Abstract
Protein prenylation is a post-translational lipid modification that governs a variety of important cellular signaling pathways, including those regulating synaptic functions and cognition in the nervous system. Two enzymes, farnesyltransferase (FT) and geranylgeranyltransferase type I (GGT), are essential for the prenylation process. Genetic reduction of FT or GGT ameliorates neuropathology but only FT haplodeficiency rescues cognitive function in transgenic mice of Alzheimer’s disease. A follow-up study showed that systemic or forebrain neuron-specific deficiency of GGT leads to synaptic and cognitive deficits under physiological conditions. Whether FT plays different roles in shaping neuronal functions and cognition remains elusive. This study shows that in contrast to the detrimental effects of GGT reduction, systemic haplodeficiency of FT has little to no impact on hippocampal synaptic plasticity and cognition. However, forebrain neuron-specific FT deletion also leads to reduced synaptic plasticity, memory retention, and hippocampal dendritic spine density. Furthermore, a novel prenylomic analysis identifies distinct pools of prenylated proteins that are affected in the brain of forebrain neuron-specific FT and GGT knockout mice, respectively. Taken together, this study uncovers that physiological levels of FT and GGT in neurons are essential for normal synaptic/cognitive functions and that the prenylation status of specific signaling molecules regulates neuronal functions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
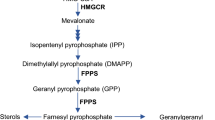
Protein prenylation is an important post-translational modification that adds lipid molecules, isoprenoids, to target proteins and governs a variety of cellular signaling pathways [1]. In the nervous system, protein prenylation mediates neurite outgrowth, dendritic morphology, and synaptic plasticity, and has been implicated in the pathogenesis of neurodegenerative diseases [2,3,4]. The substrates of protein prenylation, isoprenoid diphosphates, are products of the mevalonate/cholesterol synthesis pathway, including farnesyl pyrophosphate (FPP) and geranylgeranyl pyrophosphate (GGPP) [5]. The farnesyl and geranylgeranyl groups from FPP and GGPP, respectively, are attached to a cysteine residue in the CAAX-box motifs at the C-terminus of target proteins via an irreversible covalent bond, catalyzed by farnesyltransferase (FT) and geranylgeranyltransferase type I (GGT-I, designated as GGT throughout the paper), respectively [6]. Depending on the X amino acid of the CAAX motif, proteins are preferably or exclusively prenylated by FT or GGT, or both [7]. Another type of geranylgeranylation occurs on a family of Rab proteins catalyzed by geranylgeranyltransferase type II (GGT-II or RabGGT) and its partner Rab escort protein (REP), dually appending geranylgeranyl moieties on two proximal cysteine residues near the C-terminus [8]. Recently, another geranylgeranyltransferase type III (GGT-III) was discovered with limited substrates [9, 10].
More than a hundred proteins undergo protein prenylation, including the largest and most well-studied group, small GTP binding proteins (GTPases) [6]. Small GTPases function as molecular switches in cells and control diverse cellular activities, including receptor signaling, dendritic spine morphogenesis, cell cycle regulation, and vesicle trafficking [11, 12]. In the nervous system, the importance of small GTPases, especially the Ras family, has been underscored in neurodevelopment, synaptic plasticity, and memory formation [13, 14]. Anchoring of small GTPases to membranes is required for their normal signaling functions and protein prenylation is necessary for this process [15]. The importance of protein prenylation is further highlighted by the findings that germline homozygous deletion of FT or GGT leads to embryonic lethality and dysregulation of prenylation is involved in the pathogenesis of several diseases including Alzheimer’s disease (AD) [16].
Previous work has shown that haplodeficiency of FT but not GGT rescues cognitive deficits in a transgenic mouse model of AD, suggesting that FT and GGT may have distinct roles in regulating neuronal functions [17]. A follow-up study showed that systemic GGT haplodeficiency or forebrain neuron-specific GGT deletion is detrimental, which leads to impaired learning and memory, reduced synaptic plasticity, and decreased dendritic spine density [18]. However, it remains unclear whether FT regulates synaptic functions and cognition differentially from GGT under normal physiological conditions. The current study was undertaken to determine the impact of FT deficiency on synaptic plasticity, dendritic spine density, and cognitive function using genetically modified mouse models. The results show that systemic/germline haplodeficiency of FT does not affect hippocampal synaptic plasticity, learning and memory formation, or cortical synaptic dendritic spine density. However, forebrain neuron-specific FT deletion leads to reduced magnitude of hippocampal long-term potentiation, impaired memory retention, and reduction of dendritic spine density in the hippocampus. To explore the potential molecular mechanisms underlying different roles of FT and GGT in the brain, a novel prenylomic approach was employed to uncover changes in downstream prenylated proteins. The results show that a number of prenylated proteins, including several small GTPases, are involved. These findings suggest that certain physiological levels of FT and GGT in neurons are required for normal synaptic function and cognition, and that the prenylation status of specific signaling molecules regulates neuronal function. This study presents an innovative approach to profile prenylated proteins and downstream signaling pathways in post-mortem tissue samples as well as providing novel insights into the physiological functions of FT in the brain.
Methods
Animals
Systemic/germline FT-haplodeficient (FT+/-) mice, GGT-haplodeficient (GGT+/-) mice, and forebrain neuron-specific GGT deficient (GGTf/fCre+) mice have been described previously [17,18,19,20]. The forebrain neuron-specific FT deficiency (FTf/fCre+) was achieved by crossing the FT floxed mice (FTf/f) with the mice expressing the Cre recombinase (Cre+) under the CaMKIIα promoter [19,20,21]. All these lines of mice are on a mixed genetic background of 129/OlaHsd and C57BL/6. Specifically, FT+/- and GGT+/- were backcrossed with C57BL/6J for two generations, whereas FTf/fCre+ and GGTf/fCre+ were generated by interbreeding of offspring from crossing FTf/f or GGTf/f (129/OlaHsd/C57BL/6) and Cre+ (congenic C57BL/6) mice. Therefore, littermate controls were used whenever possible to minimize potential effects of genetic backgrounds. The mice were genotyped using DNA extracted from ear biopsies and PCR with gene-specific primers. The age of the mice ranged 9–14 months, and both males and females were included in each experiment of this study. No statistically significant sex differences were observed; therefore, data from male and female mice were combined. Experiments were conducted blinded to genotypes of the mice and all animal procedures were reviewed and approved by the Institutional Animal Care and Use Committee (IACUC) at the University of Minnesota.
Electrophysiology
Electrophysiology experiments were conducted as previously described [18, 22]. Briefly, mice were deeply anesthetized using isoflurane, followed by decapitation. Brains were dissected out and sectioned in ice-cold sucrose cutting solution containing 2.5 mM KCl, 1.25 mM NaH2PO4, 25 mM NaHCO3, 0.5 mM CaCl2, 7 mM MgCl2, 7 mM dextrose, and 240 mM sucrose. Transverse hippocampal slices were cut at 400 μm using a vibratome (Leica). Slices were then transferred and incubated for 15 min at 28 °C in artificial cerebrospinal fluid (aCSF) containing 125 mM NaCl, 2.5 mM KCl, 1.25 mM NaH2PO4, 25 mM NaHCO3, 2 mM CaCl2, 1 mM MgCl2, and 25 mM dextrose (pH 7.4) with constant bubbling of 95%O2/5% CO2. Slices were then allowed to recover for at least 1 h in aCSF at room temperature. After recovery, slices were placed into the recording chamber (Automate Scientific, Berkeley) with aCSF flowing at approximately 1.5 ml/min at 27–29 °C. Electrical stimulations were delivered to the Schaffer Collateral axon bundles of CA3 to CA1 of the hippocampus using a bipolar tungsten electrode (FHC) and evoked field excitatory post-synaptic potentials (fEPSPs) were recorded from the stratum radiatum of CA1. Stimulation was delivered at 10 s interval and the stimulation intensity was adjusted to 40–50% of each slice’s maximum input/output curve using a constant-current stimulus isolator (World Precision Instruments, Sarasota).
Following a stable baseline (at least 20 min), long-term potentiation (LTP) was induced by three trains of theta-burst stimulation (TBS) with a 20-s interval and monitored for 40 min post-induction. The fEPSP slopes were normalized to the baseline for each slice and last 5 min of response were averaged to represent the magnitude of LTP. Input/output (I/O) curves and paired-pulse facilitation (PPF) were assessed using separate slices. I/O curves were generated by delivering five stimulations per intensity ranging from 0 up to 150 μA in 10-μA intervals. Stimulation intensity was then adjusted to 40–50% of each slice’s maximum response in the I/O curve to assess PPF. Two continuous stimuli were delivered to slices with an interval of 30 ms at every 10 s. Five minutes of stable PPFs (120 PPFs) were recorded and averaged for each genotype.
Golgi Staining and Dendritic Spine Density Quantification
Golgi staining was conducted using the FD Rapid GolgiStain™ kit (Cat. #: PK401A, FD NeuroTechnologies, Inc.) as previously described [18, 23, 24]. Briefly, mice were deeply anesthetized and perfused with PBS and brains were dissected and processed following the manufacturer’s user manual. Coronal brain sections (200 μm) were prepared using a vibratome (Leica). Bright-field microscopy was done under the 100× objective using the Nikon Eclipse 55i upright microscope to capture images of dendrites in the single-plane focal field using a CCD camera (QImaging) controlled by the Image-Pro plus software. Clearly identifiable dendritic branches from randomly chosen fully stained pyramidal neurons in the layer II/III of the posterior 2/3 of the cerebral cortex and in the hippocampal CA1 region were selected. For each mouse, fifteen neurons in each area were randomly imaged and five apical and five basal dendrites were chosen from tertiary or above dendritic segments from each neuron. Imaging and spine density quantifications were achieved using Image-Pro Plus software. On average, 19 ± 0.25 μm of dendritic segment length was quantified.
Behavioral Assessment
Three behavioral functions, including locomotive response to environmental stimulus, anxiety, and spatial learning and memory, were tested as previously described [17, 18]. The open-field test and the elevated plus maze test were employed to evaluate the locomotive response and anxiety of mice. The Morris water maze was employed to evaluate the spatial learning and memory of mice.
Briefly, all behavioral functions were assessed using setups and the ANYMAZE system from San Diego Instruments. For the open field test, mice were placed into a square open box to explore freely for 5 min daily for 3 days with their path lengths recorded. The elevated plus maze test was conducted next, in which mice were placed onto a plus shape maze with open and closed arms to explore freely for 5 min daily for 2 days. In the Morris water maze test, mice were put into a water basin to find a hidden escape platform 1 cm under the water in a visual cue-rich room. The escape latency was recorded for four trials a day for five continuous days. A probe trial was carried out on the sixth day, in which the hidden platform was removed and the number of times crossing the platform location was recorded in a single 60-s trial. After a 2-h rest, a visible trial was conducted to test for visual acuity and swimming speed.
In Vitro Prenylation and Fluorescence Labeling
Brain tissue collection followed protocols previously reported [17, 18]. Right forebrain (containing cortex and hippocampus) tissue samples were homogenized in the prenylation buffer (50 mM HEPES, 50 mM NaCl, 5 mM MgCl2, 10 μM ZnCl2, 5 mM DTT (GoldBio), 2 μM phenylmethane sulfonyl fluoride (PMSF, Sigma-Aldrich), 1.5% v/v protease inhibitor cocktail, 0.02% octylglucopyranoside (OGP, Sigma-Aldrich), pH 7.2) using a Bullet Blender® homogenizer (Next Advance, Averill Park, NY, USA). Tissue lysates were centrifuged at 1500×g in 4 °C for 15 min to clear cell debris, and further centrifuged at 100,000×g in 4 °C for 60 min to pull down cell membrane fractions. Proteins from the resulting supernatants (cytosolic fractions) were quantified using BCA assay (Thermo Scientific) and 2 mg in 1 mL solutions was prepared in the prenylation buffer. Rat FT or GGT (250 to 1500 nM) and C15AlkOPP (25 μM) were added and in vitro reactions were carried out for 4 h at 37 °C. Proteins were precipitated out through chloroform precipitation (1 mL chloroform, 4 mL methanol, and 3 mL water) and washed with methanol (3 mL) twice. Protein pellets were re-dissolved in PBS + 1% SDS and concentrations were determined using BCA. Protein lysates were diluted to 100 μg/μL for click reaction. Click reaction was carried out using TAMRA-N3 by adding TAMRA-PEG4-N3 (25 μM, BroadPharm), tris(2-carboxyethyl)phosphine (TCEP, 1 mM, Sigma-Aldrich), tris[(1-benzyl-1H-1,2,3-triazol-4-yl-)methylamine (TBTA, 100 μM, Sigma-Aldrich), and CuSO4 (1 mM, Sigma-Aldrich) at room temperature for 1 h.
After the click reaction, proteins were precipitated out using ProteoExtract® protein extraction kit (Millipore Sigma) following the manufacturer’s protocol. Protein pellets were dissolved in 1x sample loading buffer containing 2% SDS, 10% glycerol, 125 mM DTT, and 0.02% bromophenol blue in 50 mM Tris-HCl pH 6.8 and resolved in a 12% SDS-PAGE gel. TAMRA fluorescence (542 nm excitation/568 nm emission) was detected and visualized on Typhoon FLA 9500 (GE Healthcare). Gel images were processed on ImageJ for visualization by adjusting contrast and brightness.
Prenylated Protein Enrichment and Sample Preparation for Proteomics
Enrichment of prenylated proteins and sample preparation was reported previously [25, 26]. Briefly, in vitro prenylated proteins (2 mg in 1 mL) were subjected to click reaction with biotin-N3 by adding biotin-PEG4-N3, TCEP, TBTA, and CuSO4. Proteins were precipitated out and dissolved in 1x PBS + 1% SDS. Protein concentrations were normalized across all samples and incubated with pre-washed Neutravidin® beads for 1.5 h, followed by washing with PBS + 1%SDS, 1X PBS, 8 M urea, and 50 mM tetraethylammonium bicarbonate (TEAB). Beads were suspended in 50 mM TEAB and digested overnight at 37 °C with trypsin (Promega). Peptides were collected from beads using 0.5% formic acid and washings (twice) were pooled and dried. Samples were dissolved in 100 mM TEAB and spiked with yeast ADH1 standard (Waters) and TMT-labeled using TMT 6plex (Thermo Fisher Scientific). Labeled samples were pooled, dried, and dissolved in 200 mM NH4HCOO pH 10, and subsequently fractionated under high pH reversed-phase conditions. The samples were dried and dissolved in 0.1% formic acid in water.
LC-MS Data Acquisition
The TMT-labeled peptides were resolved in RSLC Ultimate 3000 nano-UHPLC (Dionex) at 300 nL/min in a reversed-phase column packed in-house [25]. Each fraction ran at varying gradients of solvent B (0.1% formic acid in acetonitrile) in solvent A (0.1% formic acid in water) with amounts ranging between 7 and 34% for 80 min and sprayed directly on Orbitrap Fusion Trihybrid (Thermo Fisher Scientific) using Nanospray Flex source (Thermo Fisher Scientific). For the multinotch SPS-MS3 approach [27], MS scans were collected at 120,000 resolution at 320–2000 m/z range with max injection time (IT) of 100 ms and automatic gain control (AGC) target of 200,000. To perform SPS-MS3, subsequent data-dependent (top speed at 3 s) MS/MS scans were acquired using collision-induced dissociation (CID) at normalized collision energy (NCE) of 35% with 1.3 m/z isolation window, max IT of 100 ms, and AGC target of 5000. Dynamic exclusion was allowed for 60 s. Acquisition at MS3 was done by synchronously selecting the top 10 precursors (SPS) for fragmentation by high-collisional energy dissociation (HCD) in the orbitrap with NCE of 55% and 2.5 m/z isolation window, 120 ms max IT, and 50,000 AGC target.
Proteomic Data Analysis
The raw files were searched using Andromeda embedded in MaxQuant (version 1.6.2.10) against the non-redundant mouse (UP0000000589) database from Uniprot (EMBL-EBI, April 2018 release). The peptides identified were based on full tryptic digestion with allowed 3 maximum missed cleavages. Fixed modifications were set for the TMT labels on both the N-terminal and lysine and variable modifications on methionine oxidation and N-term acetylation. The default settings in the software for other parameters were retained.
Statistical Analysis
GraphPad Prism 8 and Perseus (version 1.6.0.7) were used to perform statistical analyses. All values are expressed as mean ± SEM. To compare differences between two genotypes, a two-tailed unpaired Student’s t test was used. For comparisons of genotype effect on mice performance over constitutive days, two-way repeated measures ANOVA was conducted. Both males and females were included in this study and no sex difference was observed; thus, male and female data were combined for analysis. For prenylomic data, the .txt file containing the list of identified proteins was imported in Perseus (version 1.6.0.7) for statistical analysis. Proteins that were identified only by site, as potential contaminant, or reversed were removed. The reporter ion intensities were log2-transformed and proteins having 3 quantified values out of 6 across TMT channels were processed. Missing values in each channel were imputed based on normal distribution. Statistical analysis was performed using two-sample t tests with FDR = 0.01 and s0 = 0.5. Data were exported and processed on Microsoft Excel to generate and format volcano plots.
Results
Systemic FT Haplodeficiency Does Not Affect Hippocampal Synaptic Function, in Contrast to the Detrimental Effect of GGT Haplodeficiency
Hippocampal LTP of CA1 is important for shaping spatial learning and memory and defined as increased signal transmission in response to an intense stimulus [28,29,30,31]. To determine whether systemic FT haplodeficiency has any effect on synaptic function, we conducted electrophysiological experiments with hippocampal slices from FT+/- mice and WT littermates at 10 to 11 months of age. The axon bundles of the Schaffer collateral from CA3 to the CA1 were stimulated and recorded from the stratum radiatum of CA1. The results show that FT haplodeficiency has no effect on the magnitude of hippocampal LTP compared with their WT littermate controls (Fig. 1a, b). The basal synaptic transmission and short-term plasticity were also assessed in FT+/- mice by constructing the I/O curve and measuring the PPF ratio, respectively. The I/O curve shows increasing fEPSPs in response to increasing stimulus intensities. No statistical difference was observed between the I/O curves from FT+/- and WT brain slices (Fig. 1c), indicating comparable basal neuronal transmission in FT+/- and WT brain slices. PPF is a form of short-term presynaptic plasticity that is defined as an increased post-synaptic response to the second stimuli of two close successive stimuli. PPF is caused by increased presynaptic Ca2+ building up when the second stimulus arrives [32]. Similarly, there was no statistically significant difference in the PPF ratio of two stimuli at a 30-ms interval between brain slices from FT+/- and WT littermates (Fig. 1d).
Systemic FT haplodeficiency does not affect hippocampal LTP, basal neuronal transmission, and PPF. a Induction of LTP. Following a 20-min stable baseline, the LTP was induced by three trains of theta-burst stimulation (TBS) protocol with a 20-s interval. The slopes of the fEPSP rising phase were calculated and normalized to baseline in hippocampal slices from WT and FT+/- mice (n = 24–26 slices/9–10 mice per genotype, 10 months of age). b The magnitude of LTP of the last 5 min (35–40 min post-induction) was averaged for each genotype. c Input/output (I/O) curves. Five stimuli were given every 10 μA ranging from 0 to 150 μA and fEPSPs were recorded and averaged at each stimulation intensity from hippocampal slices of WT and FT+/- mice (n = 12–17 slices/5–6 mice per genotype, 11 months of age). d Paired-pulse facilitation (PPF) ratios. Two continuous stimuli with an interval of 30 ms were given to WT and FT+/- hippocampal slices at every 10 s and the ratios of the second to the first fEPSP response were plotted. Five minutes of PPF (120 ratios) were recorded and averaged for each genotype (n = 12–17 slices/5 mice per genotype, 11 months of age)
Previous work has demonstrated that systemic GGT haplodeficiency leads to a significant reduction in synaptic function, including impaired LTP [18]. To compare the effects of FT and GGT haplodeficiency on synaptic plasticity side-by-side, we conducted LTP experiments on hippocampal slices from GGT+/- mice along with FT+/- mice and their respective WT littermate controls. Consistent with the results previously reported, GGT haplodeficiency leads to reduced hippocampal LTP compared with their WT controls (Fig. 2a, b), confirming the detrimental effects of GGT haplodeficiency on synaptic function. These findings demonstrate that unlike GGT haplodeficiency, FT haplodeficiency has no effects on hippocampal synaptic transmission and plasticity.
Systemic GGT haplodeficiency impairs hippocampal LTP. a Induction of LTP. Following a 20-min stable baseline, the LTP was induced by three trains of TBS protocol with a 20-s interval. The slopes of the fEPSP rising phase were calculated and normalized to baseline in hippocampal slices from WT and GGT+/- mice (n = 8–10 slices/4 mice per genotype, 13 months of age). b The magnitude of LTP of the last 5 min (35–40 min post-induction) was averaged for each genotype. **p < 0.01, Student’s t test
FT Haplodeficiency Has No Effect on Dendritic Spine Density in Cortical Neurons but Modestly Reduces Dendritic Spine Density in Hippocampal Neurons
GGT haplodeficiency has been shown to lead to remarkably reduced dendritic spine density in the cortical layer II/III pyramidal neurons [18]. Since GGT and FT differently affect synaptic plasticity, these two enzymes and downstream pathways may also differently shape synaptic morphology. To evaluate synapse morphology in FT+/- mice, we performed Golgi staining and quantified the dendritic spine density in pyramidal neurons of the layer II/III of the cortex. No statistical difference in apical or basal dendritic spine counts was observed in FT+/- mice compared with their WT littermate controls (Fig. 3a, b). This result indicates that in contrast to GGT haplodeficiency, FT haplodeficiency does not have any negative effects on dendritic spine density in cortical neurons. Since electrophysiological properties of neurons were evaluated in CA1 of the hippocampus, the dendritic spine density in pyramidal neurons of the hippocampal CA1 region was also quantified. Unexpectedly, we observed a small (~ 10%) but statistically significant decrease in the spine density on both apical and basal CA1 dendrites in FT+/- mice compared to their WT littermate controls (Fig. 3c, d). These results suggest that pyramidal neurons in hippocampus and cortex have subtle differences in tolerance towards reduction of FT.
FT haplodeficiency has no effect on dendritic spine density in cortical neurons but modestly reduces dendritic spine density in hippocampal neurons. a Representative image of apical and basal dendritic segments of cortical layer II/III pyramidal neurons in Golgi-stained brain sections from FT+/- and WT mice. b Quantification of cortical spine densities (n = 30–44 neurons/2–3 mice per genotype, 10 months of age). c Representative images of apical and basal dendrites of CA1 pyramidal neurons in Golgi-stained brain sections from FT+/- and WT mice. d Quantification of hippocampal spine densities in FT+/- and WT mice (n = 42–45 neurons/3 mice per genotype). Student’s t test, *p < 0.05; **p < 0.01
FT Haplodeficiency Does Not Affect Spatial Learning and Memory Function
Hippocampus governs spatial learning and memory and previous work has shown that GGT haplodeficiency leads to impaired spatial learning and memory formation [18]. To determine the effects of FT haplodeficiency on cognitive function, FT+/- mice and WT littermates were subjected to the Morris water maze test. As shown in Fig. 4a and b, there were no differences in the learning curve during the acquisition phase or in the platform crossover during the probe trial between FT+/- and WT mice. This result indicates FT haplodeficiency does not affect spatial memory acquisition and retention. Notably, although there was a 10% reduction of dendritic spine density in the hippocampus of FT+/- mice compared to their WT controls, there were no changes in hippocampal LTP and learning and memory performance in FT+/- mice. These findings suggest that the modest effect of FT haplodeficiency on dendritic spine density in hippocampus is not sufficient to affect synaptic plasticity and cognitive function. Furthermore, FT haplodeficiency had no significant effects on exploratory activities except for an increased entry to the center zone in the open field test, suggesting a reduced anxiety level; however, there were no differences between FT+/- and WT mice in anxiety levels assessed by the elevated plus maze test (Supplementary Table S1).
Forebrain Neuron-specific FT Deficiency Leads to Reduced Hippocampal LTP
Germline FT or GGT homozygous deletion leads to embryonic lethality, suggesting the importance of FT and GGT during development [20, 33]. To circumvent potential compensatory effects of germline FT haplodeficiency during early development and evaluate the impact of neuron-specific FT, we generated forebrain neuron-specific FT knockout (FTf/fCre+) mice by crossbreeding the FTf/f line [19] with the CAMKIIα-Cre line [21]. In these mice, Cre recombinase is expressed under the Ca2+/calmodulin-dependent protein kinase II alpha (CAMKIIα) promotor, which starts to become active late in development and drives Cre activity to increase as animals age [30, 34]. To assess the effects of neuron-specific FT deletion on synaptic function, we conducted the same sets of electrophysiological experiments on the hippocampus of these mice and their FTf/fCre− (WT) littermate controls at 9 to 10 months of age. Importantly, when we conducted hippocampal LTP experiments, FTf/fCre+ mice showed a significant decrease in the magnitude of LTP (Fig. 5a, b). Comparable I/O responses were observed in FTf/fCre+ mice and FTf/fCre− mice controls (Fig. 5c), indicating that forebrain neuron-specific FT deletion does not affect neuronal basal transmission. We next conducted PPF experiments to assess their short-term presynaptic plasticity. The PPF ratios showed a trend but not statistically significant decrease in FTf/fCre+ mice compared to their FTf/fCre− littermate controls (Fig. 5d). These results indicate that in contrast to systemic/germline FT haplodeficiency, forebrain neuronal-specific FT deletion impairs post-synaptic plasticity without affecting basal transmission and short-term presynaptic plasticity.
Forebrain neuron-specific FT deficiency leads to reduced hippocampal LTP with no significant effect on basal synaptic transmission and PPF. a Induction of LTP. Following a 20-min stable baseline, the LTP was induced by three trains of TBS protocol with a 20-s interval. The slopes of the fEPSP rising phase were calculated and normalized to baseline in hippocampal slices from FTf/fCre− and FTf/fCre+ mice (n = 14 slices/5 mice per genotype, 9 months of age). b The magnitude of LTP of the last 5 min (35–40 min post-induction) was averaged for each genotype. Student’s t test, **p < 0.01. c Input/output (I/O) curves. Five stimuli were given every 10 μA ranging from 0 to 150 μA and the fEPSPs were recorded and averaged at each stimulation intensity from hippocampal slices of FTf/fCre− and FTf/fCre+ mice (n = 16–20 slices/3 mice per genotype). d Paired-pulse facilitation (PPF) ratios. Two continuous stimuli with an interval of 30 ms were given to FTf/fCre− and FTf/fCre+ hippocampal slices at every 10 s and the ratios of the second to the first fEPSP response were plotted. Five minutes of PPF (120 ratios) were recorded and averaged for each genotype (n = 18–19 slices/3 mice per genotype, 10 months of age). Student’s t test, p = 0.051
Forebrain Neuron-specific FT Deficiency Does Not Affect Dendritic Spine Density in Cortical Neurons but Reduces Dendritic Spine Density in Hippocampal Neurons
Changes in synaptic plasticity raised the question whether FT deficiency affects neuronal structure, which may contribute to the role of FT in regulating synaptic function. We next conducted Golgi staining on brain sections of these mice and quantified the dendritic spine density in the pyramidal neurons of cortical layer II/III and hippocampus. Firstly, we found comparable cortical neuron spine density between FTf/fCre+ mice and FTf/fCre− mice (Fig. 6a, b), indicating that neuronal FT is not required for the cortex to maintain normal dendritic spine density. However, it is possible that neurons in different brain areas have different sensitivities to loss of FT. Since we evaluated electrophysiological properties in CA1 of the hippocampus, we next assessed dendritic spine density in CA1 of the hippocampus. As expected, spine density was significantly decreased on both apical (16%) and basal (13%) dendrites of hippocampal CA1 pyramidal neurons from FTf/fCre+ mice compared to their FTf/fCre− littermate controls (Fig. 6c, d). Taken together, these results indicate that neuronal FT is required to retain normal spine density in the hippocampal CA1 area but not in the cortical area. This brain regional difference suggests that FT serves distinct roles in maintaining dendritic spine density in the cortex and hippocampus.
Forebrain neuron-specific FT deficiency does not affect dendritic spine density in cortical neurons but reduces dendritic spine density in hippocampal neurons. a Representative image of apical and basal dendrites of cortical pyramidal neurons in Golgi-stained brain sections from FTf/fCre− and FTf/fCre+ mice. b Quantification of cortical spine densities (n = 45 neurons/3 mice per genotype). c Representative images of apical and basal dendrites of CA1 pyramidal neurons in Golgi-stained brain sections from FTf/fCre+ and FTf/fCre− mice. d Quantification of hippocampal spine densities (n = 25–40 neurons/3 mice per genotype, 12.5 months of age). Student’s t test, ***p < 0.001, ****p < 0.0001
Forebrain Neuron-specific FT Deficiency Does Not Affect Learning Acquisition but Leads to Impaired Memory Retention in the Morris Water Maze
Hippocampal LTP is required for normal learning and memory [31] and an apparent reduction in LTP and hippocampal spine density predicts potential learning and memory dysfunction in FTf/fCre+ mice. Therefore, we next conducted a battery of behavioral tests including the Morris water maze test to evaluate their cognitive functions. As shown in Fig. 7, FTf/fCre+ mice had a similar learning curve as their FTf/fCre− controls. However, in the probe trial, FTf/fCre+ mice displayed significantly fewer platform crossovers compared to FTf/fCre− mice. This result suggests that forebrain neuron-specific FT is required for precise memory retention but not for normal learning acquisition. No visual acuity or swimming speed difference was observed between FTf/fCre+ and FTf/fCre− mice. In addition, FTf/fCre+ mice showed similar exploratory activities and anxiety levels as FTf/fCre− mice in the open field test and the elevated plus maze test, respectively (Supplementary Table S2).
Forebrain neuron-specific FT deficiency does not affect learning acquisition but leads to impaired memory retention in the Morris water maze. a Escape latencies during the acquisition phase of the Morris water maze test (n = 11–12 mice/genotype, 9 months of age). b Platform crossover in the probe trial (n = 11–12 mice/genotype, 9 months of age). Student’s t test, *p < 0.05
Novel Prenylomic Analysis of Post-mortem Brain Tissue Reveals Distinct Unprenylated Protein Pools in Forebrain Neuron-specific FT and GGT Deficient Mice
It has been shown previously that forebrain neuron-specific GGT deletion leads to a reduced magnitude of hippocampal LTP and a reduced dendritic spine density in cortical neurons [18]. Here we showed that forebrain neuron-specific FT deletion also leads to a reduced magnitude of hippocampal LTP and a reduced dendritic spine density in hippocampal neurons but not in cortical neurons. In addition, neuron-specific FT deficiency only impairs memory retention but not learning acquisition. The difference between FT and GGT in regulating neuronal functions, morphology, and cognitive functions indicates that unprenylated protein pools might be differently affected by FT or GGT deletion. To further investigate this idea, we developed a novel prenylomic approach to profile downstream prenylated proteins in post-mortem brain tissues from FTf/fCre+ and GGTf/fCre+ mice in comparison with their respective Cre− controls. We extracted proteins from forebrain tissue lysates, which were then subjected to in vitro prenylation with an alkyne-modified isoprenoid probe analog at different concentrations of recombinant FT or GGT enzymes [35]. This single isoprenoid analogue can be efficiently incorporated by both FT and GGT in lieu of FPP or GGPP as shown in vitro and in metabolic labeling experiments [26, 36]. Subsequent “click” reaction with an azide-modified fluorophore or biotin enables visualization or enrichment, respectively, of proteins labeled by the probe.
Forebrain neuron-specific FT or GGT deletion/knockout is expected to give rise to enlarged unprenylated protein pools compared to their Cre− controls. Supplementing brain lysates with the isoprenoid probe and exogenous FT or GGT allows the incorporation of the probe into unprenylated proteins for further identification. Since Cre− controls have endogenous FT or GGT, their unprenylated protein pools should be smaller and have reduced in vitro labeling. To assess the levels of unprenylated proteins in the brain lysates, we firstly conducted fluorescence labeling using TAMRA-N3 with the knockouts and their corresponding controls. Labeled lysates were resolved by SDS-PAGE and TAMRA fluorescence was detected (Fig. 8a and b). Although background labeling was apparent in both knockouts and their corresponding controls across all concentrations of the enzyme tested, FTf/fCre+ and GGTf/fCre+ displayed differences in the banding pattern compared with their controls. For FTf/fCre+, an intense band was observed near 50 kDa region, and bands near the 20 and 25 kDa regions were present, which were not detected in the FTf/fCre− control (Fig. 8a). On the other hand, minor differences were observed between GGTf/fCre+ and GGTf/fCre−, manifested by the extra bands observed above and below the 20 kDa region in GGTf/fCre+ (Fig. 8b). These data suggest that the deletion of prenyltransferases results in enlarged pools of unprenylated proteins.
Novel prenylomic analysis reveals distinct unprenylated protein pools in forebrain neuron-specific FT and GGT deficient mice. In-gel fluorescence analysis of protein lysates labeled with C15AlkOPP probe from FTf/fCre+ and FTf/fCre− (a) or GGTf/fCre+ and GGTf/fCre− (b) at varying concentrations of supplemented FT and GGT, respectively. Arrows indicate bands observed in Cre+ but not in Cre−, indicating in vitro probe incorporation to residual unprenylated proteins from Cre+ tissues. Volcano plots (FDR = 0.01, s0 = 0.5) generated from prenylomic analysis of labeled and enriched prenylated proteins from FTf/fCre+ and FTf/fCre− (c) or GGTf/fCre+ and GGTf/fCre− (d). Fold-changes in the x-axis reflect abundance of proteins from Cre+ against their Cre− after enrichment. Known farnesylated (Far, red circles) and geranylgeranylated (GG, blue circles) proteins known to be substrates of FT and GGT are noted (n = 3 mice/genotype, 14 months of age)
After successfully labeling natively unprenylated proteins in the brain lysates with our probe, we next investigated the molecular identities of these proteins. Proteins from the knockouts and their controls that were in vitro prenylated with the isoprenoid probe in the presence of exogenous FT or GGT were subjected to “click” reaction with biotin-N3. Since there were no significant differences in labeling observed across the enzyme concentrations tested, we selected 250 nM FT or 500 nM GGT as optimal enzyme concentrations for in vitro prenylation. Biotinylated lysates were subjected to enrichment using avidin beads and the isolated proteins were digested into peptides and prepared for proteomic analysis. A multinotch SPS-MS3 approach was employed in the LC-MS analysis to accurately quantify the extent of enrichment of the prenylated proteins identified [27]. A volcano plot (FDR = 0.01, s0 = 0.5) was generated to visualize enriched proteins from FTf/fCre+ (Fig. 8c) or GGTf/fCre+ (Fig. 8d) versus their corresponding Cre− controls. Eleven and five statistically enriched known farnesylated and geranylgeranylated proteins, including several well-known small GTPases, were identified from the FTf/fCre+ and GGTf/fCre+ samples, respectively. The identities and functions of those proteins are summarized in Table 1 and Table 2. Other known prenylated proteins such as Uchl1, DiRas, Cnp, and Cdc42 were identified but were not significantly enriched (Supplementary Table S3 and S4). Most of the enriched farnesylated proteins in FTf/fCre+ have molecular weights close to 50 and 20 kDa while those from GGTf/fCre+ have 20 kDa or lower, corroborating the gel-based labeling observed in Fig. 8a, b. Moreover, the most enriched farnesylated proteins in FTf/fCre+ have molecular weights close to 50 kDa, further validating the intense labeling clearly seen in the 50 kDa region in Fig. 8a. These proteins perhaps represent the most significantly affected prenylated proteins detectable on this LC-MS-based proteomic approach upon FT or GGT deletion. As shown in the volcano plot, several proteins have been identified and there is little overlap between FTf/fCre+ and GGTf/fCre+ enriched unprenylated protein pools. These results indicate that forebrain neuron-specific FT and GGT act on different downstream prenylated protein pools, which may explain functional differences observed in these mice.
Discussion
In this study, the impact of systemic/germline haplodeficiency and forebrain neuron-specific deficiency of FT on synaptic and cognitive function has been examined. We found that FT haplodeficiency had minimal to no effects on synaptic plasticity, dendritic spine density, and learning and memory in the Morris water maze. These results are in stark contrast with those from GGT-haplodeficient mice, in which both synaptic/cognitive function and spine structure are impaired, as shown previously [18] and in the present study. These findings indicate that the physiological level of GGT is more critical than that of FT for normal brain function and suggest that farnesylated and geranylgeranylated proteins play distinct roles in shaping neuronal and cognitive function. We further showed that forebrain neuron-specific FT deficiency led to impaired hippocampal LTP, similar to the effect of forebrain neuron-specific GGT deficiency indicating that certain levels of FT and GGT in neurons are required for normal synaptic function. In addition, forebrain neuron-specific FT deficiency reduced dendritic spine density in CA1 but not in cortex, and it impaired memory retention but not learning acquisition. These results indicate that reduction of neuronal FT exerts differential effects on different areas of the brain and on different aspects of cognitive function. Alternatively, actual FT levels could be differently affected in the cortical and CA1 pyramidal neurons in FTf/fCre+ mice. Despite this possibility, the likelihood of this scenario is small, as CaMKIIα is highly expressed similarly in the pyramidal neurons of cortical layer II/III and hippocampus CA1 [74]. In addition, CaMKIIα-Cre mice have been well characterized and Cre recombinase-mediated gene deletion in the cortex and hippocampus follows the pattern of CaMKIIα expression in adult brains and displays age-dependent activity [34, 75, 76]. Therefore, FT levels are expected to reduce similarly in cortical and hippocampal neurons.
Further investigation is required to gain more insights into the molecular mechanisms underlying the region- and function-specific effects. Interestingly, although both FT-haplodeficient and forebrain neuron-specific FT deficient mice displayed reduced dendritic spine density in CA1, hippocampal LTP and memory retention were only impaired in neuron-specific FT deficient mice. It is possible that a 10% reduction of dendritic spine density in FT-haplodeficient mice was not sufficient to cause measurable functional consequence although 13–16% reduction in neuron-specific FT knockout was detrimental. It is also possible that hippocampal neurons in systemic/germline haplodeficient FT mice have developed some compensatory mechanisms to maintain synaptic plasticity and cognitive function. Additionally, it is worth noting that all cell types besides neurons have partial loss of FT in FT+/- mice, whereas FT deletion was specific to CaMKIIα expressing forebrain excitatory neurons in the FTf/fCre+ mice. Therefore, the difference in cell types affected by FT reduction and the level of FT reduction may both contribute to differential functional outcomes observed in FT+/- mice and FTf/fCre+ mice. Future studies employing other conditional cell type–specific FT deletion strategies should provide more insights into the gene dosage effect of FT and the role of FT in other brain cell types.
Functions of FT and GGT in the nervous system have been explored using pharmacological approaches, which are prone to off-target effects and limitations of not separating FT and GGT pathways [2, 77,78,79,80]. To parse out roles of FT and GGT in the brain, we utilized genetic approaches by utilizing FT and GGT haplodeficient and forebrain neuron-specific knockout mice, respectively, to better understand the role of these enzymes in shaping neuronal functions and regulating cognition. Intriguingly, GGT haplodeficiency but not FT haplodeficiency leads to a reduced magnitude of hippocampal LTP, whereas both forebrain neuron-specific GGT and FT knockouts cause hippocampal LTP impairment. These data indicate that neurons may have different levels of tolerance to the suppression of FT compared to that of GGT under the condition that the same levels of FT and GGT are reduced in neurons in these mice. Since a pool of proteins can be prenylated by both GGT and FT, it is possible that in the absence of one prenylation enzyme, proteins can be processed via the other prenylation pathway [81]. The differential effects were likely caused by the suppression of proteins that are exclusively prenylated by one enzyme. In addition, enzyme kinetics studies have shown that FT manifests higher catalytic efficiency than GGT [82, 83]; thus, it is possible that stronger suppression of FT is needed to affect signaling cascades of the farnesylated protein pool. Notably, we have shown that the levels of FT and GGT are reduced similarly in the brain of FT+/- and GGT+/- mice, and FT haplodeficiency, but not GGT haplodeficiency, is sufficient to rescue cognitive function as well as AD pathology in a mouse model of AD [17]. These results suggest that FT is dysregulated in the presence of AD pathology and reducing FT levels to the normal range but not to detrimental levels in the brain ameliorates AD pathology. Future studies are warranted to investigate the dysregulation of FT and its downstream signaling cascades that contribute to AD pathogenesis, which may lead to new treatment avenues.
To investigate prenylated protein pools and signaling cascades affected by forebrain neuron-specific FT or GGT knockout, we developed a novel prenylomic approach—an in vitro prenylation assay for use with post-mortem tissue samples. To the best of our knowledge, no prenylomic analysis on post-mortem brain tissue has ever been reported prior to this study. By extracting proteins from brain lysates and subjecting them to in vitro labeling and mass spectrometric proteomics analysis, it was possible to identify a pool of unprenylated proteins in forebrain neuron-specific FT or GGT knockouts. There was little overlap between unprenylated protein pools from these two groups of mice, consistent with the substrate specificities of FT and GGT. The results also suggest that different downstream signaling pathways may be affected by the loss of FT or GGT in neurons. Previous efforts employing in vitro prenylation of FT, GGT, and GGT-II substrates used cellular models and relied on the use of a biotinylated analogue of the isoprenoid (BGPP) [84]. Such a method works well for detecting GGT-II substrates but not for proteins prenylated by FT and GGT [85]. While that method, which employs a biotinylated analogue, circumvents the need for the subsequent derivatization step employed here (after enzymatic labeling), engineered mutants of FT and GGT were required to efficiently incorporate the bulky BGPP; the effects of those mutations on enzyme specificity remain unclear. In addition, in those studies, Western blot analysis for the biotin tag on FT and GGT substrates was used to provide qualitative assessment of in vitro labeling efficiency in cell lysates, and the molecular identities of the FT or GGT substrates were not pursued. Therefore, the prenylomic analysis reported here is the first investigation of profiling FT and GGT substrates in brain tissue facilitated by genetic deletion of these prenylation enzymes in mice [84, 86].
More than 100 proteins are known to undergo the three classes of protein prenylation [3] but a recent study reported 32 known and novel farnesylated proteins and 19 GGT substrates in a human endothelial cell line via probe metabolic labeling [81]. In the present study, it was found that subsets of these proteins (11 farnesylated and 5 geranylgeranylated) were statistically enriched as unprenylated in forebrain neuron-specific FT or GGT knockout mice. As summarized in Tables 1 and 2, some of those proteins play important roles pertinent to neuronal functions, which may contribute to the structural and functional differences observed between neuron-specific FT or GGT knockout mice and their controls. For instance, Rheb, an exclusively farnesylated small GTPase, is significantly affected by forebrain neuron-specific FT deletion. Rheb has shown to regulate neuronal morphology, synaptic plasticity, and memory functions [56,57,58, 87, 88]. It is likely that abrogation of Rheb prenylation results in reduced Rheb signaling, which may have contributed to reduced synaptic and cognitive function observed in FTf/fCre+ mice. In addition, Rac1 and RhoA are major geranylgeranylated proteins and their signaling cascades are important for shaping the neuronal cytoskeleton that determines neuronal morphology and functions [89]. Importantly, using both genetic and pharmacological approaches, recent studies have clearly demonstrated the critical roles of RhoA signaling in regulating dendritic spine structure and function in the brain [60,61,62]. Perturbation of signaling pathways mediated by these small GTPases via reduction in prenylation could contribute to structural and functional deficits in GGT knockout mice. Future studies are needed to dissect out the underlying molecular pathways of FT and GGT that regulate neuronal functions.
In this prenylomic analysis, other known prenylated proteins that did not reach statistical significance for enrichment were also identified. In particular, Uchl1, a membrane-associated protein that has been implicated in Parkinson’s disease was identified [90]. Interestingly, while a previous study supported the farnesylation of Uchl1 using mutational experiments or treatment with a farnesyltransferase inhibitor, a more recent study suggests that its C-terminal CAAX-box is not farnesylated in vitro [81]. In the current study, Uchl1 may have not been enriched due to its inability to be farnesylated in vitro or it is equally farnesylated in both FTf/fCre+ and FTf/fCre− mice. Further investigations are required to elucidate the farnesylation status of Uchl1 and its role in health and disease.
Compared to the metabolic labeling approach, which leverages the cellular machinery for probe incorporation and allows the tagging and enrichment of multiple proteins in a single experiment [25, 26, 91], the in vitro labeling used in the present study relies on the integrity of the unprenylated and soluble forms of the protein substrates in the lysate. In particular, since brain tissues with only neuron-specific FT or GGT knockouts were used, the prenylated form of proteins from other cell types may suppress the apparent abundance of the unprenylated form in soluble lysate. This limits the ability to enrich those unprenylated proteins with inherently low abundance in the lysate. Gene expression of proteins is also controlled with some having more stable expression than others [92]. The prenylation substrates detected here that remained unprenylated may have persisted at sufficient levels under steady-state conditions post-mortem compared to others. Furthermore, a significant fraction of GGT substrates belongs to the family of small GTPases, which are prenylated as complexes with chaperone inside cells [93]. While it was possible to detect some of these small GTPases using the in vitro approach, others might require these chaperone-complexed forms to be efficiently prenylated in vitro for detection. It is important to note that despite its limitations, the in vitro prenylomic approach reported here identifies differentially prenylated proteins in the brain, reflecting the effect of neuronal FT and GGT deletion in vivo. Here, we describe a tool that enables studying protein prenylation in post-mortem tissues.
In summary, the present study provides novel insights into the role of systemic and neuronal FT in regulating synaptic plasticity and cognitive function. It also offers an innovative approach for future studies to unravel physiological and pathological roles of prenylated proteins.
Data availability
Raw data files are available upon request.
References
Lane KT, Beese LS (2006) Thematic review series: lipid posttranslational modifications. Structural biology of protein farnesyltransferase and geranylgeranyltransferase type I. J Lipid Res 47(4):681–699. https://doi.org/10.1194/jlr.R600002-JLR200
Li H, Kuwajima T, Oakley D, Nikulina E, Hou J, Yang WS, Lowry ER, Lamas NJ et al (2016) Protein prenylation constitutes an endogenous brake on axonal growth. Cell Rep 16(2):545–558. https://doi.org/10.1016/j.celrep.2016.06.013
Jeong A, Suazo KF, Wood WG, Distefano MD, Li L (2018) Isoprenoids and protein prenylation: implications in the pathogenesis and therapeutic intervention of Alzheimer’s disease. Crit Rev Biochem Mol Biol 53(3):279–310. https://doi.org/10.1080/10409238.2018.1458070
Lee HJ, Kang SJ, Lee K, Im H (2011) Human α-synuclein modulates vesicle trafficking through its interaction with prenylated Rab acceptor protein 1. Biochem Biophys Res Commun 412(4):526–531
Goldstein JL, Brown MS (1990) Regulation of the mevalonate pathway. Nature 343(6257):425–430. https://doi.org/10.1038/343425a0
McTaggart SJ (2006) Isoprenylated proteins. Cell Mol Life Sci 63(3):255–267. https://doi.org/10.1007/s00018-005-5298-6
Gao J, Liao J, Yang GY (2009) CAAX-box protein, prenylation process and carcinogenesis. Am J Transl Res 1(3):312–325
Kinsella BT, Maltese WA (1992) rab GTP-binding proteins with three different carboxyl-terminal cysteine motifs are modified in vivo by 20-carbon isoprenoids. J Biol Chem 267(6):3940–3945
Kuchay S, Wang H, Marzio A, Jain K, Homer H, Fehrenbacher N, Philips MR, Zheng N et al (2019) GGTase3 is a newly identified geranylgeranyltransferase targeting a ubiquitin ligase. Nat Struct Mol Biol 26(7):628–636
Shirakawa R, Goto-Ito S, Goto K, Wakayama S, Kubo H, Sakata N, Trinh DA, Yamagata A et al (2020) A SNARE geranylgeranyltransferase essential for the organization of the Golgi apparatus. EMBO J 39(8):e104120. https://doi.org/10.15252/embj.2019104120
Hooff GP, Wood WG, Muller WE, Eckert GP (2010) Isoprenoids, small GTPases and Alzheimer’s disease. Biochim Biophys Acta 1801(8):896–905. https://doi.org/10.1016/j.bbalip.2010.03.014
Hottman DA, Li L (2014) Protein prenylation and synaptic plasticity: implications for Alzheimer’s disease. Mol Neurobiol 50(1):177–185. https://doi.org/10.1007/s12035-013-8627-z
Ryu HH, Lee YS (2016) Cell type-specific roles of RAS-MAPK signaling in learning and memory: implications in neurodevelopmental disorders. Neurobiol Learn Mem 135:13–21. https://doi.org/10.1016/j.nlm.2016.06.006
Ye X, Carew TJ (2010) Small G protein signaling in neuronal plasticity and memory formation: the specific role of ras family proteins. Neuron 68(3):340–361. https://doi.org/10.1016/j.neuron.2010.09.013
Takai Y, Sasaki T, Matozaki T (2001) Small GTP-binding proteins. Physiol Rev 81(1):153–208. https://doi.org/10.1152/physrev.2001.81.1.153
Li L, Zhang W, Cheng S, Cao D, Parent M (2012) Isoprenoids and related pharmacological interventions: potential application in Alzheimer’s disease. Mol Neurobiol 46(1):64–77. https://doi.org/10.1007/s12035-012-8253-1
Cheng S, Cao D, Hottman DA, Yuan L, Bergo MO, Li L (2013) Farnesyltransferase haplodeficiency reduces neuropathology and rescues cognitive function in a mouse model of Alzheimer disease. J Biol Chem 288(50):35952–35960. https://doi.org/10.1074/jbc.M113.503904
Hottman D, Cheng S, Gram A, LeBlanc K, Yuan LL, Li L (2018) Systemic or forebrain neuron-specific deficiency of geranylgeranyltransferase-1 impairs synaptic plasticity and reduces dendritic spine density. Neuroscience 373:207–217. https://doi.org/10.1016/j.neuroscience.2018.01.026
Liu M, Sjogren AK, Karlsson C, Ibrahim MX, Andersson KM, Olofsson FJ, Wahlstrom AM, Dalin M et al (2010) Targeting the protein prenyltransferases efficiently reduces tumor development in mice with K-RAS-induced lung cancer. Proc Natl Acad Sci U S A 107(14):6471–6476. https://doi.org/10.1073/pnas.0908396107
Sjogren AK, Andersson KM, Liu M, Cutts BA, Karlsson C, Wahlstrom AM, Dalin M, Weinbaum C et al (2007) GGTase-I deficiency reduces tumor formation and improves survival in mice with K-RAS-induced lung cancer. J Clin Invest 117(5):1294–1304. https://doi.org/10.1172/JCI30868
Tsien JZ, Chen DF, Gerber D, Tom C, Mercer EH, Anderson DJ, Mayford M, Kandel ER et al (1996) Subregion- and cell type-restricted gene knockout in mouse brain. Cell 87(7):1317–1326. https://doi.org/10.1016/s0092-8674(00)81826-7
Parent MA, Hottman DA, Cheng S, Zhang W, McMahon LL, Yuan LL, Li L (2014) Simvastatin treatment enhances NMDAR-mediated synaptic transmission by upregulating the surface distribution of the GluN2B subunit. Cell Mol Neurobiol 34(5):693–705. https://doi.org/10.1007/s10571-014-0051-z
Dumanis SB, Tesoriero JA, Babus LW, Nguyen MT, Trotter JH, Ladu MJ, Weeber EJ, Turner RS et al (2009) ApoE4 decreases spine density and dendritic complexity in cortical neurons in vivo. J Neurosci 29(48):15317–15322. https://doi.org/10.1523/JNEUROSCI.4026-09.2009
Adlard PA, Bica L, White AR, Nurjono M, Filiz G, Crouch PJ, Donnelly PS, Cappai R et al (2011) Metal ionophore treatment restores dendritic spine density and synaptic protein levels in a mouse model of Alzheimer’s disease. PLoS One 6(3):e17669. https://doi.org/10.1371/journal.pone.0017669
Brandt AC, McNally L, Lorimer EL, Unger B, Koehn OJ, Suazo KF, Rein L, Szabo A et al (2020) Splice switching an oncogenic ratio of SmgGDS isoforms as a strategy to diminish malignancy. Proc Natl Acad Sci U S A 117(7):3627–3636. https://doi.org/10.1073/pnas.1914153117
Suazo KF, Schaber C, Palsuledesai CC, John ARO, Distefano MD (2016) Global proteomic analysis of prenylated proteins in Plasmodium falciparum using an alkyne-modified isoprenoid analogue. Sci Rep 6:38615
McAlister GC, Nusinow DP, Jedrychowski MP, Wuhr M, Huttlin EL, Erickson BK, Rad R, Haas W et al (2014) MultiNotch MS3 enables accurate, sensitive, and multiplexed detection of differential expression across cancer cell line proteomes. Anal Chem 86(14):7150–7158. https://doi.org/10.1021/ac502040v
Kumar A (2011) Long-term potentiation at CA3–CA1 hippocampal synapses with special emphasis on aging, disease, and stress. Front Aging Neurosci 3:7
Okada T, Yamada N, Tsuzuki K, Horikawa HP, Tanaka K, Ozawa S (2003) Long-term potentiation in the hippocampal CA1 area and dentate gyrus plays different roles in spatial learning. Eur J Neurosci 17(2):341–349. https://doi.org/10.1046/j.1460-9568.2003.02458.x
Tsien JZ, Huerta PT, Tonegawa S (1996) The essential role of hippocampal CA1 NMDA receptor-dependent synaptic plasticity in spatial memory. Cell 87(7):1327–1338. https://doi.org/10.1016/s0092-8674(00)81827-9
McGaugh JL (2000) Memory--a century of consolidation. Science 287(5451):248–251. https://doi.org/10.1126/science.287.5451.248
Regehr WG (2012) Short-term presynaptic plasticity. Cold Spring Harb Perspect Biol 4(7):a005702. https://doi.org/10.1101/cshperspect.a005702
Mijimolle N, Velasco J, Dubus P, Guerra C, Weinbaum CA, Casey PJ, Campuzano V, Barbacid M (2005) Protein farnesyltransferase in embryogenesis, adult homeostasis, and tumor development. Cancer Cell 7(4):313–324. https://doi.org/10.1016/j.ccr.2005.03.004
Liu Q, Trotter J, Zhang J, Peters MM, Cheng H, Bao J, Han X, Weeber EJ et al (2010) Neuronal LRP1 knockout in adult mice leads to impaired brain lipid metabolism and progressive, age-dependent synapse loss and neurodegeneration. J Neurosci 30(50):17068–17078. https://doi.org/10.1523/JNEUROSCI.4067-10.2010
DeGraw AJ, Palsuledesai C, Ochocki JD, Dozier JK, Lenevich S, Rashidian M, Distefano MD (2010) Evaluation of alkyne-modified isoprenoids as chemical reporters of protein prenylation. Chem Biol Drug Des 76(6):460–471
Zhang Y, Blanden MJ, Sudheer C, Gangopadhyay SA, Rashidian M, Hougland JL, Distefano MD (2015) Simultaneous site-specific dual protein labeling using protein prenyltransferases. Bioconjug Chem 26(12):2542–2553
Mu R, Dussupt V, Jiang J, Sette P, Rudd V, Chuenchor W, Bello NF, Bouamr F et al (2012) Two distinct binding modes define the interaction of Brox with the C-terminal tails of CHMP5 and CHMP4B. Structure 20(5):887–898. https://doi.org/10.1016/j.str.2012.03.001
Marschang P, Brich J, Weeber EJ, Sweatt JD, Shelton JM, Richardson JA, Hammer RE, Herz J (2004) Normal development and fertility of knockout mice lacking the tumor suppressor gene LRP1b suggest functional compensation by LRP1. Mol Cell Biol 24(9):3782–3793. https://doi.org/10.1128/mcb.24.9.3782-3793.2004
Orthwein A, Zahn A, Methot SP, Godin D, Conticello SG, Terada K, Di Noia JM (2012) Optimal functional levels of activation-induced deaminase specifically require the Hsp40 DnaJa1. EMBO J 31(3):679–691. https://doi.org/10.1038/emboj.2011.417
Rauch JN, Gestwicki JE (2014) Binding of human nucleotide exchange factors to heat shock protein 70 (Hsp70) generates functionally distinct complexes in vitro. J Biol Chem 289(3):1402–1414. https://doi.org/10.1074/jbc.M113.521997
Imai Y, Soda M, Hatakeyama S, Akagi T, Hashikawa T, Nakayama KI, Takahashi R (2002) CHIP is associated with Parkin, a gene responsible for familial Parkinson’s disease, and enhances its ubiquitin ligase activity. Mol Cell 10(1):55–67. https://doi.org/10.1016/s1097-2765(02)00583-x
Gotoh T, Terada K, Oyadomari S, Mori M (2004) hsp70-DnaJ chaperone pair prevents nitric oxide- and CHOP-induced apoptosis by inhibiting translocation of Bax to mitochondria. Cell Death Differ 11(4):390–402. https://doi.org/10.1038/sj.cdd.4401369
Hageman J, van Waarde MA, Zylicz A, Walerych D, Kampinga HH (2011) The diverse members of the mammalian HSP70 machine show distinct chaperone-like activities. Biochem J 435(1):127–142. https://doi.org/10.1042/BJ20101247
Edwards MC, Liegeois N, Horecka J, DePinho RA, Sprague GF Jr, Tyers M, Elledge SJ (1997) Human CPR (cell cycle progression restoration) genes impart a Far- phenotype on yeast cells. Genetics 147(3):1063–1076
Pencheva N, Tran H, Buss C, Huh D, Drobnjak M, Busam K, Tavazoie SF (2012) Convergent multi-miRNA targeting of ApoE drives LRP1/LRP8-dependent melanoma metastasis and angiogenesis. Cell 151(5):1068–1082. https://doi.org/10.1016/j.cell.2012.10.028
Rual JF, Venkatesan K, Hao T, Hirozane-Kishikawa T, Dricot A, Li N, Berriz GF, Gibbons FD et al (2005) Towards a proteome-scale map of the human protein-protein interaction network. Nature 437(7062):1173–1178. https://doi.org/10.1038/nature04209
Rolland T, Tasan M, Charloteaux B, Pevzner SJ, Zhong Q, Sahni N, Yi S, Lemmens I et al (2014) A proteome-scale map of the human interactome network. Cell 159(5):1212–1226. https://doi.org/10.1016/j.cell.2014.10.050
Huttlin EL, Bruckner RJ, Paulo JA, Cannon JR, Ting L, Baltier K, Colby G, Gebreab F et al (2017) Architecture of the human interactome defines protein communities and disease networks. Nature 545(7655):505–509. https://doi.org/10.1038/nature22366
Gaudet P, Livstone MS, Lewis SE, Thomas PD (2011) Phylogenetic-based propagation of functional annotations within the Gene Ontology consortium. Brief Bioinform 12(5):449–462. https://doi.org/10.1093/bib/bbr042
O’Leary NA, Wright MW, Brister JR, Ciufo S, Haddad D, McVeigh R, Rajput B, Robbertse B et al (2016) Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res 44(D1):D733–D745. https://doi.org/10.1093/nar/gkv1189
Simon HU, Mills GB, Kozlowski M, Hogg D, Branch D, Ishimi Y, Siminovitch KA (1994) Molecular characterization of hNRP, a cDNA encoding a human nucleosome-assembly-protein-I-related gene product involved in the induction of cell proliferation. Biochem J 297(Pt 2):389–397. https://doi.org/10.1042/bj2970389
Rodriguez P, Munroe D, Prawitt D, Chu LL, Bric E, Kim J, Reid LH, Davies C et al (1997) Functional characterization of human nucleosome assembly protein-2 (NAP1L4) suggests a role as a histone chaperone. Genomics 44(3):253–265. https://doi.org/10.1006/geno.1997.4868
Jones JM, Morrell JC, Gould SJ (2004) PEX19 is a predominantly cytosolic chaperone and import receptor for class 1 peroxisomal membrane proteins. J Cell Biol 164(1):57–67. https://doi.org/10.1083/jcb.200304111
Jones JM, Morrell JC, Gould SJ (2001) Multiple distinct targeting signals in integral peroxisomal membrane proteins. J Cell Biol 153(6):1141–1150. https://doi.org/10.1083/jcb.153.6.1141
Delille HK, Schrader M (2008) Targeting of hFis1 to peroxisomes is mediated by Pex19p. J Biol Chem 283(45):31107–31115. https://doi.org/10.1074/jbc.M803332200
Sugiura H, Yasuda S, Katsurabayashi S, Kawano H, Endo K, Takasaki K, Iwasaki K, Ichikawa M et al (2015) Rheb activation disrupts spine synapse formation through accumulation of syntenin in tuberous sclerosis complex. Nat Commun 6:6842. https://doi.org/10.1038/ncomms7842
Yamagata K, Sanders LK, Kaufmann WE, Yee W, Barnes CA, Nathans D, Worley PF (1994) rheb, a growth factor- and synaptic activity-regulated gene, encodes a novel Ras-related protein. J Biol Chem 269(23):16333–16339
Brown HL, Kaun KR, Edgar BA (2012) The small GTPase Rheb affects central brain neuronal morphology and memory formation in Drosophila. PLoS One 7(9):e44888. https://doi.org/10.1371/journal.pone.0044888
Tee AR, Manning BD, Roux PP, Cantley LC, Blenis J (2003) Tuberous sclerosis complex gene products, Tuberin and Hamartin, control mTOR signaling by acting as a GTPase-activating protein complex toward Rheb. Curr Biol 13(15):1259–1268. https://doi.org/10.1016/s0960-9822(03)00506-2
Swanger SA, Mattheyses AL, Gentry EG, Herskowitz JH (2015) ROCK1 and ROCK2 inhibition alters dendritic spine morphology in hippocampal neurons. Cell Logist 5(4):e1133266. https://doi.org/10.1080/21592799.2015.1133266
Henderson BW, Greathouse KM, Ramdas R, Walker CK, Rao TC, Bach SV, Curtis KA, Day JJ et al (2019) Pharmacologic inhibition of LIMK1 provides dendritic spine resilience against beta-amyloid. Sci Signal 12(587):eaaw9318. https://doi.org/10.1126/scisignal.aaw9318
Greathouse KM, Boros BD, Deslauriers JF, Henderson BW, Curtis KA, Gentry EG, Herskowitz JH (2018) Distinct and complementary functions of rho kinase isoforms ROCK1 and ROCK2 in prefrontal cortex structural plasticity. Brain Struct Funct 223(9):4227–4241. https://doi.org/10.1007/s00429-018-1748-4
Kozma R, Sarner S, Ahmed S, Lim L (1997) Rho family GTPases and neuronal growth cone remodelling: relationship between increased complexity induced by Cdc42Hs, Rac1, and acetylcholine and collapse induced by RhoA and lysophosphatidic acid. Mol Cell Biol 17(3):1201–1211. https://doi.org/10.1128/mcb.17.3.1201
Suzukawa K, Miura K, Mitsushita J, Resau J, Hirose K, Crystal R, Kamata T (2000) Nerve growth factor-induced neuronal differentiation requires generation of Rac1-regulated reactive oxygen species. J Biol Chem 275(18):13175–13178. https://doi.org/10.1074/jbc.275.18.13175
Tashiro A, Yuste R (2004) Regulation of dendritic spine motility and stability by Rac1 and Rho kinase: evidence for two forms of spine motility. Mol Cell Neurosci 26(3):429–440. https://doi.org/10.1016/j.mcn.2004.04.001
Wiens KM, Lin H, Liao D (2005) Rac1 induces the clustering of AMPA receptors during spinogenesis. J Neurosci 25(46):10627–10636. https://doi.org/10.1523/JNEUROSCI.1947-05.2005
Schwindinger WF, Giger KE, Betz KS, Stauffer AM, Sunderlin EM, Sim-Selley LJ, Selley DE, Bronson SK et al (2004) Mice with deficiency of G protein gamma3 are lean and have seizures. Mol Cell Biol 24(17):7758–7768. https://doi.org/10.1128/MCB.24.17.7758-7768.2004
Dasgupta S, Cushman I, Kpetemey M, Casey PJ, Vishwanatha JK (2011) Prenylated c17orf37 induces filopodia formation to promote cell migration and metastasis. J Biol Chem 286(29):25935–25946. https://doi.org/10.1074/jbc.M111.254599
Maekawa M, Ishizaki T, Boku S, Watanabe N, Fujita A, Iwamatsu A, Obinata T, Ohashi K et al (1999) Signaling from Rho to the actin cytoskeleton through protein kinases ROCK and LIM-kinase. Science 285(5429):895–898. https://doi.org/10.1126/science.285.5429.895
Troca-Marin JA, Alves-Sampaio A, Tejedor FJ, Montesinos ML (2010) Local translation of dendritic RhoA revealed by an improved synaptoneurosome preparation. Mol Cell Neurosci 43(3):308–314. https://doi.org/10.1016/j.mcn.2009.12.004
Chen Y, Yang Z, Meng M, Zhao Y, Dong N, Yan H, Liu L, Ding M et al (2009) Cullin mediates degradation of RhoA through evolutionarily conserved BTB adaptors to control actin cytoskeleton structure and cell movement. Mol Cell 35(6):841–855. https://doi.org/10.1016/j.molcel.2009.09.004
Chen Y, Kramar EA, Chen LY, Babayan AH, Andres AL, Gall CM, Lynch G, Baram TZ (2013) Impairment of synaptic plasticity by the stress mediator CRH involves selective destruction of thin dendritic spines via RhoA signaling. Mol Psychiatry 18(4):485–496. https://doi.org/10.1038/mp.2012.17
Briz V, Zhu G, Wang Y, Liu Y, Avetisyan M, Bi X, Baudry M (2015) Activity-dependent rapid local RhoA synthesis is required for hippocampal synaptic plasticity. J Neurosci 35(5):2269–2282. https://doi.org/10.1523/JNEUROSCI.2302-14.2015
Wang X, Zhang C, Szabo G, Sun QQ (2013) Distribution of CaMKIIalpha expression in the brain in vivo, studied by CaMKIIalpha-GFP mice. Brain Res 1518:9–25. https://doi.org/10.1016/j.brainres.2013.04.042
Choi CI, Yoon SP, Choi JM, Kim SS, Lee YD, Birnbaumer L, Suh-Kim H (2014) Simultaneous deletion of floxed genes mediated by CaMKIIalpha-Cre in the brain and in male germ cells: application to conditional and conventional disruption of Goalpha. Exp Mol Med 46(5):e93. https://doi.org/10.1038/emm.2014.14
Minichiello L, Korte M, Wolfer D, Kuhn R, Unsicker K, Cestari V, Rossi-Arnaud C, Lipp HP et al (1999) Essential role for TrkB receptors in hippocampus-mediated learning. Neuron 24(2):401–414. https://doi.org/10.1016/s0896-6273(00)80853-3
Hernandez I, Luna G, Rauch JN, Reis SA, Giroux M, Karch CM, Boctor D, Sibih YE et al (2019) A farnesyltransferase inhibitor activates lysosomes and reduces tau pathology in mice with tauopathy. Sci Transl Med 11(485):eaat3005. https://doi.org/10.1126/scitranslmed.aat3005
Gao S, Yu R, Zhou X (2016) The role of geranylgeranyltransferase I-mediated protein prenylation in the brain. Mol Neurobiol 53(10):6925–6937. https://doi.org/10.1007/s12035-015-9594-3
Schultz BG, Patten DK, Berlau DJ (2018) The role of statins in both cognitive impairment and protection against dementia: a tale of two mechanisms. Transl Neurodegener 7:5. https://doi.org/10.1186/s40035-018-0110-3
Mans RA, McMahon LL, Li L (2012) Simvastatin-mediated enhancement of long-term potentiation is driven by farnesyl-pyrophosphate depletion and inhibition of farnesylation. Neuroscience 202:1–9
Storck EM, Morales-Sanfrutos J, Serwa RA, Panyain N, Lanyon-Hogg T, Tolmachova T, Ventimiglia LN, Martin-Serrano J et al (2019) Dual chemical probes enable quantitative system-wide analysis of protein prenylation and prenylation dynamics. Nat Chem 11(6):552–561. https://doi.org/10.1038/s41557-019-0237-6
Roskoski R Jr, Ritchie P (1998) Role of the carboxyterminal residue in peptide binding to protein farnesyltransferase and protein geranylgeranyltransferase. Arch Biochem Biophys 356(2):167–176
Winter-Vann AM, Casey PJ (2005) Post-prenylation-processing enzymes as new targets in oncogenesis. Nat Rev Cancer 5(5):405–412
Nguyen UT, Guo Z, Delon C, Wu Y, Deraeve C, Franzel B, Bon RS, Blankenfeldt W et al (2009) Analysis of the eukaryotic prenylome by isoprenoid affinity tagging. Nat Chem Biol 5(4):227–235. https://doi.org/10.1038/nchembio.149
Ali N, Jurczyluk J, Shay G, Tnimov Z, Alexandrov K, Munoz MA, Skinner OP, Pavlos NJ et al (2015) A highly sensitive prenylation assay reveals in vivo effects of bisphosphonate drug on the Rab prenylome of macrophages outside the skeleton. Small GTPases 6(4):202–211. https://doi.org/10.1080/21541248.2015.1085485
Kohnke M, Delon C, Hastie ML, Nguyen UT, Wu YW, Waldmann H, Goody RS, Gorman JJ et al (2013) Rab GTPase prenylation hierarchy and its potential role in choroideremia disease. PLoS One 8(12):e81758. https://doi.org/10.1371/journal.pone.0081758
Hanker AB, Mitin N, Wilder RS, Henske EP, Tamanoi F, Cox AD, Der CJ (2010) Differential requirement of CAAX-mediated posttranslational processing for Rheb localization and signaling. Oncogene 29(3):380–391. https://doi.org/10.1038/onc.2009.336
Buerger C, DeVries B, Stambolic V (2006) Localization of Rheb to the endomembrane is critical for its signaling function. Biochem Biophys Res Commun 344(3):869–880. https://doi.org/10.1016/j.bbrc.2006.03.220
Schwartz M (2004) Rho signalling at a glance. J Cell Sci 117(23):5457–5458
Liu Z, Meray RK, Grammatopoulos TN, Fredenburg RA, Cookson MR, Liu Y, Logan T, Lansbury PT (2009) Membrane-associated farnesylated UCH-L1 promotes α-synuclein neurotoxicity and is a therapeutic target for Parkinson’s disease. Proc Natl Acad Sci 106(12):4635–4640
Suazo KF, Hurben AK, Liu K, Xu F, Thao P, Sudheer C, Li L, Distefano MD (2018) Metabolic labeling of prenylated proteins using alkyne-modified isoprenoid analogues. Curr Protoc Chem Biol 10(3):e46
Schwanhausser B, Busse D, Li N, Dittmar G, Schuchhardt J, Wolf J, Chen W, Selbach M (2011) Global quantification of mammalian gene expression control. Nature 473(7347):337–342. https://doi.org/10.1038/nature10098
Schuld NJ, Vervacke JS, Lorimer EL, Simon NC, Hauser AD, Barbieri JT, Distefano MD, Williams CL (2014) The chaperone protein SmgGDS interacts with small GTPases entering the prenylation pathway by recognizing the last amino acid in the CAAX motif. J Biol Chem 289(10):6862–6876. https://doi.org/10.1074/jbc.M113.527192
Acknowledgments
We thank Dr. Martin Bergo for providing the original breeding pairs for FT- and GGT-haplodeficient and floxed mice and Andrea Gram for maintaining and genotyping different lines of experimental mice.
Funding
This work was supported in part by grants from the National Institute on Aging of the National Institutes of Health (AG056976 and AG058081), National Institute of General Medical Science (R01GM084152), the National Institute of Neurological Disorders and Stroke (R01NS107442), and the National Science Foundation (CHE-1308655). KFS was supported by a Doctoral Dissertation Fellowship from the University of Minnesota.
Author information
Authors and Affiliations
Contributions
WQ performed electrophysiological experiments, behavioral assays for the FT+/- mice and their controls, analyzed the data, interpreted the results, and wrote the manuscript. KS performed prenylomic analysis and wrote the manuscript. WL conducted Golgi staining and quantification of dendritic spine density. SC performed behavioral assays for the FTf/fCre+ and FTf/fCre− mice. DH assisted with electrophysiological and Golgi staining experiments. AJ processed samples for the prenylomic analysis and provided suggestions on the manuscript. MD designed and supervised the prenylomic part of the study. LY contributed to the discussion and interpretation of the data from electrophysiological experiments. LL conceived the study, supervised the progress of all experiments, interpreted the results, and edited and finalized the manuscript
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethics Approval
All animal procedures were reviewed and approved by the Institutional Animal Care and Use Committee (IACUC) at the University of Minnesota.
Consent to Participate
N/A
Consent for Publication
All authors have reviewed and approved the manuscript.
Code Availability
N/A
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
ESM 1
(PDF 259 kb).
Rights and permissions
About this article
Cite this article
Qu, W., Suazo, K.F., Liu, W. et al. Neuronal Protein Farnesylation Regulates Hippocampal Synaptic Plasticity and Cognitive Function. Mol Neurobiol 58, 1128–1144 (2021). https://doi.org/10.1007/s12035-020-02169-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-020-02169-w