which is a deubiquitinating enzyme of the UCH family, is characterized by dual hydrolase specificity for ubiquitination and Nedd8 [24]. Some previous studies found an imbalance of UCHL3 expression in pancreatic cancer [25], ovarian cancer [26], and breast cancer tissues [14]. Song et al. reported that UCHL3 promotes the proliferation, invasion and drug resistance of pancreatic cancer cells [25]. However, the function of UCHL3 in the aerobic glycolysis of pancreatic cancer has not been elucidated. In this study, we found that UCHL3 expression was significantly up-regulated in pancreatic cancer tissues and cell lines. Moreover, knocking down UCHL3 inhibited the aerobic glycolysis of pancreatic cancer cells. Those results indicated that UCHL3 plays a critical role in pancreatic cancer development.
LDHA irreversibly catalyzes the conversion of pyruvate to lactic acid through the oxidative dehydrogenation of nicotinamide adenine dinucleotide (NADH) to NAD + in the glycolytic pathway. Study also found that LDHA promotes glycolysis of oral squamous cell carcinoma cells and colorectal cancer [27, 28] and mediates the aerobic glycolysis of multiple myeloma cells [29]. Ubiquitination, an important agent in regulating FOXM1 expression, participates in the occurrence and development of tumors, and ubiquitination and degradation of FOXM1 was found to improve the proliferation and epirubicin resistance of breast cancer cells [30]. FOXM1 promotes the warburg effect and pancreatic cancer progression via transactivation of LDHA expression [31]. Knocking down UCHL3 increases FOXM1 ubiquitination to promote FOXM1 expression, subsequently enhancing the sensitivity of pancreatic cancer cells to gemcitabine [25]. Down-regulated FOXM1 increases the sensitivity of pancreatic cancer cells to gemcitabine [32]. Those results demonstrated that the regulation of UCHL3 on the aerobic glycolysis of pancreatic cancer may be correlated with LDHA and FOXM1. The current study found that UCHL3 depletion inhibited the aerobic glycolysis and proliferation of pancreatic cells in a LDHA-dependent manner via FOXM1, suggesting that targeting UCHL3/LDHA/FOXM1 has the potential to be used in treating pancreatic cancer.
It should also be pointed out that our research still has certain limitations, for example, in addition to the lack of clinical data and in vitro experiments, the mechanism of UCHL3 in pancreatic cancer, such as its effects on the proliferation, invasion and migration of the cancer cells, is unclear and therefore requires further investigation.
Our study shows that UCHL3 is a novel biomarker for predicting the progression of pancreatic cancer. Moreover, the current findings support the feasibility of a novel therapy based on targeting the UCHL3/LDHA/FOXM1 axis for the treatment of pancreatic cancer.
Data availability
The data analyzed during the current study are available from the corresponding author on reasonable request.
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin. 2017;67(1):7–30. https://doi.org/10.3322/caac.21387.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018;68(1):7–30. https://doi.org/10.3322/caac.21442.
Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, et al. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66(2):115–32. https://doi.org/10.3322/caac.21338.
Miller KD, Siegel RL, Lin CC, Mariotto AB, Kramer JL, Rowland JH, et al. Cancer treatment and survivorship statistics, 2016. CA Cancer J Clin. 2016;66(4):271–89. https://doi.org/10.3322/caac.21349.
Boulaiz H, Ramos MC, Grinan-Lison C, Garcia-Rubino ME, Vicente F, Marchal JA. What’s new in the diagnosis of pancreatic cancer: a patent review (2011-present). Expert Opin Ther Pat. 2017;27(12):1319–28. https://doi.org/10.1080/13543776.2017.1379991.
Long J, Luo GP, Xiao ZW, Liu ZQ, Guo M, Liu L, et al. Cancer statistics: current diagnosis and treatment of pancreatic cancer in Shanghai, China. Cancer Lett. 2014;346(2):273–7. https://doi.org/10.1016/j.canlet.2014.01.004.
Xie F, Ye L, Ta M, Zhang L, Jiang WG. MTSS1: a multifunctional protein and its role in cancer invasion and metastasis. Front Biosci (Scholar edition). 2011;3:621–31. https://doi.org/10.2741/s175.
Zeleniak AE, Huang W, Brinkman MK, Fishel ML, Hill R. Loss of MTSS1 results in increased metastatic potential in pancreatic cancer. Oncotarget. 2017;8(10):16473–87. https://doi.org/10.18632/oncotarget.14869.
Agarwal E, Robb CM, Smith LM, Brattain MG, Wang J, Black JD, et al. Role of Akt2 in regulation of metastasis suppressor 1 expression and colorectal cancer metastasis. Oncogene. 2017;36(22):3104–18. https://doi.org/10.1038/onc.2016.460.
Huang XY, Huang ZL, Xu B, Chen Z, Re TJ, Zheng Q, et al. Elevated MTSS1 expression associated with metastasis and poor prognosis of residual hepatitis B-related hepatocellular carcinoma. J Exp Clin Cancer Res CR. 2016;35(1):85. https://doi.org/10.1186/s13046-016-0361-8.
Guo Y, Li X, Sun X, Wang J, Yang X, Zhou X, et al. Combined aberrant expression of NDRG2 and LDHA predicts hepatocellular carcinoma prognosis and mediates the anti-tumor effect of gemcitabine. Int J Biol Sci. 2019;15(9):1771–86. https://doi.org/10.7150/ijbs.35094.
Liu YJ, Fan XY, Wang AD, Xia YZ, Fu WR, Liu JY, et al. LDHA suppression altering metabolism inhibits tumor progress by an organic arsenical. Int J Mol Sci. 2019. https://doi.org/10.3390/ijms20246239.
Woodford MR, Chen VZ, Backe SJ, Bratslavsky G, Mollapour M. Structural and functional regulation of lactate dehydrogenase-A in cancer. Future Med Chem. 2020;12(5):439–55. https://doi.org/10.4155/fmc-2019-0287.
Luo K, Li L, Li Y, Wu C, Yin Y, Chen Y, et al. A phosphorylation-deubiquitination cascade regulates the BRCA2-RAD51 axis in homologous recombination. Genes Dev. 2016;30(23):2581–95. https://doi.org/10.1101/gad.289439.116.
Wang M, Yu T, Hu L, Cheng Z, Li M. Ubiquitin carboxy-terminal hydrolasel3 correlates with human sperm count, motility and fertilization. PLoS ONE. 2016;11(10):e0165198. https://doi.org/10.1371/journal.pone.0165198.
Fekrmandi F, Wang TT, White JH. The hormone-bound vitamin D receptor enhances the FBW7-dependent turnover of NF-κB subunits. Sci Rep. 2015;5:13002. https://doi.org/10.1038/srep13002.
Song HM, Lee JE, Kim JH. Ubiquitin C-terminal hydrolase-L3 regulates EMT process and cancer metastasis in prostate cell lines. Biochem Biophys Res Commun. 2014;452(3):722–7. https://doi.org/10.1016/j.bbrc.2014.08.144.
Zhao B, Tsai YC, Jin B, Wang B, Wang Y, Zhou H, et al. Protein engineering in the ubiquitin system: tools for discovery and beyond. Pharmacol Rev. 2020;72(2):380–413. https://doi.org/10.1124/pr.118.015651.
Yu J, Qin B, Lou Z. Ubiquitin and ubiquitin-like molecules in DNA double strand break repair. Cell Biosci. 2020;10:13. https://doi.org/10.1186/s13578-020-0380-1.
Wu W, Liu X, Wei L, Li T, Zang Y, Qian Y, et al. Tp53 mutation inhibits ubiquitination and degradation of wisp1 via down-regulation of siah1 in pancreatic carcinogenesis. Front Pharmacol. 2018;9:857. https://doi.org/10.3389/fphar.2018.00857.
Hou P, Ma X, Zhang Q, Wu CJ, Liao W, Li J, et al. USP21 deubiquitinase promotes pancreas cancer cell stemness via Wnt pathway activation. Genes Dev. 2019;33(19–20):1361–6. https://doi.org/10.1101/gad.326314.119.
Chen H, Li L, Hu J, Zhao Z, Ji L, Cheng C, et al. UBL4A inhibits autophagy-mediated proliferation and metastasis of pancreatic ductal adenocarcinoma via targeting LAMP1. J Exp Clin Cancer Res CR. 2019;38(1):297. https://doi.org/10.1186/s13046-019-1278-9.
Frickel EM, Quesada V, Muething L, Gubbels MJ, Spooner E, Ploegh H, et al. Apicomplexan UCHL3 retains dual specificity for ubiquitin and Nedd8 throughout evolution. Cell Microbiol. 2007;9(6):1601–10. https://doi.org/10.1111/j.1462-5822.2007.00896.x.
Fang Y, Fu D, Shen XZ. The potential role of ubiquitin c-terminal hydrolases in oncogenesis. Biochem Biophys Acta. 2010;1806(1):1–6. https://doi.org/10.1016/j.bbcan.2010.03.001.
Song Z, Li J, Zhang L, Deng J, Fang Z, Xiang X, et al. UCHL3 promotes pancreatic cancer progression and chemo-resistance through FOXM1 stabilization. Am J Cancer Res. 2019;9(9):1970–81.
Zhang MH, Zhang HH, Du XH, Gao J, Li C, Shi HR, et al. UCHL3 promotes ovarian cancer progression by stabilizing TRAF2 to activate the NF-kappaB pathway. Oncogene. 2020;39(2):322–33. https://doi.org/10.1038/s41388-019-0987-z.
Cai H, Li J, Zhang Y, Liao Y, Zhu Y, Wang C, et al. LDHA promotes oral squamous cell carcinoma progression through facilitating glycolysis and epithelial-mesenchymal transition. Front Oncol. 2019;9:1446. https://doi.org/10.3389/fonc.2019.01446.
Wang J, Wang H, Liu A, Fang C, Hao J, Wang Z. Lactate dehydrogenase A negatively regulated by miRNAs promotes aerobic glycolysis and is increased in colorectal cancer. Oncotarget. 2015;6(23):19456–68. https://doi.org/10.18632/oncotarget.3318.
Wu H, Wang X, Wu T, Yang S. miR-489 suppresses multiple myeloma cells growth through inhibition of LDHA-mediated aerobic glycolysis. Genes Genom. 2020;42(3):291–7. https://doi.org/10.1007/s13258-019-00900-z.
Karunarathna U, Kongsema M, Zona S, Gong C, Cabrera E, Gomes AR, et al. OTUB1 inhibits the ubiquitination and degradation of FOXM1 in breast cancer and epirubicin resistance. Oncogene. 2016;35(11):1433–44. https://doi.org/10.1038/onc.2015.208.
Cui J, Shi M, Xie D, Wei D, Jia Z, Zheng S, et al. FOXM1 promotes the warburg effect and pancreatic cancer progression via transactivation of LDHA expression. Clin Cancer Res Off J Am Assoc Cancer Res. 2014;20(10):2595–606. https://doi.org/10.1158/1078-0432.Ccr-13-2407.
Liu C, Shi J, Li Q, Li Z, Lou C, Zhao Q, et al. STAT1-mediated inhibition of FOXM1 enhances gemcitabine sensitivity in pancreatic cancer. Clin Sci (London, England 1979). 2019;133(5):645–63. https://doi.org/10.1042/cs20180816.
Funding
None.
Author information
Authors and Affiliations
Contributions
DT and YF designed the study. DT, DH, DL and YT, QT and YC performed the experiments. DH, DL and YT contributed to the literature search. QT and YC wrote the initial draft of the manuscript. LD and DT reviewed and edited the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Ethical approval and consent to participate
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. No animals are involved in this research.
Patient consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Fan, Y., Hu, D., Li, D. et al. UCHL3 promotes aerobic glycolysis of pancreatic cancer through upregulating LDHA expression. Clin Transl Oncol 23, 1637–1645 (2021). https://doi.org/10.1007/s12094-021-02565-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-021-02565-1