Abstract
The human brain and its temporal behavior correlated with development, structure, and function is a complex natural system even for its own kind. Coding and automation are necessary for modeling, analyzing and understanding the 86.1 ± 8.1 billion neurons, an almost equal number of non-neuronal glial cells, and the neuronal networks of the human brain comprising about 100 trillion connections. ‘Computational neuroscience’ which is heavily dependent on biology, physics, mathematics and computation addresses such problems while the archival, retrieval and merging of the huge amount of generated data in the form of clinical records, scientific literature, and specialized databases are carried out by ‘neuroinformatics’ approaches. Neuroinformatics is thus an interface between computer science and experimental neuroscience. This article provides an introduction to computational neuroscience and neuroinformatics fields along with their state-of-the-art tools, software, and resources. Furthermore, it describes a few innovative applications of these fields in predicting and detecting brain network organization, complex brain disorder diagnosis, large-scale 3D simulation of the brain, brain–computer, and brain-to-brain interfaces. It provides an integrated overview of the fields in a non-technical way, appropriate for broad general readership. Moreover, the article is an updated unified resource of the existing knowledge and sources for researchers stepping into these fields.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
The modern neuroscience discipline academically began with the ‘Neurosciences Research Program (NRP)’ in 1962 (Adelman 2010) at MIT. The neurological disorders of human brain constitute 13% of the global disease set. It is a lot more than the cardiovascular diseases which amount to 5% of the global disease set. Different types of cancers constitute 10% of the global disease set (Kiernan 2015). Present day neuroscience researchers are a combination of physiologists, theoretical and experimental physicists, mathematicians, computer scientists, engineers, molecular biologists, doctors, clinicians, bioinformaticians, psychologists, and philosophers among others. Such a diverse mix of researchers gives a glimpse of the rich vividness of this field (figure 1).
The nervous system computes and processes information (Piccinini and Shagrir 2014) very fast. A precise and powerful mathematical theory with different functions and relations among different positions of a brain is needed for computing the activities of the nervous system. However, the detailed procedures and activities of nervous systems, as well as their underlying reasons cannot be reflected by mathematical theories alone. We need hypothetical physical systems for computing them (Copeland and Shagrir 2017; Piccinini 2011). Computation of neuronal signal is neither digital nor analog; rather it is a different kind of computation (Piccinini and Bahar 2013). There are several levels of organization in the nervous system, which can be decomposed into a number of subsystems, like cortex and brainstem (Craver 2007; Bechtel 2008). The subsystems can also be decomposed into smaller systems. These objectives have led to the formation of a distinct branch of neuroscience known as ‘computational neuroscience’. Eric L Schwartz first introduced the term in 1985. The first part of the review focuses on the role of computational neuroscience in research of brain network organization along with the description of the state-of-the-art tools, software, and databases.
Computational neuroscience research generates highly complex data in large volumes. Their proper storage, accurate knowledge extraction, and quick dispersion are a big challenge. Hence, a computer-based collation, management, and analysis of neurobiological data is a required step towards understanding several areas of neuroscience that gave rise to a new branch namely, ‘neuroinformatics’ (Young and Scannell 2000). Neuroinformatics research includes the development of neuroscience data storage infrastructure, advanced tools for data extraction and dispersion, and approaches for analyzing such data which may facilitate major advancements in understanding the structural and functional aspects of the human brain (Beltrame et al. 2000). The second part of the review describes scopes and challenges of neuroinformatics research along with a few existing languages, data sources, tools and software, simulation platforms, a few innovative applications.
2 Computational neuroscience
By an old definition (1993), ‘The expression Computational Neuroscience reflects the possibilities of generating theories of brain function in terms of the information-processing properties of structures that make up nervous systems (Schwartz 1993).’ A more recent brief definition (2010) is ‘computational neuroscience is the theoretical study of the brain used to uncover the principles and mechanisms that guide the development, organization, information-processing and mental abilities of the nervous system (Trappenberg 2009).’ In the present day scenario, computational neuroscience can also be defined as the study of brain circuits/networks to explore how the brain processes various activities according to specific information and properties of structural and functional activities with the help of computational power.
The human brain is composed of 86.1 ± 8.1 billion neurons (Azevedo et al. 2009) and almost an equal number of non-neuronal glial cells; the neurons being interconnected by about a 100 trillion inhibitory and excitatory synapses as well as above-threshold and sub-threshold synapses, which in turn constitute large neuronal networks (Goldental et al. 2014). The main motivation of computational neuroscience since the last four decades has been to explore the process of representation and manipulation of information in the brain using electrical and chemical signals. Different activities, i.e., to hear, to see, to remember and to learn among others are controlled by various regions of the brain. There are different kinds of realistic and simplified models for simulating neuronal systems and large brain networks with the advancement of computational power (Sejnowski et al. 1988).
Realistic models try to incorporate different available cellular parameters, and as a result, they become computationally expensive and complex to understand. Such an example is the Hartline-Ratliff model of the lateral inhibition and interaction in Limulus eye (Hartline and Ratcliff 1974) at the network level. But even the most successful realistic brain models often fail to predict the tissue functions. The concept of receptive field introduced first by Sherrington (Sherrington 1906) as a basic functional unit of neurons and later by Hartline (Hartline 1938) led to the classical receptive field models developed in cat and primate visual systems. These, in turn, led to the explanation of a few psychophysics based experimental results, while at the same time also failing in some (Gregory 1981, 2009). Thus it is necessary to develop simple models that can capture important principles. Though these models fail to predict the complete functional and structural features of brain circuits due to incomplete information on different parameters, they can shape future research directions with the help of experimental data and programs (Sejnowski et al. 1988).
3 Computational neuroscience and brain network organization
The scope of computation in neuroscience is enormous. One of the research areas coming under this scope is ‘brain network organization’, both structural and functional (Zhou et al. 2006; Bassett and Bullmore 2009; Achard and Bullmore 2007; Bullmore and Sporns 2012; Goñi et al. 2014). Significant progress can be found in the field of studying microscopic brain dynamics (i.e., at the level of single-neuron, single-synapse, and single-molecule) over the last 50 years. However, many research issues are yet to be addressed regarding the understanding of macroscopic/mesoscopic brain dynamics, often recorded by EEG (electroencephalography), ERPs (event related potentials), MEG (MAGNETOENCEPHALOGRAPHY), and LFPs (local field potentials) among other methods. Coherent dynamic phenomena of local brain networks (thousands of neurons) as well as the entire brain regions (millions of neurons) can be understood from electrical recordings of brain activity. There are close ties between blood flow signals measured with fMRI (functional magnetic resonance imaging) and local field potentials recorded in the cortex. Any form of synchrony, stationary/traveling, oscillatory activity can be associated with the coherent brain dynamics. It is an open problem to understand how the brain dynamics at different spatial scales are functionally interrelated. Moreover, it is quite challenging to establish a connection between the dynamics of single neurons monitored by intracellular recordings and the local or distant brain regions observed using EEG, ERPs, MEG, LFPs, and fMRI among other techniques. For the very reason, we have mentioned briefly about microscopic brain dynamics and mostly concentrated on the study of macroscopic/mesoscopic brain dynamics in the following discussion.
3.1 Microscopic brain dynamics: Single neurons and synapses
It is very difficult to form an anatomical connection matrix of human brain (connectome) at the level of single neuron because of its larger magnitude. For example, cortex alone contains 1010 neurons and 1013 connections approximately (Murre and Sturdy 1995; Braitenberg and Schuz 1991; Sporns et al. 2005). Alterations of single synapses do not reflect traceable macroscopic effects. Moreover, individual neurons and connections undergo rapid plastic changes (Mountcastle 1998). The huge number and high variability of individual neurons and synapses restrict us to include the microscopic studies as basic elements of brain network organization in this section.
3.2 Macroscopic/mesoscopic brain dynamics: Networks among brain regions as well as elementary processing units
It is a hard task to delineate brain areas and neuronal populations. There is no unique rule for parcellation of human brain regions. A previous investigation (Van Essen et al. 1998) has shown that neurons are arranged in the order of 100 or more number of anatomically distinct regions and areas in the human cerebral cortex. Different criteria for parcellation may be needed for different parts of the human brain (e.g., cerebellum, brain stem, cortex or thalamus). Thus, brain network organization among anatomically distinct brain regions and interregional pathways are most feasible to form a human neuronal matrix. However, the corticothalamic matrix at the macro scale does not provide functional informative subdivisions or segregated subcircuits within each brain region. Thus, macroscopic brain network organization is often insufficient to understand human brain’s functional dynamics and information processing capacities. Hence, mesoscopic brain network organization, i.e., the characterization of connection patterns among elementary processing units is crucial. Here, we have covered a few studies that analyze brain network organization at macro/meso scale.
Just as real-world complex systems may mathematically be modeled as graphs, in the same way, both structural (e.g. anatomical) as well as functional properties of a brain may be revealed. Identification of relations for a structural network is quite challenging. Several methods for imaging and recording from various regions of the brain have been developed for such challenges (Sporns and Kötter 2004; Bressler and Menon 2010; Sompolinsky 2014). We can predict types of possible interactions among different areas of a brain by finding out the pattern of structural network connectivity. For example, small world structure of cortical networks suggests that there are many short-range functional interactions but a few long-range interactions (Bassett et al. 2006).
Brain functional networks reflect coordination between segregation and integration, which is established by various factors, i.e., structural nodes, path length, divergence and convergence among others (Rubinov and Sporns 2010). They are often scale-free, and have small world properties along with short path length and high clustering coefficient. There is a compromised balance between network efficiency and wiring cost (Achard and Bullmore 2007). It is a challenge to predict the functional network connectivity of a brain from its corresponding structural connectivity. Often these predictions are based on analytical measurements and network communications (Goñi et al. 2014), but work has already been done with cat (Zhou et al. 2006) and monkey brain (Achard et al. 2006). Such kinds of studies help in analyzing the underlying anatomical connectivity of different portions of a brain. A normal brain functional network has been correlated with age (Chen et al. 2013), sex (Chen et al. 2013), memory (Burianova et al. 2010), education and intelligence (Bassett et al. 2011; Langer et al. 2012; Muller and Meyer 2014), occupation, environment and stress (de Quervain et al. 2000) among other factors by a stream of individual investigations. Often, more complexity arises from the fact that two hemispheres of a brain are different in terms of anatomical and functional behaviors (Chen et al. 2013; Muller and Meyer 2014).
Brain networks have a tendency to maximize functional motifs as well as minimize structural motifs (Sporns and Kötter 2004). By definition, a functional network is a set of interactions among brain regions to perform specific functions (Bressler and Menon 2010). The topology and connectivity matrix of large-scale functional networks changes through a person’s lifespan according to maturity and learning process (Bressler and Tognoli 2006). Different components of large-scale brain functional networks execute different activities with the help of other components (Miller and Cohen 2001). For example, coordination between prefrontal and posterior parietal control areas allows activities among motor and sensory areas that help in perceptuomotor processing (Corbetta and Shulman 2002; Armstrong et al. 2006; Ruff et al. 2006; Bressler et al. 2008). It explains how different parts of the brain communicate with each other due to various perceptions. There is no proper definition available for brain functional nodes (Bressler and Menon 2010). However, now-a-days, brain functional nodes can be identified by elevated metabolism in PET (Positron Emission Tomography), blood perfusion in fMRI, synchronized oscillatory activity in LFP recordings (Bressler and Menon 2010) and also by fNIRS (Functional Near Infrared Spectroscopy).
A normal brain functional network also alters with respect to body disorders and diseases, i.e., type 2 diabetes (Reijmer et al. 2013), Alzheimer’s disease (Tijms et al. 2014), attention-deficit hyperactivity disorder (ADHD) (Lee et al. 2011), autism (Lee et al. 2011), Parkinson’s disease (Feigin et al. 2007) and bipolar disorder (Leow et al. 2013) among others. Type 2 diabetic patients process information slowly due to the alteration or disruption in the white matter region (Reijmer et al. 2013). Alzheimer’s patients with more severe cognitive impairment have increased topology randomness in gray matter (Tijms et al. 2014). Bipolar patients have been characterized by impaired inter-hemispheric but relatively preserved intra-hemispheric integration. Moreover, low white matter integrity at the corpus callosum has been found in bipolar disorder patients (Leow et al. 2013).
It has also been found that lesions of many brain disorders including Alzheimer’s disease and schizophrenia are remarkably more plausible to be located in hub nodes of their respective brain networks (Crossley et al. 2014). Moreover, some previous investigations have studied various stages of different neurological disorders including autism, Alzheimer’s and epilepsy using graph theory (Bartolomei et al. 2013; Peters et al. 2013; Seo et al. 2013). Different stages of these disorders have been compared with each other and with reference normal graphs obtained from the healthy volunteers. The comparisons highlighted that the increase of local clustering index and average path length as the severity of the disease increases. There are different kind of models to determine the abnormalities in brain circuits during different psychiatric and neurological diseases, like epilepsy, Parkinson’s disease, autism, schizophrenia and disorders of consciousness (Schiff et al. 2014; Ching and Brown 2014; McCarthy et al. 2012). They consider different factors for analysis, like altered gains of neurons and synapses, abnormal oscillations, excitation-inhibition imbalance, pattern change of large-scale brain dynamics and resting state networks (Moussa et al. 2012). These kinds of relation provide the conceptual framework for the graph-theoretic analysis of large scale brain network (Smith et al. 2013; Bullmore and Sporns 2009; Battaglia et al. 2012).
Analysis of resting-state fMRI functional connectivity by graph theoretic studies (Astolfi, de Vico Fallani et al. 2007; Dosenbach et al. 2007) of large-scale human brain networks often highlights small-world network properties (Bullmore and Sporns 2009; Supekar et al. 2009). The topology of sub-networks can be described using different graph theoretic matrices (Müller-Linow et al. 2008). The analysis of resting and specific task related connectivity patterns (Smith et al. 2009) indicated that functional networks inherit small systemic change during cognition. As discussed earlier there are different models to identify the altered patterns of functional networks during various psychiatric and neurological disorders (Bhattacharya 2001; Murias et al. 2007; Seeley et al. 2009; Uhlhaas and Singer 2007; Timmermann et al. 2003; Stam et al. 2007; Ford and Mathalon 2008). However, a lot has to be done in order to explore the hierarchy of human brain networks. Thus, the field of brain network analysis explores different aspects of brain functions and aims at finding new fundamental insights during normal and abnormal cognitive states.
3.3 Existing tools, software and databases
Different tools and software are available to analyze computational neuroscience data and neuroimages. They are maintained by The Source for NeuroInformatics Tools and Resources (NITRC)Footnote 1. A few of them are discussed here. These tools and resources are freely available. Mostly, the kind of data used, other specific requirements and their sources are discussed here.
-
FSLFootnote 2: It is a library of tools for analyzing the EEG, MRI and fMRI data (freely available).
-
SPMFootnote 3: It deals with EEG, fMRI, MEG, PET and SPECT brain imaging data sequences, and their analyses (freely available).
-
BrainVoyagerQXFootnote 4: It can handle MRI, EEG and MEG data. This software incorporates statistical, numerical and image processing tools. It requires HASP license (freely available).
-
Turbo BrainVoyagerFootnote 5: It is online real-time processing software to handle fMRI data. It requires HASP license (freely available).
-
MRIcronFootnote 6: It is an image viewing toolbox to deal with MRI data. It also supports some in-built statistical functions (freely available).
-
itk-SNAPFootnote 7: This is a 3D medical image segmentation software. It handles MRI and CT scan data (freely available).
-
EEGLABFootnote 8: It is an open source environment for electrophysiological signal processing. It has interactive MATLAB toolbox for visualization, artifact removal and analysis of EEG, MEG and ECoG (ElectroCorticoGraphy) data (freely available).
-
ChronuxFootnote 9: The current release can be implemented as a MATLAB library. It preprocesses, explores, and analyses neural data, i.e., point process and continuous data (freely available).
-
FreeSurferFootnote 10: It reconstructs the brain surface from MRI data, and overlays fMRI data on the reconstructed surface (freely available).
-
BrainNet ViewerFootnote 11: It is a visualization tool and constructs structural and functional networks from filtered or processed data (freely available).
-
eConnectomeFootnote 12: It is a MATLAB package for imaging brain functional connectivity. It deals with EEG, MEG and ECoG data (freely available).
-
MNEFootnote 13: It has pre-processing tools and data conditioning utilities. It is a Linux based software. It deals with MEG and EEG data (freely available).
-
CONNFootnote 14: It is a MATLAB toolbox for computation, display and fMRI data analysis (freely available).
-
BSMacFootnote 15: It is a MATLAB based statistical and graphical visualization toolbox for fMRI data (freely available).
-
Bioelectromagnetism MATLAB ToolboxFootnote 16: It helps in visualization and measurement of Event Related Potential (ERP). It accepts EEG and MRI data (freely available).
-
The Virtual BrainFootnote 17: It simulates brain behavior by manipulating brain connectivity and some associated network parameters (freely available).
Table 1 lists a few computational neuroscience databases. It lists different types of data (image/network; EEG/MRI/fMRI; normal/diseased; real/simulated), the source of data, access type (freely available/paid), a brief description of the data source and sample properties of each listed database.
In order to maintain the huge resources, generated and used in computational neuroscience, a distinct sub-branch, called ‘neuroinformatics’ has emerged. The following two sections cover an introduction; languages, data, tools and software, simulation platforms used by the neuroinformatics community; a few new approaches; and resources.
4 Neuroinformatics
The field of neuroinformatics is an extended and elaborated application of tools and databases applied to broader, heterogeneous types of neuroscience data, which come in a wide range of spatial scales (Morse 2008). For example, a human brain must be represented at multiple levels and resolutions by properties of its neuronal and non-neuronal cells for detailed modeling and simulation. Neuronal properties include morphology, subcellular and molecular architecture, and physiology of ~1011 neurons. Properties of an equal number of non-neuronal cells (astrocytes, oligodendrocytes, and microglia among others) must also be considered. Moreover, the behavior of ~1014–1015 synapses must be included in an ideal holistic brain model. Each synapse is a complex molecular machine at the sub-micron level. In addition to that neuronal modulation, the activity of glia and synaptic activity by peptides, neurotransmitters, hormones and other molecules should be accounted for in such a model. Further complexities arise in terms of the temporal (structural and functional) variations in individual brains (environmental conditions, health, maturity and developmental stage), individualistic variations (gender and experience) and species variations (homology) (Frackowiak and Markram 2015). The size and complexity of data make it difficult to believe that a single mind or a single supercomputer or even a single country’s neuroscientists can shoulder a problem of such size without the help of neuroinformatics.
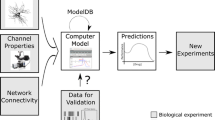
Neuroinformatics is thus becoming an integral part of neuroscience and clinical research for conducting scientific inquiry and practicing medicine (Morse 2008). Figure 2 schematically depicts the interdependence of this field with that of the neuroscience and computational neuroscience. Dedicated databases and tools of this field are required for understanding the normal as well as disordered nervous system states and clinical applications (Morse 2008). Neuroinformatics facilitates the maintenance of neuroscience databases. The field is associated with the development of analytical tools and computational models for the data sharing, knowledge integration, and analysis of big data of neuroscience (Bjaalie and Grillner 2007). It acts as a unifying framework for neuroscience towards new discoveries in human brain organization (Frackowiak and Markram 2015).
According to the International Neuroinformatics Coordinating FacilityFootnote 39 (INCF), the challenge in neuroscience is that data is generated in individual laboratories and get collected in huge volume all over the world. Neither the state-of-the-art tools/infrastructure, standard, culture, nor a community to actually bring these pieces together is still available. Neuroinformatics is about data organization, annotation, integration into atlases and models, and simulations of such knowledge in order to better understand the human brain and nervous system.
Millions of people around the globe suffer from brain diseases and disorders. New investments in neuroscience are scarce as it is too risky, complex, and highly expensive. When it comes to scientific data, data tend to be very precise. Implementing a universal standard for such precise multi-scale data is a major problem. A super data infrastructure is very much needed to address this kind of multiscale data. Brain disorders are complicated and heterogeneous. Attempting to understand a brain disorder with all its variables [genetics, imaging, clinical profiles (proteins and other blood variables)] is difficult and time-consuming. Then the prediction of next course or what happens to patients over time is the demand of the present decade. Neuroinformatics is a whole endeavor in response to such a demand which supports neuroscientists in bringing together, sharing and integrating their data to accelerate humankind’s understanding of the brain.
5 Advances in neuroinformatics
In this section, the languages, data, tools, software, and simulation platforms used by the neuroinformatics community are discussed along with a few new approaches to the field.
5.1 Languages for data storage and exchange
Generally, BrainML (Gardner et al. 2002, 2003), NeuroML (Gleeson et al. 2010), and PyNN (Davison 2015) are used for exchanging neuroscience data. Here we briefly discuss them. Moreover, PyNN being a simulator-independent language allows model comparison among multiple simulators.
BrainML provides a standard XML metaformat for neuroscience data exchange. Description formats for different biological objects (animal model/cortex/neuron) are available (Le Novère 2006) in it. BrainML is being built on the BrainMetaL metalanguage. It uses hierarchical trees of controlled-vocabulary descriptors linked to specific attributes (lexicons) (Gardner et al. 2002, 2003).
NeuroMLFootnote 40 focuses on developing an XML based language for the neuronal system models. The project provides a common data format for defining neuronal cells and network models. It is a simulator-independent neuronal model description language. It can define data-driven models of neurons as well as their networks with high degrees of biological details. This language allows the description of the complex branching structures of dendritic trees and axonal projections, and their biophysical properties. It allows the description of chemical synapses with short-term synaptic plasticity, electrical synapses, voltage and calcium gated ion channels, and both large and small scale network structure (Gleeson et al. 2010).
PyNNFootnote 41 is a simulator-independent language to build models of neuronal networks. PyNN API facilitates modeling at the level of a neuron population as well as layers, columns and the connections among them. It has a standard model library of neurons, synapses, and synaptic plasticity. A code written with PyNN API using Python can be run on any PyNN supported simulator, i.e., NEURON (Carnevale and Hines 2006), NEST (Plesser et al. 2015), PCSIM (Pecevski et al. 2009), and Brian (Goodman and Brette 2009) among others. PyNN imparts validity to modeling studies by checking results of multiple simulators (Davison et al. 2008; Davison 2015).
5.2 Data
Here we discuss a few of the data sources used by the neuroinformaticians. Brede Database (Nielsen 2014) lists 186 published neuroimaging articles and 586 experiments. Human Connectome Project (Van Essen et al. 2012) aims at identifying group differences across the lifespan by comparing imaging data in 6 age groups (4–6, 8–9, 14–15, 25–35, 45–55, 65–75 years). Brainnetome (Jiang 2013) deals with identification, characterization, network manifestation and genetic basis of brain networks.
BredeFootnote 42 lists data from functional neuroimaging related research articles with Talairach coordinates. The database has a program package written in MATLAB, called as Brede Toolbox. It structures data from each article into one or multiple experiments which are supported by simple ontologies for brain regions, journals, persons and topics (Nielsen 2014).
Human Connectome ProjectFootnote 43 (HCP) characterizes brain connectivity, function, and variability in normal human beings. It aims to build a network map among the functional and anatomical connectivity of normal and disordered brains. It has multiple imaging modalities (diffusion-weighted MRI, resting-state fMRI, task-based fMRI, T1- and T2- weighted MRI for structural and myelin mapping and combined MEG and EEG). It is acquiring and sharing pilot multimodal imaging data collected across different age range of human lifespanFootnote 44 (in 6 age groups, i.e., 4–6, 8–9, 14–15, 25–35, 45–55, 65–75 years) for identifying group differences across the lifespan and compare data across scanner platforms (Van Essen et al. 2012).
BrainnetomeFootnote 45 investigates human brain hierarchy from genetics and neuronal circuits point-of-view. It aims at identification, study of dynamics and characteristics, network manifestation of functions and malfunctions, analysis of genetic basis, simulation and modeling of brain networks (Jiang 2013).
5.3 Tools and software
There exist a lot of tools and software for neuroinformatics research. A few of them (Gleeson et al. 2007; Glaser and Glaser 1990; Aguiar et al. 2013; Stalling et al. 2005; Ermentrout 2002) are discussed here.
neuroConstructFootnote 46 is used in developing realistic 3D neuronal networks. It facilitates models to incorporate dendritic morphologies and realistic cell membrane conductances. It has been developed in Java. Its script files can be used in many neuronal simulation platforms. It uses the latest NeuroML (Gleeson et al. 2010) specifications (ChannelML, MorphML and NetworkML). neuroConstruct can create, visualize, and analyze multi-compartmental neuronal networks in 3D space. It allows modeling brain function by building, visualizing, and analyzing network models in 3D space using a user-friendly GUI (Gleeson et al. 2007).
NeurolucidaFootnote 47 is used for creating and analyzing neuron reconstructions from microscope images. It can map and analyze the distribution of cells in a region of interest along with quantification of axons, dendrites, nodes, spines and synapses. It can quantify volume, proximity of one object to another, and distance between the object and an anatomical boundary (Glaser and Glaser 1990).
Py3DNFootnote 48 analyzes 3D data collected with Neurolucida (Glaser and Glaser 1990) with its in-house tools. It facilitates mathematical representation of neuronal topology, visualization, and analysis of the neurons. It is a Python-based application and uses the open-source Blender program to create 3D representations of raw and processed data. A user can import Neurolucida reconstruction data, access it using intuitive data structures and Python, do morphometric analysis, and construct 3D graphical representations of the results with Py3DN (Aguiar et al. 2013).
AmiraFootnote 49 is a multifaceted 3D software platform for visualization, manipulation of microscopy, computed tomography, MRI and other imaging modality data. It focuses on visualization and analysis of volumetric data generated in medicine, biology and microscopy. Amira is designed with the goals of ease of use, flexibility, interactivity, extensibility, scripting interface, multi-platform support, and state-of-the-art algorithms (Stalling et al. 2005).
X Windows Phase Plane plus Auto (XPP-AUT)Footnote 50 solves difference, differential, delay, functional and stochastic equations along with boundary value problems. It has the code for the popular bifurcation program, called as AUTOFootnote 51. It was originally developed for complete numerical analysis of the dependence of solutions on parameters. However, later it has been applied in computational and theoretical neuroscience (Ermentrout 2002). The open source program has the capabilities for handling up to 590 differential equations (Brette et al. 2007; Ermentrout 2012).
5.4 Simulation platforms
Simulation platforms in general are needed for building neuronal computational models. They provide tools to build numerically sound and computationally efficient models. These platforms help users to solve biological issues rather than get gimmicked by the related computational difficulties. Often, these platforms are used for education in neurobiology. Some are designed with a focus on dynamics, size, and structure of large networks of neurons. A few others deal with heterogeneous network simulations with different model neurons and their synapses as components.
NEURONFootnote 52 is a free open source simulation environment. It is well-suited to problems of complex experimental data (anatomical and biophysical properties). It has a user-friendly interface and a library of biophysical mechanisms that can be extended by a user (Carnevale and Hines 2006). It facilitates efficient network modeling and offers a user scope of customizable initialization and simulation flow control (Brette et al. 2007).
GEneral NEural SImulation System (GENESIS)Footnote 53 and its version for parallel and networked computers (PGENESIS) are the first macro scale modeling systems in computational neuroscinece. GENESIS can simulate biological neuronal systems at different levels, i.e., a single neuron, large neuronal network, subcellular components and biochemical reactions, and system-level models (Bower and Beeman 1998). GENESIS version 3 (development version) will enable import and export of model descriptions in a common simulator-independent XML format.
NEural Simulation Tool (NEST)Footnote 54 can model networks of spiking neurons of varying size (Plesser et al. 2015). It can model laminar cortical networks, the auditory or visual cortex of mammals and models of learning and plasticity among others (Gewaltig and Diesmann 2007; Brette et al. 2007). NEST can be used with SLI (NEST’s native simulation language interpreter), PyNEST (Eppler et al. 2008) and PyNN (Davison 2015).
PyNESTFootnote 55 is a user interface to NEST. It combines the simulation kernel of NEST with Python. PyNEST facilitates easy set up for simulations than SLI (Eppler et al. 2008; Plesser et al. 2015).
CSIM: A neural Circuit SIMulatorFootnote 56 simulates heterogeneous networks with neurons and synapses as components. It can simulate up to a few thousand neurons with 1,000,000 synapses. A parallel (distributed) version of CSIM is also available for researchers (Pecevski et al. 2009).
Parallel Circuit SIMulator (PCSIM)Footnote 57 is the successor of CSIM. It simulates networks containing up to millions of neurons with billions of synapses. It allows distributed (via MPI) and multithreaded simulation of large neuronal networks (Pecevski et al. 2009).
NeoCortical Simulator (NCS)Footnote 58 is a parallel (MPI-based) spiking neuronal network simulator capable of large discrete-time simulations (million neurons with billion synapses). The simulator is well versed with cell membrane voltage dynamics and customizable ion channels of neuron models. It supports multi-compartment cells (Drewes et al. 2009). Currently, a web-based application, known as ‘Neocortical Builder (NCB)’, is also available (Berlinski et al. 2014). NCB is used for creating simulation input, building brain models, and output parameters with NCS. It provides a graphical interface for streamlining the brain simulations. It also provides real-time applications for neurobotics along with the integrate-and-fire neurons and conductance-based synapses.
Multiscale Object-Oriented Simulation Environment (MOOSE)Footnote 59 simulates neuronal systems at multiple levels (Ray et al. 2008). It operates at many levels of detail, i.e., stochastic chemical computations, a single neuron, spiking neuron network and multi-compartment single-neuron models (Dudani et al. 2013). It supports many model formats including SBMLFootnote 60, NeuroML (Gleeson et al. 2010), HDF5Footnote 61, GENESIS KinetikitFootnote 62 and Neuroscience Simulation Data Format (NSDF) for writing data.
Parallel Stochastic Ion Channel Simulator (PSICS)Footnote 63 computes the behavior of neurons with ion channel position and stochasticity. It uses population statistics for electrically compact homogeneous channel populations. Input models to PSICS can be created with any XML-aware text editor (Cannon et al. 2010). Specification of channel distributions and the positioning of stimuli and recorders are achieved with ICINGFootnote 64.
MvaspikeFootnote 65 is involved with event-based modeling and simulation. It handles the events with an event-driven simulation algorithm which sorts the events, updates the neurons and propagates the spikes. The tool maintains a good balance between modeling freedom and simulation efficiency (Kaabi et al. 2011).
Brian spiking neuronal network simulator (Brian)Footnote 66 is a simulator for spiking neuronal networks. It defines neuronal models by differential equations. It is a Python-based simulator and uses vector-based computation (Goodman and Brette 2009). It is currently available in two versions, i.e., Brian 1.x and Brian 2.x (Stimberg et al. 2014).
5.5 A few innovative applications
Here we discuss a few interesting and innovative neuroinformatics applications like BrainX3 (Arsiwalla et al. 2015), Brain-Gene Ontology (Kasabov et al. 2008), Brain Cartography (Frackowiak and Markram 2015), and Brain Cartography and Connectomics (Sporns 2015). These applications use computation to explore and analyze various behaviors of human brain networks. They hold greater scope of upliftment for human health and life, albeit with a need for more improvement and fine-tuning.
BrainX3 (Arsiwalla et al. 2015) is a 3D simulation of the human cerebral neuronal matrix. It uses biophysical dynamics and anatomical structure for activity reconstruction and function prediction. Interestingly, it can be used in a virtual reality chamber for real-time interactions using natural gestures (figure 3). Presently, it can process networks of up to 4000 nodes. A laptop version of BrainX3 is under development. This technology has many future applications. One among them is the virtual brain surgery. A surgeon can access several virtual surgical procedures along with their pros and cons on models based on the patients’ data via BrainX3. However, such an application requires optimization of BrainX3 with parallel computing.
BrainX3 (Arsiwalla et al. 2015), a 3D simulation of the human cerebral connectome.
Brain-Gene Ontology (BGO) includes data (animations, concepts, experimental publications, genetic, knowledge, facts, graphs, software simulations, theories, and visualizations among others) of mammalian brain functions, diseases, and their inter-relationships. It also lists brain organization, gene regulatory networks, and simulation models (Kasabov et al. 2006, 2008).
Brain Cartography (Frackowiak and Markram 2015) represents the science or practice of drawing maps for the brain. It identifies brain regions and localizes them for neurosurgical uses. It provides an anatomical framework for the brain’s structural and functional architecture. The future endeavors aim for inter- and intra-individual variability and representation of brain organization across different spatial and temporal scales.
Brain Cartography and Connectomics (Sporns 2015) represents a brain as a complex network of neurons and their interconnecting synapses. A connectome is simply a wiring diagram of neuronal connections in the brain. However, cartographists face many tough challenges while representing a connectome, because varieties of data at different scales of resolution have to be associated with it. They have to incorporate and arrange multi-level information (temporal, dynamic, definition, hierarchical organization, structure, function, relational, and models among others) at one go for construction of a connectome. For building a human connectome, these challenges have to be overcome. Table 2 lists a few worldwide databases in the area of neuroinformatics.
6 Discussion and conclusions
This review along with its compilation of research resources tries to encompass the journey of neuroscience towards technology. Neuroscientists are always up against the task of understanding the function of human brain. It is a very daunting task with billions of brain cells and approximately 100 trillion connections among them. Leave alone the overall view; some believe we are still scratching the surface of this enigma called a brain. Recently the contents of three cylindrical chunks of mouse brain tissue, each of the size of a grain of salt are analyzed by VASTFootnote 76. VAST is a manual computer program that can automatically label neuronal structures by space-filling segmentation and annotation. It labels individual dendrites, glia, mitochondria, neurons, and blood vessels with different colors. The researchers (Kasthuri et al. 2015) created an inventory of 1700 annotated synapses. The data refuted ‘the Peters’ rule’ that physical proximity is enough to predict synaptic connectivity. Thus the very basis of estimation of neuronal connectivity is shaken now. This is just one facet of neuroscience that deals with small-scale neuroscience and advanced imaging technology.
Another experiment compared the way monkey and human brains respond to abstract information. According to it, monkeys recognize a pattern but do not realize it and take it no further. However, humans take it on to the next level of analysis collectively. The researchers (Wang et al. 2015) found that inferior frontal gyrus of the cortex was intensely activated only in human brains for such kind of analysis that made them unique in processing abstract information analysis. A reader cannot refrain from self-wonder with such kind of precise discoveries. There are many other equally astounding facets of neuroscience, which remain to be solved/refuted with coming age technology. Many-a-times computational neuroscience comes to aid in these scenarios.
Computational neuroscience approaches further our understanding of brain function, and help in translating the acquired knowledge into technological applications. Be it cellular and synaptic dynamics or biophysical basis of neuronal computation or algorithms for the analysis of neuronal data, the field has provided analytical and computational skills for understanding the neuronal systems. Let us take two instances of research where computational neuroscience plays an upper hand for extraordinary developments. It gives power to a disabled musician to control a musical performance via a Brain Computer Music Interface (BCMI)Footnote 77. A BCMI system (figure 4) detects brainwave signals of a disabled musician and allows him to control musical systems.
On a very different note of computational neuroscience related development, Brainnets, (Pais-Vieira et al. 2015; Ramakrishnan et al. 2015) have emerged as an alternative to super-computation and hyper-performance. A Brainnet is a system of multiple interconnected animal brains. It has a distributed and parallel computing architecture. Brainnets add an immense opportunity for the upliftment of human life. How good it will be if a crucial surgery can be performed by a Brainnet of multiple eminent surgeons around the world (figure 5). Some non-invasive ethically approved first steps have already been taken in this direction. A few researchers of University of Washington, USA first reported about the direct human brain-to-brain interface. It records EEG signals of one human brain and uses transcranial magnetic stimulation (TMS) to deliver information to another human brain. Some researchers of Starlab Barcelona, Spain (Grau et al. 2014) have used brain-computer interfaces (BCI) and non-invasive computer-brain interfaces (CBI) to achieve B2B (computer-assisted brain-to-brain) communication between human subjects (hyperinteraction) (Rao et al. 2014). These are a few of the wonders made possible by computational neuroscience.
With the advent of computational neuroscience, generation and analysis of the enormous amount of data became easy. On the other hand, such kind of big data need proper handling, a common standard/protocol of data generation, storage, appropriate retrieval and requirement-specific merging for rich analysis. The field of Neuroinformatics handles such kind of requirements. A few hand-picked recent developments in the field of Neuroinformatics can help a reader to get a grasp of its power. BrainBrowser (Sherif et al. 2014) is a visualization library which provides on-demand visualization of remote datasets. It enables a user to visualize volumetric neuroimaging data and 3D surface in any modern web browser without any further requirements. The Brain Genomics Superstruct Project (GSP) (Holmes et al. 2015) enables exploration of inter-links among behavior, brain function, and genetic variation.
Functional analysis of human brain invariably leads towards creation and analysis of connectomes, a relatively new term researchers started using from 2005 (Sporns 2015). A connectome is simply a wiring diagram of neuronal connections in the brain. Generating connectomes even in part is complex, assembling them is tough, and analyzing them is yet new to the research fraternity. It is a good thing that the projects dedicated to connectomics of human brain follow open data policies which generate scope for many future discoveries. A disorder-specific human connectome holds new hope for solving the complex brain disorders. Computing techniques like probabilistic clustering can identify communities and hubs in the human connectome (Hinne et al. 2015). Such methods can be used to study the connectomics of brain disorders, and generate predictive models for disease spread pattern and consequences (Fornito et al. 2015). Function-specific connectomes, i.e., ‘functional connectome of speech control’ has already been discovered (Fuertinger et al. 2015). The connectome of the human brain is getting constructed even as we are writing this review, which can revolutionize the whole human perception regarding behavior, intelligence, memory, and diseases.
Advanced experimental methods, imaging techniques and support from neuroinformatics infrastructure have illuminated many areas of neuroscience, and yet many other questions lay unanswered. This article is an introductory guide to a few of the already existing knowledge bases and research in computational neuroscience and neuroinformatics. It aims to help many of the scientific fraternity who put their nascent steps into these fields to solve these unanswered questions.
Notes
http://fcon_1000.projects.nitrc.org/indi/retro/cobre.html
http://fcon_1000.projects.nitrc.org/indi/adhd200/
http://sbml.org/Main_Page [Systems Biology Markup Language (SBML) is a representation format, based on XML, for communicating and storing computational models of biological processes.]
https://support.hdfgroup.org/HDF5/ [Hierarchical Data Format, version 5.]
http://genesis-sim.org/ [GENESIS version 2.3 and higher contain Kinetikit. Kinetikit is an interface and utility for developing simulations of chemical kinetics.]
http://www.psics.org/icing/index.html [A stand-alone graphical tool for creating and visualizing ion channel distributions.]
http://fcon_1000.projects.nitrc.org/ [International Neuroimaging Data-sharing Initiative.]
https://dataverse.harvard.edu/dataverse/GSP [Brain Genomics Superstruct Project.]
https://ndar.nih.gov/ [National Database for Autism Research.]
http://fcon_1000.projects.nitrc.org/indi/enhanced/
https://preprocessed-connectomes-project.github.io/ [Preprocessed Connectomes Project.]
http://fcon_1000.projects.nitrc.org/indi/abide/ [Autism Brain Imaging Data Exchange.]
References
Achard S and Bullmore E 2007 Efficiency and cost of economical brain functional networks. PLoS Comput. Biol. 3 e17
Achard, S, Salvador R, Whitcher B, Suckling J and Bullmore E 2006 A resilient, low-frequency, small-world human brain functional network with highly connected association cortical hubs. J. Neurosci. 26 63–72
Adelman G 2010 The neurosciences research program at MIT and the beginning of the modern field of neuroscience. J. Hist. Neurosci. 19 15–23
Aguiar P, Sousa M and Szucs P 2013 Versatile morphometric analysis and visualization of the three-dimensional structure of neurons. Neuroinformatics 11 393–403
Andrzejak RG, Schindler K and Rummel C 2012 Nonrandomness, nonlinear dependence, and nonstationarity of electroencephalographic recordings from epilepsy patients. Phys. Rev. E Stat. Nonlin. Soft Matr. Phys. 86 046206
Armstrong KM, Fitzgerald JK and Moore T 2006 Changes in visual receptive fields with microstimulation of frontal cortex. Neuron 50 791–798
Arsiwalla X, Zucca R, Betella A, Martinez E, Dalmazzo D, Omedas P, Deco G and Verschure P 2015 Network dynamics with BrainX(3) a large-scale simulation of the human brain network with real-time interaction. Front. Neuroinform. 9 02
Astolfi L, de Vico Fallani F, Cincotti F, Mattia D, Marciani MG, Bufalari S, Salinari S, Colosimo A, Ding L, et al. 2007 Imaging functional brain connectivity patterns from high-resolution EEG and fMRI via graph theory. Psychophysiology 44 880–893
Azevedo FA, Carvalho LR, Grinberg LT, Farfel JM, Ferretti RE, Leite RE, Jacob W Filho, Lent R et al. 2009 Equal numbers of neuronal and nonneuronal cells make the human brain an isometrically scaled-up primate brain. J. Comp. Neurol. 513 532–541
Bartolomei F, Bettus G, Stam CJ and Guye M 2013 Interictal network properties in mesial temporal lobe epilepsy a graph theoretical study from intracerebral recordings. Clin. Neurophysiol. 124 2345–2353
Bassett DS, Andreas M-L, Achard S, Duke T and Bullmore E 2006 Adaptive reconfiguration of fractal small-world human brain functional networks. Proc. Natl. Acad. Sci. USA 103 19518–19523
Bassett DS and Bullmore ET 2009 Human brain networks in health and disease. Curr. Opin. Neurol. 22 340–347
Bassett DS, Wymbs NF, Porter MA, Mucha PJ, Carlson JM and Grafton ST 2011 Dynamic reconfiguration of human brain networks during learning. Proc. Natl. Acad. Sci. USA 108 7641–7646
Battaglia, D, Witt A, Wolf F and Geisel T 2012 Dynamic effective connectivity of inter-areal brain circuits. PLoS Comput. Biol. 8 e1002438
Bechtel W 2008 Mental mechanisms philosophical perspectives on cognitive neuroscience (Hove, Psychology)
Beltrame F, Koslow S, Gardner D, Ascoli G and Usai S 2000 Neuroinformatics as a mega science-a panel discussion. IEEE EMBS International Conference on Information Technology Applications in Biomedicine 314
Berlinski J, Rowe C, Chavez DM, Jordan NM, Tanna D, Hoang RV, Dascalu SM, Bray L CJ et al. 2014 NeoCortical Builder A Web Based Front End for NCS. Proceedings. of the 27th International Conference on Computer Applications in Industry and Engineering (CAINE-2014)
Bhattacharya J 2001 Reduced degree of long-range phase synchrony in pathological human brain. Acta. Neurobiol. Exp. 61 309–318
Birn RM, Cox RW and Bandettini PA 2002 Detection versus estimation in event-related fMRI choosing the optimal stimulus timing. Neuroimage 15 252–264
Bjaalie JG and Grillner S 2007 Global neuroinformatics the International Neuroinformatics Coordinating Facility. J. Neurosci. 27 3613–3615
Bota, M, Dong HW and Swanson LW 2003 From gene networks to brain networks. Nat. Neurosci. 6 795–799
Bower JM and Beeman D 1998 The. book of GENESIS exploring realistic neural models with the GEneral. NEural. SImulation. System (Santa Clara, Calif., TELOS)
Braitenberg V and Schuz A A 1991 Anatomy of the cortex statistics and geometry (Berlin, Springer)
Bressler SL and Menon V 2010 Large-scale brain networks in cognition emerging methods and principles. Trends. Cogn. Sci. 14 277–290
Bressler SL, Tang W, Sylvester CM, Shulman GL and Corbetta M 2008 Top-down control of human visual cortex by frontal and parietal cortex in anticipatory visual spatial attention. J. Neurosci. 28 10056–10061
Bressler SL and Tognoli E 2006 Operational principles of neurocognitive networks. Int. J. Psychophysiol. 60 139–148
Brette, R, Rudolph M, Carnevale T, Hines M, Beeman D, Bower JM, Diesmann M, Morrison A, et al. 2007 Simulation of networks of spiking neurons a review of tools and strategies. J. Comput. Neurosci. 23 349–398
Brown MR, Sidhu GS, Greiner R, Asgarian N, Bastani M, Silverstone PH, Greenshaw AJ and Dursun SM 2012 ADHD-200 Global Competition diagnosing ADHD using personal characteristic data can outperform resting state fMRI measurements. Front. Syst. Neurosci. 6 69
Bullmore E and Sporns O 2009 Complex brain networks graph theoretical analysis of structural and functional systems. Nat. Rev. Neurosci. 10 186–198
Bullmore E and Sporns O 2012 The economy of brain network organization. Nat. Rev. Neurosci. 13 336–349
Burianova HAR McIntosh and Grady CL 2010 A common functional brain network for autobiographical, episodic, and semantic memory retrieval. Neuroimage 49 865–874
Cannon RC, O'Donnell C and Nolan MF 2010 Stochastic ion channel gating in dendritic neurons: morphology dependence and probabilistic synaptic activation of dendritic spikes. PLoS Comput. Biol. 6(8) e1000886. https://doi.org/10.1371/journal.pcbi.1000886
Carnevale N and Hines M 2006 The. NEURON book (Cambridge University Press)
Chen Z, Liu M, Gross DW and Beaulieu C 2013 Graph theoretical analysis of developmental patterns of the white matter network. Front. Hum. Neurosci. 7 716
Ching S and Brown EN 2014 Modeling the dynamical effects of anesthesia on brain circuits. Curr. Opin. Neurobiol. 25 116–122
Cocosco CA, Kollokian VR, Kwan K-S, Pike GB and Evans AC 1997 BrainWeb: Online interface to a 3D MRI simulated brain database. Neuroimage 5 S425
Copeland BJ and Shagrir O 2017 Physical computation how general are Gandys principles for mechanisms? Minds Machines 17 217–231
Corbetta M and Shulman GL 2002 Control of goal-directed and stimulus-driven attention in the brain. Nat. Rev. Neurosci. 3 201–215
Craver CF 2007 Explaining the brain (Oxford, Oxford University Press)
Crossley NA, Mechelli A, Scott J, Carletti F, Fox PT P McGuire and Bullmore ET 2014 The hubs of the human connectome are generally implicated in the anatomy of brain disorders. Brain 137 2382–2395
da Fontoura Costa L and Bollt E 2006 Fast and accurate nonlinear spectral method for image recognition and registration. Appl. Phys. Lett. 89 174102
Davison AP 2015 PyNN A Python API for neural network modeling; In Encyclopedia of Computational Neuroscience pp 2548–2550
Davison AP, Brüderle D, Eppler J, Kremkow J, Muller E, Pecevski D, Perrinet L and Yger P 2008 PyNN A common interface for neuronal network simulators. Front. Neuroinform. 2 11
de Quervain DJ, Roozendaal B, Nitsch RM JL McGaugh and Hock C 2000 Acute cortisone administration impairs retrieval of long-term declarative memory in humans. Nat. Neurosci. 3 313–314
Delorme A, Makeig S, Fabre M-Thorpe and Sejnowski T 2002 From single-trial EEG to brain area dynamics. Neurocomputing 44 1057–1064
Dosenbach NU, Fair DA, Miezin FM, Cohen AL, Wenger KK, Dosenbach RA, Fox MD, Snyder AZ, et al. 2007 Distinct brain networks for adaptive and stable task control in humans. Proc. Natl. Acad. Sci. USA 104 11073–11078
Drewes R, Zou Q and Goodman PH 2009 Brainlab A Python toolkit to aid in the design, simulation, and analysis of spiking neural networks with the neocortical simulator. Front. Neuroinform. 3 16
Dudani N, Bhalla U and Ray S 2013 MOOSE, the multiscale object-oriented simulation environment; in Encyclopedia of Computational Neuroscience pp 1–4
Eppler JM, Helias M, Muller E, Diesmann M and Gewaltig MO 2008 PyNEST A Convenient Interface to the NEST Simulator. Front. Neuroinform. 2 12
Ermentrout B 2002 Simulating, analyzing, and animating dynamical systems: A guide to XPPAUT for researchers and students (Philadelphia, Pa., Society for Industrial & Applied Mathematics; Sunbury-on-Thames Electronica Books & Media)
Ermentrout B 2012 XPPAUT. Comput. Syst. Neurobiol. 519–531
Feigin A, Kaplitt MG, Tang C, Lin T, Mattis P, Dhawan V, During MJ and Eidelberg D 2007 Modulation of metabolic brain networks after subthalamic gene therapy for Parkinsons disease. Proc. Natl. Acad. Sci. USA 104 19559–19564
Ford JM and Mathalon DH 2008 Neural synchrony in schizophrenia. Schizophr. Bull. 34 904–906
Fornito A, Zalesky A and Breakspear M 2015 The connectomics of brain disorders. Nat. Rev. Neurosci. 16 159–172
Frackowiak R and Markram H 2015 The future of human cerebral cartography a novel approach. Philos. Trans. R Soc. Lond. B Biol. Sci. 3701668
Fuertinger S, Horwitz B and Simonyan K 2015 The functional Connectome of speech control. PLoS Biol. 13 e1002209
Gardner D, Abato M, Knuth KH and Xiao Y 2003 BrainML: Neuroinformatics for data sharing. Biophys. J. 84(2) 305A
Gardner D, Xiao Y, Abato M, Knuth K and Gardner E 2002 BrainML and GENIE Neuroinformatics schemas for neuroscience data sharing. Soc. Neurosci. Abstr. 28
Gewaltig M.-O. and Diesmann M 2007 NEST (neural simulation tool). Scholarpedia 2 1430
Glaser JR and Glaser EM 1990 Neuron imaging with Neurolucida–a PC-based system for image combining microscopy. Comput. Med. Imaging. Graph. 14 307–317
Gleeson P, Crook S, Cannon RC, Hines ML, Billings GO, Farinella M, Morse TM, Davison AP et al. 2010 NeuroML a language for describing data driven models of neurons and networks with a high degree of biological detail. PLoS Comput. Biol. 6 e1000815
Gleeson P, Steuber V and Silver RA 2007 neuroConstruct a tool for modeling networks of neurons in 3D space. Neuron 54 219–235
Goldental A, Guberman S, Vardi R and Kanter I 2014 A computational paradigm for dynamic logic-gates in neuronal activity. Front. Comput. Neurosci. 8 52
Gollub RL, Shoemaker JM, King MD, White T, Ehrlich S, Sponheim SR, Clark VP, Turner JA et al. 2013 The MCIC collection a shared repository of multi-modal, multi-site brain image data from a clinical investigation of schizophrenia. Neuroinformatics 11 367–388
Goodman DF and Brette R 2009 The brian simulator. Front. Neurosci. 3 192–197
Goñi J, van den Heuvel MP, Avena A-Koenigsberger, Velez N de Mendizabal, Betzel RF, Griffa A, Hagmann P, Corominas B-Murtra et al. 2014 Resting-brain functional connectivity predicted by analytic measures of network communication. Proc. Natl. Acad. Sci. USA 111 833–838
Grau C, Ginhoux R, Riera A, Nguyen TL, Chauvat H, Berg M, Amengual JL, Pascual A-Leone et al. 2014 Conscious brain-to-brain communication in humans using non-invasive technologies. PLoS One 9 e105225
Gregory RL 1981 Mind in science – a history of explanations in psychology and physics (Harmondsworth, Penguin)
Gregory RL 2009 Seeing. through illusions (Oxford, Oxford University Press)
Hanlon FM, Houck JM, Pyeatt CJ, Lundy SL, Euler MJ, Weisend MP, Thoma RJ, Bustillo JR et al. 2011 Bilateral hippocampal dysfunction in schizophrenia. Neuroimage 58 1158–1168
Hartline H 1938 The response of single optic nerve fibers of the vertebrate eye to illumination of the retina. Am. J. Physiol. 121 400–415
Hartline HK and Ratcliff F 1974 Studies on excitation and inhibition in the retina a collection of papers from the laboratories of H. Keffer. Hartline (London, Chapman and Hall)
Hawrylycz MJ, Lein ES, Guillozet AL-Bongaarts, Shen EH, Ng L, Miller JA LN van de Lagemaat, Smith KA, Ebbert A et al. 2012 An anatomically comprehensive atlas of the adult human brain transcriptome. Nature 489 391–399
Hinne M, Ekman M, Janssen RJ, Heskes T and M. A. van Gerven 2015 Probabilistic clustering of the human connectome identifies communities and hubs. PLoS One 10 e0117179
Holmes AJ, Hollinshead MO TM OKeefe, Petrov VI, Fariello GR, Wald LL, Fischl B, Rosen BR et al. 2015 Brain Genomics Superstruct Project initial data release with structural, functional, and behavioral measures. Sci. Data 2 150031
Jiang T 2013 Brainnetome a new -ome to understand the brain and its disorders. Neuroimage 80 263–272
Kaabi MG, Tonnelier A and Martinez D 2011 On the performance of voltage stepping for the simulation of adaptive, nonlinear integrate-and-fire neuronal networks. Neural. Comput. 23 1187–1204
Kasabov N, Jain V and Benuskova L 2008 Integrating evolving brain-gene ontology and connectionist-based system for modeling and knowledge discovery. Neural. Netw. 21 266–275
Kasabov N, Jain V, Gottgtroy PC, Benuskova L and Joseph F 2006 Brain-gene ontology Integrating bioinformatics and neuroinformatics data, information and knowledge to enable discoveries Sixth International Conference on Hybrid IntelligentSystems 2006 (HIS06) 13
Kasthuri N, Hayworth KJ, Berger DR, Schalek RL, Conchello JA, Knowles S-Barley, Lee D, Vázquez A-Reina et al. 2015 Saturated reconstruction of a volume of neocortex. Cell 162 648–661
Kempton MJ, Geddes JR, Ettinger U, Williams SC and Grasby PM 2008 Meta-analysis, database, and meta-regression of 98 structural imaging studies in bipolar disorder. Arch. Gen. Psychiatry 65 1017–1032
Kiernan MC 2015 A fine neuroscience vintage. J. Neurol. Neurosurg. Psychiatry 86 1–2
Kuklisova-Murgasova M, Aljabar P, Srinivasan L, Counsell SJ, Doria V, Serag A, Gousias IS et al. 2011 A dynamic 4D probabilistic atlas of the developing brain. Neuroimage 54 2750–2763
Langer N, Pedroni A, Gianotti LR, Hänggi J, Knoch D and Jäncke L 2012 Functional brain network efficiency predicts intelligence. Hum. Brain. Mapp. 33 1393–1406
Le Novère N 2006 Model storage, exchange and integration. BMC Neurosci. 7 Suppl 1 S11
Lee,H, Chung MK, Kang H, Kim BN and Lee DS 2011 Computing the shape of brain networks using graph filtration and Gromov-Hausdorff metric. Med. Image. Comput. Comput. Assist. Interv. 14 302–309
Lein ES, Hawrylycz MJ, Ao N, Ayres M, Bensinger A, Bernard A, Boe AF, Boguski MS et al. 2007 Genome-wide atlas of gene expression in the adult mouse brain. Nature 445 168–176
Leow, A, Ajilore O, Zhan L, Arienzo D J GadElkarim, Zhang A, Moody T, Van J Horn et al. 2013 Impaired inter-hemispheric integration in bipolar disorder revealed with brain network analyses. Biol. Psychiatry 73 183–193
MacKenzie-Graham A, Jones ES, Shattuck DW, Dinov ID, Bota M and Toga AW 2003 The informatics of a C57BL/6J mouse brain atlas. Neuroinformatics 1 397–410
Marcus DS, Fotenos AF, Csernansky JG, Morris JC and Buckner RL 2010 Open access series of imaging studies longitudinal MRI data in nondemented and demented older adults. J. Cogn. Neurosci. 22 2677–2684
Marcus DS, Wang TH, Parker J, Csernansky JG, Morris JC and Buckner RL 2007 Open Access Series of Imaging Studies (OASIS) cross-sectional MRI data in young, middle aged, nondemented, and demented older adults. J. Cogn. Neurosci. 19 1498–1507
Mason EE, Tang S, Renquist KE, Barnes DT, Cullen JJ, Doherty C and Maher JW 1997 A decade of change in obesity surgery. National Bariatric Surgery Registry (NBSR) Contributors. Obes. Surg. 7 189–197
McCarthy MM, Ching S, Whittington MA and Kopell N 2012 Dynamical changes in neurological diseases and anesthesia. Curr. Opin. Neurobiol. 22 693–703
Mikula S, Trotts I, Stone JM and Jones EG 2007 Internet-enabled high-resolution brain mapping and virtual microscopy. Neuroimage 35 9–15
Miller EK and Cohen JD 2001 An integrative theory of prefrontal cortex function. Annu. Rev. Neurosci. 24 167–202
Morse TM 2008 Neuroinformatics from bioinformatics to databasing the brain. Bioinform. Biol. Insights 253–264
Mountcastle VB 1998 Perceptual neuroscience: The cerebral cortex (Boston, Harvard University Press)
Moussa MN, Steen MR, Laurienti PJ and Hayasaka S 2012 Consistency of network modules in resting-state FMRI connectome data. PLoS One 7 e44428
Mueller SG, Weiner MW, Thal LJ, Petersen RC, Jack CR, Jagust W, Trojanowski JQ, Toga AW et al. 2005 Ways toward an early diagnosis in Alzheimers disease the Alzheimers Disease Neuroimaging Initiative (ADNI). Alzheimers Dement. 1 55–66
Muller AM and Meyer M 2014 Language in the brain at rest new insights from resting state data and graph theoretical analysis. Front. Hum. Neurosci. 8 228
Murre JMJ and Sturdy DPF 1995 The connectivity of the brain Multi-level quantitative analysis. Biol. Cybern. 73 529–545
Murias M, Webb SJ, Greenson J and Dawson G 2007 Resting state cortical connectivity reflected in EEG coherence in individuals with autism. Biol. Psychiatry. 62 270–273
Müller-Linow M, Hilgetag CC and Hütt MT 2008 Organization of excitable dynamics in hierarchical biological networks. PLoS Comput. Biol. 4 e1000190
Nielsen F 2014 Brede tools and federating online neuroinformatics databases. Neuroinformatics 12 27–37
Pais-Vieira M, Chiuffa G, Lebedev M, Yadav A and Nicolelis MA 2015 Building an organic computing device with multiple interconnected brains. Sci. Rep. 5 11869
Pecevski D, Natschläger T and Schuch K 2009 PCSIM a parallel simulation environment for neural circuits fully integrated with Python. Front. Neuroinform. 3 11
Peters JM, Taquet M, Vega C, Jeste SS, Fernández IS, Tan J, Nelson CA, Sahin M et al. 2013 Brain functional networks in syndromic and non-syndromic autism a graph theoretical study of EEG connectivity. BMC Med. 11 54
Piccinini G 2011 The physical Church-Turing thesis Modest or bold?. Brit. J. Philos. Sci. 62(4) 733–769
Piccinini G and Bahar S 2013 Neural computation and the computational theory of cognition. Cogn. Sci. 37 453–488
Piccinini G and Shagrir O 2014 Foundations of computational neuroscience. Curr. Opin. Neurobiol. 25 25–30
Plesser HE, Diesmann M, M.-Gewaltig O and Morrison A 2015 NEST The Neural Simulation Tool. Encyclopedia of Computational Neuroscience pp 1849–1852
Ramakrishnan A, Ifft PJ, Pais M-Vieira, Byun YW, Zhuang KZ, Lebedev MA and Nicolelis MA 2015 Computing arm movements with a monkey brainet. Sci. Rep. 5 10767
Rao RP, Stocco A, Bryan M, Sarma D, Youngquist TM, Wu J and Prat CS 2014 A direct brain-to-brain interface in humans. PLoS One 9 e111332
Ray S, Deshpande R, Dudani N and Bhalla US 2008 A general biological simulator the multiscale object oriented simulation environment, MOOSE BMC Neurosci. 9 P93
Reijmer YD, Leemans A, Brundel M, Kappelle LJ and Biessels GJ 2013 Disruption of the cerebral white matter network is related to slowing of information processing speed in patients with type 2 diabetes. Diabetes DB_121644
Robinson EC, Hammers A, Ericsson A, Edwards AD and Rueckert D 2010 Identifying population differences in whole-brain structural networks a machine learning approach. Neuroimage 50 910–919
Rubinov M and Sporns O 2010 Complex network measures of brain connectivity uses and interpretations. Neuroimage 52 1059–1069
Ruff CC, Blankenburg F, Bjoertomt O, Bestmann S, Freeman E, Haynes JD, Rees G, Josephs O, Deichmann R and Driver J 2006 Concurrent TMS-fMRI and psychophysics reveal frontal influences on human retinotopic visual cortex. Curr. Biol. 16 1479–1488
Sajda P, Gerson A, Müller KR, Blankertz B and Parra L 2003 A data analysis competition to evaluate machine learning algorithms for use in brain-computer interfaces. IEEE Trans. Neural. Syst. Rehabil. Eng. 11 184–185
Schiff ND, Nauvel T and Victor JD 2014 Large-scale brain dynamics in disorders of consciousness. Curr. Opin. Neurobiol. 25 7–14
Schwartz E 1993 Computational. Neuroscience (MIT Press)
Seeley WW, Crawford RK, Zhou J, Miller BL and Greicius MD 2009 Neurodegenerative diseases target large-scale human brain networks. Neuron 62 42–52
Sejnowski TJ, Koch C and Churchland PS 1988 Computational neuroscience. Science. 241 1299–1306
Seo EH, Lee DY, Lee JM, Park JS, Sohn BK, Lee DS, Choe YM and Woo JI 2013 Whole-brain functional networks in cognitively normal, mild cognitive impairment, and Alzheimers disease. PLoS One 8 e53922
Sherif T, Kassis N, Rousseau M, Adalat R and Evans AC 2014 BrainBrowser distributed, web-based neurological data visualization. Front. Neuroinform. 8 89
Sherrington CSS 1906 Integrative. action of.the nervous system (New Haven, Yale U.P.)
Shoeb AH 2009 Application of machine learning to epileptic seizure onset detection and treatment (Massachusetts Institute of Technology)
Smith SM, Fox PT, Miller KL, Glahn DC, Fox PM, Mackay CE, Filippini N, Watkins KE et al. 2009 Correspondence of the brains functional architecture during activation and rest. Proc. Natl. Acad. Sci. USA 106 13040–13045
Smith SM, Vidaurre D, Beckmann CF, Glasser MF, Jenkinson M, Miller KL, Nichols TE, Robinson EC et al. 2013 Functional connectomics from resting-state fMRI. Trends. Cogn. Sci. 17 666–682
Sompolinsky H 2014 Computational neuroscience beyond the local circuit. Curr. Opin. Neurobiol. 25 xiii-xviii
Sporns O 2015 Cerebral cartography and connectomics. Philos. Trans. R Soc. Lond. B Biol. Sci. 370 1668
Sporns O and Kötter R 2004 Motifs in brain networks. PLoS Biol. 2 e369
Sporns O, Tononi G, and Kötter R 2005 The human connectome a structural description of the human brain. PLoS Comput. Biol. 1 e42
Stalling D, Westerhoff M and H.-Hege C 2005 Amira a highly interactive system for visual data analysis. Visualiz. Handbook 38 749–767
Stam CJ, Jones BF, Nolte G, Breakspear M and Scheltens P 2007 Small-world networks and functional connectivity in Alzheimers disease. Cereb. Cortex. 17 92–99
Stimberg M, Goodman DF, Benichoux V and Brette R 2014 Equation-oriented specification of neural models for simulations. Front. Neuroinform. 8 6
Supekar K, Musen M and Menon V 2009 Development of large-scale functional brain networks in children. PLoS Biol. 7 e1000157
Sutton D 1999 The whole brain atlas. BMJ 319 1507
Tijms BM, Yeung HM, Sikkes SA, Möller C, Smits LL, Stam CJ, Scheltens P WM van der Flier et al. 2014 Single-subject gray matter graph properties and their relationship with cognitive impairment in early- and late-onset Alzheimers disease. Brain. Connect. 4 337–346
Timmermann L, Gross J, Dirks M, Volkmann J, Freund HJ and Schnitzler A 2003 The cerebral oscillatory network of parkinsonian resting tremor. Brain. 126 199–212
Trappenberg T 2009 Fundamentals. of computational neuroscience (OUP Oxford)
Turner JA, Lane SR, Bockholt HJ and Calhoun VD 2011 The clinical assessment and remote administration tablet. Front. Neuroinform. 5 31
Uhlhaas PJ and Singer W 2007 What do disturbances in neural synchrony tell us about autism? Biol. Psychiatry 62 190–191
Van Essen DC and Drury HA and Joshi S and Miller MI 1998 Functional and structural mapping of human cerebral cortex Solutions are in the surfaces. Proc. Natl. Acad. USA 95 788–795
Van Essen D C, Ugurbil K, Auerbach E, Barch D, Behrens TE, Bucholz R, Chang A, Chen L et al. 2012 The Human Connectome Project a data acquisition perspective. Neuroimage 62 2222–2231
Wang L, Kogan A, Cobia D, Alpert K, Kolasny A, Miller MI and Marcus D 2013 Northwestern University Schizophrenia Data and Software Tool (NUSDAST). Front. Neuroinform. 7 25
Wang L, Uhrig L, Jarraya B and Dehaene S 2015 Representation of numerical and sequential patterns in macaque and human brains. Curr. Biol. 25 1966–1974
Young MP and Scannell JW 2000 Brain structure-function relationships advances from neuroinformatics. Philos. Trans. R Soc. Lond. B Biol. Sci. 355 3–6
Zhou C, Zemanová L, Zamora G, Hilgetag CC and Kurths J 2006 Hierarchical organization unveiled by functional connectivity in complex brain networks. Phys. Rev. Lett. 97 238103
Acknowledgements
LN acknowledges University Grants Commission, India for a UGC Post-Doctoral Fellowship (No. F.15-1/2013-14/PDFWM-2013-14-GE-ORI-19068(SA-II)). AD acknowledges Digital India Corporation (formerly Media Lab Asia), Ministry of Electronics and Information Technology, Government of India, for providing him a Senior Research Fellowship under the Visvesvaraya Ph.D. scheme for Electronics and IT. RD acknowledges Council of Scientific and Industrial Research, India, for providing him a Senior Research Fellowship (No. 09/093(0182)/2018 EMR-I).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nayak, L., Dasgupta, A., Das, R. et al. Computational neuroscience and neuroinformatics: Recent progress and resources. J Biosci 43, 1037–1054 (2018). https://doi.org/10.1007/s12038-018-9813-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12038-018-9813-y