Abstract
In plants, the transcription factor families have been implicated in many important biological processes. These processes include morphogenesis, signal transduction and environmental stress responses. Proteins containing the lateral organ boundaries domain (LBD), which encodes a zinc finger-like domain are only found in plants. This finding indicates that this unique gene family regulates only plant-specific biological processes. LBD genes play crucial roles in the growth and development of plants such as Arabidopsis, Oryza sativa, Zea mays, poplar, apple and tomato. However, relatively little is known about the LBD genes in grape (Vitis vinifera). In this study, we identified 40 LBD genes in the grape genome. A complete overview of the chromosomal locations, phylogenetic relationships, structures and expression profiles of this gene family during development in grape is presented here. Phylogenetic analysis showed that the LBD genes could be divided into classes I and II, together with LBDs from Arabidopsis. We mapped the 40 LBD genes on the grape chromosomes (chr1–chr19) and found that 37 of the predicted grape LBD genes were distributed in different densities across 12 chromosomes. Grape LBDs were found to share a similar intron/exon structure and gene length within the same class. The expression profiles of grape LBD genes at different developmental stages were analysed using microarray data. Results showed that 21 grape LBD genes may be involved in grape developmental processes, including preveraison, veraison and ripening. Finally, we analysed the expression patterns of six LBD genes through quantitative real-time polymerase chain reation analysis. The six LBD genes showed differential expression patterns among the three representative grape tissues, and five of these genes were found to be involved in responses to mannitol, sodium chloride, heat stress and low temperature treatments. To our knowledge, this is the first study to analyse the LBD gene family in grape and provides valuable information for classification and functional investigation of this gene family.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Transcription factor (TF) families play important roles in several biological processes in plants including growth and development, signal transduction and environmental stress responses. The lateral organ boundaries domain (LBD) gene family encodes plant-specific TFs that function in lateral organ development (Husbands et al. 2007). The LBD gene family can be divided into classes I and II on the basis of the structure of the lateral organ boundaries (LOB) domain in the N-terminus (Iwakawa et al. 2002; Shuai et al. 2002; Yang et al. 2006; Zhu et al. 2007; Majer and Hochholdinger 2011). The LOB domain, which is approximately 100 amino acids in length, contains a completely conserved CX 2 CX 6 CX 3C zinc finger-like domain and a LX 6 LX 3 LX 6L leucine zipper-like domain (Shuai et al. 2002; Lee et al. 2014; Zhang et al. 2014). The LBD gene family has various temporal and tissue expression patterns, indicating that these genes play different functions in diverse processes (Shuai et al. 2002; Wang et al. 2013a).
Previous studies investigated the roles of LBD family in organ development, nitrogen metabolism and anthocyanin biosynthesis in Arabidopsis thaliana (Okushima et al. 2005, 2007; Naito et al. 2007; Rubin et al. 2009; Mangeon et al. 2011; Fan et al. 2012; Feng et al. 2012; Kim and Kim 2012; Thatcheret al. 2012; Kim and Lee 2013; Lee et al. 2013, 2014; Cabrera et al. 2014). For example, ASL9, a unique member of the LBD family (LBD3) was rapidly induced by cytokinin in Arabidopsis and identified as a primary target of cytokinin receptor-mediated phosphorelay signal transduction (Naito et al. 2007). AtLBD16, AtLBD18 and AtLBD29 are the direct targets of AtARF7 and AtARF19 which play a common role in lateral root formation (Okushima et al. 2005, 2007; Fan et al. 2012; Feng et al. 2012). ASL18/LBD16 contains two nuclear localization signal (NLS) domains comprising an atypical NLS that resides in the coiled coil motif and the monopartite-like NLS in the C-terminal region (Kim and Kim 2012). AtLBD16 is also a key component of the auxin pathway which leads to divisions in the xylem pole pericycle for lateral root formation (Cabrera et al. 2014). AtLBD18 activates EXPANSIN14 (EXP14) expression by binding to a specific region of the EXP14 promoter in the gene regulatory network controlling lateral root formation in Arabidopsis (Kim and Lee 2013; Lee et al. 2013). G-box binding factor interacting protein 1, an LBD18 transcription coactivator in the EXP14 promoter, was isolated using a two-hybrid system in yeast and Arabidopsis protoplasts (Lee et al. 2014). As the first identified member that regulates jasmonic acid (JA) signalling and plant–pathogen interactions, AtLBD20 is a novel negative regulator of a subset of JA-dependent responses and a susceptibility gene for Fusarium wilt that is highly expressed in roots (Thatcher et al. 2012). The Arabidopsis LBD gene DOWN IN DARK AND AUXIN1, formerly LBD25/ASL3, is involved in both auxin signalling and photomorphogenesis (Mangeon et al. 2011). In addition to being developmental regulators, LBD genes also function in metabolic regulation. AtLBD37, AtLBD38 and AtLBD39 serve as negative regulators of anthocyanin biosynthesis in Arabidopsis (Rubin et al. 2009).
The functions of some LBD proteins have been identified in different species such as Oryza sativa, Zea mays, poplar and apple (Inukai et al. 2005; Phelps-Durr et al. 2005; Vollbrecht et al. 2005; Bortiri et al. 2006; Yordanov et al. 2010; Majer et al. 2012; Majer et al. 2013a). CROWN ROOTLESS 1/ADVENTITIOUS ROOTLESS 1, a homologue of AtLBD29, is essential for crown root formation in rice and is regulated by auxin response factor (ARF) (Inukai et al. 2005; Liu et al. 2005). Rootless concerning crown and seminal roots encode an LBD protein that regulates shoot-borne root initiation in maize (Majer et al. 2012). RA2 and ZmLBD19 have been shown to regulate reproductive growth and female gametophyte development in maize, respectively (Theodoris et al. 2003; Phelps-Durr et al. 2005; Vollbrecht et al. 2005; Bortiri et al. 2006). PtaLBD1, a novel member of the LBD gene family in Populus, regulates secondary phloem development in poplar stems (Yordanov et al. 2010).
Grape (Vitis vinifera) is an important fruit crop and economically valuable model crop worldwide. Sequencing of the highly homozygous grapevine PN40024 genome has provided a great opportunity to analyse the grapevine genome and gene family evolution (Jaillon et al. 2007; Wang et al. 2014a, 2014b). Recent studies have conducted genomewide analyses of the MIKC gene family, respiratory burst oxidase homologue gene family, mitogen-activated protein kinase kinase kinase gene family, ARF gene family, WRKY gene family and subtilase gene family in grape (Díaz-Riquelme et al. 2009; Cheng et al. 2013; Cao et al. 2014; Wan et al. 2014; Wang et al. 2014a, 2014b). A total of 42, 35, 44, 57, 58 and 46 LBD genes have been identified in Arabidopsis, rice, maize, poplar, apple and tomato, respectively (Shuai et al. 2002; Yang et al. 2006; Zhu et al. 2007; Wang et al. 2013a, 2013b, Zhang et al. 2014). However, genomewide information on the grape LBD gene family is currently lacking.
In the current study, we identified 40 LBD genes in the grape genome. Considering the importance of LBD in diverse biological and physiological processes, and their potential application in the development of transgenic plants, we conducted a systematic analysis of the grape LBD family. On the basis of the structural features of the conserved motifs, the grape LBD genes were divided into classes I and II, corresponding to the presence of CX 2 CX 6 CX 3C and LX 6 LX 3 LX 6L motifs, respectively. The phylogenetic relationships, chromosomal locations and structures of these LBD genes were also studied. Expression profiles in microarray analysis demonstrated that 21 putative LBD genes in grape may be involved in developmental processes, including preveraison, veraison and ripening. Six LBD genes showed differential expression patterns among grape tissues, such as root, stem and leaf, while several genes may be involved in response to abiotic stress treatments. Overall, this study provides valuable information for future cloning and functional studies of LBD genes in grape.
Materials and methods
Identification of the LBD genes in grape
Three different approaches were used to identify the LBD proteins in the grape genome (Wang et al. 2013a; Zhang et al. 2014). First, all of the known Arabidopsis LBD gene sequences that were downloaded from the Arabidopsis genome TAIR 9.0 release (The Arabidopsis Information Resource, http://www.arabidopsis.org/) (Lamesch et al. 2012) were used as query sequences to perform multiple database searches against the proteome and genome files downloaded from the Phytozome database (http://www.phytozome.net/) (Goodstein et al. 2012). Stand-alone versions of BLASTP and TBLASTN (http://blast.ncbi.nlm.nih.gov/), which are available from the NCBI were used with an e-value cut-off set to 1e-003 (Altschul et al. 1990; Mount 2007). All of the protein sequences derived from the collected candidate LBD genes were examined using the domain analysis programs Pfam (protein family: http://pfam.sanger.ac.uk/) (Finn et al. 2014) and SMART (simple modular architecture research tool: http://smart.embl-heidelberg.de/) (Letunic et al. 2012) with the default cut-off parameters. Second, we analysed the domains of all the maize peptide sequences by using the hidden Markov model (Jeanmougin et al. 1998; Wu et al. 2002) analysis with Pfam searching. Subsequently, we obtained the sequences with the PF03195 Pfam number which contained a typical LBD domain from the grape genome sequences by using a Perl-based script. Third, all of the protein sequences were compared with known LBD sequences using ClustalX (http://www.clustal.org/) to verify whether or not the sequences are candidate LBDs (Jeanmougin et al. 1998).
The isoelectric points (PI) and molecular weights of the LBD proteins were obtained with the help of proteomics and sequence analysis tools on the ExPASy proteomics server (http://expasy.org/) (Artimo et al. 2012). The exon/intron information was obtained from the Phytozome database using a Perl-based program (Goodstein et al. 2012).
Chromosomal location and gene structure of the LBD genes in grape
To determine the locations of LBD genes on grape chromosomes, the chromosomal locations were retrieved from the genome data downloaded from the Phytozome database using a Perl-based program (Goodstein et al. 2012). The predicted grape LBD genes were mapped to the chromosomes by using a revised version of MapDraw tools (Liu and Meng 2003), and the gene structure of the LBD genes was generated using the gene structure display server (http://gsds.cbi.pku.edu.cn/) (Guo et al. 2007).
Sequence alignment and phylogenetic analysis of the LBD genes
The LBD sequences were aligned using ClustalX program with BLOSUM30 as the protein-weight matrix (Jeanmougin et al. 1998). Multiple sequence alignments were also performed using ClustalX (http://www.clustal.org/) and confirmed using multiple sequence comparison by log-expectation (ver. 3.52) (Chenna et al. 2003; Edgar 2004). To obtain clues about the evolutionary history of the LBD gene family in grape, phylogenetic trees were constructed using the neighbor-joining (NJ) method of MEGA 5.0 (http://www.megasoftware.net/) with p-distance and complete deletion option parameters (Tamura et al. 2011). The reliability of the obtained trees was tested using a bootstrapping method with 1000 replicates. Images of the phylogenetic trees were drawn using MEGA 5.0.
Microarray data analysis of LBD genes in grape
To understand the spatial and temporal expression patterns of LBD genes during the life cycle of grape, the expression profiles of the LBD genes were analysed on the basis of published high-throughput microarray data (Fasoli et al. 2012). Five developmental stages were included in the datasets: preveraison, veraison, grape ripening 1, grape ripening 2 and grape ripening 3.
Plant materials and growth conditions
Grape ecotype Guifei was obtained from the Institute of Vegetables and Flowers, Chinese Academy of Agricultural Sciences, Beijing, China. Seeds were stratified at 4 ∘C for two days and then transferred to a 16/8 h photoperiod (100 μmol m −2 s −1) at 25 ∘C in the growing chamber for four weeks. For the different stresses, seedlings with six growing leaves were transferred to a blotting paper without hormone treatment or treated with 200 mM mannitol, 300 mM NaCl, heat stress (39 ∘C) or low temperature (4 ∘C) for 6 h. Each treated sample had a corresponding regularly watered control. Three independent biological replications (three independent plants) were sampled for each point. For RNA extraction, whole plants were frozen, stored in liquid nitrogen immediately after harvest, and then stored at –80 ∘C until used.
RNA extraction
For RNA isolation, the plant tissues were separately harvested, frozen in liquid nitrogen and then stored at −80 ∘C until used. Total RNA was isolated from different grape tissues and seedlings with TRIzol reagent (Invitrogen, Carlsbad, USA).
Quantitative real-time polymerase chain reaction (qRT-PCR) analysis
Total RNAs were extracted from the leaves of grape seedlings with different treatments by using TRIzol reagent in accordance with the manufacturer’s instructions. Contaminated DNA was removed using RNase-free DNase I. Firststrand cDNAs were synthesized using FirstStrand cDNA Synthesis kit (Fermentas, USA). qRT-PCR reactions were performed in the MyiQ TM RT-PCR detection system (Biorad, USA) using the TransStart Top Green qPCR SuperMix (TransGen, China) in accordance with the manufacturer’s instructions. Each PCR reaction (25 μL) contained 12.5 μL of 2 ×real-time PCR Mix (containing SYBR Green I), 0.5 μL of each primer, and appropriately diluted cDNA. The thermal cycling conditions were 95 ∘C for 30 s, followed by 48 cycles of 95 ∘C for 20 s, 53–57 ∘C for 30 s and 72 ∘C for 30 s. The qRT-PCR experiment was carried out at least thrice under identical conditions using actin as an internal control. Details of the primers used are listed in table ?? in electronic supplementary material at http://www.ias.ac.in/jgenet/.
Results
Identification of LBD genes in grape
We used a local BLAST program, Hidden Markov Model of the SMART and Pfam tools to identify members of the LBD family in grape. A total of 40 transcripts in the grape genome sequence were identified as possible members of the LBD family. The 40 LBD genes were subsequently renamed from VvLBD1 to VvLBD40 on the basis of their locations in chr1– chr19 (table 1). As shown in table 1, the open reading frame (ORF) was 381 (VvLBD15) to 1158 bp (VvLBD02) long, and the average length was 633 bp. All identified LBD genes encode proteins with lengths varying from 126 (VvLBD15) to 385 (VvLBD02) amino acids, but some of the encoded proteins were exceptionally long or short. The predicted PI of the proteins ranged from 4.77 (VvLBD04) to 9.28 (VvLBD01). The predicted LBD genes were further analysed using a local BLAST alignment program to identify the relevant homologues in Arabidopsis (table 1). Detailed information on the grape LBD gene family is shown in table 2 in electronic supplementary material.
Chromosomal location of LBD genes in the grape genome
We analysed the chromosomal location of the 40 LBD genes by using MapDraw to determine the genomic distribution of the LBD genes. Thirty-seven of the 40 LBD genes were mapped on the grape chromosomes and were subsequently named VvLBD01–VvLBD37 on the basis of their locations in chr1–chr19; the three genes that were not conclusively mapped to any particular chromosome were named VvLBD38–VvLBD40 (figure 1). Chr13 had the highest number of VvLBDs (11 out of 40 members or 27.5%), but no gene was found in chr2, chr4, chr5, chr9, chr10, chr11 and chr12. Relatively high gene distribution densities were observed in chr6, chr7 and chr15, whereas three LBD genes or less were located in chr1, chr3, chr8, chr14, chr16, chr17 and chr18. The grape LBD genes were distributed across the majority of 19 chromosomes with different densities (figure 1). Four duplicated pairs of LBD genes were identified in the grape genome and investigated. Interestingly, two pairs of VvLBDs (VvLBD04/VvLBD05 and VvLBD30/VvLBD31) were located in the same chromosome, while another two sister pairs (linked by red line) were located on different chromosomes (chr6 and chr13). These results suggest that segmental duplication and transposition events are involved in the evolution of the LBD gene family in grape.
Phylogenetic and gene structure analysis of the LBD genes in grape
We performed phylogenetic analysis based on the full-length amino acid sequences of the LBD genes in grape to evaluate their evolutionary relationships. A phylogenetic tree was generated using the NJ method in MEGA 5.0, and an alignment of full-length LBD proteins was constructed using ClustalX (Thompson et al. 1997; Tamura et al. 2011). On the basis of the phylogenetic analysis, the grape LBD proteins were divided into two monophyletic subfamilies (classes I and II) (figure 2, right). Class I included 33 LBD proteins, whereas class II included only seven (figure 2, right). Eight sister pairs of paralogous LBDs (marked by red shadow) had well-supported bootstrap values (>97%). Gene duplication events are considered the most important drivers of evolution of gene families in higher plants (Cannon et al. 2004). Our results suggest a clear paralogous pattern of LBD gene divergence through gene duplication in grape. We speculate that paralogous genes may have similar functions in grape, as in Arabidopsis and apple (Shuai et al. 2002; Wang et al. 2013a). In addition, we performed exon–intron analysis of the grape LBD genes using the gene structure display server (GSDS, http://gsds.cbi.pku.edu.cn/) to generate gene structure diagrams (figure 2, left). Most members in the same subfamilies had similar exon–intron structures. The number of exons in VvLBDs ranged from 1 to 4. Most VvLBD genes shared two exons, two genes (VvLBD13 and VvLBD03) had three exons, and three genes did not contain any introns (VvLBD02, VvLBD34 and VvLBD39). These results suggest that the close evolutionary relationships of the LBD genes are reliable.
Phylogenetic tree and gene structure analysis of the LBD gene family in grape. The amino acid sequences of the LBD proteins were aligned with ClustalX and a phylogenetic tree was constructed using the NJ method in MEGA 5.0. Each node is represented by a number that indicates the bootstrap value for 1000 replicates. The scale bar represents 0.1 substitution per sequence position (left). The right side illustrates the exon–intron organization of the corresponding LBD genes. The exons CDS, UTR and intron are represented by the green boxes, blue boxes and black lines, respectively. The scale bar represents 1 kb (right).
Phylogenetic relationships of LBD genes in grape and Arabidopsis
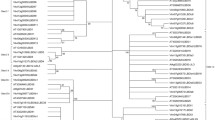
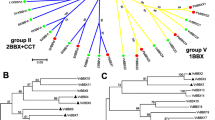
We generated an unrooted NJ phylogenetic tree by using the full-length amino acid sequences of 40 and 42 LBDs from grape and Arabidopsis, respectively (figure 3). The LBD proteins were also divided into two classes (I and II) based on previous studies. Classes I and II were further divided into four (classes Ia–Id) and two (classes IIa and IIb) subgroups, respectively. A total of 33 and seven separate genes in grape and 36 and six genes in Arabidopsis were observed in classes I and II, respectively (figure 3). Similarly, classes Ia, Ib, Ic and Id contained 2, 6, 18 and 7 VvLBD members, respectively, in grape and 8, 10, 9 and 9 members, respectively, in Arabidopsis. However, class IIa and IIb subgroups had similar numbers of LBD genes, namely, 4 and 3 in grape, and 3 and 3 in Arabidopsis (figure 3). These results indicate that class II VvLBDs has similar functions as AtLBDs.
Phylogenetic tree of the LBD proteins in Arabidopsis and grape. The amino acid sequences of the LBD proteins were aligned with ClustalX, and a phylogenetic tree was constructed using the NJ method in MEGA 5.0. Each node is represented by a number that indicates the bootstrap value for 1000 replicates. The subgroups are marked by colourful background.
Sequence alignment and conserved motifs of VvLBD genes
We performed sequence alignments of the 40 grape LBD proteins and generated a WebLogo to identify the sequence characteristics of their conserved motifs (Crooks et al. 2004). The results confirmed that the LBD motif is highly conserved in the LBD gene family (figure 4). The WebLogo indicated that four Cys amino acids were completely conserved among all the CCCC motifs and that more than 70% of the LBD motifs contained alanine (A), lysine (K), leucine (L) and arginine (R) (figure 4a).
Sequence logos for the conserved motifs of all LBD proteins in grape. The numbers on the x-axis represent the sequence positions in zinc finger-like motifs. The numbers on the y-axis represent the information content measured in bits. (a) CX 2 CX 6 CX 3C zinc finger-like domain sequence logos. (b) LX 6 LX 3 LX 6L leucine zipper-like domain sequences logos. Sequence alignment of two domains by ClustalX and conserved motif logos was performed using WebLogo.
Expression profiles of the VvLBD genes during different growth processes from the microarray database
We searched the published literature for relevant grape probes and microarray data to investigate the potential functions of VvLBD genes in grape development. We then analysed the expression patterns of VvLBD genes under normal growth conditions during different developmental phases, including preveraison, veraison and grape ripening (figure 5). Of the 40 predicted genes, 21 were detected in the microarray analysis, whereas the remaining 19 were not. These results indicate that these 21VvLBDs play major roles at different developmental stages.
Expression profiles of six selected VvLBD genes in grape using qRT-PCR
We used qRT-PCR to validate the expression patterns of six genes in different tissues, including root, stem and leaf (4-week old). VvLBD01, VvLBD02 and VvLBD04 exhibited high expression levels in stems (figure 6; table 3 in electronic supplementary material), whereas VvLBD10, VvLBD08 and VvLBD18 showed high expression levels in roots and leaves, respectively. These results suggest that the six VvLBD genes differentially expressed in the three detected tissues and play key roles in grape tissue development.
To determine whether LBD genes are involved in responses to mannitol, NaCl, heat stress and low temperature treatments, we performed qRT-PCR analysis using total RNA extracted from 4-week-old grape seedlings. VvLBD04 and VvLBD18 were significantly upregulated under mannitol and heat stress treatments but obviously downregulated under NaCl and low temperature treatments. By contrast, VvLBD08 was highly expressed under NaCl and low temperature treatments. VvLBD02 was significantly upregulated after heat stress treatment but showed almost no expression under mannitol treatment. The expression level of VvLBD01 significantly increased under mannitol, NaCl and heat stress treatments but markedly decreased under low temperature treatment. VvLBD10 displayed no evident changes (figure 7; table 3 in elecroinc supplementary material). These findings suggest that these LBD genes are involved in abiotic stress responses during seedling development in grape.
qRT-PCR analysis of six genes under different treatments; CK, contrast check; Man, mannitol; HT, high temperature (heat stress); LT, low temperature. Gene expression was normalized to the control unstressed expression level, which was assigned with a value of 1. Data represent the average of three independent experiments ± SD. Standard errors are shown as bars above the columns.
Discussion
Characterization of grape LBD gene family
Recent research has shown that LBD genes belong to a new plant-specific transcription factor family (Iwakawa et al. 2002; Shuai et al. 2002; Yang et al. 2006; Zhu et al. 2007; Lee et al. 2013, 2014). In the past decade, LBD genes have attracted considerable attention because of their roles in diverse plant processes. Several LBD genes have been identified and functionally characterized in the growth and development of plants, particularly Arabidopsis, rice and maize (Phelps-Durr et al. 2005; Bortiri et al. 2006; Okushima et al. 2007; Kim and Kim 2012; Thatcher et al. 2012; Cabrera et al. 2014). Many LBD genes have been intensively studied for their functions in important plant biological processes (Okushima et al. 2005; Phelps-Durr et al. 2005; Vollbrecht et al. 2005; Bortiri et al. 2006; Jaillon et al. 2007; Naito et al. 2007; Okushima et al. 2007; Rubin et al. 2009; Yordanov et al. 2010; Mangeon et al. 2011; Fan et al. 2012; Feng et al. 2012; Kim and Kim 2012; Thatcher et al. 2012; Kim and Lee 2013; Lee et al. 2013). However, to the best of our knowledge, only a few LBD genes have been identified in grape, and genomewide information on grape LBD genes is currently lacking. Therefore, a comprehensive investigation is necessary to unravel the biological functions of grape LBD genes.
The current study is the first to identify 40 VvLBD genes in the grape genome through genomewide analysis. The number of LBD genes in grape is similar to that reported in other plant species, i.e. 42 in Arabidopsis, 35 in rice, 44 in maize, 57 in poplar, 58 in apple and 46 in tomato (Shuai et al. 2002; Zhu et al. 2007; Wang et al. 2013a, 2013b; Zhang et al. 2014). This finding indicates that the LBD gene family is highly conserved in plants.
The LBD genes account for ∼0.131% of the predicted protein coding genes in grape. This percentage is similar to that in tomato (∼0.131%), higher than those in rice, poplar and apple (∼0.0933, 0.127 and 0.101%, respectively), slightly lower than that in maize (∼0.138%), and considerably lower than that in Arabidopsis (∼0.165%) (table 4 in electronic supplementary material). Thus, the total number of LBD genes is higher in dicots than in the monocots. Moreover, a phylogenetic tree was constructed by aligning the full-length protein sequences of the LBD genes in grape and Arabidopsis. These genes clustered into class I (Ia–Id) and class II (IIa and IIb), which is in agreement with the previous studies (Shuai et al. 2002; Zhu et al. 2007; Wang et al. 2013a, 2013b; Zhang et al. 2014). Chromosomal location analyses showed that 37 VvLBD genes were located in 12 chromosomes at different densities, but no LBD gene was found in one-third of grape chromosomes (figure 1). Results of sequence alignment showed that the LBD motif is highly conserved in the LBD gene family despite some discrepancies (figure 4).
Phylogenetic analysis and evolution of LBD proteins in grape and Arabidopsis and functional prediction of VvLBD genes
Several LBD genes play important roles in the regulation of development processes, specifically in leaf and lateral root development (Chalfun-Junior et al. 2005; Okushima et al. 2005, 2007; Fan et al. 2012; Feng et al. 2012; Kim and Lee 2013; Lee et al. 2013; Cabrera et al. 2014). Previous functional analyses of other LBD proteins revealed diverse functions of LBD genes which may provide important insights for future research of VvLBD gene functions in grape. AtLBD1 is a nuclear protein that as a potential transcription factor, may be involved in sustaining the indeterminate cell fate of shoot apical meristems (SAMs) in Arabidopsis (Sun et al. 2010). VvLBD04, VvLBD05, VvLBD18, VvLBD19, VvLBD23, VvLBD24 and VvLBD30 may have similar functions in SAM development processes because of their high homology with AtLBD1 (table 1 and figure 3). VvLBD08, VvLBD09, VvLBD28 and VvLBD26, which are homologues of AtLBD29, AtLBD16, AtLBD18 and AtLBD25, respectively, share similar functions in lateral root initiation (Okushima et al. 2005, 2007; Mangeon et al. 2011; Fan et al. 2012; Feng et al. 2012; Kim and Lee 2013; Lee et al. 2013; Cabrera et al. 2014) AtLBD20 plays a role in JA signalling in response to Fusarium wilt, whereas AtLBD37, AtLBD38 and AtLBD39 are induced by nitrate and involved in anthocyanin synthesis and nitrate metabolism in Arabidopsis (Rubin et al. 2009; Thatcher et al. 2012). Hence, VvLBD03, VvLBD12, VvLBD13 and VvLBD36 with high homology to the above four AtLBD genes may be involved in hormone or nutrient metabolism. In addition, VvLBD16 may be involved in SAM regulation owing to the functions of ASL11/LBD15 that affect cellular differentiation in SAM and regulate WUS expression (Sun et al. 2013).
Expression patterns of VvLBD genes at different grape developmental stages
We utilized Genevestigator analysis to determine the expression profiles of LBD genes at different developmental stages. Under normal growth conditions, 21 of 40 LBD genes in grape (more than 50%) were regulated in at least one of the five developmental stages tested (figure 5). During fleshy fruit ripening, genes with a wide range of biological functions are recruited and their expression is coordinately modulated in a phase-specific manner (Gapper et al. 2013; Gouthu et al. 2014). In several plants, TF families play important roles in fruit development. Therefore, identifying other TF that function in the regulation of fruit ripening is highly important. In addition, expressions of about half of the predicted LBD genes were found to be regulated to different degrees during fruit ripening in grape (figure 5). Based on these results, we speculate that some VvLBD genes regulate fruit development in general, while others participate in specific developmental stages of fruit ripening. However, further research is needed to determine the functions of the key LBD genes in grape.
Expression level of VvLBD genes under different abiotic stresses
In recent years, increasing evidence has indicated that LBD proteins are involved in the development of boundary organs, regulation of nitrogen metabolism and biosynthesis of anthocyanins in Arabidopsis (Okushima et al. 2007; Rubin et al. 2009; Mangeon et al. 2011; Fan et al. 2012; Feng et al. 2012; Kim and Kim 2012; Thatcher et al. 2012; Kim and Lee 2013). However, limited studies have focussed on the roles of LBD genes in response to abiotic stresses. In the present study, qRT-PCR results showed that VvLBD01, VvLBD02, VvLBD04, VvLBD08, VvLBD10 and VvLBD18 were detected in all tissues tested, and several genes may be involved in response to abiotic stresses (figure 7). AtLBD17 and AtLBD29 were upregulated by auxin treatment (Feng et al. 2012), whereas VvLBD08 was upregulated by heat stress treatment and NaCl, respectively. Considering its high homology with AtLBD17 and AtLBD29, VvLBD08 may also exhibit a similar response to auxin treatment (figures 3 and 7). Moreover, VvLBD02, which is a homologue of AtLBD40 andAtLBD41 might be involved in leaf dorsoventral patterning and gibberellin signal transduction, respectively (Chalfun-Junior et al. 2005; Zentella et al. 2007). These data serve as a useful reference for future studies on the involvement of several grape LBD genes in responses to particular abiotic stresses. Taken together, our results indicate that LBD genes play important roles in grape growth and development in response to abiotic stresses.
Conclusions
The LBD gene family has been implicated in many developmental processes in plants. Understanding their regulatory roles in plant growth and development is highly important. In this study, 40 LBD genes were identified in grape and a comprehensive analysis of this gene family was performed. A total of 37 LBD genes were localized to 12 out of 19 chromosomes (i.e. excluding chr2, chr4, chr5, chr9, chr10, chr11 and chr12). These genes were divided into classes I and II, and further subdivided into classes Ia–Id and classes IIa and IIb, respectively. The relationship of 40 grape LBDs with their Arabidopsis counterparts was also observed using an unrooted NJ phylogenetic tree. Most of the VvLBD members within the same clade shared a similar intron/exon structure and gene length. Expression profile analysis showed that 21 VvLBD genes may be involved in controlling different developmental stages. Finally, six LBD genes showed differential expression patterns among three representative grape tissues and these genes were mostly involved in response to some abiotic stresses. The results of this study may serve as a basis in selecting candidate genes related to grape development and in further elucidating the functional roles of these VvLBD genes.
References
Altschul S. F., Gish W., Miller W., Myers E. W. and Lipman D. J. 1990 Basic local alignment search tool. J. Mol. Biol. 215, 403–410.
Artimo P., Jonnalagedda M., Arnold K., Baratin D., Csardi G, de Castro E. et al. 2012 ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res. 40, W597–W603.
Bortiri E., Chuck G., Vollbrecht E., Rocheford T., Martienssen R. and Hake S. 2006 ramosa2 encodes a LATERAL ORGAN BOUNDARY domain protein that determines the fate of stem cells in branch meristems of maize. Plant Cell 18, 574–585.
Cabrera J., Díaz-Manzano F. E., Sanchez M., Rosso M. N., Melillo T., Goh T. et al. 2014 A role for LATERAL ORGAN BOUNDARIES-DOMAIN 16 during the interaction of ArabidopsisMeloidogyne spp. provides a molecular link between lateral root and root-knot nematode feeding site development. New Phytol. 203, 632–645.
Cannon S. B., Mitra A., Baumgarten A., Young N. D. and May G. 2004 Te roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 4, 10.
Cao J., Han X., Zhang T. C., Yang Y. P., Huang J. L. and Hu X. Y. 2014 Genome-wide and molecular evolution analysis of the subtilase gene family in Vitis vinifera. BMC Genomics 15, 1116–1134.
Chalfun-Junior A., Franken J., Mes J. J., Marsch-Martinez N., Pereira A. and Angenent G. C. 2005 ASYMMETRIC LEAVES2-LIKE1 gene, a member of the AS2/LOB family, controls proximal-distal patterning in Arabidopsis petals. Plant Mol. Biol. 57, 559–575.
Cheng C. X., Xu X. Z., Gao M., Li J., Guo C. L., Song J. Y. et al. 2013 Genome-wide analysis of respiratory burst oxidase homologs in grape (Vitis vinifera L.). Int. J. Mol. Sci. 14, 24169–24186.
Chenna R., Sugawara H., Koike T., Lopez R., Gibson T. J., Higgins D. G. et al. 2003 Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Res. 31, 3497–3500.
Crooks G. E., Hon G., Chandonia J. M. and Brenner S. E. 2004 WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190.
Díaz-Riquelme J., Lijavetzky D., Martínez-Zapater J. M. and Carmona M. J. 2009 Genome-wide analysis of MIKC Ctype MADS box genes in grapevine. Plant Physiol. 149, 354–369.
Edgar R. C. 2004 MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797.
Fan M., Xu C., Xu K. and Hu Y. 2012 LATERAL ORGAN BOUNDARIES DOMAIN transcription factors direct callus formation in Arabidopsis regeneration. Cell Res. 22, 1169– 1180.
Fasoli M., Dal Santo S., Zenoni S., Tornielli G. B., Farina L., Zamboni A. et al. 2012 The grapevine expression atlas reveals a deep transcriptome shift driving the entire plant into a maturation program. Plant Cell 24, 3489–3505.
Feng Z., Zhu J., Du X. and Cui X. 2012 Effects of three auxin-inducible LBD members on lateral root formation in Arabidopsis thaliana. Planta 236, 1227–1237.
Finn R. D., Bateman A., Clements J., Coggill P., Eberhardt R. Y., Eddy S. R. et al. 2014 Pfam: the protein families database. Nucleic Acids Res. 42, D222–D230.
Gapper N. E., McQuinn R. P. and Giovannoni J. J. 2013 Molecular and genetic regulation of fruit ripening. Plant Mol. Biol. 82, 575–591.
Goodstein D. M., Shu S., Howson R., Neupane R., Hayes R. D., Fazo J. et al. 2012 Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res. 40, D1178–D1186.
Gouthu S., O’Neil S. T., Di Y. M., Ansarolia M., Megraw M. and Deluc L. G. 2014 A comparative study of ripening among berries of the grape cluster reveals an altered transcriptional programme and enhanced ripening rate in delayed berries. J. Exp. Bot. 65, 5889–5902.
Guo A. Y., Zhu Q. H., Chen X. and Luo J. C. 2007 GSDS: a gene structure display server. Yi Chuan 29, 1023–1026.
Husbands A., Bell E. M., Shuai B., Smith H. M. and Springer P. S. 2007 LATERAL ORGAN BOUNDARIES defines a new family of DNA-binding transcription factors and can interact with specific bHLH proteins. Nucleic Acids Res. 35, 6663–6671.
Inukai Y., Sakamoto T., Ueguchi-Tanaka M., Shibata Y., Gomi K., Umemura I. et al. 2005 Crown rootless1, which is essential for crown root formation in rice, is a target of an AUXIN RESPONSE FACTOR in auxin signaling. Plant Cell 17, 1387–1396.
Iwakawa H., Ueno Y., Semiarti E., Onouchi H., Kojima S., Tsukaya H. et al. 2002 The ASYMMETRIC LEAVES2 gene of Arabidopsis thaliana, required for formation of a symmetric flat leaf lamina, encodes a member of a novel family of proteins characterized by cysteine repeats and a leucine zipper. Plant Cell Physiol. 43, 467–478.
Jaillon O., Aury J. M., Noel B., Policriti A., Clepet C., Casagrande A. et al. 2007 French–Italian Public Consortium for Grapevine Genome Characterization. The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449, U463–U467.
Jeanmougin F., Thompson J. D., Gouy M., Higgins D. G. and Gibson T. J. 1998 Multiple sequence alignment with Clustal X. Trends Biochem. Sci. 23, 403–405.
Kim J. and Lee H. W. 2013 Direct activation of EXPANSIN14 by LBD18 in the gene regulatory network of lateral root formation in Arabidopsis. Plant Signal. Behav. 8, e22979.
Kim M. J. and Kim J. 2012 Identification of nuclear localization signal in ASYMMETRIC LEAVES2-LIKE18/LATERAL ORGAN BOUNDARIES DOMAIN16 (ASL18/LBD16) from Arabidopsis. J. Plant Physiol. 169, 1221–1226.
Lamesch P., Berardini T. Z., Li D., Swarbreck D., Wilks C., Sasidharan R. et al. 2012 The Arabidopsis Information Resource (TAIR): improved gene annotation and new tools. Nucleic Acids Res. 40, D1202–D1210.
Lee H. W., Kim M. J., Kim N. Y., Lee S. H. and Kim J. 2013 LBD18 acts as a transcriptional activator that directly binds to the EXPANSIN14 promoter in promoting lateral root emergence of Arabidopsis. Plant J. 73, 212–224.
Lee H. W., Park J. H., Park M. Y. and Kim J. 2014 GIP1 may act as a coactivator that enhances transcriptional activity of LBD18 in Arabidopsis. J. Plant Physiol. 171, 14–18.
Letunic I., Doerks T. and Bork P. 2012 SMART 7: recent updates to the protein domain annotation resource. Nucleic Acids Res. 40, D302–D305.
Liu H. J., Wang S. F., Yu X. B., Yu J., He X. W., Zhang S. L. et al. 2005 ARL1, a LOB-domain protein required for adventitious root formation in rice. Plant J. 43, 47–56.
Liu R. H. and Meng J. L. 2003 MapDraw: a microsoft excel macro for drawing genetic linkage maps based on given genetic linkage data. Yi Chuan 25, 317–321.
Majer C. and Hochholdinger F. 2011 Defining the boundaries: structure and function of LOB domain proteins. Trends Plant Sci. 16, 47–52.
Majer C., Xu C., Berendzen K. W. and Hochholdinger F. 2012 Molecular interactions of ROOTLESS CONCERNING CROWN AND SEMINAL ROOTS, a LOB domain protein regulating shoot-borne root initiation in maize (Zea mays L.). Phil. Trans. R. Soc. B 367, 1542–1551.
Mangeon A., Bell E. M., Lin W. C., Jablonska B. and Springer P. S. 2011 Misregulation of the LOB domain gene DDA1 suggests possible functions in auxin signalling and photomorphogenesis. J. Exp. Bot. 62, 221–233.
Mount D. W. 2007 Using the basic local alignment search tool (BLAST). CSH Protoc pdb.top17 (doi:10.1101/pdb.top17).
Naito T., Yamashino T., Kiba T., Koizumi N., Kojima M., Sakakibara H. et al. 2007 A link between cytokinin and ASL9 (ASYMMETRIC LEAVES 2 LIKE 9) that belongs to the AS2/LOB (LATERAL ORGAN BOUNDARIES) family genes in Arabidopsis thaliana. Biosci. Biotechnol. Biochem. 71, 1269– 1278.
Okushima Y., Fukaki H., Onoda M., Theologis A. and Tasaka M. 2007 ARF7 and ARF19 regulate lateral root formation via direct activation of LBD/ASL genes in Arabidopsis. Plant Cell 19, 118–130.
Okushima Y., Overvoorde P. J., Arima K., Alonso J. M., Chan A., Chang C. et al. 2005 Functional genomic analysis of the AUXIN RESPONSE FACTOR gene family members in Arabidopsis thaliana: unique and overlapping functions of ARF7 and ARF19. Plant Cell 17, 444–463.
Phelps-Durr T. L., Thomas J., Vahab P. and Timmermans M. C. 2005 Maize rough sheath2 and its Arabidopsis orthologue ASYMMETRIC LEAVES1 interact with HIRA, a predicted histone chaperone, to maintain knox gene silencing and determinacy during organogenesis. Plant Cell 17, 2886–2898.
Rubin G., Tohge T., Matsuda F., Saito K. and Scheible W. R. 2009 Members of the LBD family of transcription factors repress anthocyanin synthesis and affect additional nitrogen responses in Arabidopsis. Plant Cell 21, 3567–3584.
Shuai B., Reynaga-Peña C. G. and Springer P. S. 2002 The lateral organ boundaries gene defines a novel, plant-specific gene family. Plant Physiol. 129, 747–761.
Sun S. B., Meng L. S., Sun X. D. and Feng Z. H. 2010 Using high competent shoot apical meristems of cockscomb as explants for studying function of ASYMMETRIC LEAVES2-LIKE11 (ASL11) gene of Arabidopsis. Mol. Biol. Rep. 37, 973–982.
Sun X. D., Feng Z. H., Meng L. S., Zhu J. and Geitmann A. 2013 Arabidopsis ASL11/LBD15 is involved in shoot apical meristem development and regulates WUS expression. Planta 237, 1367–1378.
Tamura K., Peterson D., Peterson N., Stecher G., Nei M. and Kumar S. 2011 MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739.
Thatcher L. F., Powell J. J., Aitken E. A., Kazan K. and Manners J. M. 2012 The lateral organ boundaries domain transcription factor LBD20 functions in Fusarium wilt susceptibility and jasmonate signaling in Arabidopsis. Plant Physiol. 160, 407–418.
Theodoris G., Inada N. and Freeling M. 2003 Conservation and molecular dissection of ROUGH SHEATH2 and ASYMMETRIC LEAVES1 function in leaf development. Proc. Natl. Acad. Sci. USA 100, 6837–6842.
Thompson J. D., Gibson T. J., Plewniak F., Jeanmougin F. and Higgins D. G. 1997 The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 25, 4876–4882.
Vollbrecht E., Springer P. S., Goh L., Buckler 4th E. S. and Martienssen R. 2005 Architecture of floral branch systems in maize and related grasses. Nature 436, 1119–1126.
Wan S. B., Li W. L., Zhu Y. Y., Liu Z. M., Huang W. D. and Zhan J. C. 2014 Genome-wide identification, characterization and expression analysis of the auxin response factor gene family in Vitis vinifera. Plant Cell Rep. 33, 1365–1375.
Wang G., Lovato A., Polverari A. and Ma L. Y. H. 2014a Genome-wide identification and analysis of mitogen activated protein kinase kinase kinase gene family in grapevine (Vitis vinifera). BMC Plant Biol. 14, 219–237.
Wang L. N., Zhu W., Fang L. C., Sun X. M., Su L. Y., Liang Z. C. et al. 2014b Genome-wide identification of WRKY family genes and their response to cold stress in Vitis vinifera. BMC Plant Biol. 14, 103–116.
Wang X. F., Zhang S. Z., Su L., Liu X. and Hao Y. J. 2013a A genome-wide analysis of the LBD (LATERAL ORGAN BOUNDARIES Domain) gene family in Malus domestica with a functional characterization of MdLBD11. PLoS One 8, e57044.
Wang X. F., Liu X., Su L., Sun Y. J., Zhang S. Z., Hao Y. J. et al. 2013b Identification, evolution and expression analysis of the LBD gene family in tomato. Sci. Agric. Sin. 46, 2501–2513.
Wu X., Song C., Wang B. and Cheng J. 2002 Hidden Markov model used in protein sequence analysis. Sheng Wu Yi Xue Gong Cheng Xue Za Zhi 19, 455–458.
Yang Y., Yu X. B. and Wu P. 2006 Comparison and evolution analysis of two rice subspecies LATERAL ORGAN BOUNDARIES domain gene family and their evolutionary characterization from Arabidopsis. Mol. Phylogenet. Evol. 39, 248–262.
Yordanov Y. S., Regan S and Busov V. 2010 Members of the LATERAL ORGAN BOUNDARIES DOMAIN transcription factor family are involved in the regulation of secondary growth in Populus. Plant Cell 22, 3662–3677.
Zentella R., Zhang Z. L., Park M., Thomas S. G., Endo A., Murase K. et al. 2007 Global analysis of della direct targets in early gibberellin signaling in Arabidopsis. Plant Cell 19, 3037–3057.
Zhang Y. M., Zhang S. Z. and Zheng C. C. 2014 Genomewide analysis of LATERAL ORGAN BOUNDARIES Domain gene family in Zea mays. J. Genet. 93, 79–91.
Zhu Q. H., Guo A. Y., Gao G., Zhong Y. F., Xu M., Huang M. et al. 2007 DPTF: a database of poplar transcription factors. Bioinformatics 23, 1307–1308.
Acknowledgements
This work was supported by the National Natural Science Foundation (grant no. 31400225), Shandong Province Natural Science Foundation (grant no. ZR2011CM032), and the Shandong Province Young and Middle-aged Scientists Research Awards Fund (grant no. BS2014SW014) in China.
Author information
Authors and Affiliations
Corresponding authors
Additional information
[Cao H., Liu C.-Y., Liu C.-X., Zhao Y.-L. and Xu R.-R. 2016 Genomewide analysis of the lateral organ boundaries domain gene family in Vitis vinifera. J. Genet. 95, xx–xx]
RX led and coordinated the project and carried out the bioinformatics analyses, CL conducted the RNA extraction and qRT-PCR experiments, RX and YZ wrote the manuscript, HC revised the manuscript. All authors read and agreed with the final manuscript.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
CAO, H., LIU, CY., LIU, CX. et al. Genomewide analysis of the lateral organ boundaries domain gene family in Vitis vinifera . J Genet 95, 515–526 (2016). https://doi.org/10.1007/s12041-016-0660-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12041-016-0660-z