Abstract
Plasma microRNAs (miRNAs) have been proposed as potential biomarkers in Alzheimer’s disease (AD). Here, we explored their use as early sensors of the preclinical phase of the disease, when brain pathology is being developed and no cognitive loss is detected. For this purpose, we analyzed a set of ten mature plasma miRNAs in symptomatic patients with AD from a cohort that also included healthy controls (HC) and patients with preclinical Alzheimer’s disease (PAD) (cohort 1). Plasmas from subjects with Parkinson’s disease (PD) were used to control for disease specificity. We found that miR-15b-5p, miR-34a-5p, miR-142-3p, and miR-545-3p levels significantly distinguished AD from PD and HC subjects. We next examined the expression of these four miRNAs in plasma from subjects with PAD. Among these, miR-34a-5p and miR-545-3p presented good diagnostic accuracy to distinguish both AD and PAD from HC subjects, according to the receiver operating characteristic (ROC) curve analysis. Both miRNAs also demonstrated a significant positive correlation with Aβ1–42 levels in cerebrospinal fluid (CSF). Taking into account the clinical potential of these findings, we decided to validate the diagnostic accuracy of miR-34a-5p and miR-545-3p in plasma samples from an independent cohort (cohort 2), in which we did not observe the alterations described by us and others in AD and PAD samples. Although miR-34a-5p and miR-545-3p might be promising early biomarker candidates for AD, our study highlights possible sources of variability in miRNA analysis across hospitals, which currently prevents their use as reliable clinical tools.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Converging evidence from both genetic at-risk cohorts and clinically normal older individuals suggests that the pathophysiological process of Alzheimer’s disease (AD) is characterized by a first, preclinical stage of slow and asymptomatic development that can last for decades, during which biological changes associated with the emerging pathology are already present. This stage is followed by clinical symptoms of mild cognitive impairment (MCI) and a final stage of AD dementia. As a result, the pathology is diagnosed too late, when effective therapeutic intervention is difficult [1, 2]. Early detection and early intervention in preclinical AD subjects (PAD, defined as a biomarker positive while asymptomatic) would help provide the basis for preventive approaches involving modifiable and lifestyle risk factors of disease progression [3, 4]. Within this context, there is an increasing interest in secondary prevention trials in persons with preclinical AD (e.g., A4 study, which is a 3-year pharmacological trial for testing an anti-Aβ antibody [5]).
Recent studies have reported encouraging data on the possibility of diagnosis of AD pathophysiology in MCI stages based on the analysis of proteins, antibodies, or miRNA in human blood [6–8]. However, there are only few biomarker changes currently known to precede the appearance of clinical symptoms that can be used to define preclinical Alzheimer’s disease (PAD): (1) brain accumulation of amyloid-β (Aβ) detectable by the positron emission tomography (PET) imaging technique and (2) low-Aß1–42 levels in cerebrospinal fluid (CSF), elevated CSF total tau (t-tau), and phosphorylated tau (p-tau). Invasiveness, complexity, and/or the economic burden of PET and CSF collection render them unsuitable for routine clinical screening [2, 9–13]. Thus, the finding of biomarkers to be measured by non-invasive and relatively inexpensive techniques remains a pending and extremely necessary task. Quantification of biomarkers in the bloodstream complies with these requirements and can be performed from different biological samples (i.e., whole blood, plasma, and serum). Among them, EDTA-treated plasma samples seem to be more reliable for some analyses since coagulation is not required and less blood cell lysis is observed [14–17].
Recently, microRNAs (miRNAs) have drawn attention as potential peripheral biomarkers of brain and systemic physiological disturbances [18]. These small molecules (approximately 22 nucleotides) bind to their target messenger RNA (mRNA), inhibiting its translation or promoting its degradation; thus, these represent one of the main epigenetic mechanisms for the regulation of gene expression [19]. Since miRNAs play an important role in the developing nervous system and in the physiology of high-order brain functions, their alteration can contribute to the manifestation of neurological disorders [20–26]. Particularly, during the last years, many studies in both AD animal models and patients have highlighted the impact of miRNA dysregulation on AD pathophysiology [27–31]. miRNAs can reach the blood circulation through different mechanisms, including being exported from the cell through exosomes, through shedding vesicles, or in association with RNA-binding proteins. It is thought that they can mediate paracrine, endocrine, and/or juxtacrine signaling, as exosomes and shedding vesicles can donate their miRNA to recipient cells by the process of endocytosis, phagocytosis, direct fusion with the plasma membrane, or uptake of the RNA-binding protein-miRNA complex through specific cell membrane receptors [32–35]. Circulating miRNA might represent biomarkers of several age-associated diseases [36]. In particular, some circulating miRNAs in CSF, whole blood, plasma, and serum appear to be altered in both AD animal models and patients [6, 7, 27, 37–52]. However, methodological heterogeneity among studies and other uncontrolled factors (e.g., bodily fluid collection and storage, miRNA purification protocols, miRNA normalization strategies) has led to contradictory results in the literature; thus, there is a great need for validation studies and standardization of procedures [27, 53, 54].
Here, we have analyzed whether plasma levels of miRNA characterized as AD candidate biomarkers are already altered in PAD. For this, we examined miRNA expression in plasma from subjects classified as having AD compared with PAD subjects and healthy control (HC) subjects. Moreover, we compared data from two independent cohorts to examine the replicability of the results in samples obtained at different hospitals. Findings and limitations for the use of miRNA as biomarkers of AD are discussed.
Materials and Methods
Subjects
Subjects were recruited at Hospital Clínic de Barcelona (cohort 1 n = 20–21/group; groups HC, AD, PAD, and PD) and at Hospital de Sant Pau de Barcelona (cohort 2 n = 15/group; groups HC, AD, PAD). All participants gave informed written consent. Subjects were selected based on their clinical and neuropsychological characteristics and their levels of AD CSF biomarkers Aβ1-42, tau phosphorylated at the threonine 181 position (p-tau), and total tau protein (t-tau). All participants were Caucasians from Barcelona, Spain.
Selection Criteria and Demographics
HC subjects were healthy individuals with cognitive performance within the normal range without biomarker-level abnormalities. PAD subjects were defined according to National Institute on Aging–Alzheimer’s Association (NIA-AA) criteria [2], including cognitive performance within the normal range (performance within 1.5 standard deviations [SD]) in all tests from a specific test series, no significant psychiatric symptoms or previous neurological disease, and decreased CSF Aβ1-42.
AD subjects were selected according to International Working Group (IWG) consensus criteria (Dubois B. et al. 2007) and NIA-AA criteria [2]. They presented decreased CSF Aβ42 levels (<500 pg/ml for cohort 1, <550 pg/ml for cohort 2) plus high CSF t-tau (>450 pg/ml for cohort 1, >350 pg/ml for cohort 2) and CSF p-tau (>75 pg/ml for cohort 1; >61 pg/ml for cohort 2) [55–57].
Demographic and clinical characteristics of cohorts 1 and 2 are summarized in Table 1 .
Apolipoprotein E (APOE) genotyping was performed by PCR-based restriction enzyme digestion and restriction length polymorphism analysis (RLPA) on polyacrylamide gels as described previously [58].
Regarding patients with PD, all individuals were characterized with clinically definite PD [59]. Uncertain PD diagnosis, severe depression, and dementia were exclusion criteria.
Total RNA Isolation from Human Plasma
Blood and CSF samples were obtained from all subjects at the same visit. Blood samples were processed within 2 h after collection with EDTA-containing tubes (BD Vacutainer) and centrifuged at 2000×g at 4 °C for 10 min. Plasma was then separated and stored in 1 ml aliquots at −80 °C until further use.
Samples were thawed on ice and total RNA was isolated from 300 μl of plasma utilizing the mirVana™ PARIS™ Isolation Kit (Applied Biosystems) according to the manufacturer’s instructions for liquid samples. To allow for normalization of sample-to-sample variation in RNA isolation, 12.5 fmols of synthetic Caenorhabditis elegans miRNA cel-miR-39 (Qiagen) was added to each sample during the first step of the RNA isolation protocol. RNA was then eluted with 100 μl of H2O according to the manufacturer’s protocol.
Measurement of Plasma miRNA Levels Using TaqMan qRT-PCR Assays
RNA was reverse transcribed using the TaqMan miRNA Reverse Transcription Kit and miRNA-specific stem-loop primers (Applied Biosystems) in a 5-μl reverse transcription (RT) reaction containing 1.67 μl of the RNA extract, 0.05 μl of 100 mM dNTP (with dTTP), 0.33 μl of MultiScribe™ reverse transcriptase (50 U/uL), 0.5 μl of 10× RT buffer, 0.063 μl of RNase inhibitor (20 U/μl), 1.387 μl of RNase-free water, and 1uL of 5× miRNA-specific stem-loop RT primer (Applied Biosystems, Foster, CA, USA), as previously described [60, 61] (Online resource 1). RT reactions were performed in triplicate for each specific miRNA and were incubated at 16 °C for 30 min, at 42 °C for 30 min, and at 85 °C for 5 min and then maintained at 4 °C. The complementary DNA (cDNA) products were stored at −20 °C until analysis.
For relative gene expression analysis of both miR-34a-5p and miR-301a-3p, the RT products were preamplified prior to real-time quantitative PCR to enhance sensitivity, in that the corresponding fluorescence signal was undetectable without the preamplification step. The preamplification step was carried out in a small-scale (5 μl) reaction comprised of 2.5 μl Taqman PreAmp Master Mix (2×) (Applied Biosystems), 1.25 μl Taqman miRNA assay (0.2× in TE) (Applied Biosystems), and 1.25 μl undiluted RT product. The preamplification reactions were incubated at 95 °C for 10 min, followed by 14 cycles of 95 °C for 15 s and at 60 °C for 4 min and then maintained at 4 °C.
For real-time quantitative PCR (qRT-PCR), 1 μl of the cDNA product was employed as a template in a 10-μl reaction containing 0.5 μl of TaqMan miRNA Assay (Applied Biosystem), 3.5 μl of RNase-free water, and 5 μl of QuantiMix Easy Probes Kit Master Mix (Biotools, Spain). The qRT-PCR was performed with Bio-Rad CFX96 real-time PCR system (Bio-Rad) at 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min. Duplicates were run for each sample on a 96-well plate. Data were then analyzed with Bio-Rad CFX Manager (Bio-Rad) with the automatic Ct setting for assigning baseline and threshold for Ct determination. The relative expression level of each specific miRNA was calculated using the 2-∆∆Ct method after normalization to the mean of internal housekeeping miRNA (hsa-miR-106a-5p and hsa-miR-17-5p) and to the spiked cel-miR-39.
Statistical Analysis
Comparisons among groups were performed by the Student t test or non-parametric statistical tests for independent samples, the Mann–Whitney U test, and Kruskal–Wallis test, together with the corresponding post-hoc tests, as indicated. Receiver operating characteristic (ROC) curve analyses were conducted to assess the diagnostic value of the miRNA for distinguishing patients with AD or subjects with PAD from HC. Area under the curve (AUC) values and the associated maximal values of specificity and sensibility are indicated throughout the manuscript. Partial correlations (controlling for group effect), Spearman’s bivariate correlations, and multiple linear regression models were utilized to analyze the relationship between miRNA plasma levels and physiological and/or neuropsychological and clinical variables (CSF Aβ1–42 levels, p-tau, t-tau, age, gender, the APOE allele variant, the mini mental state examination [MMSE] and the Geriatric Depression Scale [GDS]). Data were normalized (square root or log10) for performing partial correlation analysis. P values of <0.05 were considered statistically significant. Statistical outliers (≥2 SD from the mean) were removed from the analyses. The statistical analysis was performed using the Statistical Package for Social Sciences (SPSS, ver. 15.0) and MedCalc (ver. 14.12.0) [62].
Results
Expression and Specificity of Plasma miRNA in Alzheimer’s Disease
Ten mature microRNAs have been previously reported to be altered in plasma from AD patients compared with HC (i.e., miR-34a-5p, miR-146a-5p, miR-34c-5p, let-7d-5p, let-7g-5p, miR-15b-5p, miR-142-3p, miR-191-5p, miR-301a-3p, and miR-545-3p) [38, 41, 43] (Online resource 2).
Here, we evaluated the levels of these miRNAs in plasma samples from HC and AD and determined the disease specificity of the validated miRNA using plasma samples from sex- and age-matched patients with PD. We did not include in our study miRNAs reported to be altered in biological samples other than plasma. Since autopsy studies demonstrate that a substantial percentage of MCI patients does not develop AD pathology, we did not include miRNAs described to be altered in MCI (see Online resource 2).
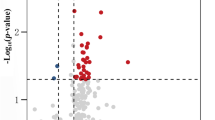
We found that only six of the set of ten miRNAs showed alterations that were consistent with previous reports. Plasma miR-15b-5p, miR-34a-5p, miR-142-3p, and miR-545-3p levels were significantly lower in AD samples than in HC (U(39) = 96.0, Z = −2.62, P < 0.01; U(42) = 100.0, Z = −3.03, P < 0.01; U(40) = 121.0, Z = −2.13, P < 0.05; U(36) = 80.0, Z = −2.58, P < 0.01, respectively). AD plasma samples also presented reduced levels of let-7d-5p and miR-146a-5p compared with HC (U(40) = 128, Z = −1.94, P = 0.053; U(35) = 100.0, Z = −1.91, P = 0.056). In contrast to previous studies [41, 43], we did not find alterations in miR-34c-5p, let-7g-5p, miR-191-5p, and miR-301a-3p plasma levels in patients with AD compared with HC. Data are summarized in Table 2 (HC and AD).
The diagnostic accuracy of miRNA differentially expressed between patients with AD and HC subjects was further tested by the ROC-curve analysis (Fig. 1). The AUC for miR-34a-5p was 0.77, indicating good diagnostic value for AD, with 76.19 % sensitivity and 71.43 % specificity. Similarly, the AUC for both miR-545-3p and miR-15b-5p was 0.75, indicating clinical relevance [63], with 94.12 and 72.22 % of sensitivity and 76.01 and 71.43 % of specificity, respectively. On the other hand, the AUC for miR-142-3p, let-7d-5p, and miR-146a-5p were lower than 0.70, indicating poor diagnostic value for AD.
Receiver operating characteristic (ROC) plots for plasma miRNAs differentially expressed between HC and AD from cohort 1. The true positive rate (sensitivity %) is plotted as a function of the false positive rate (100 %—specificity %) for the six miRNAs individually. Area under the curve (AUC) values, % sensitivity, % specificity, and % accuracy at cut-off points are indicated in each plot as AUC, %Sens., %Spec., and %Acc., respectively
In order to check for disease specificity, we measured those miRNAs found to be altered in AD samples in plasma samples from PD (Table 2, PD).
Kruskal–Wallis test analysis showed that miR-15b-5p, miR-34a-5p, miR-142-3p, and miR-545-3p levels were different among HC, AD, and PD groups (χ 2(2) = 22.92, P < 0.001; χ 2(2) = 12.86, P < 0.01; χ 2(2) = 21.54, P < 0.001; χ 2(2) = 13.87, P < 0.001). The same statistical analysis showed that let-7d-5p and miR-146a-5p levels were similar among groups (χ 2(2) = 3.53, P = 0.17); χ 2(2) = 4.10, P = 0.13, thus discarding these two miRNAs as candidates as specific AD biomarkers.
The corresponding post-hoc pairwise comparison tests revealed that miR-15b-5p, miR-34a-5p, miR-142-3p, and miR-545-3p levels were significantly different between patients with AD and patients with PD (P < 0.01). While, miR-34a-5p and miR-545-3p did not show different levels between HC and patients with PD (P = 0.800, P = 0.322, respectively), miR-15b-5p and miR-142-3p presented higher levels in patients with PD compared with HC (P < 0.05, P < 0.01, respectively).
Expression of miRNA miR-15b-5p, miR-34a-5p, miR-142-3p, and miR-545-3p in Plasma Samples from Subjects in the Preclinical Phase of AD
In order to determine whether the validated miRNA (those differentiating AD from HC and PD) could represent early biomarkers of AD, we measured their plasma levels in 21 sex- and age-matched samples from PAD subjects [2, 64] (Table 2, PAD).
As determined by the Kruskal–Wallis non-parametric test, all four miRNAs showed differences among the HC, AD, PD, and PAD groups (miR-15b-5p χ 2(3) = 26.46, P < 0.001; miR-34a-5p χ 2(3) = 21.83, P < 0.01; miR-142-3p χ 2(3) = 26.56, P < 0.001; miR-545-3p χ 2(3) = 21.97, P < 0.001).
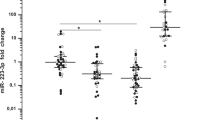
Post-hoc pairwise comparison tests showed that miR-34a-5p and miR-545-3p presented reduced levels in PAD subjects compared to both HC and patients with PD (PAD subjects vs. HC P < 0.001, P < 0.01; PAD subjects vs. PD patients P < 0.001, P < 0.001, respectively), while their expression levels were similar to those detected in AD samples (P = 0.785, P = 0.870, respectively). In contrast, miR-15b-5p and miR-142-3p expression levels did not differ between HC and PAD subjects (P = 0.760, P = 0.122, respectively) (Online resource 3, Fig. 2).
Expression levels of validated miRNAs in cohort 1. Mean ± standard error (SE) are indicated. Cel-39-mir (spike-in external control) and the means of hsa-miR-106a-5p and hsa-miR-17-5p (endogenous housekeeping) were employed for normalization. Values were normalized relative to the HC. Kruskal–Wallis test and corresponding post-hoc paired-group comparisons were performed. The results are indicated as ***P value <0.001, **P value <0.01, *P value <0.05, and +P value <0.1. Healthy control (HC), Alzheimer disease (AD), preclinical Alzheimer disease (PAD), and Parkinson’s disease (PD)
ROC-curve analysis revealed good diagnostic values for both miR-34a-5p and miR-545-3p (AUC values of 0.80 and 0.81, diagnostic accuracy of 78.34 and 76.78 %, respectively). Most importantly, our results indicate that both miR-34a-5p and miR-545-3p present good early diagnostic value for distinguishing HC from subjects presenting decreased CSF Aβ1-42 (<500 pg/ml) (including patients with AD and PAD subjects) (Fig. 3).
Receiver operating characteristic (ROC) plots of miR-34a-5p and miR-545-3p for HC vs. individuals with decreased CSF Aβ1–42 levels (AD + PAD) from cohort 1. The true positive rate (sensitivity %) is plotted as a function of the false positive rate (100 %—specificity %) for each miRNA individually. Area under the curve (AUC) values, % sensitivity, % specificity, and % accuracy at cut-off points are indicated in each plot as AUC, %Sens., %Spec., and %Acc., respectively
We assessed the potential effect of gender or the presence of the APOE ε4 allele for both miR-34a-5p and miR-545-3p by two-way analysis of variance (ANOVA) analysis. Our results showed no significant differences between females and males (miR-34a-5p n = 33 vs. 29, F(1, 58) = 1.25, P = 0.27; miR-545-3p n = 29 vs. 26, F(1, 51) = 0.018, P = 0.89), as well as between APOE ε4 allele non-carriers and APOE ε4 allele carriers (miR-34a-5p n = 39 vs. 23, F(1, 58) = 0.349, P = 0.56; miR-545-3p n = 36 vs. 19, F(1, 51) = 1.707, P = 0.20).
Associations Between Validated miRNA and Other Parameters
Interestingly, miR-15b-5p, miR34a-5p, miR-142-3p, and miR-545-3p plasma levels positively correlate with each other (P < 0.01) as well as with Aβ1–42 levels in CSF (ρr = 0.452, P < 0.001; ρr = 0.384, P < 0.01; ρr = 0.293 and P < 0.05; ρr = 0.309 and P < 0.05, respectively). On the other hand, miR-34a-5p was the only miRNA associated with t-tau, exhibiting a negative correlation (ρr = −0.266, P < 0.05). None of the miRNA was correlated with p-tau.
In order to determine the strongest factors predicting Aβ1–42 levels in CSF, we entered the data from the four validated miRNA for AD (miR-15b-5p, miR-34a-5p, miR-142-3p, and miR-545-5p) together with gender, age, and APOE ε4 allele variables into a multiple linear regression model. Our results highlighted miR-34a-5p as the best predicting miRNA for Aβ1–42 levels in CSF, also influenced significantly by age and the presence of APOE ε4 allele, F(3, 49) = 10.488, P < 0.001 (Table 3).
Validation of Plasma miR-34a-5p and miR-545-3p as Biomarkers of AD and PAD Across Hospitals
In order to test the replicability of our results, we measured the two miRNAs identified here as potential early biomarkers of AD in an independent cohort (cohort 2; Hospital de Sant Pau de Barcelona, Spain).
While the majority of sample characteristics were comparable between cohorts 1 and 2, they presented some significant differences. Mean age was significantly lower in all groups from cohort 2 (t(126) = 6.36, P < 0.001), and mean levels of CSF Aβ, t-tau, and p-tau in the PAD group were significantly different between the cohorts (U(36) = 74.5, Z = −2.663, P < 0.01; U(36) = 71.0, Z = −2.78, P < 0.01; U(36) = 57.0, Z = −3.23, P < 0.01, respectively) (Table 1).
As an internal control, we compared the plasma levels of two housekeeping miRNAs between cohorts. As expected, both miR-106a-5p and miR-17 were similarly expressed between cohorts (coefficient of variation (CV) = 12 and 11 %, respectively) (Online resource 4).
However, in contrast to previous data, including our own results from cohort 1, the miR-34a-5p and miR-545-3p plasma levels from cohort 2 were not diminished in patients with AD as compared with HC (P = 1.00). Moreover, PAD subjects demonstrated higher plasma levels of miR-545-3p and a tendency toward higher levels of miR-34a-5p compared with HC (P < 0.05, P = 0.07, respectively), as well as higher plasma levels of miR-545-3p compared with AD (P < 0.05) (Table 4, Online resource 5). As a consequence, in this case, ROC curves did not show clinically relevant diagnostic values for distinguishing HC from subjects presenting decreased CSF Aβ1-42 (<500 pg/ml) (both in patients with AD and PAD subjects) (miR-34a-5p AUC = 0.624, 46.67 % sensitivity, 86.67 % specificity; miR-545-3p AUC = 0.648, 55.17 % sensitivity, 80 % specificity).
Discussion
Previous studies have identified a set of ten plasma miRNAs as potentially involved in the pathogenesis of AD. The aims of this study were as follows: (i) to validate the alteration of these miRNAs in plasma from patients with AD, using samples of PD subjects as controls for disease specificity (cohort 1), (ii) to explore the potential use of the validated miRNA as early biomarkers for the preclinical phase of AD (cohort 1), and (iii) to explore the consistency of our results across hospitals from the same city (cohort 2). Our data showed that only six of the ten previously described miRNAs were validated in patients with AD, four of whom demonstrated disease specificity utilizing PD samples as control (miR-15b-5p, miR-34a-5p, miR-142-3p, and miR-545-3p). Moreover, two of these miRNAs (miR-34a-5p and miR-545-3p) presented good diagnostic accuracy for distinguishing both PAD subjects and patients with AD from HC and PD controls. Although these findings might be promising, we did not reproduce these results in samples from cohort 2, highlighting some clinical and methodological issues in the use of plasma miRNA as biomarkers for AD.
In the first cohort, we were able to reproduce previously reported differences between patients with AD and HC in plasma levels of let-7d-5p, miR-15b-5p, miR-34a-5p, miR-142-3p, miR-146a-5p, and miR-545-3p [38, 43]. Thereby, these six miRNAs may represent a reliable miRNA signature of AD because they maintained their diagnostic potential across different studies and countries. In particular, miR-15b-5p, miR-34a-5p, and miR-545-3p presented the best diagnostic value among the six validated miRNA, reaching AUC values with high clinical relevance (>0.75) [63]. In contrast with previous reports [41, 43], we did not find significant changes in patients with AD compared with HC for let-7g-5p, miR-34c-5p, miR-191-5p, and miR-301a-5p plasma levels.
A limitation of the previous studies that have investigated the potential of plasma miRNA as biomarkers for AD is the lack of control of disease specificity in age- and sex-matched subjects with an unrelated neurodegenerative disorder. In our study, we included samples of patients with PD, which showed that some of the miRNAs found to be altered in AD samples were not specific for AD pathology. Indeed, let-7d-5p and miR-146a-5p showed reduced plasma levels in PD plasmas compared with HC, thus suggesting common mechanisms concerning the deregulation of these specific miRNA in both AD and PD pathologies. Contrariwise, patients with PD presented significantly higher plasma levels of miR-15b-5p, miR-34a-5p, miR-142-3p, and miR-545-3p compared with patients with AD, thus supporting the specificity of the downregulation of these four miRNAs in AD. Moreover, according to our analyses in PAD subjects from cohort 1, our data indicate that miR-34a-5p and miR-545-3p represent potential early plasma biomarkers for AD, which are already altered in PAD when compared with HC. Both miRNA exhibited good diagnostic values for predicting decreased CSF Aβ1-42 in AD and PAD subjects. Particularly, the multiple linear regression model revealed that the best miRNA for predicting CSF Aβ1–42 levels is miR-34a-5p and that both AD and PAD subjects with lower CSF Aβ1–42 levels also display lower miR-34a-5p plasma levels. Our data are consistent with several studies that have also found upregulation of miR-34a-5p in brains from AD mouse models, as well as its downregulation in CSF and plasma samples from patients with AD [38, 65–68].
Contrary to our findings in plasma, miR-34a-5p and miR-545-3p have been not identified by other authors as altered in serum from AD subjects [36, 40, 51]. Indeed, it is frequent that miRNAs detected in plasma are not detected in serum and vice versa. Tiberio et al. [69] reviewed numerous studies comparing different plasma/serum preparation protocols and analyzing differences between the two types of biological fluids in terms of circulating miRNAs. McDonald et al. [70], Mitchell et al. [61], and Wang et al. [71] reported inconsistencies between miRNA levels in plasma and serum. To explain this difference, it has been postulated that the variability may be due, at least in part, to miRNA release from blood cells (such as platelets) during the coagulation and centrifugation processes [71] [60, 72].
From a mechanistic point of view, miR-34a-5p is upregulated in affected brain regions of AD patients and several AD mouse models [66, 67, 73]. In the brain, the upregulation of miR-34a-5p correlates with concurrent repression of its target genes involved in synaptic plasticity, such as presynaptic-related proteins VAMP2 and SYT1, the NR2A subunit of the NMDA glutamate receptor, the potassium/sodium channel HCN1, and the glutamate-gated ion channel family member GLUR1 [74]. In addition, miR-34a can inhibit the pentose phosphate pathway (PPP) in neurons, which is essential for its long-term survival (it increases oxidative stress contributing to neurodegeneration), and disrupt mitochondrial oxidative phosphorylation by repressing the expression of electron transport chain components [74]. Other important targets of this miRNA are the antiapoptotic protein BCL-2, which inhibits caspases and promotes neuroprotection [38, 67], and the nicotinamide adenine dinucleotide-dependent deacetylase SIRT1 [75]. The latter is a well characterized neuroprotective factor involved in the regulation of Aβ production and downstream targets such as ROCK1, AMPK, NF-kB, or ADAM10, and it is also downregulated in brains of AD patients [76, 77, 78]. Intriguingly, miR-34a-5p is also altered in age-related macular degeneration (AMD), a late-onset neurodegenerative retinal disease which shares several clinical and pathological features with AD such as detrimental intra- and extracellular iron and lipofuscin deposits, increased oxidative stress, and mitochondrial and lysosomal dysfunction [79, 80]. Overall, mir-34a-5p represses several targets involved in synaptic, energy, and Aβ metabolism. Regarding miR-545-3p, one of its targets is the mRNA of LDL receptor-related protein 1 (LRP1), a plasma membrane receptor of APOE involved in Aβ clearance across the blood brain barrier (BBB) and cholesterol importation into neurons [81]. Patients with AD exhibit reduced levels of LRP1 and it has been proposed as a potential target for AD therapy [82, 83]. Another target of this miRNA is the presynaptic vesicle protein synaptophysin [81]. AD patients show reduced levels of synaptophysin in the hippocampus, and this correlates with their cognitive decline [84, 85].
In summary, a downregulation of miR-34a-5p and miR-545-3p in AD plasma samples has been reported in different studies, including ours [38, 43, 65, 68]. Moreover, a downregulation of some of their targets together with an upregulation of miR-34a has been observed in the brain from AD patients. Notably, inversed alterations in biological fluids compared with brain levels have also been reported for other AD and other neurological disorder biomarkers [38, 86, 87]. The possibility of a mechanism involving disruption in the secretion or clearance systems for these miRNAs leading to their accumulation in brain and the reduction of their levels in plasma warrants further studies. This is the first time that two miRNAs are reported to be altered in preclinical AD subjects; nevertheless, these findings need further validation due to the small number of participants included in our study and the fact that all the samples were collected in the Barcelona metropolitan area.
Regarding the second cohort, our results demonstrated no alterations in miR-34a-5p and miR-545-3p expression levels in AD or in PAD plasma samples. These data are especially difficult to explain in AD samples, in that we validated in cohort 1 low levels of these two miRNAs previously reported by other authors [38, 43]. This lack of reproducibility across hospitals evidences the need to identify and standardize the sources of variability that might be causing the inconsistencies between cohorts.
Different factors can account for the variability across miRNA studies including methodological heterogeneity, conditions of sample collection and storage, differences in miRNA purification protocols and miRNA normalization strategies, and environmental influences [27, 53, 54]. In our study, the methodology for collection and sample processing employed by the two centers was similar. The fact that the expression levels of two housekeeping miRNAs (miR-106a-5p and miR-17) were stable between cohorts suggests that the lack of reproducibility observed for miR-34a-5p and miR-545-3p miRNAs may not be related with technical issues such as sample collection and processing.
One of the main differences between cohorts was their mean age, which in cohort 2 was significantly younger than in all previous studies examining miRNA alterations in AD, including our cohort 1. Several evidences indicate that age is a critical variable to consider when defining thresholds or cut-off values for some biomarkers. Van Harten et al. have recently proposed that differences in age could be underlying non-reproducible results among miRNA biomarker studies in AD [88, 89], and Kester et al. found that the association of Aβ42, t-tau, and p-tau CSF levels with APOE genotype is modified by age in both controls and AD patients [90]. Similarly, biomarkers such as t-tau or prostate-specific antigen (PSA) are adjusted for age [91, 92]. On the other hand, as reviewed by many authors [93–98], multiple studies have shown that miRNA expression is altered following exposure to environmental factors. Some of these factors include sensorial elements such as light, noise or odors, physical activity, diet, environmental contaminants such as lead or aluminum, drugs such as alcohol or nicotine, and psychological stressors such as chronic social stress. Figure 4 summarizes findings, proposed mechanisms, and current limitations discussed here.
miR-34a-5p and miR-545-3p as potential biomarkers for AD: a summary of proposed mechanisms and limitations. Both miR-34a-5p and miR-545-3p showed decreased levels in plasma from clinical and preclinical AD subjects in cohort 1, indicating their potential value as early AD biomarkers. Both miRNAs are involved in synaptic and energy metabolism, Aβ production and clearance, and apoptosis (summary of the corresponding targets indicated in squares). A downregulation of both miRNAs in AD plasma samples and in CSF in the case of miR-34a-5p has been reported in different studies, including ours [38, 43, 65, 68]. A downregulation of its targets together with an upregulation of the miRNA itself in the case of miR-34a-5p has been observed in brain from AD patients. Notably, inversed alterations in biological fluids compared with brain levels have also been reported for other AD and other neurological disorder biomarkers [38, 86, 87]. The lack of reproducibility in cohort 2 indicates that the conditions of sample processing across hospitals as well as the contribution of several individual variables (such as age, genetic, health, and environmental factors) need to be further characterized to determine the accuracy of these miRNAs as early biomarkers for AD. The possibility of a mechanism involving disruption in the secretion or clearance systems for these miRNAs leading to their accumulation in brain and the reduction of their levels in plasma warrants further studies. PPP, pentose phosphate pathway; ETC, electron transport chain
Secondary prevention is based on low specificity and low cost biomarkers that allow the detection of preclinical subjects [3, 4, 99]. Accordingly, here, we propose plasma levels of miR-34a-5p and miR-545-3p, both involved in AD pathogenesis, as potential early biomarkers of the disease. Further studies are required to better explore the influence of individual and environmental factors on the alterations of these miRNAs. Advances in this field will help develop blood-based biomarker tests for early detection of AD as valuable tools for the delay of the disease progression [3, 4, 99].
References
Morris JC (2005) Early-stage and preclinical Alzheimer disease. Alzheimer Dis Assoc Disord 19(3):163–165
Sperling RA et al. (2011) Toward defining the preclinical stages of Alzheimer’s disease: recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement 7(3):280–292
Molinuevo JL et al. (2016) Ethical challenges in preclinical Alzheimer’s disease observational studies and trials: results of the Barcelona summit. Alzheimers Dement 12(5):614–622
Dubois B et al. (2016) Preclinical Alzheimer’s disease: definition, natural history, and diagnostic criteria. Alzheimers Dement 12(3):292–323
Sperling RA et al. (2014) The A4 study: stopping AD before symptoms begin? Sci Transl Med 6(228):228fs13
Cheng, L., et al. (2014) Prognostic serum miRNA biomarkers associated with Alzheimer’s disease shows concordance with neuropsychological and neuroimaging assessment. Mol Psychiatry 20(10):1188–1196
Sheinerman KS et al. (2013) Plasma microRNA biomarkers for detection of mild cognitive impairment: biomarker validation study. Aging (Albany NY) 5(12):925–938
Ray S et al. (2007) Classification and prediction of clinical Alzheimer’s diagnosis based on plasma signaling proteins. Nat Med 13(11):1359–1362
Jack CR Jr et al. (2010) Hypothetical model of dynamic biomarkers of the Alzheimer’s pathological cascade. Lancet Neurol 9(1):119–128
Jack CR Jr et al. (2013) Tracking pathophysiological processes in Alzheimer’s disease: an updated hypothetical model of dynamic biomarkers. Lancet Neurol 12(2):207–216
Hansson O et al. (2006) Association between CSF biomarkers and incipient Alzheimer’s disease in patients with mild cognitive impairment: a follow-up study. Lancet Neurol 5(3):228–234
Villemagne VL et al. (2013) Amyloid beta deposition, neurodegeneration, and cognitive decline in sporadic Alzheimer’s disease: a prospective cohort study. Lancet Neurol 12(4):357–367
Mistur R et al. (2009) Current challenges for the early detection of Alzheimer’s disease: brain imaging and CSF studies. J Clin Neurol 5(4):153–166
Guder WG, Nayaranan S, Wisser H, Zawta B (2014) Diagnostic samples: from the patient to the laboratory: the impact of preanalytical variables on the quality of laboratory results. Wiley
Banfi G, Deom A, Fräser CG, Hagemann P, Henny J, Kaliner A, Leppänen EA, Narayanan S, Neumaier M, Gomes MPA, Probst R (2009) Serum, plasma or whole blood? Which anticoagulants to use? Journal of Laboratory Medicine:297–312
Lehmann S et al. (2009) Preanalytical guidelines for clinical proteomics investigation of biological fluids. Ann Biol Clin (Paris) 67(6):629–639
Pritchard CC et al. (2012) Blood cell origin of circulating microRNAs: a cautionary note for cancer biomarker studies. Cancer Prev Res (Phila) 5(3):492–497
Kim YK (2015) Extracellular microRNAs as biomarkers in human disease. Chonnam Med J 51(2):51–57
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136(2):215–233
Wang W, Kwon EJ, Tsai LH (2012) MicroRNAs in learning, memory, and neurological diseases. Learn Mem 19(9):359–368
Mastroeni D et al. (2011) Epigenetic mechanisms in Alzheimer’s disease. Neurobiol Aging 32(7):1161–1180
Van den Hove DL et al. (2014) Epigenetically regulated microRNAs in Alzheimer’s disease. Neurobiol Aging 35(4):731–745
Xu B et al. (2012) MicroRNA dysregulation in neuropsychiatric disorders and cognitive dysfunction. Neurobiol Dis 46(2):291–301
Eacker SM, Dawson TM, Dawson VL (2009) Understanding microRNAs in neurodegeneration. Nat Rev Neurosci 10(12):837–841
Dwivedi Y (2016) Pathogenetic and therapeutic applications of microRNAs in major depressive disorder. Prog Neuro-Psychopharmacol Biol Psychiatry 64:341–348
Cao DD, Li L, Chan WY (2016) MicroRNAs: key regulators in the central nervous system and their implication in neurological diseases. Int J Mol Sci 17(6)
Femminella GD, Ferrara N, Rengo G (2015) The emerging role of microRNAs in Alzheimer’s disease. Front Physiol 6:40
Maes OC et al. (2009) MicroRNA: implications for Alzheimer disease and other human CNS disorders. Curr Genomics 10(3):154–168
Delay C, Mandemakers W, Hebert SS (2012) MicroRNAs in Alzheimer’s disease. Neurobiol Dis 46(2):285–290
Cosin-Tomas M et al. (2014) Epigenetic alterations in hippocampus of SAMP8 senescent mice and modulation by voluntary physical exercise. Front Aging Neurosci 6:51
Noh H et al. (2014) Prediction of miRNA-mRNA associations in Alzheimer’s disease mice using network topology. BMC Genomics 15:644
Weilner S et al. (2013) Secretion of microvesicular miRNAs in cellular and organismal aging. Exp Gerontol 48(7):626–633
Gibbings DJ et al. (2009) Multivesicular bodies associate with components of miRNA effector complexes and modulate miRNA activity. Nat Cell Biol 11(9):1143–1149
Iguchi H, Kosaka N, Ochiya T (2010) Secretory microRNAs as a versatile communication tool. Commun Integr Biol 3(5):478–481
Chen X et al. (2012) Secreted microRNAs: a new form of intercellular communication. Trends Cell Biol 22(3):125–132
Dorval V, Nelson PT, Hebert SS (2013) Circulating microRNAs in Alzheimer’s disease: the search for novel biomarkers. Front Mol Neurosci 6:24
Sheinerman KS et al. (2012) Plasma microRNA biomarkers for detection of mild cognitive impairment. Aging (Albany NY) 4(9):590–605
Kiko T et al. (2014) MicroRNAs in plasma and cerebrospinal fluid as potential markers for Alzheimer’s disease. J Alzheimers Dis 39(2):253–259
Tan L et al. (2014) Circulating miR-125b as a biomarker of Alzheimer’s disease. J Neurol Sci 336(1–2):52–56
Tan L et al. (2014) Genome-wide serum microRNA expression profiling identifies serum biomarkers for Alzheimer’s disease. J Alzheimers Dis 40(4):1017–1027
Bhatnagar S et al. (2014) Increased microRNA-34c abundance in Alzheimer’s disease circulating blood plasma. Front Mol Neurosci 7:2
Geekiyanage H et al. (2012) Blood serum miRNA: non-invasive biomarkers for Alzheimer’s disease. Exp Neurol 235(2):491–496
Kumar P et al. (2013) Circulating miRNA biomarkers for Alzheimer’s disease. PLoS One 8(7):e69807
Bekris LM, Leverenz JB (2015) The biomarker and therapeutic potential of miRNA in Alzheimer’s disease. Neurodegener Dis Manag 5(1):61–74
Bekris LM et al. (2013) MicroRNA in Alzheimer’s disease: an exploratory study in brain, cerebrospinal fluid and plasma. Biomarkers 18(5):455–466
Hu YB et al. (2016) Diagnostic value of microRNA for Alzheimer’s disease: a systematic review and meta-analysis. Front Aging Neurosci 8:13
Zi Y et al. (2015) Circulating MicroRNA as potential source for neurodegenerative diseases biomarkers. Mol Neurobiol 52(3):1494–1503
Tan L, Yu JT, Hu N, Non-coding RNA (2013) In Alzheimer’s disease. Mol Neurobiol 47(1):382–393
Sorensen SS, Nygaard AB, Christensen T (2016) miRNA expression profiles in cerebrospinal fluid and blood of patients with Alzheimer’s disease and other types of dementia—an exploratory study. Transl Neurodegener 5:6
Muller M et al. (2016) Validation of microRNAs in cerebrospinal fluid as biomarkers for different forms of dementia in a multicenter study. J Alzheimers Dis 52(4):1321–1333
Danborg PB et al. (2014) The potential of microRNAs as biofluid markers of neurodegenerative diseases—a systematic review. Biomarkers 19(4):259–268
Leidinger P et al. (2013) A blood based 12-miRNA signature of Alzheimer disease patients. Genome Biol 14(7):R78
Sheinerman KS, Umansky SR (2013) Circulating cell-free microRNA as biomarkers for screening, diagnosis and monitoring of neurodegenerative diseases and other neurologic pathologies. Front Cell Neurosci 7:150
Moldovan L et al. (2014) Methodological challenges in utilizing miRNAs as circulating biomarkers. J Cell Mol Med 18(3):371–390
Alcolea D et al. (2014) Relationship between beta-secretase, inflammation and core cerebrospinal fluid biomarkers for Alzheimer’s disease. J Alzheimers Dis 42(1):157–167
Alcolea D et al. (2015) Amyloid precursor protein metabolism and inflammation markers in preclinical Alzheimer disease. Neurology 85(7):626–633
Antonell A et al. (2011) Different profiles of Alzheimer’s disease cerebrospinal fluid biomarkers in controls and subjects with subjective memory complaints. J Neural Transm (Vienna) 118(2):259–262
Zivelin A et al. (1997) Improved method for genotyping apolipoprotein E polymorphisms by a PCR-based assay simultaneously utilizing two distinct restriction enzymes. Clin Chem 43(9):1657–1659
Hughes AJ et al. (1992) Accuracy of clinical diagnosis of idiopathic Parkinson’s disease: a clinico-pathological study of 100 cases. J Neurol Neurosurg Psychiatry 55(3):181–184
Kroh EM et al. (2010) Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qRT-PCR). Methods 50(4):298–301
Mitchell PS et al. (2008) Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci U S A 105(30):10513–10518
Stephan C et al. (2003) Comparison of eight computer programs for receiver-operating characteristic analysis. Clin Chem 49(3):433–439
Fan J, Upadhye S, Worster A (2006) Understanding receiver operating characteristic (ROC) curves. CJEM 8(1):19–20
Dubois B et al. (2010) Revising the definition of Alzheimer’s disease: a new lexicon. Lancet Neurol 9(11):1118–1127
Rohn TT et al. (2008) Lack of pathology in a triple transgenic mouse model of Alzheimer’s disease after overexpression of the anti-apoptotic protein Bcl-2. J Neurosci 28(12):3051–3059
Li X et al. (2011) Circulatory miR34a as an RNAbased, noninvasive biomarker for brain aging. Aging (Albany NY) 3(10):985–1002
Wang X et al. (2009) miR-34a, a microRNA up-regulated in a double transgenic mouse model of Alzheimer’s disease, inhibits bcl2 translation. Brain Res Bull 80(4–5):268–273
Ferreiro E et al. (2007) Bcl-2 overexpression protects against amyloid-beta and prion toxicity in GT1-7 neural cells. J Alzheimers Dis 12(3):223–228
Tiberio P et al. (2015) Challenges in using circulating miRNAs as cancer biomarkers. Biomed Res Int 2015:731479
McDonald JS et al. (2011) Analysis of circulating microRNA: preanalytical and analytical challenges. Clin Chem 57(6):833–840
Wang K et al. (2012) Comparing the MicroRNA spectrum between serum and plasma. PLoS One 7(7):e41561
Cheng HH et al. (2013) Plasma processing conditions substantially influence circulating microRNA biomarker levels. PLoS One 8(6):e64795
Cogswell JP et al. (2008) Identification of miRNA changes in Alzheimer’s disease brain and CSF yields putative biomarkers and insights into disease pathways. J Alzheimers Dis 14(1):27–41
Sarkar S et al. (2016) Expression of microRNA-34a in Alzheimer’s disease brain targets genes linked to synaptic plasticity, energy metabolism, and resting state network activity. Brain Res 1646:139–151
Yamakuchi M, Ferlito M, Lowenstein CJ (2008) miR-34a repression of SIRT1 regulates apoptosis. Proc Natl Acad Sci U S A 105(36):13421–13426
Julien C et al. (2009) Sirtuin 1 reduction parallels the accumulation of tau in Alzheimer disease. J Neuropathol Exp Neurol 68(1):48–58
Ng F, Wijaya L, Tang BL (2015) SIRT1 in the brain-connections with aging-associated disorders and lifespan. Front Cell Neurosci 9:64
Pallas M et al. (2008) Modulation of SIRT1 expression in different neurodegenerative models and human pathologies. Neuroscience 154(4):1388–1397
Kaarniranta K et al. (2011) Age-related macular degeneration (AMD): Alzheimer’s disease in the eye? J Alzheimers Dis 24(4):615–631
Hill JM, Pogue AI, Lukiw WJ (2015) Pathogenic microRNAs common to brain and retinal degeneration; recent observations in Alzheimer’s disease and age-related macular degeneration. Front Neurol 6:232
Hsu SD et al. (2014) miRTarBase update 2014: an information resource for experimentally validated miRNA-target interactions. Nucleic Acids Res 42(Database issue):D78–D85
Martiskainen H et al. (2013) Targeting ApoE4/ApoE receptor LRP1 in Alzheimer’s disease. Expert Opin Ther Targets 17(7):781–794
Donahue JE et al. (2006) RAGE, LRP-1, and amyloid-beta protein in Alzheimer’s disease. Acta Neuropathol 112(4):405–415
Sze CI et al. (1997) Loss of the presynaptic vesicle protein synaptophysin in hippocampus correlates with cognitive decline in Alzheimer disease. J Neuropathol Exp Neurol 56(8):933–944
Yuki D et al. (2014) DHA-PC and PSD-95 decrease after loss of synaptophysin and before neuronal loss in patients with Alzheimer’s disease. Sci Rep 4:7130
Glatt SJ et al. (2005) Comparative gene expression analysis of blood and brain provides concurrent validation of SELENBP1 up-regulation in schizophrenia. Proc Natl Acad Sci U S A 102(43):15533–15538
Ma X et al. (2014) Expression, regulation and function of microRNAs in multiple sclerosis. Int J Med Sci 11(8):810–818
van Harten AC et al. (2013) Cerebrospinal fluid Abeta42 is the best predictor of clinical progression in patients with subjective complaints. Alzheimers Dement 9(5):481–487
van Harten AC et al. (2015) Differential expression of microRNA in cerebrospinal fluid as a potential novel biomarker for Alzheimer’s disease. J Alzheimers Dis 47(1):243–252
Kester MI et al. (2009) CSF biomarkers in Alzheimer’s disease and controls: associations with APOE genotype are modified by age. J Alzheimers Dis 16(3):601–607
Humpel C (2011) Identifying and validating biomarkers for Alzheimer’s disease. Trends Biotechnol 29(1):26–32
Moul JW et al. (2007) Age adjusted prostate specific antigen and prostate specific antigen velocity cut points in prostate cancer screening. J Urol 177(2):499–503 discussion 503–4
Hollins SL, Cairns MJ (2016) MicroRNA: small RNA mediators of the brains genomic response to environmental stress. Prog Neurobiol 143:61–81
Codocedo JF, Inestrosa NC (2016) Environmental control of microRNAs in the nervous system: implications in plasticity and behavior. Neurosci Biobehav Rev 60:121–138
Flowers E, Won GY, Fukuoka Y (2015) MicroRNAs associated with exercise and diet: a systematic review. Physiol Genomics 47(1):1–11
Xu T et al. (2015) Circulating microRNAs in response to exercise. Scand J Med Sci Sports 25(2):e149–e154
Zacharewicz E, Lamon S, Russell AP (2013) MicroRNAs in skeletal muscle and their regulation with exercise, ageing, and disease. Front Physiol 4:266
Palmer JD et al. (2014) MicroRNA expression altered by diet: can food be medicinal? Ageing Res Rev 17:16–24
Vanderstichele H et al. (2012) Standardization of preanalytical aspects of cerebrospinal fluid biomarker testing for Alzheimer’s disease diagnosis: a consensus paper from the Alzheimer’s biomarkers standardization initiative. Alzheimers Dement 8(1):65–73
Acknowledgments
This work is supported by grants CSD2010-00045 and SAF2012-39852 from the Spanish Ministry of Economy and Competitiveness (MINECO) and the European Regional Development Fund (ERDF), Instituto de Salud Carlos III (PI11/03035, PI11/02425, PI14/01561), and a grant from the Fundació la Marató de TV3 (20142610). Marta Cosín-Tomás is supported by a predoctoral fellowship from MINECO (FPU 2013).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Subjects were recruited at Hospital Clínic de Barcelona (cohort 1 n = 20–21/group; groups HC, AD, PAD, and PD) and at Hospital de Sant Pau de Barcelona (cohort 2 n = 15/group; groups HC, AD, PAD). All participants gave informed written consent.
Electronic Supplementary Material
Online resource 1
(PDF 66 kb)
Online resource 2
(PDF 88 kb)
Online resource 3
(PDF 177 kb)
Online resource 4
(PDF 64 kb)
Online resource 5
(PDF 102 kb)
Rights and permissions
About this article
Cite this article
Cosín-Tomás, M., Antonell, A., Lladó, A. et al. Plasma miR-34a-5p and miR-545-3p as Early Biomarkers of Alzheimer’s Disease: Potential and Limitations. Mol Neurobiol 54, 5550–5562 (2017). https://doi.org/10.1007/s12035-016-0088-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-016-0088-8