Abstract
Lack of regulated expression and tissue specificity are the major drawbacks of plant and virus-derived constitutive promoters. A precise tissue or site-specific expression, facilitate regulated expression of proteins at the targeted time and site. Publically available microarray data on whitefly and aphid infested Arabidopsis thaliana L. was used to identify whitefly and aphid-inducible genes. The qRT-PCR further validated the inducible behaviour of these genes under artificial infestation. Promoter sequences of genes were retrieved from the Arabidopsis Information Resources database with their corresponding \(5^\prime \hbox {UTR}\) and cloned from the A. thaliana genome. Promoter reporter transcriptional fusions were developed with the beta-glucuronidase (GUS) gusA gene in a binary expression vector to validate the inducible behaviour of these promoters in eight independent transgenic Nicotiana tabaccum lines. Histochemical analysis of the reporter gene in \(\hbox {T}_{2}\) transgenic tobacco lines confirmed promoter driven expression at the sites of aphid and whitefly infestation. The qRT-PCR and GUS expression analysis of transgenic lines revealed that abscisic acid largely influenced the expression of both aphid and whitefly inducible promoters. Further, whitefly-specific promoter respond to salicylic acid and jasmonic acid (JA), whereas aphid-specific promoters to JA and 1-aminocyclopropane carboxylic acid. The response of promoters to phytohormones correlated to the presence of corresponding conserved cis-regulatory elements.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants have evolved a wide range of adaptations in response to many biotic and abiotic stresses to improve their survival and reproduction. Among various biotic interactions, plant–pest interactions are the key factors in stabilizing both natural and man-managed ecosystem (Couldridge et al. 2007). More than one million phytophagus insects utilize aerial and underground plant parts as their food source, either by chewing or by sucking the plant’s phloem sap. Phytophagus insects such as phloem sap-sucking aphids (Myzus persicae) (Bai et al. 2010) and whiteflies (Bemisia tabaci) (Nikos et al. 2011) are the pests of many temperate and glasshouse grown crops. These insects not only assimilate the nutritious sap of phloem but also transmit viral diseases (Kempema et al. 2007) resulting in yield losses in almost all crops. To recover from these yield losses, genetically modified crops, by incorporating accurate tissue-specific expression system for the desired agronomic important gene(s), is the best alternate strategy (Potenza et al. 2004). Thus, the regulated transgene expression at the site and time of insect attack is required to minimize the potential adverse effects of transgene expression on nontargeted organisms and physiology of plants themselves.
The most commonly used promoter in developing transgenic plants is a virus-derived CaMV35S promoter, which is expressed constitutively in all the plant parts (Odell et al. 1985). Similarly, other virus-derived constitutive promoters are also commonly used, such as Cassava vein mosaic virus (CsVMV) (Verdaguer et al. 1998), Australian banana streak virus (BSV) (Schenk et al. 2001), Mirabilis mosaic virus (MMV) (Dey and Maiti 1999) and Figwort mosaic virus (FMV) (Sanger et al. 1990; Maiti et al. 1997). Unnecessary, the overexpression of the transgene in all plant parts, all the time, may lead to unexpected consequences on plant growth and development. Sometimes, transgene silencing occurs for the foreign promoters result in the stable suppression of gene activity throughout the plant or affect the specificity of a promoter (Kloti et al. 2002). The silencing process was observed to be less prevalent when constitutive promoters from a plant were used (Potenza et al. 2004). Several groups have used different plant promoters for transgenes expression such as actin (Act2) of A. thaliana (An et al. 1996), rice actin 1 (McElroy et al. 1991; Zhang et al. 1991), maize ubiquitin 1 (pUbi) (Christensen et al. 1992), and UbiU4 of Nicotiana sylvestris (Plesse et al. 2001). Similarly, maize ubiquitin 1 and rice sucrose synthase (RSs1) gene promoter were used to express the snowdrop lectin Galanthus nivalis leaf agglutinin (GNA) against the brown plant hopper (Nilaparvata lugens) (Sudhakar et al. 1998).
Phloem specific expression of insecticidal protein would be the better strategy to control and enhance the expression of protein in the phloem sap for higher activity against sucking pests. Researchers already used phloem-specific promoters such as RolC (Chakraborti et al. 2009), maize sucrose synthase-1 promoter (Yang and Russell 1990); these promoters were also used to express Alium sativum leaf agglutinin (ASAL) against aphids (Saha et al. 2007). The promoters of pumpkin PP2 (Guo et al. 2004) and Commelina yellow mottle virus (Matsuda et al. 2002) were also reported to be phloem-specific promoters. These promoters are phloem specific, but their heterologous source and constitutive nature are still the concerns. Thus, the identification of sap-sucking insect-inducible plant promoters to target transgene expression specifically in phloem sap may ideally prove to be a powerful technique for sap-sucking insect resistant transgenic plant development. In the present study, we identified three promoters in A. thaliana and conceptually demonstrated their expression in transgenic plants after infestation with aphids and whitefly.
Materials and methods
Selection of candidate genes and construct preparation
The .cel files of the GEO database (GSE6516 and GSE5525) were selected and analysed as explained in the result section. A number of genes were significantly affected by infestations of aphids and whitefly (figure 1 and table 1 (sheets 1–6) in electronic supplementary material at http://www.ias.ac.in/jgenet/). About 549 genes were upregulated in whitefly infestation while 372 and 480 genes were upregulated at 48 and 72 h at postaphid infestation respectively (figure 1 in electronic supplementary material). About 599 genes were downregulated in whitefly infestation while 490 and 960 genes were downregulated at 48 and 72 h postaphid infestation respectively (figure 1 in electronic supplementary material). List of significantly affected genes after whitefly and aphid infestation at different time points is provided in table 1 in electronic supplementary material, includes genes upregulated after 21 days of whitefly infestation (sheet 1 in table 1 of electronic supplementary material), downregulated genes (sheet 2 in table 1 of electronic supplementary material), aphid 48 h infestation upregulated genes (sheet 3 in table 1 of electronic supplementary material), downregulated genes (sheet 4 in table 1 of electronic supplementary material), aphid 72 h infestation upregulated genes (sheet 5 in table 1 of electronic supplementary material), downregulated genes (sheet 6 in table 1 of electronic supplementary material) and commonly deferentially expressed gene between aphid and whitefly infestation (sheet 7 in table 1 of electronic supplementary material).
Three genes, namely W250 (AT1G19250, flavin-containing monooxygenase), A360 (AT3G48360, speckle-type POZ protein), and A080 (AT2G40080, early flowering 4), were selected (table 2 in electronic supplementary material), validated with quantitative reverse transcription polymerase chain reaction (qRT-PCR), and used for promoter cloning and transgenic plant development. The 1 kb promoter regions of the selected genes with their respective \(5^\prime \hbox {UTRs}\) were retrieved from the TAIR database (http://www.arabidopsis.org/). All three selected promoters were amplified by using A. thaliana genomic DNA as a template and AccuPrime Pfx DNA polymerase (cat. no. 12344-024, Invitrogen, Carlsbad, USA). Primer sequences used for the amplification of promoters and used elsewhere in this study are listed in table 3 of electronic supplementary material. PCR products containing SalI and BamHI restriction sites at their ends were first cloned into \(\hbox {pSK}^{+}\) for sequencing. These promoters from \(\hbox {pSK}^{+}\) clone were subcloned into a binary vector pBI 101 upstream to a GUS reporter gene (Clontech, http://www.clontech.com). Tobacco transformation was done as established by Horsch et al. (1985). Transgenic tobacco seeds (\(\hbox {T}_{1}\)) were collected and grown on kanamycin (300 mg/L) for positive plant selection. Seeds of the eight independent transgenic lines of \(\hbox {T}_{2}\) generation were grown for a maximum of 8 weeks for further experiments.
Plant growth conditions and insect infestation
A. thaliana plants of a Col-0 background were grown on vermiculite and solarite TC mix (Keltech, Bengaluru, India) in a 10-inch plastic pot and kept at \(4{^{\circ }}\hbox {C}\) for 3 days. After 3 days, the sown seeds were transferred to a culture room that was maintained at 16 h light / 8 h dark cycle at \(22{^{\circ }}\hbox {C}\). Three to four week-old plants were selected for the experiments. Transgenic tobacco plants were grown in a greenhouse under standard field irrigation and photoperiod.
Expression patterns of selected genes established by qRT-PCR during whitefly and aphid infestation. Leaves of A. thaliana (Col-0) plants were challenged with whiteflies and aphids for 7, 14 and 21 d and 2, 24, 48, 72 and 96 h, respectively. Leaves of unchallenged plants were considered as controls. All the experiments were performed in biological and experimental triplicate condition.
The culture of aphids (M. persicae and M. nicotinae) and whiteflies (B. tabaci) was maintained on potted A. thaliana and N. tabaccum plants in the insectry at \(26\pm 2{^{\circ }}\hbox {C}\) and 70% relative humidity (Upadhyay et al. 2011). Newly emerged whiteflies and second instars aphid nymphs were used for the experiments.
Ten aphids (M. persicae) per plant were released on A. thaliana. After 2, 24, 48, 72 and 96 h of aphid infestation, aphids were removed with a fine brush and the leaves were used for RNA isolation.
For aphid (M. nicotinae) infestation treatment of the \(\hbox {T}_{2}\) transgenic tobacco plant, leaf disks of a 10-mm diameter were cut, transferred to agar plate, and challenged with 15 aphids for 48 h in parallel to noninfested control leaf disks. After 48 h of aphid infestation on leaf disks, aphids were removed and leaves were used in GUS analysis (both fluorimetric and histochemical).
Fifteen whiteflies (B. tabaci) per plant were released on A. thaliana. After 7, 14 and 21 days of whitefly infestation, whiteflies were removed and RNA was isolated at each time point with control noninfested plants in biological triplicates. Time points of the highest induction of genes were identified in A. thaliana leaves, and they were selected for the infestation experiment in transgenic tobacco leaf.
For whitefly infestation, the leaf disk of the transgenic plant was challenged for 7 days with 10 to 15 whiteflies in parallel to the control (Upadhyay et al. 2011). All the experiments were performed in biological triplicate and experimental duplicate conditions.
Phytohormone treatment
Hundred milligrams of leaves from A. thaliana plants were cut and dipped in Hoagland media containing 1 mM salicylic acid (SA) (Onate-Sanchez and Singh 2002), \(100~\mu \hbox {M}\) meJA (Onate-Sanchez and Singh 2002), \(100~\mu \hbox {M}\) abscisic acid (ABA) (Zhang et al. 2008a), and \(5\,\mu \hbox {M}\) 1-aminocyclopropane carboxylic acid (ACC) (Staal et al. 2011) in parallel to the control Hoagland media. After 2, 6, 12, 18, 24 and 48 h of treatment, the RNA was isolated. Time points of the highest expression of selected genes in A. thaliana leaves were selected for the phytohormone treatment in transgenic tobacco leaf disks. All the experiments were performed in the biological triplicate condition.
Complementary DNA (cDNA) preparation and qRT-PCR
Ten \(\mu \hbox {g}\) RNA was used for DNaseI treatment (Ambion). DNaseI-treated RNA (\(2~\mu \hbox {g}\)) was used for the cDNA preparation by using SuperScript cDNA Synthesis Kit (Invitrogen). The qRT-PCR reaction was performed on the ABI 7500 Real-Time PCR Detection System (Applied Biosystems, Foster City, USA) by using SYBR Green PCR Master Mix (Applied Biosystems). Relative fold changes in expression were measured by using the \(2^{-\Delta \Delta {\mathrm{CT}}}\) method. The expression of actin gene (AT3G18780.2) was used as an internal control for data normalization of selected genes at all time points (Livak and Schmittgen 2001). The primer sequences used in this study are given in table 3 in electronic supplementary material.
Identification of cis-regulatory element
The promoter sequences were analysed by the PLACE analysis (PLACE, plant cis-acting regulatory DNA elements, http://www.dna.affrc.go.jp/PLACE/) for the theoretical identification of the cis-regulatory element-binding sites against the PLACE database (Higo et al. 1999).
Fluorimetric GUS assay and histochemical GUS analysis
Fluorimetric GUS assays were performed as described by Chaturvedi et al. (2006). For histochemical analysis, insect challenged and control leaf disks were coincubated at \(37{^{\circ }}\hbox {C}\) with 50 mM X-Gluc (5-bromo-4-chloro-3-indolylglucuronide) that was buffered overnight with 50 mM sodium phosphate buffer pH 7, 0.2% Triton X100, 3 mM potassium ferricyanide, 3 mM potassium ferrocyanide and 20% methanol.
Results
Identification of inducible genes and their validation with qRT-PCR
To identify aphid and whitefly-inducible genes, microarray expression profiles in the GEO database, namely GSE6516 for whitefly infestation of A. thaliana (Kempema et al. 2007) and GSE5525 for aphid infestation of A. thaliana (De Vos et al. 2005) were selected. The analysis was carried out with ArrayAssist software 5.2.2 (Stratagene, Santa Clara, USA) to select highly inducible genes in comparison to the control. We considered only those differentially expressed genes which showed fold change \({\ge } 2.0\) and \(P \le 0.05\). Further, the results were also confirmed with MeV software (https://sourceforge.net/projects/mev-tm4/). Based on our analysis, one gene, namely flavin-containing monooxygenase (AT1G19250), coded as W250 from the whitefly dataset, and two genes, namely speckle-type POZ protein (AT3G48360), coded as A360 and early flowering 4 (AT2G40080), coded as A080 from the aphid-inducible dataset, were selected for further evaluation (table 2 in electronic supplementary material).
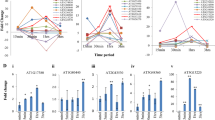
To evaluate the aphid and whitefly-inducible behaviour of selected genes, RNA prepared from A. thaliana-infested plants were subjected to qRT-PCR analysis. More than a 50-fold induction of the W250 gene was observed in leaves of A. thaliana plants after 21 days of whitefly infestation (figure 1a). Thus, the qRT-PCR analysis validated the expression of the W250 gene, as determined earlier in the microarray result (table 2 in electronic supplementary material). In the case of aphid infestation, the expression of A360 was induced about eight-fold at 24 h whereas the expression of A080 was induced about four-fold at 48 h of infestation (figure 1b). The expression of A360 was also induced at a later time point of infestation, i.e. 96 h, which was more than six-fold compared with noninfested control plants. Thus, the expression of all the selected genes by qRT-PCR was found similar to that of microarray result (table 2 in electronic supplementary material).
Validation of selected promoters in transgenic tobacco plants
Promoters of all the three selected genes were cloned upstream to the gusA gene in the pBI101 vector, and 12–14 independent transgenic tobacco lines were generated by using agrobacterium-mediated transformation. Tobacco was selected as a model plant system for studying the promoters isolated from A. thaliana. Evaluation of the inducible behaviour of promoters in a heterologous plant system will validate the instant feasibility of its use of the expression of insect-resistant proteins in crop plants. The expression pattern of selected promoters in tobacco will also reflect their utility in the crop improvement programme. Eight-week-old transgenic tobacco plants from eight independent transgenic (\(\hbox {T}_{2}\)) lines were selected for the fluorimetric experiments. All the assays were performed in biological triplicate and experimental duplicate in parallel to the control. Transgenic plants expressing the GUS protein driven by \(\hbox {P}_{\mathrm{W250}}\) showed average GUS activity 91 pmol/mg/min protein after seven days of whitefly infestation (figure 2a), which was significantly higher than the noninfested control. The histochemical staining of whitefly-infested transgenic leaves showed that the GUS expression of promoter \(\hbox {P}_{\mathrm{W250}}\) was specifically at the site of whitefly infestation (figure 2d) in agreement with the quantitative data. Whereas in \(\hbox {P}_{\mathrm{A360},}\) the average GUS activity of 13 pmol/mg/min protein was observed after 48 h of aphid infestation (figure 2b), which was significantly higher compared with the noninfested control. Also, in the case of \(\hbox {P}_{\mathrm{A360}}\), histochemical staining for GUS activity was restricted on the site of aphid infestation (figure 2 in electronic supplementary material). However, transgenic lines of \(\hbox {P}_{\mathrm{A080}}\) showed a high background level GUS activity, which was induced nonsignificantly as much as 417 pmol/mg/min protein after 48 h of aphid infestation (figure 2c). Further, significant GUS staining was observed in both aphid-infested and noninfested areas in the stained leaves of transgenic lines of the promoter \(\hbox {P}_{\mathrm{A080}}\) (figure 2, e&f). Thus, these results validated the inducible expression of selected promoters of the site of aphid and whitefly infested transgenic leaves.
GUS expression pattern of selected promoters \(\hbox {P}_{\mathrm{W250},}\) \(\hbox {P}_{\mathrm{A360}}\) and \(\hbox {P}_{\mathrm{A080}}\) (fluorimetric: a, b, c; histochemical: d (\(\hbox {P}_{\mathrm{W250}}\)), e, f (\(\hbox {P}_{\mathrm{A080}}\)) in \(\hbox {T}_{2}\) transgenic tobacco plants after 7 days of whitefly (\(\hbox {P}_{\mathrm{W250}}\)) and 48 h of aphid (\(\hbox {P}_{\mathrm{A360}}\) and \(\hbox {P}_{\mathrm{A080}}\)) infestation). Mean ± SE were obtained from eight independent transgenic lines with biological triplicate and experimental duplicate. Bars labelled with stars indicate the significant differences as determined by t-test analysis (\(P \le 0.05\)).
Expression profile of selected genes in response to SA, jasmonic acid (JA), ACC and ABA treatment
The expression of insect-inducible genes is regulated by phytohormones such as SA, JA, ABA and ethylene. Thus, phytohormones SA, JA, ABA and ACC (a nonvolatile precursor of ethylene) were selected to understand the regulatory nature of the selected promoters. The influence of these hormones on the expression of selected genes (W250, A360 and A080) was assessed by qRT-PCR after treatment with phytohormones in A. thaliana. The expression of selected genes was assessed at 2, 6, 12, 18, 24 and 48 h after phytohormone treatment.
The expression of W250 was significantly induced after 48 h of SA treatment (figure 3a). In the case of JA treatment, the expression of W250 was found to be substantially higher after 12 h of treatment (figure 3b). The response of W250 toward ABA was gradual, and it attains maximum expression after 48 h of treatment (figure 3c). The treatment of ACC did not show any significant induction at any time points when we examined (figure 3d). Thus, we selected 48 h for SA, 12 h for JA, and 48 h for ABA as optimal time points for phytohormonal treatment of transgenic lines expressing \(\hbox {P}_{\mathrm{W250}}\). We observed significant induction in the GUS activity in transgenic lines treated with SA and JA (figure 3e). However, in the case of ABA treatment, although transgenic lines showed an elevated level of expression, it was not statistically significant. The expression of A360 was also examined post-treatment of SA, JA, ABA and ACC at different time points. The expression of A360 was found to be the highest at 24 h in all the treatments, including JA (figure 4a), ACC (figure 4b), and ABA (figure 4c). However, in the case of SA treatment, we did not observe any significant induction at any time points analysed (figure 4d). Thus, the 24-h time point was selected to evaluate the expression of \(\hbox {P}_{\mathrm{A360}}\) in transgenic tobacco lines expressing GUS post-treatment of JA, ACC and ABA. The expression of \(\hbox {P}_{\mathrm{A360}}\) was elevated in SA and JA treatments, but it was statistically significant only in ABA treatment (figure 4e). The expression of A080 was also evaluated after treatment of SA, JA, ABA and ACC at different time points. The expression of A080 was not significantly affected by treatment of SA and JA; however, A080 does respond to ACC and ABA (figure 5, a–d). The expression of A080 was found highest after 6 h of ACC treatment and 48 h of ABA treatment; thus, these hormones and time points were further selected to evaluate transgenic lines expressing GUS under the control of \(\hbox {P}_{\mathrm{A080}}\). Transgenic lines showed elevated expression in response to 6 h of ACC treatment; however, this induction was not statistically significant considering the large variation in independent transgenic lines in response to ACC (figure 5e). However, ABA treatment for 48 h resulted in induced expression of GUS, which was statistically significant compared with the untreated control (figure 5e).
Expression pattern of the W250 gene in response to SA, JA, ABA and ACC treatment in A. thaliana (Col-0) leaves at different time points (a–d). The time point of maximum induction was selected for further hormone treatment to transgenic tobacco leaves. GUS activity of \(\hbox {P}_{\mathrm{W250}}\) after SA (48 h), JA (12 h), and ABA (48 h) treatment (e). Bars labelled with * indicate the significant differences as determined by t-test analysis (\(P \le 0.05\)).
Expression pattern of the A360 gene in response to JA, ACC, ABA and SA treatment in A. thaliana (Col-0) leaves at different time points (a–d). The time point of maximum induction was selected for further hormone treatment to transgenic tobacco leaves. GUS expression of \(\hbox {P}_{\mathrm{A360}}\) after JA (24 h), ACC (24 h), and ABA (24 h) treatment (e). Bars labelled with stars indicate the significant differences as determined by t-test analysis (\(P \le 0.05\)).
Expression pattern of A080 gene in response to ACC, ABA, SA and JA treatment in A. thaliana (Col-0) leaves (a–d). The time point of maximum induction was selected for further hormone treatment to transgenic tobacco leaves. GUS activity of \(\hbox {P}_{\mathrm{A080}}\) after ACC (6 h) and ABA (48 h) treatment (e). Bars labelled with stars indicate the significant differences as determined by t-test analysis (\(P \le 0.05\)).
Cis-regulatory motif analysis of cloned promoter
All the cloned promoter sequences were subjected to the cis-regulatory motif analysis by using the PLACE (http://www.dna.affrc.go.jp/PLACE/) database (figure 6). The maximum numbers of conserved cis motifs identified were of MYCCONSENSUSAT (found in dehydration responsive genes) origin in all three cloned promoters (figure 6). All the cloned promoters also showed a higher number of ABA-responsive elements such as ABRELATERD1, ACGTATERD1, ABRERATCAL, DPBFCOREDCDC3, MYB1AT, MYB2CONSENSUSAT and MYCCONSENSUSAT, and probably responsible for ABA-induced expression either in A. thaliana (figures 3c, 4c and 5b) or in transgenic tobacco lines (figures 3e, 4e and 5e). \(\hbox {P}_{\mathrm{W250}}\) also showed conservation of SA-responsive cis-regulatory element WBOXATNPR1 (figure 6) that agrees with its SA-induced expression (figure 3, a&c), thus corroborating our results. We did not find any JA-responsive cis-regulatory element in the cloned promoter fragment of \(\hbox {P}_{\mathrm{W250}}\) but the induction of the native gene was observed in JA at 12 h in A. thaliana (figure 3b) and transgenic tobacco lines (figure 3e). Other than the ABREs mentioned earlier, three more motifs, ABREATCONSENSUS, ACGTABREMOTIF2OSEM and DRECRTCOREAT, were found to present exclusively in \(\hbox {P}_{\mathrm{A360}}\). In accordance, \(\hbox {P}_{\mathrm{A360}}\) showed relatively early ABA induction in native A. thaliana (figure 4c) and transgenic tobacco lines (figure 4e). Along with several ABREs found in the two promoters mentioned earlier, an additional MYB recognition sequence MYB 2AT was found to be present in \(\hbox {P}_{\mathrm{A080}}\). The abundance of ABREs in \(\hbox {P}_{\mathrm{A080}}\) was also mirrored in its ABA-inducible expression in the case of both A. thaliana (figure 5b) and transgenic tobacco lines (figure 5e).
Discussion
In this study, we cloned and characterized one whitefly and two aphid-inducible promoters from A. thaliana. The whitefly and aphid inducible genes were identified by analysing publically available datasets (GSE6516 and GSE5525). All the three selected genes were well validated for their expression by qRT-PCR in A. thaliana (figure 1), indicating the selection of genes by microarray profiles represents their true expression patterns. The selected gene W250 (AT1G19250) encodes FMO, which is reported to be involved in the plant–pathogen interaction (Mishina and Zeier 2006). The gene A360 (AT3G48360), known as BT2, a protein with BTB, TAZ, and calmodulin-binding domains, is an essential component of the TAC1-mediated telomerase activation pathway (Ren et al. 2007) and it is localized in the nucleus and cytosol (Robert et al. 2009). The gene A080 (AT2G40080) encoded a small protein known as ELF4 and is essential for circadian clock function, seedling de-etiolation, photoperiod perception, and flowering (Khanna et al. 2003; Doyle et al. 2005; Kim et al. 2013). The expression of A360 and A080, in response to aphid infestation, is interesting and suggests crosstalk of hormonal pathways that are responsible for telomerase activation and flowering with that of plant insect interactions.
The GUS estimation showed that \(\hbox {P}_{\mathrm{A080}}\) is the strongest among all the selected promoters, as GUS activity driven by this promoter reaches a maximum of 400 pmol/mg/min protein after aphid infestation (figure 2c). The highest expression of \(\hbox {P}_{\mathrm{A080}}\) might also be due to its higher level of background expression. Since histochemical analysis of aphid-infested leaves of transgenic lines indicated that the expression of \(\hbox {P}_{\mathrm{A080}}\) was not strictly restricted on the site of infestation (figure 2, e&f), which was in sharp contrast to that observed in the case of \(\hbox {P}_{\mathrm{W250}}\) and \(\hbox {P}_{\mathrm{A360}}\) (figure 2d; figure 2 in electronic supplementary material). We identified that fold induction in all three selected genes in response to whitefly or aphid infestation was significantly higher (figure 1a) in A. thaliana as compared with their respective promoters in transgenic tobacco lines (figure 2, a, b&c). This higher gene expression may be due to better sensitivity and linearity of qRT-PCR than the enzymatic GUS activity. It is also noteworthy that for each promoter, several transgenic lines are evaluated; thus, the expression of promoters is strongly influenced by their position of integration in the genome (Horner et al. 1995; Kiran et al. 2006; Chaturvedi et al. 2007; Lodhi et al. 2008; Tiwari et al. 2008; Srivastava et al. 2014).
Insect attacks lead to the modulation of plants phytohormonal pathways to cope with attacking insects (Kempema et al. 2007). SA-related and JA-related pathways play an important role during the infestation of whitefly and aphid, respectively (Kempema et al. 2007). Our previous result showed that the aphids and whiteflies modulated the expression of phytohormonal pathway related genes in cotton plants (Dubey et al. 2013). The expression of W250 (FMO1) responds to 12 h of JA treatment; whereas its expression increased in the later stage of SA and ABA treatments (figure 3). An enhancement in the expression of W250 in response to SA treatment complements the earlier published report, where the authors reported it as an SA marker gene for cell death, involvement in lesion and SAR development (Pieterse et al. 1996; Brodersen et al. 2005; Graaff et al. 2006; Mishina and Zeier 2006; Zhang et al. 2008b). The expression of \(\hbox {P}_{\mathrm{A360}}\) (BT2) was reported to be downregulated after infection with the Plasmodiophora brassicae in A. thaliana (Siemens et al. 2006) and induced by other stresses such as drought, high salt and cold (Fujita et al. 2007). Exogenous sugars decreased A360 expression, whereas exogenous nitrogen increased its expression (Mandadi et al. 2009). Sap-sucking insects, including aphids, are secondary sinks of sugar in plants and deprivation of sugar through aphids may be the reason of induction of A360 genes in aphid-infested plants (figure 1b). Induction of A360 by both ACC and ABA at the same time point, i.e. 24 h (figure 4, b&c), and induction of A080 by both ACC and ABA at 6 h (figure 5, a&b) indicate the involvement of ABA and ACC cross-talk in the regulation of A360 and A080 genes. The expression profile of selected genes in response to SA, JA, ACC and ABA indicates that both whitefly and aphid can induce selected genes via different phytohormone pathways.
In our study, we identified potential cis-regulatory elements conserved in the cloned promoters (figure 3 in electronic supplementary material). These cis-acting elements were earlier reported to play their respective role in regulating the expression of those genes. One of the important cis-regulatory motifs identified in all the cloned promoters was ‘MYCCONSENSUSAT’; this particular motif was previously reported to regulate ABA-induced expression in the rd22 gene in A. thaliana (Abe et al. 2003). We also identified additional ABA-responsive elements such as ABREATCONSENSUS and ACGTABREMOTIFA2OSE in \(\hbox {P}_{\mathrm{A360}}\); both these motifs are a part of ABA and drought response (Choi et al. 1999; Huang et al. 2008). Drought-induced upregulation of ABA signalling decreases SA-dependent response but increases the JA-dependent response in wild-type Medicago truncatula after the infestation of the pea aphid (Guo et al. 2016). Other than ABREs, ‘MYCATERD1’ and ‘MYCATRD22’ are the two MYC recognition sequences exclusively present in the \(\hbox {P}_{\mathrm{W250}}\) that are also known to have a distinct role in ABA and drought inducible expression (Abe et al. 1997; Simpson et al. 2003). In the case of \(\hbox {P}_{\mathrm{A080}}\), an extra ABA-responsive motif MYB2AT was identified that was reported for the regulation of ABA response in the Atmyb2 gene in A. thaliana (Urao et al. 1993). Besides, we also identified conserved WBOXATNPR1 in the \(\hbox {P}_{\mathrm{W250}}\) promoter (figure 6), which explains its SA-induced expression (figure 3, c&e); this particular motif was previously reported for SA-induced expression of the NPR1 gene in A. thaliana (Yu et al. 2001). However, further detailed investigations involving site-directed mutagenesis of these elements and expression of the mutant version of these promoters in plants will be needed to substantiate the role of these cis-regulatory motifs in the regulation.
In conclusion, well characterized inducible promoter identified in this study can be used to regulate the expression of the insecticidal protein in fine-tune manner. These promoters are strictly expressed at the site of whitefly and aphid infestation, making them suitable for biotechnological interventions. Both the whitefly and aphid inducible promoters are regulated by ABA, whereas whitefly inducible promoters specifically induced by SA and aphid induced promoters are regulated by JA.
References
Abe H., Yamaguchi-Shinozaki K., Urao T., lwasaki T., Hosokawa C. D. and Shinozaki K. 1997 Role of arabidopsis MYC and MYB homologs in drought and abscisic acid-regulated gene expression. Plant Cell 9, 1859–1868.
Abe H., Urao T., Ito T., Seki M., Shinozaki K. and Yamaguchi-Shinozaki K. 2003 Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signaling. Plant Cell 15, 63–78.
An Y. Q., McDowell J. M., Huang S., McKinney E. C., Chambliss S. and Meagher R. B. 1996 Strong, constitutive expression of the Arabidopsis ACT2/ACT8 actin subclass in vegetative tissues. Plant J. 10, 107–121.
Bai X., Zhang W., Orantes L., Jun T. H., Mittapalli O., Mian M. A. R. and Michel A. P. 2010 Combining next-generation sequencing strategies for rapid molecular resource development from an invasive aphid species, Aphis glycines. PLoS One 5, e11370.
Brodersen P., Malinovsky F. G., Hematy K., Newman M. A. and Mundy J. 2005 The role of salicylic acid in the induction of cell death in Arabidopsis acd11. Plant Physiol. 138, 1037–1045.
Chakraborti D., Sarkar A., Mondal H. A. and Das S. 2009 Tissue specific expression of potent insecticidal, Allium sativum leaf agglutinin (ASAL) in important pulse crop, chickpea (Cicer arietinum L.) to resist the phloem feeding Aphis craccivora. Transgenic Res. 18, 529–544.
Chaturvedi C. P., Sawant S. V., Kiran K., Mehrotra R., Lodhi N., Ansari S. A. and Tuli R. 2006 Analysis of polarity in the expression from a multifactorial bidirectional promoter designed for high-level expression of transgenes in plants. J. Biotechnol. 123, 1–12.
Chaturvedi C. P., Lodhi N., Ansari S. A., Tiwari S., Srivastava R., Sawant S. V. and Tuli R. 2007 Mutated TATA-box/TATA binding protein complementation system for regulated transgene expression in tobacco. Plant J. 50, 917–925.
Christensen A. H., Sharrock R. A. and Quail P. H. 1992 Maize poly ubiquitin genes: structure, thermal perturbation of expression and transcript splicing, and promoter activity following transfer to protoplasts by electroporation. Plant Mol. Biol. 18, 675–689.
Choi H., Hong J. H., Ha J., Kang J. Y. and Kim S. Y. 1999 ABFs, a family of ABA-responsive element binding factors. J. Biol. Chem. 275, 1723–1730.
Couldridge C., Newbury H. J., Ford-Lloyd B., Bale J. and Pritchard J. 2007 Exploring plant responses to aphid feeding using a full Arabidopsis microarray reveals a small number of genes with significantly altered expression. Bull. Entomol. Res. 97, 523–532.
De Vos M., Oosten V. R., Van Poecke R. M. P., Van Pelt J. A., Pozo M. J., Mueller M. J. et al. 2005 Signal signature and transcriptome changes of Arabidopsis during pathogen and insect attack. Mol. Plant Microbe Interact. 18, 923–937.
Dey N. and Maiti I. B. 1999 Structure and promoter/leader deletion analysis of mirabilis mosaic virus (MMV) full-length transcript promoter in transgenic plants. Plant Mol. Biol. 40, 771–782.
Doyle M. R., Bizzell C. M., Keller M. R., Michaels S. D., Song J., Noh Y. S. and Amasino R. M. 2005 HUA2 is required for the expression of floral repressors in Arabidopsis thaliana. Plant J. 41, 376–385.
Dubey N. K., Goel R., Ranjan A., Idris A., Singh S. K., Bagh S. K., et al. 2013 Comparative transcriptome analysis of Gossypium hirsutum L. in response to sap sucking insects: aphid and whitefly. BMC Genomics 14, 241.
Fujita M., Mizukado S., Fujita Y., Ichikawa T., Nakazawa M., Seki M. et al. 2007 Identification of stress tolerance related transcription factor genes via mini scale Full length cDNA over expressor (FOX) gene hunting system. Biochem. Biophys. Res. Commun. 364, 250–257.
Graaff V. D. E., Schwacke R., Schneider A., Desimone M., Flogge U. I. and Kunze R. 2006 Transcription analysis of Arabidopsis membrane transporters and hormone pathways during developmental and induced leaf senescence. Plant Physiol. 141, 776–792.
Guo H., Chen X., Zhang H., Fang R., Yuan Z., Zhang Z. and Tian Y. 2004 Characterization and activity enhancement of the phloem-specific pumpkin PP2 gene promoter. Transgenic Res. 13, 559–566.
Guo H., Sun Y., Peng X., Wang Q., Harris M. and Ge F. 2016 Up-regulation of abscisic acid signaling pathway facilitates aphid xylem absorption and osmoregulation under drought stress. J. Exp. Bot. 67, 681–693.
Higo K., Ugawa Y., Iwamoto M. and Korenaga T. 1999 Plant cis-acting regulatory DNA elements (PLACE) database. Nucleic Acids Res. 27, 297–300.
Horner G. W., Tham K. M., Orr D., Ralston J., Rowe S. and Houghton T. 1995 Comparison of an antigen capture enzyme-linked assay with reverse transcription polymerase chain reaction and cell culture immunoperoxidase tests for the diagnosis of ruminant pestivirus infections. Vet. Microbiol. 43, 75–84.
Horsch R. B., Fry J. E., Hoffmann N. L., Eichholtz D., Rogers S. G. and Fraley R. T. 1985 A simple and general method for transferring genes into plants. Science 227, 1229–1231.
Huang D., Wu W., Abrams S. R. and Cutler A. J. 2008 The relationship of drought-related gene expression in Arabidopsis thaliana to hormonal and environmental factors. J. Exp. Bot. 59, 2991–3007.
Kempema L. A., Cui X., Holzer F. M. and Walling L. L. 2007 Arabidopsis transcriptome changes in response to phloem-feeding silver leaf whitefly nymphs. Similarities and distinctions in responses to aphids. Plant Physiol. 143, 849–865.
Khanna R., Kikis E. A. and Quail P. H. 2003 EARLY FLOWERING 4 Functions in phytochrome B regulated seedling de-etiolation. Plant Physiol. 133, 1530–1538.
Kim Y., Lim J., Yeom M., Kim H., Kim J., Wang L., et al. 2013 ELF4 regulates GIGANTEA chromatin access through sub nuclear sequestration. Cell Rep. 28, 671–677.
Kiran K., Ansari S. A., Srivastava R., Lodhi N., Chaturvedi C. P., Sawant S. V. et al. 2006 The TATA-box sequence in the basal promoter contributes to determining light-dependent gene expression in plants. Plant Physiol. 142, 364–376.
Kloti A., He X., Potrykus I., Hohn T. and Futterer J. 2002 Tissue-specific silencing of a transgene in rice. Proc. Natl. Acad. Sci. USA 99, 10881–10886.
Livak K. J. and Schmittgen T. D. 2001 Analysis of relative gene expression data using real-time quantitative PCR and the 2\(^{-\Delta \Delta \text{ CT }}\) method. Methods 25, 402–408.
Lodhi N., Ranjan A., Singh M., Srivastava R., Singh S. P., Chaturvedi C. P. et al. 2008 Interactions between upstream and core promoter sequences determine gene expression and nucleosome positioning in tobacco PR-1a promoter. Biochim. Biophys. Acta. 1779, 634–644.
Maiti I. B., Gowda S., Kiernan J., Ghosh S. K. and Shepherd R. J. 1997 Promoter/leader deletion analysis and plant expression vectors with the figwort mosaic virus (FMV) full length transcript (FLt) promoter containing single or double enhancer domains. Transgenic Res. 6, 143–156.
Mandadi K. K., Misra A., Ren S. and McKnight T. D. 2009 BT2, a BTB protein, mediates multiple responses to nutrients, stresses, and hormones in Arabidopsis. Plant Physiol. 150, 1930–1939.
Matsuda Y., Liang G., Zhu Y., Ma F., Nelson R. S. and Ding B. 2002 The Commelina yellow mottle virus promoter drives companion-cell-specific gene expression in multiple organs of transgenic tobacco. Protoplasma 220, 51–58.
McElroy D., Blowers A. D., Jenes B. and Wu R. 1991 Construction of expression vectors based on the rice actin 1 (Act1) 5\(^\prime \) region for use in monocot transformation. Mol. Gen. Genet. 231, 150–160.
Mishina T. E. and Zeier J. 2006 The Arabidopsis flavin-dependent mono oxygenase FMO1 is an essential component of biologically induced systemic acquired resistance. Plant Physiol. 141, 1666–1675.
Nikos K., Yannick P., Ritika C., Ian D., David N. and Chris B. 2011 Pyro-sequencing the transcriptome of the greenhouse whitefly, Trialeurodes vaporariorum reveals multiple transcripts encoding insecticide targets and detoxifying enzymes. BMC Genomics 12, 1–14.
Odell J. T., Nagy F. and Chua N. H. 1985 Identification of DNA sequences required for activity of the cauliflower mosaic virus 35S promoter. Nature 313, 810–812.
Onate-Sanchez L. and Singh K. B. 2002 Identification of Arabidopsis ethylene-responsive element binding factors with distinct induction kinetics after pathogen infection. Plant Physiol. 128, 1313–1322.
Pieterse C. M. J., VanWees S. C. M., Hoffland E., vanPelt J. A. and vanLoon L. C. 1996 Systemic resistance in Arabidopsis induced by biocontrol bacteria is independent of salicylic acid accumulation and pathogenesis-related gene expression. Plant Cell 8, 1225–1237.
Plesse B., Criqui M. C., Durr A., Parmentier Y., Fleck J. and Genschik P. 2001 Effects of the polyubiquitin gene Ubi.U4 leader intron and first ubiquitin monomer on reporter gene expression in Nicotiana tabacum. Plant Mol. Biol. 45, 655–667.
Potenza C., Aleman L. and Sengupta-Gopalan C. 2004 Targeting transgene expression in research, agricultural, and environmental applications: promoters used in plant transformation. In Vitro Cell Dev. Biol. Plant. 40, 1–22.
Ren S., Mandadi K. K., Boedeker A. L., Rathore K. S. and McKnight T. D. 2007 Regulation of telomerase in Arabidopsis by BT2, an apparent target of telomerase activator1. Plant Cell 19, 23–31.
Robert H. S., Quint A., Brand D., Viviana-Smith A. and Offringa R. 2009 BTB and TAZ DOMAIN scaffold proteins perform a crucial function in Arabidopsis development. Plant J. 58, 109–121.
Saha P., Chakraborti D., Sarkar A., Dutta I., Basu D. and Das S. 2007 Characterization of vascular-specific RSs1 and rolC promoters for their utilization in engineering plants to develop resistance against hemipteran insect pests. Planta 226, 429–442.
Sanger M., Daubert S. and Goodman R. M. 1990 Characteristics of a strong promoter from figwort mosaic virus: comparison with the analogous 35S promoter from cauliflower mosaic virus and the regulated mannopine synthase promoter. Plant Mol. Biol. 14, 433–443.
Schenk P. M., Remans T., Saqi L., Elliott A. R., Dietzgen R. G., Swennen R. et al. 2001 Promoters for pre-genomic RNA of banana streak badna virus are active for transgene expression in monocot and dicot plants. Plant Mol. Biol. 47, 399–412.
Siemens J., Keller I., Sarx J., Kunz S., Schuller A., Nagel W. et al. 2006 Transcriptome analysis of Arabidopsis clubroots indicate a key role for cytokinins in disease development. Mol. Plant Microbe Interact. 19, 480–494.
Simpson S. D., Nakashima K., Narusaka Y., Seki M., Shinozaki K. and Yamaguchi-Shinozaki K. 2003 Two different novel cis-acting elements of erd1, a clpA homologous Arabidopsis gene function in induction by dehydration stress and dark-induced senescence. Plant J. 33, 259–270.
Srivastava R., Rai K. M., Srivastava M., Kumar V., Pandey B., Singh S. P. et al. 2014 Distinct role of core promoter architecture in regulation of light-mediated responses in plant genes. Mol. Plant 7, 626–641.
Staal M., De Cnodder T., Simon D., Vandenbussche F., Van Der Straeten D., Verbelen J. P. et al. 2011 Apoplastical kalinization is instrumental for the inhibition of cell elongation in the Arabidopsis root by the ethylene precursor 1-aminocyclopropane-1-carboxylic acid. Plant Physiol. 155, 2049–2055.
Sudhakar D., Fu X., Stoger E., Williams S., Spence J., Brown D. P. et al. 1998 Expression and immune-localisation of the snowdrop lectin, GNA in transgenic rice plants. Transgenic Res. 7, 371–378.
Tiwari S., Mishra D. K., Singh A., Singh P. K. and Tuli R. 2008 Expression of a synthetic cry1EC gene for resistance against Spodoptera litura in transgenic peanut (Arachis hypogaea L.). Plant Cell Rep. 27, 1017–1025.
Upadhyay S. K., Chandrashekar K., Thakur N., Verma P. C., Borgio J. F., Singh P. K., and Tuli R. 2011 RNA interference for the control of whiteflies (Bemisia tabaci) by oral route. J. Biosci. 36, 153–161.
Urao T., Yamaguchi-Shinozaki K., Urao S. and Shinozaki K. 1993 An arabidopsis myb homolog is induced by dehydration stress and its gene product binds to the conserved MYB recognition sequence. Plant Cell 5, 1529–1539.
Verdaguer B., de Kochko A., Fux C. I., Beachy R. N. and Fauquet C. 1998 Functional organization of the cassava vein mosaic virus (CsVMV) promoter. Plant Mol. Biol. 37, 1055–1067.
Yang N. S. and Russell D. 1990 Maize sucrose synthase-1 promoter directs phloem cell-specific expression of Gus gene in transgenic tobacco plants. Proc. Natl. Acad. Sci. USA 87, 4144–4148.
Yu D., Chen C. and Chen Z. 2001 Evidence for an important role of WRKY DNA binding proteins in the regulation of NPR1 gene expression. Plant Cell 13, 1527–1539.
Zhang W., McElroy D. and Wu R. 1991 Analysis of rice Act1 5\(^\prime \) region activity in transgenic rice plants. Plant Cell 3, 1155–1165.
Zhang Y., Xu W., Li Z., Deng X. W., Wu W. and Xue Y. 2008a F-box protein DOR functions as a novel inhibitory factor for abscisic acid-induced stomatal closure under drought stress in Arabidopsis. Plant Physiol. 148, 2121–2133.
Zhang Z., Lenk A., Andersson M. X., Gjetting T., Pedersen C., Nielsen M. E et al. 2008b A lesion-mimic syntaxin double mutant in Arabidopsis reveals novel complexity of pathogen defense signaling. Mol. Plant 1, 510–527.
Acknowledgements
NKD, DKM and DN are grateful to the Council of Scientific and Industrial Research (CSIR), India, for providing the research fellowship. This work was supported under the Council of Scientific and Industrial Research Network Project (NWP03).
Author information
Authors and Affiliations
Corresponding author
Additional information
Corresponding editor: Arun Joshi
NKD performed qRT-PCR, promoter cloning and transgenic analysis, and drafted this article. DN performed microarray data analysis. DKM and AI helped in transgenic development and lab experiments. PKS revised the manuscript. SVS conceived the study, participated in its design and coordination of work, data analysis and interpretation, and revised the article.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dubey, N.K., Mishra, D.K., Idris, A. et al. Whitefly and aphid inducible promoters of Arabidopsis thaliana L.. J Genet 97, 109–119 (2018). https://doi.org/10.1007/s12041-018-0887-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12041-018-0887-y