Abstract
Late-onset Alzheimer’s disease (LOAD) is a high-occurrence neurological disorder but the difficulty in identifying precise and early biomarkers has complicated the understanding of the disease and the development of new treatments. In this sense, important knowledge is emerging regarding novel molecular and biological candidates with diagnostic potential, including microRNAs (miRNAs), which have a key role in gene repression. The aim of this systematic review was to define the role of miRNAs’ expression as biomarkers for LOAD both in brain tissues, which could help understand the biology of the disease, and circulating tissues, which could serve as non-invasive markers of the pathology. A systematic search was performed in Web of Science and PubMed using the keywords ((Alzheimer or Alzheimer’s) and (microRNA or microRNAs or miRNA or miRNAs or miR)) until August 2018 to retrieve all articles that presented independent original data evaluating the impact of miRNA expression on the development of LOAD in human population. A total of 90 studies investigating the role of miRNAs’ expression in the development of LOAD were identified. While other widely studied miRNAs such as hsa-miR-146a presented contradictory results among studies, deregulation in brain tissue of seven miRNAs, hsa-miR-16-5p, hsa-miR-34a-5p, hsa-miR-107, hsa-miR-125-5p, hsa-miR-132-3p, hsa-miR-181-3p, and hsa-miR-212-3p, was consistently identified in LOAD patients. Their role in the disease could be mediated through the regulation of key pathways, such as axon guidance, longevity, insulin, and MAPK signaling pathway. However, regarding their role as non-invasive biomarkers of LOAD in fluids, although the limited results available are promising, further studies are required.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Alzheimer’s disease (AD) is a progressive neurodegenerative disorder with a prevalence of almost 50 million people around the world. In fact, it is the most common cause of dementia worldwide [1, 2]. AD can be classified in two categories depending on onset age: early-onset Alzheimer’s disease (EOAD), the development of which starts before the patients are 65 years old, and late-onset Alzheimer’s disease (LOAD), which begins at a higher age [3, 4].
These disorders have a heterogeneous origin. While several markers have been established as risk factors for EOAD [4, 5], identifying precise and early biomarkers for LOAD has proven to be more difficult, which complicates the understanding of the disease and the development of new treatments [6].
In this sense, important knowledge is emerging regarding novel molecular and biological candidates with diagnostic potential, including microRNAs (miRNAs). miRNAs conform a subclass of small non-coding RNAs that play a significant regulatory role appointing mRNAs for cleavage or translational repression [7]. Actually, more than 60% of human genes are regulated by miRNAs, which are present in most body tissues, including brain tissue, cerebrospinal fluid (CSF), or serum [8, 9]. As a result, deregulation of miRNAs is crucial in the pathogenesis of neurodegenerative diseases [10]. Therefore, the expression of miRNAs collected from brain tissue biopsies could serve as a new biomarker and help understand the biology of LOAD [10]. Furthermore, recent findings indicate that miRNAs present in biofluids such as blood or CSF, known as circulating miRNAs [11], could reflect the composition of the brain extracellular space fluid and serve as non-invasive biomarkers of the disease [12].
Here, we have conducted a systematic review aiming to define a specific profile of deregulated miRNAs, both non-circulating and circulating, associated with the development of LOAD in order to identify precise biomarkers for the disease, as well as possible explanations for their involvement in the disorder. This may contribute to clarify the current state of this topic and establish the basis for future research.
Material and Methods
Systematic Review
Search Strategy
In order to retrieve all the published evidence related to miRNA expression and their association with LOAD, a systematic search was performed in Web of Science, using Web of Science Core Collection [13], and in PubMed database [14] using the Best Match algorithm. The keywords employed in the current search strategy were: Alzheimer or Alzheimer’s and microRNA or microRNAs or miRNA or miRNAs or miR. The search covered articles published until August 2018.
Inclusion and Exclusion Criteria
Articles were included if they presented independent original data and evaluated the impact of miRNA expression on the development of LOAD in human population. Hence, reviews or meta-analyses, case reports, abstracts, letters or comments, methodological studies, and articles not published in English were excluded. Studies were also excluded if they did not analyze miRNAs, did not include data from human populations or control groups, and were focused on other diseases or had different aims. After full-text evaluation, articles were also excluded if they did not specify miRNA expression, did not provide the mean age of participants, did not focus on LOAD, or did not compare miRNA expression in LOAD patients with a control group.
Data Extraction
Eligibility for inclusion of each evaluated study was independently assessed by two reviewers (SH and BS) and disagreements were resolved by consensus. After study selection, reviewers extracted the following information from each study: publication year, type of sample analyzed, characteristics of the study population, methodology used, number of miRNAs assessed, list of significant miRNAs provided, and their regulation status (upregulated, downregulated, or unchanged) in LOAD patients vs. controls. The significance level was set at p value < 0.05, according to the statistical method established in each article. In order to analyze the data, a unification of tissue sources was carried out in non-circulating studies. The following areas were classified as the frontal cortex lobe: Brodmann area 6, Brodmann area 9, Brodmann area 10, frontal cortex, frontal gyrus, medial frontal gyrus, and gray and white matter; the following areas were classified as the temporal cortex lobe: Brodmann area 20, Brodmann area 22, temporal cortex, temporal lobe cortex, temporal gyrus, and temporal lobe neocortex; the following areas were classified as the parietal cortex lobe: parietal lobe and parietal cortex; the CA 1 region was classified as the hippocampus.
On a first step, miRNAs that presented the same regulation status in four or more articles were selected. In order to minimize publication bias, selected miRNAs were searched in the supplementary material of the rest of the included articles to determine whether they had been analyzed in those articles with non-significant results. Then, miRNAs to be considered for further analyses were selected based on the following criteria:( a) that they did not present contradictory results, that is, they did not present an upregulated status and a downregulated status in different studies performed with the same kind of sample source; and b) that the ratio for that specific miRNA were 2:1 or higher taking into account the number of studies finding statistically significant differences in expressions comparing LOAD patients and controls, and those that did not find any differences.
Data Analysis
Selection of Target Genes
To predict the putative target genes of the selected miRNAs, miRWalk 2.0 database [15] was used. Only those genes predicted by 7 or more of the 12 miRNA-target prediction algorithms available at miRWalk were considered.
Pathway Enrichment Analysis
Pathway enrichment analyses were performed with the overrepresentation analysis module of the ConsensusPathDB web tool (CPdB) [16]. KEGG [17], Reactome [18], and BioCarta [19] pathway databases were used to analyze the lists of predicted target genes and determine the overrepresented pathways, assuming a conservative p value cutoff of 0.0001.
Venn Diagram
In order to study and establish concordances in the regulation status of miRNAs in Alzheimer’s patients regarding analyses in brain tissues and fluids, InteractiVenn tool [20] and Bioinformatics & Evolutionary Genomics [21] were used.
Results
Search Results
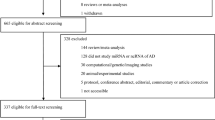
A total of 1727 records were discovered following the search parameters, 884 in PubMed and 843 in Web of Science (Main Collection). After removing duplicated articles, 1007 remained. Of these, 878 were excluded after abstract review because they did not met the inclusion criteria. Full texts of the 129 remaining studies focused on miRNAs in human LOAD were evaluated thoroughly. As a result, 39 additional studies were excluded at this step because they did not compare miRNA expressions between LOAD patients and healthy controls. Finally, 90 studies investigating the role of miRNAs expression in the development of LOAD were included. These articles were classified according to the source of sample analyzed (brain tissue or fluids): 42 of them provided data on non-circulating miRNAs (brain tissue), whereas 54 studied circulating miRNAs (fluids) (Fig. 1). Results regarding non-circulating and circulating miRNAs are presented separately in the following sections.
Non-circulating miRNAs
Forty-two studies analyzed non-circulating miRNA to search for differences in expression between LOAD cases and healthy controls (Online Resource Supplementary Table 1) [22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63]. As a result, a total of 319 different miRNAs were found to be deregulated in LOAD patients in at least one study.
Then, we identified 11 miRNAs that presented the same regulation status in at least 4 studies (Table 1) although some of them presented contradictions in additional studies. Seven miRNAs were mainly donwregulated: hsa-miR-9-5p [25, 27, 32, 42, 49, 51, 54, 59,60,61, 63], hsa-miR-16-5p [27, 30, 40, 42, 60], hsa-miR-29-3p [36, 40, 54, 58, 60, 64], hsa-miR-107 [27, 30, 42, 62], hsa-miR-132-3p [23, 27, 31, 35, 38, 39, 44,45,46, 59,60,61, 63] and hsa-miR-181a/c/d-5p [26, 27, 39, 54, 58, 60], and hsa-miR-212-3p [23, 27, 31, 39, 46, 60]; while, four miRNAs were mostly upregulated: hsa-miR-34a-5p [25, 42, 48, 51, 52], hsa-miR-125a/b-5p [25, 27, 40, 48, 49, 51, 59, 60, 63], hsa-miR-146a-5p [25, 27, 34, 42, 48, 49, 51, 56, 59,60,61], and hsa-miR-155-5p [25, 27, 34, 48, 49, 51, 53, 60]. The only miRNA that did not show any contradiction was hsa-miR-212-3p, which was downregulated in the 6 studies in which it was analyzed [23, 27, 31, 39, 46, 60].
Since the studies were performed analyzing miRNAs in different cerebral areas, mainly temporal, frontal or parietal lobe cortex, among other, we looked for correlations between specific brain regions and precise regulation status. However, due to the limited number of studies within each category, it was not possible to identify a clear correlation and, as a result, all results were evaluated together.
Therefore, we focused on the miRNAs that did not present contradictions (upregulated and downregulated in different studies) and that had a high ratio (2:1 or more) of significant vs. non-significant results. As a result, 7 miRNAs were selected for further analyses: hsa-miR-16-5p, hsa-miR-107, hsa-miR-132-3p, hsa-miR-181a/c/d-5p, and hsa-miR-212-3p met the established criteria for downregulation, while hsa-miR-34a-5p and hsa-miR-125a/b-5p were mainly upregulated (Table 2).
Predicted target genes for hsa-miR-16-5p, hsa-miR-34a-5p, hsa-miR-107, hsa-miR-125a/b-5p, hsa-miR-132-3p, hsa-miR-181a/c/d-5p, and hsa-miR-212-3p were searched in order to identify significantly overrepresented pathways regulated by those miRNAs and linked with LOAD. The 9 pathways that were most significantly overrepresented are shown in Online Resource Supplementary Table 2.
Among the most significant pathways, axon guidance (Online Resource Supplementary Fig. 1), longevity regulating pathway (Online Resource Supplementary Fig. 2), insulin signaling pathway (Online Resource Supplementary Fig. 3), and MAPK (Online Resource Supplementary Fig. 4) signaling pathway were the ones that were associated with a highest number of the selected miRNAs and had a plausible mechanistical connection with LOAD (Table 3 and Online Resource Supplementary Table 2).
Axon guidance was overrepresented among the predicted target genes of hsa-miR-34a-5p, hsa-miR-125a/b-5p, and hsa-miR-132-3p, which represented 41.1% of the genes in the pathway. Longevity regulating pathway was associated with hsa-miR-132-3p and hsa-miR-212-3p, with coverage of 22.6% of the pathway. Insulin signaling pathway, which is associated with the longevity process, was associated with hsa-miR-16-5p and hsa-miR-107, their predicted target genes covering a 35% of the pathway. Finally, MAPK signaling pathway was overrepresented among hsa-miR-16-5p and hsa-miR-125a/b-5p target genes (30.5% coverage).
Circulating miRNAs
Fifty-four studies compared the expression of circulating miRNAs in LOAD cases vs. healthy controls using different tissue sources (CSF, serum, plasma, blood, monocytes, and lymphocytes) [27, 28, 33, 42, 49, 51, 60, 64,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93,94,95,96,97,98,99,100,101,102,103,104,105,106,107,108,109,110,111]. Characteristics of those studies are shown in Online Resource Supplementary Table 3.
These 54 studies identified a total of 271 different miRNAs that presented significant changes in expressions between patients and controls. However, none of the miRNAs was found to present the same deregulation status in four or more studies (Online Resource Supplementary Table 3).
When the seven miRNAs selected as the most promising non-circulating biomarkers (hsa-miR-16-5p, hsa-miR-34-5p, hsa-miR-107, hsa-miR-125a/b-5p, hsa-miR-132-3p, hsa-miR-181a/c/d-5p, and hsa-miR-212-3p) were studied in fluids, the obtained results were heterogeneous among miRNAs. On the one hand, hsa-miR-212-3p, which was the miRNA showing the most consistent results in brain tissue, has not been studied in any circulating tissues, and hsa-miR-16-5p, which was mainly downregulated in LOAD brain tissues, was found as unchanged in CSF in the 3 studies in which it was analyzed [42, 69, 75]. Additionally, while hsa-miR-34a-5p was mainly upregulated in brain tissue, the results are contradictory among studies regarding circulating tissues, both in CSF and blood-derived samples [72, 100].
On the other hand, while hsa-miR-181a/c/d-5p was found equally downregulated or unchanged in blood-derived samples from LOAD patients, it was mainly found downregulated in CSF, in accordance with the results found in brain tissue [60, 64, 99]. Likewise, hsa-miR-132-3p expression in CSF followed the same downregulation observed in brain tissue in the only study in which it was analyzed [99], while results in plasma and serum were contradictory [65, 112]. A similar tendency could be observed for hsa-miR-125-5p, blood-derived samples showing contradictory results, while it was mainly upregulated in CSF, especially when only hsa-miR-125a-5p was considered, which showed no contradiction among studies [60, 65]. By contrast, hsa-miR-107 followed the opposite tendency, with the two studies performed in CSF showing non-significant differences between LOAD and healthy individuals and the studies performed in blood-derived tissues mirroring the downregulation observed in brain tissues [86, 95, 108].
Finally, we performed an analysis to study the correlation between the miRNAs reported in brain tissue and fluids. miRNAs identified as differentially expressed in both circulating and non-circulating tissues were compared taking into account their regulation status (Online Resource Supplementary Fig. 5a and 5b), 20 and 61 commonly upregulated and downregulated miRNAs were identified, respectively (Online Resource Supplementary Tables 4 and 5). On the one hand, among the commonly upregulated miRNAs, in addition to those mentioned above, we could highlight hsa-miR-455-3p, upregulated in the three studies in which it was analyzed [24, 28, 107]. On the other hand, regarding the downregulated miRNAs, hsa-miR-191-5p and hsa-miR-495-3p presented concordant results in the four [27, 103, 107] and three [27, 45, 99] studies in which they were analyzed, respectively.
Discussion
The main aim of this systematic review was to define the role of miRNAs’ expression as biomarkers for LOAD, both in brain tissues, which could help understand the biology of the disease, and circulating tissues, which could serve as non-invasive markers of the pathology.
It must be noted that we have specifically focused on those studies performed with LOAD patients, due to the difficulty in identifying biomarkers associated with this category of AD patients. Our extensive search strategy has allowed us to identify a total of 90 articles that met the inclusion criteria, 42 of them providing data on non-circulating miRNAs and 54 studies performed on circulating tissues, which represents a larger number of studies than the most recent systematic reviews in AD [113, 114].
Non-circulating miRNAs
We have identified 11 miRNAs that presented the same regulation status in at least 4 studies. However, four of them presented contradictory results among studies: hsa-miR-9-5p, hsa-miR-29-3p, hsa-miR-155-5p, and hsa-miR-146a-5p, the latter having been widely studied for its implication in neurodegeneration. Further studies would be needed in order to further explore the role of these miRNAs in LOAD and the reasons for the differences observed. In this review, we focused on the 7 miRNAs that were significantly deregulated in LOAD brain tissue in a mostly consistent way among studies.
First, hsa-miR-16-5p was analyzed in five different studies, being found downregulated in at least one brain region in LOAD patients in four of them [27, 30, 42, 60]. The only study that did not find any difference between patients and controls was the only one to be performed with samples from frontal cortex, which raised the question whether this difference in sample source was responsible for the difference in result. In fact, in one of the studies, mir-16-5p expression was measured in three different brain regions, being downregulated in LOAD temporal cortex and hippocampus but not in cerebellum, when compared with the same regions in healthy individuals [30]. Although, in the context of this review, it was difficult to establish associations between miRNA expression and brain regions, these results may indicate that miRNA expression related to LOAD may vary among certain specific brain regions and, thus, choosing the most appropriate sample source may be of great relevance.
Accordingly, miRNAs of the miR-181 family were downregulated in five out of the six studies in which they were analyzed [26, 27, 39, 54, 58, 60]. Once again, the only study that presented a contradicting result was performed considering a different brain region, the hippocampus, which might be to blame for the difference in result.
hsa-miR-107 was downregulated in LOAD patients in at least one of the brain regions analyzed in the four studies in which it was analyzed [27, 30, 42, 62]. Interestingly, it must be noted again that in one study it was analyzed in three different brain regions (temporal cortex, hippocampus, and cerebellum), being only unchanged in the cerebellum [30]. Therefore, this contributes to the idea that establishing an appropriate and standardized brain region for sample collection would be relevant for miRNA expression studies.
hsa-miR-132-3p was downregulated in nine out of thirteen articles in which it was analyzed [23, 27, 31, 35, 38, 39, 44,45,46, 59,60,61, 63], and hsa-miR-212-3p was downregulated in the six studies in which it was analyzed [23, 27, 31, 39, 46, 60]. In agreement with these results, the absence of these miRNAs has been shown to lead to programmed cell death [46], which would be a plausible mechanism for their involvement in LOAD.
By contrast, hsa-miR-34a-5p was upregulated in four out of five studies [25, 48, 51, 52]. It must be taken into account that in the study in which no difference in hsa-miR-34a-5p expression was observed between patients and controls, the control population [42] showed some initial neurodegeneration evidence (an average of I and II Braak stages), which could have contributed to an underestimation of differences in miRNA expression. Furthermore, hsa-miR-34a-5p has been previously reported to regulate neuronal differentiation and neurite outgrowth, and it has been shown to inhibit the expression of human tau by binding to its long 3′ UTR isoform [115], which supports a possible role in LOAD.
Finally, hsa-miR-125-5p was also upregulated in ten out of eleven studies [25, 27, 29, 40, 48, 49, 59, 60, 63]. In agreement with this result, upregulation of hsa-miR-125b-5p has been previously proposed to contribute to the degeneration of brain tissues [116] and it has been reported to play an important role in the mammalian neuronal development and differentiation [117, 118], which would explain its potential role in LOAD.
In order to further explore the role of these miRNAs in LOAD pathology, we performed an in silico analysis to identify the pathways regulated by these miRNAs that could be more related to the disease. To do so, we must take into consideration that, histopathologically, AD brains are characterized by two main hallmarks. On the one hand, amyloid β peptides (AβPs) are accumulated extracellularly generating plaques [119]. On the other hand, an intracellular accumulation of hyperphosphorylated tau protein leads to the formation of neurofibrillary tangles (NFTs) [120]. In addition, other modifications which are common to other neurodegenerative diseases could contribute to cognitive impairment, such as neuron and synapses loss or neuroinflammation [121, 122].
Our results indicate that the main target pathways for the seven miRNAs selected in the brain tissue with a putative role in LOAD are axon guidance, longevity, insulin, and MAPK signaling pathway.
First, axon guidance pathway is involved in axon outgrowth, repulsion, or attraction. This pathway could be directly implicated in LOAD development due to the relevance of those processes in the formation of neuritic plaques (NPs). NPs are a specific subtype of amyloid β plaques, of relevance in LOAD, formed by extracellular accumulation of well-known AβPs and degenerating axons and dendrites [123]. This pathway is overrepresented among the putative targets of hsa-miR-34a-5p, hsa-miR-125a/b-5p, and hsa-miR-132-3p. Therefore, changes in the regulation of genes in those pathways as a result of the deregulation of those miRNAs could be of relevance in the context of LOAD. For instance, glycogen synthase kinase (GSK3B) is a specific predicted target gene of hsa-miR-132-3p, which is downregulated in LOAD patients. GSK3B has been previously associated with LOAD, considering its ability to promote tau hyperphosphorylation [124, 125]. Tau is a microtubule-associated protein; when it is hyperphosphorylated, its binding affinity to microtubules decreases [126] and, thus, NFTs, a main hallmark of LOAD brains, are formed [120, 127,128,129,130]. Therefore, GSK3B upregulation as a consequence of hsa-miR-132-3p downregulation could be a relevant process for LOAD development.
Secondly, longevity pathway is overrepresented among the predicted target genes of hsa-miR-212-3p and hsa-miR-132-3p. Both miRNAs are downregulated in LOAD patients, which would lead to the overexpression of their target genes. Alzheimer is a well-known age-related disorder; thus, longevity process was expected to have a main function and to be altered in patients.
In addition, insulin pathway is enriched among the predicted targets of hsa-miR-16-5p and hsa-miR-107, which are downregulated in LOAD patients. Several articles have focused on the putative relationship between insulin pathway and LOAD, pointing to the existence of alterations in insulin signaling in the LOAD brain, which can be both a cause and consequence of LOAD [131, 132]. In fact, many studies have shown that LOAD brains present resistance to insulin [133,134,135,136], and there is substantial evidence that brain insulin resistance can increase Aβ and tau, which would promote LOAD development [133, 137,138,139]. Additionally, it was hypothesized that this insulin resistance in brain could trigger oxidative stress, inflammation, neuronal cell death, etc. [140, 141]. Therefore, changes in the regulation of this pathway, via changes in miRNA expression, could contribute to the development of LOAD, which would be an additional plausible explanation for the role of miRNAs in LOAD.
Finally, MAPK signaling pathway is overrepresented among the predicted target genes of hsa-miR-125-5p and hsa-miR-16-5p. This pathway is mainly related to cell cycle, apoptosis, or inflammation, processes of relevance for LOAD development,, which could explain the role of those miRNAs in the disease. In fact, previous studies have concluded that alterations in MAPK signaling pathways are essential for the development of some AD symptoms, such as neuroinflammation and tau hyperphosphorilation [142]. Target genes of hsa-miR-125-5p are mainly represented in a subpathway responsible for anti-apoptotic processes. Therefore, if genes such as NFATC3, MAPK8, PLA2G2A, or MAX are repressed by hsa-miR-125-5p upregulation, apoptosis would be increased. Alternatively, hsa-miR-16-5p is downregulated in LOAD patients, which implies an overexpression of its targets in this pathway, such as Ras or NF-ĸB, the upregulation of both having been previously related to AD development [143, 144]. On the one hand, earlier studies of Ras levels in AD neurons concluded that an increased expression of Ras leads to phosphorylation of APP and tau, which is associated with Aβ levels in AD brains [144]. On the other hand, NF-ĸB, has been reported to induce neuroinflammation in the CNS of neurodegenerative diseases in animal models and in patients [143, 145]. In addition, NF-ĸB activation has been linked to Aβ-induced neurotoxicity [146], further connecting this pathway to LOAD [147]. All of this together could explain the role of those miRNAs in LOAD.
Circulating miRNAs
We considered the studies analyzing circulating miRNAs, with the aim of identifying miRNAs that could be used as non-invasive biomarkers of LOAD, alone or in combination with traditional markers [148]. However, none of the miRNAs presented the same regulation status in four or more articles, which could be a result of the limited number of studies analyzing the same miRNAs and the heterogeneity in sample sources among studies.
Therefore, we focused on the seven miRNAs that were identified as deregulated in brain tissues of LOAD patients.
On the one hand, some miRNAs did not seem to be good markers in circulating tissues, i.e., hsa-miR-34a-5p, which presented contradictory results among studies [60, 65, 72, 98, 100, 111], and hsa-miR-16-5p, which seems to be unchanged in circulating samples of LOAD patients [42, 69, 75]. Furthermore, one of the most consistent miRNAs in brain tissues, hsa-miR-212-3p, has not been studied in liquid tissues to date. Thus, the study of this miRNA would be of great relevance.
On the other hand, we observed that hsa-miR-107, hsa-miR-132-3p, hsa-miR-125-5p, and hsa-miR-181-5p showed a certain potential and could be useful as biomarkers of LOAD in circulating tissues. hsa-miR-107 was measured in 6 different studies [60, 65, 68, 86, 95, 108] and it was downregulated in three out of the four studies analyzing blood-derived tissues but not in the two studies carried out in CSF. By contrast, hsa-miR-132-3p, hsa-miR-125-5p, and hsa-miR-181-5p only mirrored the results obtained in the brain tissues in CSF and not in blood-derived tissues. First, hsa-miR-132-3p was downregulated, as in brain tissue, in the only studies performed in CSF and serum, respectively, while the study performed in plasma showed a contradictory upregulation [112], further studies are required. Second, hsa-miR-181-5p was found downregulated in only two out of five studies performed in blood-derived samples, while it was downregulated in most studies (three out of four) [60, 65, 99] which analyzed this miRNA in CSF, replicating the result obtained in brain tissues. Lastly, hsa-miR-125-5p family was widely studied in fluid samples, the results showing different regulation status between CSF and blood. Blood-derived samples showed mainly an unchanged status for the miRNA, while it was mainly upregulated in CSF, especially when only hsa-miR-125a-5p was considered, which showed no contradiction among studies [60, 65]. These promising results warrant the need for more studies to identify accurate non-invasive biomarkers of LOAD.
Finally, both categories, studies in non-circulating and circulating tissues, were analyzed together in order to identify miRNAs that could have been studied in a more limited set of studies but that showed consistent results. This way, we observed that all results of hsa-miR-455-3p agree on its upregulation [24, 28]. By contrast, other miRNAs such as hsa-miR-191-5p and hsa-miR-495-3p were concluded as downregulated [27, 103, 107]. Since these miRNAs were analyzed in a limited number of studies, this implies the need to perform additional large scale studies to confirm these results and identify new markers of the disease.
Several limitations were faced while performing this systematic review. A determinant limitation of this study was the heterogeneity in sample source among studies. Additionally, in this review, neighboring brain areas were unified into larger categories in order to be able to compare the data. Different specimen types can set up a wide range of effects on miRNA concentrations and this biological variance may play an important role in clinical utility [10]. Furthermore, LOAD diagnosis was a key limitation, specifically on studies performed in circulating tissues. Unequivocal AD diagnosis is based on a scale that considers brain affectation, which is measured by mediating a histopathological brain autopsy [149]. In the studies in which patients were alive, though, LOAD was diagnosed trusting neurological tests (Mini-Mental State Exam (MMSE), Mini-Cognitive, National Institute of Neurological and Communicative Disorders and Stroke and the Alzheimer’s Disease and Related Disorders Association (NINCDS-ADRDA) criteria, or Diagnostic and Statistical Manual of Mental Disorders IV (DSM-IV)) [150, 151]. Different tests and ranges of punctuation were found among articles, which may have had an impact on the results obtained, implying difficulties in conclusion extraction. Furthermore, miRNA detection is a complex process that can be affected by technical variability such as collection and storage, miRNA extraction protocol, detection platforms, or normalization [152]. Since the effect of such differences is difficult to determine in the context of a review, it would be of great relevance to reach a consensus and standardize the methodology of study used for future studies in order to facilitate reproducibility and comparisons among studies.
Additionally, most studies analyzed a limited number of miRNAs by qRT-PCR or array, which could underestimate the effect of other miRNAs that might be involved in LOAD. Using methods such as next-generation sequencing (NGS) could help in identifying a larger range of miRNAs, including those that had not been previously described. Furthermore, it is possible that some additional results have been missed because of tendency to only publish statistically significant results, which may lead to a bias. All of these limitations may contribute to lack of consistency in many of the results, which makes it difficult to draw final conclusions about the role of some of the miRNAs analyzed as biomarkers in LOAD. Finally, we have to take into account the inaccuracy of the prediction algorithms of the databases used to determine the target gene and pathways but nowadays this limitation has to be assumed.
Conclusions
Deregulation in brain tissue of seven miRNAs, hsa-miR-16-5p, hsa-miR-34a-5p, hsa-miR-107, hsa-miR-125-5p, hsa-miR-132-3p, hsa-miR-181-5p, and hsa-miR-212-3p, has been consistently identified in LOAD patients. Their role in the disease could be mediated by their putative role in the regulation of key pathways, such as axon guidance, longevity, insulin, and MAPK signaling. Despite the consistent results regarding non-circulating miRNAs as putative LOAD biomarkers, the results regarding circulating miRNAs are more limited and heterogeneous. In this context, preliminary results have shown that the expression of some of those miRNAs, such as hsa-miR-125-5p, hsa-miR-132-3p, and hsa-miR-181-5p, in CSF could serve as a biomarker of LOAD. However, further homogeneously performed studies are required to establish a reliable relationship between miRNAs’ expression in fluids and LOAD. This knowledge could allow the identification of better diagnostic markers for LOAD.
References
Pan Y, Liu R, Terpstra E, Wang Y, Qiao F, Wang J et al (2016) Dysregulation and diagnostic potential of microRNA in Alzheimer’s disease. J Alzheimers Dis 49(1):1–12
Disease AAs (2018). Association Alzheimer’s Disease. https://www.alz.org/global/overview.asp. Accessed 2018.
Wattmo C, Wallin AK, Londos E, Minthon L (2011) Predictors of long-term cognitive outcome in Alzheimer’s disease. Alzheimers Res Ther 3(4):23
Ossenkoppele R, Mattsson N, Teunissen CE, Barkhof F, Pijnenburg Y, Scheltens P et al (2015) Cerebrospinal fluid biomarkers and cerebral atrophy in distinct clinical variants of probable Alzheimer’s disease. Neurobiol Aging 36(8):2340–2347
Cui L, Li Y, Ma G, Wang Y, Cai Y, Liu S et al (2014) A functional polymorphism in the promoter region of microRNA-146a is associated with the risk of Alzheimer disease and the rate of cognitive decline in patients. PLoS One 9(2):e89019
Mattsson N, Rosén E, Hansson O, Andreasen N, Parnetti L, Jonsson M et al (2012) Age and diagnostic performance of Alzheimer disease CSF biomarkers. Neurology. 78(7):468–476
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 116(2):281–297
Friedman RC, Farh KK, Burge CB, Bartel DP (2009) Most mammalian mRNAs are conserved targets of microRNAs. Genome Res 19(1):92–105. https://doi.org/10.1101/gr.082701.108
Johanson TM, Skinner JP, Kumar A, Zhan Y, Lew AM, Chong MM (2014) The role of microRNAs in lymphopoiesis. Int J Hematol 100(3):246–253. https://doi.org/10.1007/s12185-014-1606-y
Shah SZA, Zhao D, Hussain T, Sabir N, Yang L (2018) Regulation of microRNAs-mediated autophagic flux: a new regulatory avenue for neurodegenerative diseases with focus on prion diseases. Front Aging Neurosci 10:139
Gilad S, Meiri E, Yogev Y, Benjamin S, Lebanony D, Yerushalmi N et al (2008) Serum microRNAs are promising novel biomarkers. PLoS One 3(9):e3148
Edsbagge M, Andreasson U, Ambarki K, Wikkelsø C, Eklund A, Blennow K et al (2017) Alzheimer’s disease-associated cerebrospinal fluid (CSF) biomarkers do not correlate with CSF volumes or CSF production rate. J Alzheimers Dis 58(3):821–828. https://doi.org/10.3233/JAD-161257
Web, Science o. Web of Science core collection. https://apps.webofknowledge.com/WOS_GeneralSearch_input.do?product=WOS&SID=F3C19V4ITCwyVekjgNr&search_mode=GeneralSearch. Accessed 2019.
PubMed. https://www.ncbi.nlm.nih.gov/pubmed/ Accessed 2018.
Dweep H, Sticht C, Pandey P, Gretz N (2011) miRWalk--database: prediction of possible miRNA binding sites by “walking” the genes of three genomes. J Biomed Inform 44(5):839–847. https://doi.org/10.1016/j.jbi.2011.05.002
Kamburov A, Wierling C, Lehrach H, Herwig R (2009) ConsensusPathDB--a database for integrating human functional interaction networks. Nucleic Acids Res 37(Database issue):D623–D628. https://doi.org/10.1093/nar/gkn698
Kanehisa M, Goto S (2000) KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28(1):27–30
Croft D, O'Kelly G, Wu G, Haw R, Gillespie M, Matthews L et al (2011) Reactome: a database of reactions, pathways and biological processes. Nucleic Acids Res 39(Database issue):D691–D697. https://doi.org/10.1093/nar/gkq1018
Biocarta. http://cgap.nci.nih.gov/Pathways/BioCarta_Pathways Accessed 2019.
Heberle H, Meirelles GV, da Silva FR, Telles GP, Minghim R (2015) InteractiVenn: a web-based tool for the analysis of sets through Venn diagrams. BMC Bioinformatics 16:169. https://doi.org/10.1186/s12859-015-0611-3
Lam F, Lalansingh CM, Babaran HE, Wang Z, Prokopec SD, Fox NS et al (2016) VennDiagramWeb: a web application for the generation of highly customizable Venn and Euler diagrams. BMC Bioinformatics. 17(1):401. https://doi.org/10.1186/s12859-016-1281-5
Akhter R, Shao Y, Shaw M, Formica S, Khrestian M, Leverenz JB et al (2018) Regulation of ADAM10 by miR-140-5p and potential relevance for Alzheimer’s disease. Neurobiol Aging 63:110–119
Annese A, Manzari C, Lionetti C, Picardi E, Horner DS, Chiara M et al (2018) Whole transcriptome profiling of late-onset Alzheimer’s disease patients provides insights into the molecular changes involved in the disease. Sci Rep 8(1):4282–018-22701-2
Kumar S, Reddy PH (2018) MicroRNA-455-3p as a potential biomarker for Alzheimer’s disease: an update. Front Aging Neurosci 10:41
Pogue AI, Lukiw WJ (2018) Up-regulated pro-inflammatory microRNAs (miRNAs) in Alzheimer’s disease (AD) and age-related macular degeneration (AMD). Cell Mol Neurobiol 38(5):1021–1031
Zumkehr J, Rodriguez-Ortiz CJ, Medeiros R, Kitazawa M (2018) Inflammatory cytokine, IL-1beta, regulates glial glutamate transporter via microRNA-181a in vitro. J Alzheimers Dis 63(3):965–975
Hara N, Kikuchi M, Miyashita A, Hatsuta H, Saito Y, Kasuga K et al (2017) Serum microRNA miR-501-3p as a potential biomarker related to the progression of Alzheimer’s disease. Acta Neuropathol Commun 5(1):10-017-0414-z
Kumar S, Vijayan M, Reddy PH (2017) MicroRNA-455-3p as a potential peripheral biomarker for Alzheimer’s disease. Hum Mol Genet 26(19):3808–3822
Ma X, Liu L, Meng J (2017) MicroRNA-125b promotes neurons cell apoptosis and tau phosphorylation in Alzheimer’s disease. Neurosci Lett 661:57–62
Moncini S, Lunghi M, Valmadre A, Grasso M, Vescovo VD, Riva P et al (2017) The miR-15/107 family of microRNA genes regulates CDK5R1/p35 with implications for Alzheimer’s disease pathogenesis. Mol Neurobiol 54(6):4329–4342
Pichler S, Gu W, Hartl D, Gasparoni G, Leidinger P, Keller A et al (2017) The miRNome of Alzheimer’s disease: consistent downregulation of the miR-132/212 cluster. Neurobiol Aging 50(167):e1–e10
Jesko H, Wilkaniec A, Cieslik M, Hilgier W, Gassowska M, Lukiw WJ et al (2016) Altered arginine metabolism in cells transfected with human wild-type beta amyloid precursor protein (beta APP). Curr Alzheimer Res 13(9):1030–1039
Moon J, Lee ST, Kong IG, Byun JI, Sunwoo JS, Shin JW et al (2016) Early diagnosis of Alzheimer’s disease from elevated olfactory mucosal miR-206 level. Sci Rep 6:20364
Zhao Y, Alexandrov PN, Jaber V, Lukiw WJ (2016) Deficiency in the ubiquitin conjugating enzyme UBE2A in Alzheimer’s disease (AD) is linked to deficits in a natural circular miRNA-7 sponge (circRNA; ciRS-7). Genes. 7:12. https://doi.org/10.3390/genes7120116
Zhu QB, Unmehopa U, Bossers K, Hu YT, Verwer R, Balesar R et al (2016) MicroRNA-132 and early growth response-1 in nucleus basalis of Meynert during the course of Alzheimer’s disease. Brain 139(Pt 3):908–921
Lei X, Lei L, Zhang Z, Cheng Y (2015) Downregulated miR-29c correlates with increased BACE1 expression in sporadic Alzheimer’s disease. Int J Clin Exp Pathol 8(2):1565–1574
Santa-Maria I, Alaniz ME, Renwick N, Cela C, Fulga TA, Vactor DV et al (2015) Dysregulation of microRNA-219 promotes neurodegeneration through post-transcriptional regulation of tau. J Clin Invest 125(2):681–686
Smith PY, Hernandez-Rapp J, Jolivette F, Lecours C, Bisht K, Goupil C et al (2015) miR-132/212 deficiency impairs tau metabolism and promotes pathological aggregation in vivo. Hum Mol Genet 24(23):6721–6735
Weinberg RB, Mufson EJ, Counts SE (2015) Evidence for a neuroprotective microRNA pathway in amnestic mild cognitive impairment. Front Neurosci 9:430
Banzhaf-Strathmann J, Benito E, May S, Arzberger T, Tahirovic S, Kretzschmar H et al (2014) MicroRNA-125b induces tau hyperphosphorylation and cognitive deficits in Alzheimer’s disease. EMBO J 33(15):1667–1680
Long JM, Ray B, Lahiri DK (2014) MicroRNA-339-5p down-regulates protein expression of beta-site amyloid precursor protein-cleaving enzyme 1 (BACE1) in human primary brain cultures and is reduced in brain tissue specimens of Alzheimer disease subjects. J Biol Chem 289(8):5184–5198
Muller M, Kuiperij HB, Claassen JA, Kusters B, Verbeek MM (2014) MicroRNAs in Alzheimer’s disease: differential expression in hippocampus and cell-free cerebrospinal fluid. Neurobiol Aging 35(1):152–158
Absalon S, Kochanek DM, Raghavan V, Krichevsky AM (2013) MiR-26b, upregulated in Alzheimer’s disease, activates cell cycle entry, tau-phosphorylation, and apoptosis in postmitotic neurons. J Neurosci 33(37):14645–14659
Hebert SS, Wang W-X, Zhu Q, Nelson PT (2013) A study of small RNAs from cerebral neocortex of pathology-verified Alzheimer’s disease, dementia with Lewy bodies, hippocampal sclerosis, frontotemporal lobar dementia, and non-demented human controls. J Alzheimers Dis 35(2):335–348
Lau P, Bossers K, Janky RS, Salta E, Frigerio CS, Barbash S et al (2013) Alteration of the microRNA network during the progression of Alzheimer’s disease. EMBO Mol Med 5(10):1613–1634
Wong HK, Veremeyko T, Patel N, Lemere CA, Walsh DM, Esau C et al (2013) De-repression of FOXO3a death axis by microRNA-132 and -212 causes neuronal apoptosis in Alzheimer’s disease. Hum Mol Genet 22(15):3077–3092. https://doi.org/10.1093/hmg/ddt164
Yan H, Xu T, Zhao H, Lee KC, Wang HY, Zhang Y (2013) Isoflurane increases neuronal cell death vulnerability by downregulating miR-214. PLoS One 8(2):e55276
Zhao Y, Bhattacharjee S, Jones BM, Dua P, Alexandrov PN, Hill JM et al (2013) Regulation of TREM2 expression by an NF-small ka, CyrillicB-sensitive miRNA-34a. Neuroreport. 24(6):318–323
Alexandrov PN, Dua P, Hill JM, Bhattacharjee S, Zhao Y, Lukiw WJ (2012) MicroRNA (miRNA) speciation in Alzheimer’s disease (AD) cerebrospinal fluid (CSF) and extracellular fluid (ECF). Int J Biochem Mol Biol 3(4):365–373
Lee ST, Chu K, Jung KH, Kim JH, Huh JY, Yoon H et al (2012) miR-206 regulates brain-derived neurotrophic factor in Alzheimer disease model. Ann Neurol 72(2):269–277
Lukiw WJ, Alexandrov PN, Zhao Y, Hill JM, Bhattacharjee S (2012) Spreading of Alzheimer’s disease inflammatory signaling through soluble micro-RNA. Neuroreport. 23(10):621–626
Agostini M, Tucci P, Killick R, Candi E, Sayan BS, di Val Cervo PR et al (2011) Neuronal differentiation by TAp73 is mediated by microRNA-34a regulation of synaptic protein targets. Proc Natl Acad Sci U S A 108(52):21093–21098
Culpan D, Kehoe PG, Love S (2011) Tumour necrosis factor-alpha (TNF-alpha) and miRNA expression in frontal and temporal neocortex in Alzheimer’s disease and the effect of TNF-alpha on miRNA expression in vitro. Int J Mol Epidemiol Genet 2(2):156–162
Geekiyanage H, Chan C (2011) MicroRNA-137/181c regulates serine palmitoyltransferase and in turn amyloid beta, novel targets in sporadic Alzheimer’s disease. J Neurosci 31(41):14820–14830
Zovoilis A, Agbemenyah HY, Agis-Balboa RC, Stilling RM, Edbauer D, Rao P et al (2011) microRNA-34c is a novel target to treat dementias. EMBO J 30(20):4299–4308
Cui JG, Li YY, Zhao Y, Bhattacharjee S, Lukiw WJ (2010) Differential regulation of interleukin-1 receptor-associated kinase-1 (IRAK-1) and IRAK-2 by microRNA-146a and NF-kappaB in stressed human astroglial cells and in Alzheimer disease. J Biol Chem 285(50):38951–38960
Faghihi MA, Zhang M, Huang J, Modarresi F, der Brug MPV, Nalls MA et al (2010) Evidence for natural antisense transcript-mediated inhibition of microRNA function. Genome Biol 11(5) R56–2010-11-5-r56
Nunez-Iglesias J, Liu CC, Morgan TE, Finch CE, Zhou XJ (2010) Joint genome-wide profiling of miRNA and mRNA expression in Alzheimer’s disease cortex reveals altered miRNA regulation. PLoS One 5(2):e8898
Sethi P, Lukiw WJ (2009) Micro-RNA abundance and stability in human brain: specific alterations in Alzheimer’s disease temporal lobe neocortex. Neurosci Lett 459(2):100–104
Cogswell JP, Ward J, Taylor IA, Waters M, Shi Y, Cannon B et al (2008) Identification of miRNA changes in Alzheimer’s disease brain and CSF yields putative biomarkers and insights into disease pathways. J Alzheimers Dis 14(1):27–41
Lukiw WJ, Zhao Y, Cui JG (2008) An NF-kappaB-sensitive micro RNA-146a-mediated inflammatory circuit in Alzheimer disease and in stressed human brain cells. J Biol Chem 283(46):31315–31322
Wang WX, Rajeev BW, Stromberg AJ, Ren N, Tang G, Huang Q et al (2008) The expression of microRNA miR-107 decreases early in Alzheimer’s disease and may accelerate disease progression through regulation of beta-site amyloid precursor protein-cleaving enzyme 1. J Neurosci 28(5):1213–1223
Lukiw WJ (2007) Micro-RNA speciation in fetal, adult and Alzheimer’s disease hippocampus. Neuroreport. 18(3):297–300
Geekiyanage H, Jicha GA, Nelson PT, Chan C (2012) Blood serum miRNA: non-invasive biomarkers for Alzheimer’s disease. Exp Neurol 235(2):491–496
Denk J, Oberhauser F, Kornhuber J, Wiltfang J, Fassbender K, Schroeter ML et al (2018) Specific serum and CSF microRNA profiles distinguish sporadic behavioural variant of frontotemporal dementia compared with Alzheimer patients and cognitively healthy controls. PLoS One 13(5):e0197329
Derkow K, Rossling R, Schipke C, Kruger C, Bauer J, Fahling M et al (2018) Distinct expression of the neurotoxic microRNA family let-7 in the cerebrospinal fluid of patients with Alzheimer’s disease. PLoS One 13(7):e0200602
Dias IHK, Brown CL, Shabir K, Polidori MC, Griffiths HR (2018) miRNA 933 expression by endothelial cells is increased by 27-hydroxycholesterol and is more prevalent in plasma from dementia patients. J Alzheimers Dis 64(3):1009–1017
Manzine PR, Pelucchi S, Horst MA, Vale FAC, Pavarini SCI, Audano M et al (2018) MicroRNA 221 targets ADAM10 mRNA and is downregulated in Alzheimer’s disease. J Alzheimers Dis 61(1):113–123
McKeever PM, Schneider R, Taghdiri F, Weichert A, Multani N, Brown RA et al (2018) MicroRNA expression levels are altered in the cerebrospinal fluid of patients with young-onset Alzheimer’s disease. Mol Neurobiol
Piscopo P, Grasso M, Puopolo M, D'Acunto E, Talarico G, Crestini A et al (2018) Circulating miR-127-3p as a potential biomarker for differential diagnosis in frontotemporal dementia. J Alzheimers Dis
Yang TT, Liu CG, Gao SC, Zhang Y, Wang PC (2018) The serum exosome derived microRNA-135a, −193b, and −384 were potential Alzheimer’s disease biomarkers. Biomed Environ Sci 31(2):87–96
Cosin-Tomas M, Antonell A, Llado A, Alcolea D, Fortea J, Ezquerra M et al (2017) Plasma miR-34a-5p and miR-545-3p as early biomarkers of Alzheimer’s disease: potential and limitations. Mol Neurobiol 54(7):5550–5562
Dangla-Valls A, Molinuevo JL, Altirriba J, Sanchez-Valle R, Alcolea D, Fortea J et al (2017) CSF microRNA profiling in Alzheimer’s disease: a screening and validation study. Mol Neurobiol 54(9):6647–6654
Guo R, Fan G, Zhang J, Wu C, Du Y, Ye H et al (2017) A 9-microRNA signature in serum serves as a noninvasive biomarker in early diagnosis of Alzheimer’s disease. J Alzheimers Dis 60(4):1365–1377
Lusardi TA, Phillips JI, Wiedrick JT, Harrington CA, Lind B, Lapidus JA et al (2017) MicroRNAs in human cerebrospinal fluid as biomarkers for Alzheimer’s disease. J Alzheimers Dis 55(3):1223–1233
Nagaraj S, Laskowska-Kaszub K, Debski KJ, Wojsiat J, Dabrowski M, Gabryelewicz T et al (2017) Profile of 6 microRNA in blood plasma distinguish early stage Alzheimer’s disease patients from non-demented subjects. Oncotarget. 8(10):16122–16143
Riancho J, Vazquez-Higuera JL, Pozueta A, Lage C, Kazimierczak M, Bravo M et al (2017) MicroRNA profile in patients with Alzheimer’s disease: analysis of miR-9-5p and miR-598 in raw and exosome enriched cerebrospinal fluid samples. J Alzheimers Dis 57(2):483–491
Wu Y, Xu J, Cheng J, Jiao D, Zhou C, Dai Y et al (2017) Lower serum levels of miR-29c-3p and miR-19b-3p as biomarkers for Alzheimer’s disease. Tohoku J Exp Med 242(2):129–136
Jia LH, Liu YN (2016) Downregulated serum miR-223 servers as biomarker in Alzheimer’s disease. Cell Biochem Funct 34(4):233–237
Keller A, Backes C, Haas J, Leidinger P, Maetzler W, Deuschle C et al (2016) Validating Alzheimer’s disease micro RNAs using next-generation sequencing. Alzheimers Dement 12(5):565–576
Muller M, Jakel L, Bruinsma IB, Claassen JA, Kuiperij HB, Verbeek MM (2016) MicroRNA-29a is a candidate biomarker for Alzheimer’s disease in cell-free cerebrospinal fluid. Mol Neurobiol 53(5):2894–2899
Muller M, Kuiperij HB, Versleijen AA, Chiasserini D, Farotti L, Baschieri F et al (2016) Validation of microRNAs in cerebrospinal fluid as biomarkers for different forms of dementia in a multicenter study. J Alzheimers Dis 52(4):1321–1333
Ragusa M, Bosco P, Tamburello L, Barbagallo C, Condorelli AG, Tornitore M et al (2016) miRNAs plasma profiles in vascular dementia: biomolecular data and biomedical implications. Front Cell Neurosci 10:51
Ren RJ, Zhang YF, Dammer EB, Zhou Y, Wang LL, Liu XH et al (2016) Peripheral blood microRNA expression profiles in Alzheimer’s disease: screening, validation, association with clinical phenotype and implications for molecular mechanism. Mol Neurobiol 53(8):5772–5781
Xing H, Guo S, Zhang Y, Zheng Z, Wang H (2016) Upregulation of microRNA-206 enhances lipopolysaccharide-induced inflammation and release of amyloid-beta by targeting insulin-like growth factor 1 in microglia. Mol Med Rep 14(2):1357–1364
Yilmaz SG, Erdal ME, Ozge AA, Sungur MA (2016) Can peripheral microRNA expression data serve as epigenomic (upstream) biomarkers of Alzheimer’s disease? OMICS 20(8):456–461
Zhang Y, Liu C, Wang J, Li Q, Ping H, Gao S et al (2016) MiR-299-5p regulates apoptosis through autophagy in neurons and ameliorates cognitive capacity in APPswe/PS1dE9 mice. Sci Rep 6:24566
Zhang Y, Xing H, Guo S, Zheng Z, Wang H, Xu D (2016) MicroRNA-135b has a neuroprotective role via targeting of beta-site APP-cleaving enzyme 1. Exp Ther Med 12(2):809–814
Cheng L, Doecke JD, Sharples RA, Villemagne VL, Fowler CJ, Rembach A et al (2015) Prognostic serum miRNA biomarkers associated with Alzheimer’s disease shows concordance with neuropsychological and neuroimaging assessment. Mol Psychiatry 20(10):1188–1196
Denk J, Boelmans K, Siegismund C, Lassner D, Arlt S, Jahn H (2015) MicroRNA profiling of CSF reveals potential biomarkers to detect Alzheimer’s disease. PLoS One 10(5):e0126423
Dong H, Li J, Huang L, Chen X, Li D, Wang T et al (2015) Serum microRNA profiles serve as novel biomarkers for the diagnosis of Alzheimer’s disease. Dis Markers 2015:625659
Guedes JR, Santana I, Cunha C, Duro D, Almeida MR, Cardoso AM et al (2015) MicroRNA deregulation and chemotaxis and phagocytosis impairment in Alzheimer’s disease. Alzheimers Dement (Amst) 3:7–17
Gui Y, Liu H, Zhang L, Lv W, Hu X (2015) Altered microRNA profiles in cerebrospinal fluid exosome in Parkinson disease and Alzheimer disease. Oncotarget. 6(35):37043–37053
van Harten AC, Mulders J, Scheltens P, van der Flier WM, Oudejans CB (2015) Differential expression of microRNA in cerebrospinal fluid as a potential novel biomarker for Alzheimer’s disease. J Alzheimers Dis 47(1):243–252
Wang T, Chen K, Li H, Dong S, Su N, Liu Y et al (2015) The feasibility of utilizing plasma MiRNA107 and BACE1 messenger RNA gene expression for clinical diagnosis of amnestic mild cognitive impairment. J Clin Psychiatry 76(2):135–141
Yang G, Song Y, Zhou X, Deng Y, Liu T, Weng G et al (2015) MicroRNA-29c targets beta-site amyloid precursor protein-cleaving enzyme 1 and has a neuroprotective role in vitro and in vivo. Mol Med Rep 12(2):3081–3088
Zhu Y, Li C, Sun A, Wang Y, Zhou S (2015) Quantification of microRNA-210 in the cerebrospinal fluid and serum: Implications for Alzheimer’s disease. Exp Ther Med 9(3):1013–1017
Bhatnagar S, Chertkow H, Schipper HM, Yuan Z, Shetty V, Jenkins S et al (2014) Increased microRNA-34c abundance in Alzheimer’s disease circulating blood plasma. Front Mol Neurosci 7:2
Burgos K, Malenica I, Metpally R, Courtright A, Rakela B, Beach T et al (2014) Profiles of extracellular miRNA in cerebrospinal fluid and serum from patients with Alzheimer’s and Parkinson’s diseases correlate with disease status and features of pathology. PLoS One 9(5):e94839
Kiko T, Nakagawa K, Tsuduki T, Furukawa K, Arai H, Miyazawa T (2014) MicroRNAs in plasma and cerebrospinal fluid as potential markers for Alzheimer’s disease. J Alzheimers Dis 39(2):253–259
geng Liu C, ling Wang J, Li L, xiang Xue L, qi Zhang Y, chang Wang P (2014) MicroRNA-135a and-200b, potential biomarkers for Alzheimer’s disease, regulate beta secretase and amyloid precursor protein. Brain Res 1583:55–64
Tan L, Yu JT, Liu QY, Tan MS, Zhang W, Hu N et al (2014) Circulating miR-125b as a biomarker of Alzheimer’s disease. J Neurol Sci 336(1–2):52–56
Tan L, Yu JT, Tan MS, Liu QY, Wang HF, Zhang W et al (2014) Genome-wide serum microRNA expression profiling identifies serum biomarkers for Alzheimer’s disease. J Alzheimers Dis 40(4):1017–1027
Tiribuzi R, Crispoltoni L, Porcellati S, Lullo MD, Florenzano F, Pirro M et al (2014) miR128 up-regulation correlates with impaired amyloid beta(1-42) degradation in monocytes from patients with sporadic Alzheimer’s disease. Neurobiol Aging 35(2):345–356
Bekris LM, Lutz F, Montine TJ, Yu CE, Tsuang D, Peskind ER et al (2013) MicroRNA in Alzheimer’s disease: an exploratory study in brain, cerebrospinal fluid and plasma. Biomarkers. 18(5):455–466. https://doi.org/10.3109/1354750X.2013.814073
Frigerio CS, Lau P, Salta E, Tournoy J, Bossers K, Vandenberghe R et al (2013) Reduced expression of hsa-miR-27a-3p in CSF of patients with Alzheimer disease. Neurology. 81(24):2103–2106
Kumar P, Dezso Z, MacKenzie C, Oestreicher J, Agoulnik S, Byrne M et al (2013) Circulating miRNA biomarkers for Alzheimer’s disease. PLoS One 8(7):e69807
Leidinger P, Backes C, Deutscher S, Schmitt K, Mueller SC, Frese K et al (2013) A blood based 12-miRNA signature of Alzheimer disease patients. Genome Biol 14(7):R78
Lehmann SM, Kruger C, Park B, Derkow K, Rosenberger K, Baumgart J et al (2012) An unconventional role for miRNA: let-7 activates toll-like receptor 7 and causes neurodegeneration. Nat Neurosci 15(6):827–835
Villa C, Fenoglio C, Riz MD, Clerici F, Marcone A, Benussi L et al (2011) Role of hnRNP-A1 and miR-590-3p in neuronal death: genetics and expression analysis in patients with Alzheimer disease and frontotemporal lobar degeneration. Rejuvenation Res 14(3):275–281
Schipper HM, Maes OC, Chertkow HM, Wang E (2007) MicroRNA expression in Alzheimer blood mononuclear cells. Gene Regul Syst Biol 1:263–274
Sheinerman KS, Tsivinsky VG, Abdullah L, Crawford F, Umansky SR (2013) Plasma microRNA biomarkers for detection of mild cognitive impairment: biomarker validation study. Aging (Albany NY) 5(12):925–938. https://doi.org/10.18632/aging.100624
Swarbrick S, Wragg N, Ghosh S, Stolzing A (2019) Systematic review of miRNA as biomarkers in Alzheimer’s disease. Mol Neurobiol. https://doi.org/10.1007/s12035-019-1500-y
Fransquet PD, Ryan J (2018) Micro RNA as a potential blood-based epigenetic biomarker for Alzheimer’s disease. Clin Biochem 58:5–14. https://doi.org/10.1016/j.clinbiochem.2018.05.020
Dickson JR, Kruse C, Montagna DR, Finsen B, Wolfe MS (2013) Alternative polyadenylation and miR-34 family members regulate tau expression. J Neurochem 127(6):739–749. https://doi.org/10.1111/jnc.12437
Pogue AI, Cui JG, Li YY, Zhao Y, Culicchia F, Lukiw WJ (2010) Micro RNA-125b (miRNA-125b) function in astrogliosis and glial cell proliferation. Neurosci Lett 476(1):18–22. https://doi.org/10.1016/j.neulet.2010.03.054
Basavaraju M, de Lencastre A (2016) Alzheimer’s disease: presence and role of microRNAs. Biomol Concepts 7(4):241–252. https://doi.org/10.1515/bmc-2016-0014
Lukiw WJ, Andreeva TV, Grigorenko AP, Rogaev EI (2012) Studying micro RNA function and dysfunction in Alzheimer’s disease. Front Genet 3:327. https://doi.org/10.3389/fgene.2012.00327
Selkoe DJ (2008) Biochemistry and molecular biology of amyloid beta-protein and the mechanism of Alzheimer’s disease. Handb Clin Neurol 89:245–260. https://doi.org/10.1016/S0072-9752(07)01223-7
Goedert M (2004) Tau protein and neurodegeneration. Semin Cell Dev Biol 15(1):45–49
Slota JA, Booth SA (2019) MicroRNAs in neuroinflammation: implications in disease pathogenesis, biomarker discovery and therapeutic applications. Noncoding RNA 5:2. https://doi.org/10.3390/ncrna5020035
Gaudet AD, Fonken LK, Watkins LR, Nelson RJ, Popovich PG (2018) MicroRNAs: roles in regulating neuroinflammation. Neuroscientist. 24(3):221–245. https://doi.org/10.1177/1073858417721150
Nelson PT, Alafuzoff I, Bigio EH, Bouras C, Braak H, Cairns NJ et al (2012) Correlation of Alzheimer disease neuropathologic changes with cognitive status: a review of the literature. J Neuropathol Exp Neurol 71(5):362–381. https://doi.org/10.1097/NEN.0b013e31825018f7
Spittaels K, Van den Haute C, Van Dorpe J, Geerts H, Mercken M, Bruynseels K et al (2000) Glycogen synthase kinase-3beta phosphorylates protein tau and rescues the axonopathy in the central nervous system of human four-repeat tau transgenic mice. J Biol Chem 275(52):41340–41349. https://doi.org/10.1074/jbc.M006219200
Shal B, Ding W, Ali H, Kim YS, Khan S (2018) Anti-neuroinflammatory potential of natural products in attenuation of Alzheimer’s disease. Front Pharmacol 9:548. https://doi.org/10.3389/fphar.2018.00548
Piedrahita D, Hernández I, López-Tobón A, Fedorov D, Obara B, Manjunath BS et al (2010) Silencing of CDK5 reduces neurofibrillary tangles in transgenic Alzheimer's mice. J Neurosci 30(42):13966–13976
Masliah E (1995) Mechanisms of synaptic dysfunction in Alzheimer’s disease. Histol Histopathol 10(2):509–519
Masliah E, Mallory M, Hansen L, DeTeresa R, Terry RD (1993) Quantitative synaptic alterations in the human neocortex during normal aging. Neurology. 43(1):192–197
Nunomura A, Castellani RJ, Zhu X, Moreira PI, Perry G, Smith MA (2006) Involvement of oxidative stress in Alzheimer disease. J Neuropathol Exp Neurol 65(7):631–641
Arriagada PV, Growdon JH, Hedley-Whyte ET, Hyman BT (1992) Neurofibrillary tangles but not senile plaques parallel duration and severity of Alzheimer’s disease. Neurology. 42(3 Pt 1):631–639
Stanley M, Macauley SL, Holtzman DM (2016) Changes in insulin and insulin signaling in Alzheimer’s disease: cause or consequence? J Exp Med 213(8):1375–1385. https://doi.org/10.1084/jem.20160493
de la Monte SM, Tong M, Daiello LA, Ott BR (2019) Early-stage Alzheimer’s disease is associated with simultaneous systemic and central nervous system dysregulation of insulin-linked metabolic pathways. J Alzheimers Dis. https://doi.org/10.3233/JAD-180906
Talbot K, Wang HY, Kazi H, Han LY, Bakshi KP, Stucky A et al (2012) Demonstrated brain insulin resistance in Alzheimer’s disease patients is associated with IGF-1 resistance, IRS-1 dysregulation, and cognitive decline. J Clin Invest 122(4):1316–1338. https://doi.org/10.1172/JCI59903
de la Monte SM (2012) Contributions of brain insulin resistance and deficiency in amyloid-related neurodegeneration in Alzheimer’s disease. Drugs. 72(1):49–66. https://doi.org/10.2165/11597760-000000000-00000
Rivera EJ, Goldin A, Fulmer N, Tavares R, Wands JR, de la Monte SM (2005) Insulin and insulin-like growth factor expression and function deteriorate with progression of Alzheimer’s disease: link to brain reductions in acetylcholine. J Alzheimers Dis 8(3):247–268
Schubert M, Gautam D, Surjo D, Ueki K, Baudler S, Schubert D et al (2004) Role for neuronal insulin resistance in neurodegenerative diseases. Proc Natl Acad Sci U S A 101(9):3100–3105. https://doi.org/10.1073/pnas.0308724101
Craft S (2007) Insulin resistance and Alzheimer’s disease pathogenesis: potential mechanisms and implications for treatment. Curr Alzheimer Res 4(2):147–152
de la Monte SM (2014) Type 3 diabetes is sporadic Alzheimer’s disease: mini-review. Eur Neuropsychopharmacol 24(12):1954–1960. https://doi.org/10.1016/j.euroneuro.2014.06.008
Luchsinger JA (2010) Type 2 diabetes, related conditions, in relation and dementia: an opportunity for prevention? J Alzheimers Dis 20(3):723–736. https://doi.org/10.3233/JAD-2010-091687
Newsholme P, Morgan D, Rebelato E, Oliveira-Emilio HC, Procopio J, Curi R et al (2009) Insights into the critical role of NADPH oxidase(s) in the normal and dysregulated pancreatic beta cell. Diabetologia. 52(12):2489–2498. https://doi.org/10.1007/s00125-009-1536-z
Ho L, Qin W, Pompl PN, Xiang Z, Wang J, Zhao Z et al (2004) Diet-induced insulin resistance promotes amyloidosis in a transgenic mouse model of Alzheimer’s disease. FASEB J 18(7):902–904. https://doi.org/10.1096/fj.03-0978fje
Kitazawa M, Cheng D, Tsukamoto MR, Koike MA, Wes PD, Vasilevko V et al (2011) Blocking IL-1 signaling rescues cognition, attenuates tau pathology, and restores neuronal β-catenin pathway function in an Alzheimer’s disease model. J Immunol 187(12):6539–6549. https://doi.org/10.4049/jimmunol.1100620
Kempuraj D, Thangavel R, Natteru PA, Selvakumar GP, Saeed D, Zahoor H et al (2016) Neuroinflammation induces neurodegeneration. J Neurol Neurosurg Spine 1:1
Kirouac L, Rajic AJ, Cribbs DH, Padmanabhan J (2017) Activation of Ras-ERK signaling and GSK-3 by amyloid precursor protein and amyloid beta facilitates neurodegeneration in Alzheimer’s disease. eNeuro. 4(2). https://doi.org/10.1523/ENEURO.0149-16.2017
Bassani TB, Vital MA, Rauh LK (2015) Neuroinflammation in the pathophysiology of Parkinson's disease and therapeutic evidence of anti-inflammatory drugs. Arq Neuropsiquiatr 73(7):616–623. https://doi.org/10.1590/0004-282X20150057
Longpré F, Garneau P, Christen Y, Ramassamy C (2006) Protection by EGb 761 against beta-amyloid-induced neurotoxicity: involvement of NF-kappaB, SIRT1, and MAPKs pathways and inhibition of amyloid fibril formation. Free Radic Biol Med 41(12):1781–1794. https://doi.org/10.1016/j.freeradbiomed.2006.08.015
O’Neill LA, Kaltschmidt C (1997) NF-kappa B: a crucial transcription factor for glial and neuronal cell function. Trends Neurosci 20(6):252–258
Marchegiani F, Matacchione G, Ramini D, Marcheselli F, Recchioni R, Casoli T et al (2019) Diagnostic performance of new and classic CSF biomarkers in age-related dementias. Aging (Albany NY) 11(8):2420–2429. https://doi.org/10.18632/aging.101925
Braak H, Braak E (1991) Neuropathological stageing of Alzheimer-related changes. Acta Neuropathol 82(4):239–259
Folstein MF, Folstein SE, McHugh PR (1975) “Mini-mental state”. A practical method for grading the cognitive state of patients for the clinician. J Psychiatr Res 12(3):189–198
Nelson-Gray RO (1991) DSM-IV: empirical guidelines from psychometrics. J Abnorm Psychol 100(3):308–315
Kopkova A, Sana J, Fadrus P, Slaby O (2018) Cerebrospinal fluid microRNAs as diagnostic biomarkers in brain tumors. Clin Chem Lab Med 56(6):869–879. https://doi.org/10.1515/cclm-2017-0958
Funding
This study was funded by the Basque Government (IT989-16).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Soraya Herrera-Espejo and Borja Santos-Zorrozua share first authorship
Elixabet Lopez-Lopez and África Garcia-Orad share last authorship
Electronic supplementary material
ESM 1
(PDF 856 kb)
Supplementary Table 1
(XLSX 41.4 kb)
Supplementary Table 2
(XLSX 15.6 kb)
Supplementary Table 3
(XLSX 34.9 kb)
Supplementary Table 4
(XLSX 63.1 kb)
Supplementary Table 5
(XLSX 80 kb)
Rights and permissions
About this article
Cite this article
Herrera-Espejo, S., Santos-Zorrozua, B., Álvarez-González, P. et al. A Systematic Review of MicroRNA Expression as Biomarker of Late-Onset Alzheimer’s Disease. Mol Neurobiol 56, 8376–8391 (2019). https://doi.org/10.1007/s12035-019-01676-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-019-01676-9