Abstract
The retina is a delicate neural tissue responsible for light signal capturing, modulating, and passing to mid-brain. The brain then translated the signals into three-dimensional vision. The mature retina is composed of more than 50 subtypes of cells, all of which are developed from a pool of early multipotent retinal progenitors, which pass through sequential statuses of oligopotent, bipotent, and unipotent progenitors, and finally become terminally differentiated retinal cells. A transitional progenitor model is proposed here to describe how intrinsic developmental programs, along with environmental cues, control the step-by-step differentiation during retinogenesis. The model could elegantly explain many current findings as well as predict roles of intrinsic factors during retinal development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
In the adult vertebrate retina, there are six major classes of neurons and one class of Müller glial cells. The retinal neurons include cone and rod photoreceptors and horizontal, amacrine, bipolar, and ganglion cells. Photons from the light are first caught by cone or rod photoreceptors, which convert them into signals and relay the signals to the interneurons, the bipolar cells. The horizontal cells modulate the signals from photoreceptors before they are relayed to the bipolar cells. Again, the amacrine cells integrate and modulate the signals from bipolar cells before they are relayed to retinal ganglion cells (RGCs). Finally, the RGCs transferred the signals to the specific brain regions and the brain generated the three-dimensional vision. As the only major non-neuronal cell type, Müller cells provide scaffolding supports, nutrients, metabolite recycling, etc. for the neurons. A special cell type, microglia, is not from retinal origin but arises from circulating monocytes/macrophages. Microglia is involved in immune surveillance and cleaning of the retina.
Retinal cells are highly diversified. Most of the major class of retinal cells can be further categorized into subgroups according to their morphology and function. For example, RGCs have more than 30 subgroups [1]; amacrine cells have over 29 subgroups [2, 3]. All these retinal cells are originally developed from a small group of multipotent progenitor cells in the optic vesicle. This complicated process is delicately controlled. A popular intrinsic model has been proposed to describe this process [4–8]. The main point of the model is that progenitors go through intrinsically determined competence states, during which they are capable of giving rise to a limited subset of retinal cell types under the influence of extrinsic signals. The model could gracefully explain how the intrinsic and extrinsic factors together control retinal cell genesis. However, details of how multipotent progenitors develop into each cell type are still murky. Here, I summarized the recent advances on retinal and neural development, focusing on topics such as cell division and differentiation, retinal intrinsic programs, intrinsic program versus stochastic mechanism, transitional progenitor model, early-born progenitors (EBPs) versus late-born progenitors (LBPs), and so on.
Cell division and differentiation
Retinal progenitor cell (RPC) pool contains a limited number of cells. To guarantee the normal size of the retina, the multipotent progenitors undergo fast cycles of self-replication before initiating cell differentiation. The self-replicating process is under the tight control of Rax, Meis1&2, Pax6, Notch1, Shh, and other factors [9–14]. Mutations in this group of factors generally cause microphthalmia or more serious phenotypes in human and animal models, due to far fewer progenitor cells available in the progenitor pool.
There are two scenarios when a progenitor cell undergoes mitosis (Fig. 1). One scenario is that the progenitor is evenly divided into two identical daughter cells, either two identical progenitors (Fig. 1a) or two neurons/glias of the same type (Fig. 1c). Two progenitors are usually generated at the early developmental stage when the progenitor pool needs to be expanded rapidly. On the other hand, two differentiated daughter cells will be produced at the late stage of organogenesis. The second scenario is that the progenitor divides asymmetrically and gives rise to one progenitor cell and one neuron/glia (Fig. 1b), or two neuron/glial cells of different types (Fig. 1d). Examples of these scenarios were provided in the study of Ath5 in the wild-type and lakritz mutant mice [15] and in a work by Cayouette and colleague [16, 17].
Four cell division modes for symmetric and asymmetric divisions. (A) A progenitor cell divided symmetrically and gave rise to two undifferentiated progenitor cells. (B) A progenitor cell divided asymmetrically and resulted in one undifferentiated daughter cell and one differentiated daughter cell. (C) A progenitor cell divided symmetrically and two differentiated daughter cells of same type were born. (D) A progenitor cell divided asymmetrically, and two differentiated daughter cells of different types were generated. In E&F, cleavage plane aligns in parallel or vertical to the apical-basal axis, namely horizontal or vertical division, respectively. (E) Horizontal division usually leads to symmetric division of two identical daughter cells, each inheriting equal amount of cell fate determinants. (F) Vertical division tends to asymmetrically divide a progenitor into two daughter cells of different size and morphology, each inheriting unequal amount of cell fate determinants
At the onset of neurogenesis, the progenitors gradually switch from proliferative, symmetric to neurogenic, asymmetric divisions. The shift is shown to be associated with a change of the orientation of the mitotic spindles in the dividing progenitors. It is proposed in CNS development that if the cleavage plane aligned in parallel with apical-basal axis, it is termed horizontal division; whereas the cleavage plane is perpendicular to apical-basal axis, it is named vertical division (Fig. 1e, f). It is found that horizontal division of a mitotic cell usually divides symmetrically and gives two identical daughter cells, while the vertical division tends to be asymmetric and gives two daughter cells of different size and morphology. This may partially attribute to the unequal inheritance of cell surface molecule Notch and intracellular cell fate determinants Numb and etc. [16, 18]. Observations by Kosodo et al. suggest that the vertical division is mostly asymmetric, but it can be either proliferative, symmetric or neurogenic, asymmetric, depending on the equal or unequal inheritance of cell constitutes [19]. Interestingly, the definition of horizontal versus vertical division of neural progenitors is similar to Dr. Hans Spemann’s definition of sagittal versus frontal division of fertilized eggs, and the apical side is counterpart of the gray crescent region.
The key question is, what is the molecular machinery determining symmetric versus asymmetric inheritance? The findings from drosophila to mammals show that the polarity is controlled by evolutionarily conserved protein complexes: the Par proteins (Par3-Par6-aPKC), the heterotrimeric G protein complex (Gαi-Pins (LGN/Gpsm2 in mammals)-Mud (NuMA in mammals)), and Insc that can bind Par3 and Pins [20, 21]. The Notch signaling pathway is also critical in regulating the division polarity. Notch is found to promote the asymmetric localization of the protein Numb and the positioning of the cleavage furrow [22]. Eya1 and Gαs also controls spindle orientation and asymmetrical division by regulating Notch signaling [23, 24]. However, it is still unclear how each fate-determining transcription factor specifically controls the asymmetrical cell division by regulating these polarity proteins during cell cycle. Mostly, the regulation was initiated at G1 phase, as G1 phase cells have permissive epigenetic environment that allows developmental programs to be activated, and G1 phase cells favorably respond to fate-determining extrinsic cues [25].
Retinal intrinsic programs specifying cell fates
Emphasis has been put on intrinsic developmental programs in specifying cell fates. However, what composes an intrinsic program is not clearly defined. An intrinsic program is established in evolution and is mainly encoded by a cascade of intrinsic factors, especially transcription factors. An intrinsic program can be initiated by a signal from intrinsic or extrinsic factors, or another program. Once initiated, a program is hardly reversible in vivo; however, it can be forcibly aborted by another program, which sometimes leads to cell apoptosis. A cell in homeostasis is in a balanced state of a series of programs. Once the balance is destroyed in the progenitor, the cell either changes its status (i.e., dividing or differentiation) or commits to apoptosis. A program may have two or more alternative subprograms at a branching point. Selective execution of one of the subprograms depends on the inputs of extrinsic signals, which underlies the basis that cell fate decision relies on both intrinsic and extrinsic factors.
In general, there are three major types of intrinsic programs in retinal or other tissue development (Fig. 2). The first type is the linear program, which usually occurs at the late stages of development (Fig. 2a). The linear program reflects the simplest biological causal relationship, and one program specifies only one cell-type fate. Disruption of the program leaves the cell nowhere to go, and it usually commits to cell apoptosis. For instance, Bhlhb4 is specifically expressed in the rod bipolar cells and guides the terminal differentiation of the cells. Deletion of Bhlhb4 causes the cells take the path to apoptosis in the mouse retina (Fig. 2a, [26]).
Typical intrinsic programs in retinal and other tissue development. a The linear program specifies only one cell type fate, and disruption of the program usually results in cell apoptosis. b The branching program has two or more subprograms to choose at the branching point. Which subprogram(s) to execute depends on the extrinsic signal input and will lead to different cell fate(s). For example, the extrinsic factor x will force the progenitor to take the RGC path while inhibiting Amacrine and Horizontal path. Disruption of one subprogram will result in more execution of the other programs. In this case, more amacrine and horizontal cells will be generated when RGC subprogram is destroyed during development. c The converged program is the contrary to the branching program. Two or more subprograms merged together to define one cell type fate. Block of one of subprograms will generate less target cells to some extent
The second type is the branching program (Fig. 2b), which mostly takes place in the early and peak stage development. The progenitor cell taking the branching program has two to several subprograms to choose, and it depends on the extrinsic signals to pick the right subprogram to continue. The lack of complete reliance on intrinsic signals ensures that there are branching choices and not a linear program. Again, this is how extrinsic and intrinsic factors together determine cell fate. Taken as an example, the early-born progenitor is capable of differentiating into a RGC, horizontal or amacrine cell (Fig. 2b). The fate choices depend on whether the extrinsic signals to activate the Foxn4 or Brn3b subprograms. If Brn3b is activated, the progenitor will terminally differentiate into a RGC. If Foxn4 is triggered, the progenitor will become a horizontal or amacrine cell depending on further choice. If Brn3b gene is knocked out, there would be a temporal increase of horizontal and amacrine cells [27]. Alternatively, if Foxn4 expression is abolished, RGC population would temporally get boosted [28, 29].
The last major type is the converged program (Fig. 2c). Contrary to the branching program, the converged program has two or more subprograms merged together to specify one cell type. Evolutionally considered, this is important since it provides a redundant mechanism to guarantee the generation of a crucial cell type. Interruption of one of the subprograms usually results in a decrease but not a complete loss of target cells. For instance, Brn3b, Isl1, Sox4, and Sox11 are all important for RGC development. Deletion of any one of them would cause partial loss of RGCs, but compound knockout of two factors would lead to a near-complete loss of RGCs [30–32].
In conclusion, three types of programs are all important in fate decision during development. It should be emphasized that none of the programs is isolated in the cell, and they work synergistically in sequential or in parallel to activate/inhibit downstream genes, since conflicting programs usually lead to cell death, which were eliminated during evolution. For instance, Foxn4, Ptf1a [33], and Tfap2α/Tfap2β [34, 35] operate in a sequential cascade to determine horizontal and amacrine cell fates, while RORβ1 seems to work in parallel with Foxn4 to turn on Ptf1a expression [36]. These programs work coordinately to form functional horizontal and amacrine neurons in the retina.
Intrinsic programs versus stochastic mechanism
It is believed that intrinsic developmental programs are the most decisive forces in specifying cell fates [37, 38]. Slater [39] and Gomes [40] et al. found that stochastic mechanism also plays an important role in the process. However, Gomes’s conclusion that stochasticity is the major role during retinal development is in question. If it is largely a stochastic force, one would expect that an approximately equal number of rods, amacrine, bipolar, and Müller cells would be generated during the experiment, obviously this is not the case. Then, it raises the questions: (i) what is underlying the stochastic mechanism, and (ii) when does the stochastic mechanism take over the control? One plausible answer to (i) is random fate choice due to the balance of forces from two or more opposite programs, per se, the balance due to the dose-dependent effect of antagonistic transcription factors. As illustrated in Fig. 3, in postmitotic Otx2+ progenitors, Blimp1 and Vsx2 (Chx10) were the two intrinsic determinants to choose a bipolar or a rod cell fate [41, 42]. There exists a point which I called balance point when Blimp1 and Vsx2 reach equivalence. A small region flanking the balance point, named the stochastic zone, is the key to question (ii). Beyond the left side of stochastic zone, the Blimp1 dominates the Vsx2 and the cell will definitely become a rod, and vice versa. Only in the stochastic zone, the stochastic force has some effects on cell fate choices; however, the ratio should be close to 50:50 for rod and bipolar fates in the zone. As a result, stochasticity causes variations in cell fate determination, but will not affect the overall cell ratio statistically. We may conclude that the intrinsic program(s) is the major driving force, while the stochastic mechanism also plays a minor role and causes variations in the development.
Illustration of stochastic mechanism in the rod versus bipolar cell fate decision. The rod versus bipolar cell fate decision was determined by the dose-dependent effect of Blimp1 and Vsx2 protein. When Blimp1 dominates Vsx2, the cell will become a rod cell; otherwise, it will become a bipolar cell if Vsx2 is dominant. There is a balance point when Blimp1 and Vsx2 reach equivalence. Flanking the balance point is the stochastic zone where the cell fate was decided by stochastic mechanism. In the stochastic zone, the progenitor has equal chances to differentiate into a rod or a bipolar cell
Transitional progenitor model for retinogenesis
A pluripotent progenitor cell usually does not give rise to terminally differentiated cells directly; instead, it generates the multipotent progenitors. The multipotent progenitors lose the pluripotency, are partially determined cells, and can only produce a limited set of cell types. The multipotent progenitors also give birth to more restricted intermediate progenitors, the oligopotent progenitors, which are capable of generating only three or more cell types. The oligopotent cells then differentiate into bipotent and finally into unipotent progenitors that can only produce terminally differentiated progeny. This multi-step commitment seems to be the case in the retinogenesis as well as in other tissue development, such as in hematopoietic genesis. Each step is controlled by one or a serial of intrinsic programs. The intermediate oligopotent, bipotent, and unipotent retinal progenitors are defined as transitional progenitors in this model. The transitional progenitor model emphasizes the heterogeneity of RPCs, reflecting the fact that the retina, at any given developing point, is a bag of mixed multipotent and transitional RPCs with terminal cells.
Here, a model is proposed that the intrinsic programs define the competence state of transitional progenitors and specify retinal cell types and subtypes. Previous popular models describing retinal development were reviewed in details recently [43]. These models suggest that the RPCs gradually lose the competence during the developmental process [4, 5, 43]; however, the transitional progenitor model here proposes that the multipotent RPCs keep their competence to generate all potential retinal cell types. The ‘restricted’ competence is due to a gradual decrease of multipotent RPCs and a steady increase of the transitional progenitors and terminal cells over the developing course. The model is supported by discoveries that the late RPC pool is comprised of mixed population with stem cell-like multipotent progenitors and heterogeneous lineage-restricted progenitors [44, 45]. Even in the mature retina, there are adult stem cells (equivalent to multipotent RPCs by definition) that have the full potential to generate all retinal cell types and maintain the integrity of the retina. Another difference is that previous models suggest that differentiating RPCs follow a linear route during the retinal development (Fig. 4a), while the transitional progenitor model proposes a tree-structure route which is more reasonable (Fig. 4b). For example, readers could be misled by the linear route to believe that amacrine cells are generated from the RGC-fate-incompetent progenitors and are born later than RGCs. As a matter of fact, the early progenitors are competent for RGC, amacrine, horizontal, and cone cell fates, and birthdays of these cells are interweaved.
Models of RPC differentiation and fate determination. a One representative intrinsic model is shown. According to the model, multipotent RPCs pass through temporal order of states of competence over the course of development, and get more and more restricted in cell fate choices from one state to another. During each competence state, the RPC is capable of terminally differentiating into one or several specific subtypes of cells (for simplicity, only one type of cell is depicted in each state in the cartoon). b The model proposed here emphasizes transitional changes and complex heterogeneity among progenitors. The cell fate determination follows an order from multipotent progenitor to oligopotent then bipotent and unipotent progenitor, and the RPCs finally differentiated into particular cells. At any given developmental stages, the retina is composed of mixed population of multipotent, oligopotent, bipotent, and unipotent progenitors and terminally differentiated cells. The multipotent RPCs are getting fewer and fewer in number, but they keep their competence and are not restricted in fate choices. The fates of multipotent progenitors were determined by both the environmental cues and intrinsic programs
The crucial transitional progenitors, including the early-born and late-born oligopotent progenitors, bipotent progenitors, unipotent progenitors, and the associated transcription factors defining the intrinsic programs in these progenitors, are listed in Fig. 5 and discussed below.
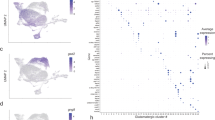
A hypothetical model of intrinsic programs for hierarchy lineage specification in the mouse retina. The multipotent progenitors give rise to transitional progenitors from oligopotent to bipotent and unipotent. RGC, horizontal, amacrine, and early-born cone photoreceptor cells originate from a common pool of intermediate progenitors (EBPs). Bipolar, Müller, and late-born photoreceptor cells share a pool of common intermediate progenitors (LBPs). The differentiated cells can give negative feedback signals to its progenitor pool. Critical factors are listed on each pathway
EBP groups
There is evidence to suggest that there is a pool of common oligopotent progenitors for early-born cell types, such as horizontal, amacrine, RGC, and cone photoreceptor cells (Fig. 5). All EBPs are Pax6-positive. Pax6 is a transcription factor containing paired box and homeodomain and is highly conserved in vertebrates. In the chick, the onset of Pax6 starts at Hamburger and Hamilton stage 8.5 in primordial eye field and persists at high levels throughout optic cup morphogenesis [46]. The continuous expression of Pax6 activates Math5 expression in some of the EBPs [47]. Math5 endorses the progenitors to differentiate into RGCs, and Math5-negative progenitors become amacrine, horizontal, and photoreceptor cells. In the Math5 null retina, there is an increase of amacrine and cone cells but not bipolar cells [48], supporting the idea that RGCs, cones, and horizontal and amacrine cells share common EBPs, since promotion of one cell type will inhibit the other cell types as they share the same pool of progenitors. Continual expression of Pax6 can also activate the expression of Otx2 and Trβ2 in some of the progenitors and leads to generation of cone but not rod photoreceptors [49], which explains why cone photoreceptors are generated earlier than rod photoreceptors. It also lends evidence to support the idea that rods were developed from cones evolutionarily, since the embryonic development is a reflection of evolution history to some extent.
Onecut1 (OC1) and Onecut2 (OC2) are initiated at E11.5 which is later than Pax6. They seem to be specifically expressed in EBPs at the stage. Compound deletion of both will result in complete absence of horizontal cells and early-born cholinergic amacrine cells, and partial losses of RGCs and cone photoreceptors [50, 51]. Other than expressed in EBPs, OC1/2 also expresses in RGCs and horizontal cells at late stages [50]. During RGC development, OC1/2 seems to function independently of Math5.
The intrinsic programs defining unipotent RGC progenitors seem to be complicated. The RGCs have more than 30 subtypes with morphological and functional differences. The specification of the subtypes is decided by differential expression of Brn3b, Dlx1, Dlx2, Islet1, and others [30, 32, 52, 53]. For example, Brn3b (Pou4f2) is a Pou domain and homeobox domain transcription factor expressed in the postmitotic precursors. Expression of Brn3b promotes RGC fate at the expense of amacrine, horizontal, and cone cells [27], which is another evidence supporting the existence of common EBPs.
Horizontal and amacrine cells share a pool of bipotent progenitors that are progenies of EBPs. This is strongly supported by evidence from the Foxn4 and Ptf1a mutant mice. Foxn4, a forkhead box domain transcription factor, is critical in the retinal and other tissue development [28, 54]. In the Foxn4 null retina, all horizontal cells and the majority of amacrine cells are lost; instead, more RGCs and photoreceptors are temporarily generated [28], showing a fate switch from amacrine and horizontal cells to RGCs and photoreceptors. Similar phenotypes are found in the Ptf1a knockout mice [33, 55]. These evidences support the ideas that these four cell types share the common pool of EBPs and that horizontal and amacrine cells rise from the same bipotent progenitors. The activating protein-2 (AP-2) transcription factors Tfap2α and Tfap2β act downstream of Ptf1a and regulate the genesis of horizontal and amacrine cells as well [34, 35].
Horizontal cells are interneurons situated between photoreceptors and bipolar cells and form synaptic connections with both cell types. There are two kinds of horizontal cells in the vertebrates, the axon-bearing and the axon-less ones, though there are only axon-bearing ones in some vertebrates, i.e., rodents. The terminal differentiation of horizontal cell depends on the homeodomain protein Lim1, which is exclusively expressed in the unipotent horizontal progenitors [56]. Loss of Lim1 causes the horizontal cell precursors stuck in the wrong laminar position. As a result, the ectopic horizontal cells adopt a morphology more reminiscent of amacrine cells.
Unlike horizontal cell, the amacrine cell has more than 29 subtypes and its terminal differentiation is more complicated, most possibly controlled by many terminal programs. Transcription factors NeuroD and Math3 are genetically downstream of Ptf1a and redundantly control the genesis of amacrine cells. In NeuroD and Math3 double knockout retinae, all amacrine cells fail to differentiate; however, ganglion and horizontal cells are increased [57–60], which is also consistent with the EBP hypothesis. Similar to NeuroD or Math3, Prdm13 is also a downstream gene of Ptf1a and regulates the development and function of a subset of glycinergic and GABAergic amacrine neurons [61]. Only a few genes are known to control the development of one specific subtype of amacrine cells. For instance, the dopaminergic amacrine cell fate is controlled by the orphan nuclear receptor Nr4a2 (Nurr1) [62], and the development of cholinergic amacrine cells depends on LIM-homeodomain factor Isl1 [63].
LBP groups
Evidence also suggests existence of a pool of common oligopotent progenitors for late-born cell types, including Müller, bipolar, and photoreceptor cells (Fig. 5). The common transcription factors controlling LBPs were unclear; however, LBPs might oscillatorily express Notch1 [64, 65] and temporally express the paired-type homeodomain transcription factor Vsx2 [66]. The onset of Vsx2 expression starts no earlier than Hamburger and Hamilton stage 12 [67]. Vsx2 expression at the early stage of development controls proliferation of RPCs, and Vsx2 mutation causes ocular retardation due to RPC proliferation defects [66].
The progeny fate of LBPs depends on differential expression of Vsx2, Otx2, Blimp1, and Notch1. It has been shown that Vsx2 directly controls bipolar cell genesis by inhibiting rod differentiation [68]. Mutation of Vsx2 results in the total loss of bipolar cells; however, the genesis of horizontal and amacrine cells is pretty much normal in the Vsx2 mutants [66], implying that only LBPs but not EBPs are affected. By regulating Vsx2 and Otx2, Blimp1 promotes photoreceptor cell fate while inhibiting bipolar and Müller cell fates [41, 42, 69]. Ablation of Blimp1 results in far fewer photoreceptors, more bipolar and Müller cells, but unchanged amacrine, horizontal, and RGCs [41, 42], which also supports our hypothesis that Müller, bipolar, and photoreceptor cells share the same pool of LBPs, and consistent with the notion that promotion of one cell fate will inhibit the other cell fates. LBP fate determination is also controlled by transcription factor Math3 and Mash1. Compound knockout of Math3 and Mash1 also results in loss of all bipolar cells, but Müller cells are significantly increased at the cost of bipolar cells [60], suggesting bipolar and Müller cells originate from the same progenitor pool.
Knockout of Otx2 results in loss of all photoreceptor cells in the mouse retina [70], which indicates that there is a repertoire of common bipotent progenitors for cone and rod photoreceptor cells (Fig. 5). It is supported that the default fate of the cone and rod progenitors is S-cone cell, since mutations of numerous genes in the common progenitors unanimously lead to enhanced S-cone syndromes, such as mutations in Trβ2 and Nr2E3 [71, 72]. Expression of wild-type Nrl or Nr2E3 blocks this default pathway in the progenitor and initiates rod cell genesis, while expression of Trβ2 or Rxrγ blocks the default pathway and leads to generation of M-cones [71]. Conditional deletion of Otx2 also leads to a vast increase of GABAergic and glycinergic amacrine cells that span into the space where a normal ONL layer should be. The Müller and bipolar cells also increased due to cell fate switches from photoreceptor cells [70, 73]. Cone-rod homeobox gene Crx works at the downstream of Otx2, it seems not involved in photoreceptor fate specification, but rather to enhance the expression of photoreceptor-related terminal genes [70, 74, 75]. Similarly, transcription factor Mef2d works with Crx to drive the retina-specific gene expression in photoreceptor, but not involved in the fate decision [76, 77].
As one of the most important pathways affecting the development of many tissues, the Notch pathway is not only involved in early RPC maintenance but also in the rod and Müller cell fate determination. Overexpression of Notch1 promotes Müller cell fate [78]. Overexpression of its downstream gene Hesr2 or Hes1 also promotes Müller genesis at the expense of rod cells [78, 79]. Retroviral-mediated conditional ablation of Notch1 at postnatal stage induces the generation of rod photoreceptors at the expense of bipolar and Müller but not amacrine cells [80]. These findings together strongly support that bipolar, Müller, and photoreceptor cells share a pool of common LBPs.
The terminal differentiation of Müller cells involves in Sry-related HMG box gene Sox9. Specific deletion of Sox9 from developing retina resulted in loss of Müller cells [81]. Apparently, other Sry-related HMG box genes were involved in Müller cell development as well, such as Sox2 [82, 83], Sox8 [84], etc. The cone bipolar cells have around nine subsets, and specification programs for the subsets depend on Vsx1 [85], Irx5 [86], Bhlhb5 [87], etc.
Genesis of EBPs and LBPs
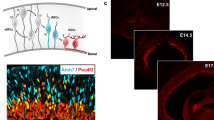
Multipotent RPCs were maintained in the collaborated network of Rx [9, 13], Notch1 [64, 78], Meis1/2 [10], Pten [88], Pax6 [14, 49], etc. A multipotent progenitor’s decision to be either an EBP or a LBP was mostly controlled by extracellular environmental cues, including cell-cell contacts (neighboring effect), interaction between Notchs and ligands [29, 80, 89–91], extrinsic factors such as the BMPs gradient [92–95], hormones and growth factors [96–98], ephrins and receptors [99, 100], etc., which initiated intrinsic developmental programs. Early developmental environment favorably promotes the EBP fates, and late developmental environment prefers LBP fates. Genesis of EBPs, LBPs, and major retinal cell types is in a timely overlapping manner. RGCs initiated at about E11 and were the first cell type differentiated, followed by amacrine, cone, and horizontal cells at about E12 (Fig. 6, [101]). Rod photoreceptors first appeared at around E14 and then joined by bipolar and Müller cells. We could deduct that EBPs arise from E10 to P4 and peak at E16, while LBPs appear from E13 to P10 and peak at P2 (Fig. 6).
Developmental stages for transitional EBPs and LBPs in the mouse retina. EBPs initiated at about E10 and ended at about P3. LBPs started around E13 and ended around P10. The development of EBPs and LBPs overlapped spatiotemporally. The specification timing of RGC, amacrine, horizontal, bipolar, Müller, cone, and rod photoreceptor cells was also illustrated
Conclusion
The transitional progenitor model defines progenitor status from multipotent to oligopotent, bipotent, and unipotent stepwisely. In each transitional status, the competence is controlled by intrinsic developmental programs as well as environmental cues. The multipotent RPCs become fewer and fewer with the ongoing differentiation and are almost depleted in the adult retina. The terminally differentiated retinal cells usually are not capable of proliferating, except some stem cell-like Müller glial cells [102–105] and Lgr5+ amacrine cells [106]. The epithelial cells in the ciliary margin zone were previously identified as retinal stem cells [107, 108]; however, they are unable to differentiate into retinal neurons in vivo or in vitro [109, 110] and thus disqualified as stem cells.
It needs to be pointed out that there are many mechanisms, besides transcriptional control, regulating retinal development, i.e., RNA-level (microRNAs, ncRNAs, etc.) regulation [111, 112], protein modification and degradation [113, 114], epigenetic modification [115, 116], cell death regulation [117, 118], and etc. Neighbor tissues, such as RPE and lens, certainly have indispensable effects on retinal development as well, mostly through the mechanism of mutual induction and inhibition. We will not discuss these topics in further details.
Understanding the programs underlying the developmental process not only will provide insights into how each individual cell type is formed and what function it may have, but also will give clues on how to treat retinal diseases in the future. Recent advances in stem cell researches illustrate the importance of understanding such developmental processes. Many efforts have been made worldwide to induce retina-specific cells from adult somatic cells or induced pluripotent stem cells (iPSCs) [119–121], which shed some light on cell replacement therapies to treat patients in the near future.
References
Baden T, Berens P, Franke K, Roman Roson M, Bethge M, Euler T (2016) The functional diversity of retinal ganglion cells in the mouse. Nature 529(7586):345–350. doi:10.1038/nature16468
Voinescu PE, Kay JN, Sanes JR (2009) Birthdays of retinal amacrine cell subtypes are systematically related to their molecular identity and soma position. J Comp Neurol 517(5):737–750. doi:10.1002/cne.22200
Masland RH (2001) The fundamental plan of the retina. Nat Neurosci 4(9):877–886. doi:10.1038/nn0901-877
Livesey FJ, Cepko CL (2001) Vertebrate neural cell-fate determination: lessons from the retina. Nat Rev Neurosci 2(2):109–118. doi:10.1038/35053522
Cepko CL, Austin CP, Yang X, Alexiades M, Ezzeddine D (1996) Cell fate determination in the vertebrate retina. Proc Natl Acad Sci U S A 93(2):589–595
Watanabe T, Raff MC (1990) Rod photoreceptor development in vitro: intrinsic properties of proliferating neuroepithelial cells change as development proceeds in the rat retina. Neuron 4(3):461–467
Marquardt T (2003) Transcriptional control of neuronal diversification in the retina. Prog Retin Eye Res 22(5):567–577
Cayouette M, Barres BA, Raff M (2003) Importance of intrinsic mechanisms in cell fate decisions in the developing rat retina. Neuron 40(5):897–904
Zhang L, Mathers PH, Jamrich M (2000) Function of Rx, but not Pax6, is essential for the formation of retinal progenitor cells in mice. Genesis 28(3–4):135–142
Heine P, Dohle E, Bumsted-O’Brien K, Engelkamp D, Schulte D (2008) Evidence for an evolutionary conserved role of homothorax/Meis1/2 during vertebrate retina development. Development 135(5):805–811. doi:10.1242/dev.012088
Wall DS, Mears AJ, McNeill B, Mazerolle C, Thurig S, Wang Y, Kageyama R, Wallace VA (2009) Progenitor cell proliferation in the retina is dependent on Notch-independent Sonic hedgehog/Hes1 activity. J Cell Biol 184(1):101–112. doi:10.1083/jcb.200805155
Furukawa T, Kozak CA, Cepko CL (1997) rax, a novel paired-type homeobox gene, shows expression in the anterior neural fold and developing retina. Proc Natl Acad Sci U S A 94(7):3088–3093
Mathers PH, Grinberg A, Mahon KA, Jamrich M (1997) The Rx homeobox gene is essential for vertebrate eye development. Nature 387(6633):603–607. doi:10.1038/42475
Marquardt T, Ashery-Padan R, Andrejewski N, Scardigli R, Guillemot F, Gruss P (2001) Pax6 is required for the multipotent state of retinal progenitor cells. Cell 105(1):43–55
Poggi L, Vitorino M, Masai I, Harris WA (2005) Influences on neural lineage and mode of division in the zebrafish retina in vivo. J Cell Biol 171(6):991–999. doi:10.1083/jcb.200509098
Cayouette M, Raff M (2003) The orientation of cell division influences cell-fate choice in the developing mammalian retina. Development 130(11):2329–2339
Cayouette M, Whitmore AV, Jeffery G, Raff M (2001) Asymmetric segregation of Numb in retinal development and the influence of the pigmented epithelium. J Neurosci 21(15):5643–5651
Chenn A, McConnell SK (1995) Cleavage orientation and the asymmetric inheritance of Notch1 immunoreactivity in mammalian neurogenesis. Cell 82(4):631–641
Kosodo Y, Roper K, Haubensak W, Marzesco AM, Corbeil D, Huttner WB (2004) Asymmetric distribution of the apical plasma membrane during neurogenic divisions of mammalian neuroepithelial cells. EMBO J 23(11):2314–2324. doi:10.1038/sj.emboj.7600223
Siller KH, Doe CQ (2009) Spindle orientation during asymmetric cell division. Nat Cell Biol 11(4):365–374. doi:10.1038/ncb0409-365
Culurgioni S, Mapelli M (2013) Going vertical: functional role and working principles of the protein Inscuteable in asymmetric cell divisions. Cell Mol Life Sci 70(21):4039–4046. doi:10.1007/s00018-013-1319-z
Bhat KM (2014) Notch signaling acts before cell division to promote asymmetric cleavage and cell fate of neural precursor cells. Sci Signal 7(348):ra101. doi:10.1126/scisignal.2005317
El-Hashash AH, Turcatel G, Al Alam D, Buckley S, Tokumitsu H, Bellusci S, Warburton D (2011) Eya1 controls cell polarity, spindle orientation, cell fate and Notch signaling in distal embryonic lung epithelium. Development 138(7):1395–1407. doi:10.1242/dev.058479
Liu K, Lin Q, Wei Y, He R, Shao X, Ding Z, Zhang J, Zhu M et al (2015) Galphas regulates asymmetric cell division of cortical progenitors by controlling Numb mediated Notch signaling suppression. Neurosci Lett 597:97–103. doi:10.1016/j.neulet.2015.04.034
Dalton S (2015) Linking the cell cycle to cell fate decisions. Trends Cell Biol 25(10):592–600. doi:10.1016/j.tcb.2015.07.007
Bramblett DE, Pennesi ME, Wu SM, Tsai MJ (2004) The transcription factor Bhlhb4 is required for rod bipolar cell maturation. Neuron 43(6):779–793. doi:10.1016/j.neuron.2004.08.032
Qiu F, Jiang H, Xiang M (2008) A comprehensive negative regulatory program controlled by Brn3b to ensure ganglion cell specification from multipotential retinal precursors. J Neurosci 28(13):3392–3403. doi:10.1523/JNEUROSCI.0043-08.2008
Li S, Mo Z, Yang X, Price SM, Shen MM, Xiang M (2004) Foxn4 controls the genesis of amacrine and horizontal cells by retinal progenitors. Neuron 43(6):795–807. doi:10.1016/j.neuron.2004.08.041
Luo H, Jin K, Xie Z, Qiu F, Li S, Zou M, Cai L, Hozumi K et al (2012) Forkhead box N4 (Foxn4) activates Dll4-Notch signaling to suppress photoreceptor cell fates of early retinal progenitors. Proc Natl Acad Sci U S A 109(9):E553–E562. doi:10.1073/pnas.1115767109
Gan L, Xiang M, Zhou L, Wagner DS, Klein WH, Nathans J (1996) POU domain factor Brn-3b is required for the development of a large set of retinal ganglion cells. Proc Natl Acad Sci U S A 93(9):3920–3925
Jiang Y, Ding Q, Xie X, Libby RT, Lefebvre V, Gan L (2013) Transcription factors SOX4 and SOX11 function redundantly to regulate the development of mouse retinal ganglion cells. J Biol Chem 288(25):18429–18438. doi:10.1074/jbc.M113.478503
Pan L, Deng M, Xie X, Gan L (2008) ISL1 and BRN3B co-regulate the differentiation of murine retinal ganglion cells. Development 135(11):1981–1990. doi:10.1242/dev.010751
Fujitani Y, Fujitani S, Luo H, Qiu F, Burlison J, Long Q, Kawaguchi Y, Edlund H et al (2006) Ptf1a determines horizontal and amacrine cell fates during mouse retinal development. Development 133(22):4439–4450. doi:10.1242/dev.02598
Jin K, Jiang H, Xiao D, Zou M, Zhu J, Xiang M (2015) Tfap2a and 2b act downstream of Ptf1a to promote amacrine cell differentiation during retinogenesis. Mol Brain 8(1):28. doi:10.1186/s13041-015-0118-x
Bassett EA, Korol A, Deschamps PA, Buettner R, Wallace VA, Williams T, West-Mays JA (2012) Overlapping expression patterns and redundant roles for AP-2 transcription factors in the developing mammalian retina. Dev Dyn 241(4):814–829. doi:10.1002/dvdy.23762
Liu H, Kim SY, Fu Y, Wu X, Ng L, Swaroop A, Forrest D (2013) An isoform of retinoid-related orphan receptor beta directs differentiation of retinal amacrine and horizontal interneurons. Nat Commun 4:1813. doi:10.1038/ncomms2793
Shen Q, Wang Y, Dimos JT, Fasano CA, Phoenix TN, Lemischka IR, Ivanova NB, Stifani S et al (2006) The timing of cortical neurogenesis is encoded within lineages of individual progenitor cells. Nat Neurosci 9(6):743–751. doi:10.1038/nn1694
Kao CF, Lee T (2010) Birth time/order-dependent neuron type specification. Curr Opin Neurobiol 20(1):14–21. doi:10.1016/j.conb.2009.10.017
Slater JL, Landman KA, Hughes BD, Shen Q, Temple S (2009) Cell lineage tree models of neurogenesis. J Theor Biol 256(2):164–179. doi:10.1016/j.jtbi.2008.09.034
Gomes FL, Zhang G, Carbonell F, Correa JA, Harris WA, Simons BD, Cayouette M (2011) Reconstruction of rat retinal progenitor cell lineages in vitro reveals a surprising degree of stochasticity in cell fate decisions. Development 138(2):227–235. doi:10.1242/dev.059683
Brzezinski JA, Lamba DA, Reh TA (2010) Blimp1 controls photoreceptor versus bipolar cell fate choice during retinal development. Development 137(4):619–629. doi:10.1242/dev.043968
Katoh K, Omori Y, Onishi A, Sato S, Kondo M, Furukawa T (2010) Blimp1 suppresses Chx10 expression in differentiating retinal photoreceptor precursors to ensure proper photoreceptor development. J Neurosci 30(19):6515–6526. doi:10.1523/JNEUROSCI.0771-10.2010
Cepko C (2014) Intrinsically different retinal progenitor cells produce specific types of progeny. Nat Rev Neurosci 15(9):615–627. doi:10.1038/nrn3767
Jensen AM, Raff MC (1997) Continuous observation of multipotential retinal progenitor cells in clonal density culture. Dev Biol 188(2):267–279. doi:10.1006/dbio.1997.8645
Turner DL, Cepko CL (1987) A common progenitor for neurons and glia persists in rat retina late in development. Nature 328(6126):131–136. doi:10.1038/328131a0
Hsieh YW, Yang XJ (2009) Dynamic Pax6 expression during the neurogenic cell cycle influences proliferation and cell fate choices of retinal progenitors. Neural Dev 4:32. doi:10.1186/1749-8104-4-32
Riesenberg AN, Le TT, Willardsen MI, Blackburn DC, Vetter ML, Brown NL (2009) Pax6 regulation of Math5 during mouse retinal neurogenesis. Genesis 47(3):175–187. doi:10.1002/dvg.20479
Wang SW, Kim BS, Ding K, Wang H, Sun D, Johnson RL, Klein WH, Gan L (2001) Requirement for math5 in the development of retinal ganglion cells. Genes Dev 15(1):24–29
Oron-Karni V, Farhy C, Elgart M, Marquardt T, Remizova L, Yaron O, Xie Q, Cvekl A et al (2008) Dual requirement for Pax6 in retinal progenitor cells. Development 135(24):4037–4047. doi:10.1242/dev.028308
Wu F, Sapkota D, Li R, Mu X (2012) Onecut 1 and Onecut 2 are potential regulators of mouse retinal development. J Comp Neurol 520(5):952–969. doi:10.1002/cne.22741
Sapkota D, Chintala H, Wu F, Fliesler SJ, Hu Z, Mu X (2014) Onecut1 and Onecut2 redundantly regulate early retinal cell fates during development. Proc Natl Acad Sci U S A 111(39):E4086–E4095. doi:10.1073/pnas.1405354111
Mu X, Fu X, Beremand PD, Thomas TL, Klein WH (2008) Gene regulation logic in retinal ganglion cell development: Isl1 defines a critical branch distinct from but overlapping with Pou4f2. Proc Natl Acad Sci U S A 105(19):6942–6947. doi:10.1073/pnas.0802627105
de Melo J, Du G, Fonseca M, Gillespie LA, Turk WJ, Rubenstein JL, Eisenstat DD (2005) Dlx1 and Dlx2 function is necessary for terminal differentiation and survival of late-born retinal ganglion cells in the developing mouse retina. Development 132(2):311–322. doi:10.1242/dev.01560
Xiang M, Li S (2013) Foxn4: a multi-faceted transcriptional regulator of cell fates in vertebrate development. Sci China Life Sci 56(11):985–993. doi:10.1007/s11427-013-4543-8
Nakhai H, Sel S, Favor J, Mendoza-Torres L, Paulsen F, Duncker GI, Schmid RM (2007) Ptf1a is essential for the differentiation of GABAergic and glycinergic amacrine cells and horizontal cells in the mouse retina. Development 134(6):1151–1160. doi:10.1242/dev.02781
Poche RA, Kwan KM, Raven MA, Furuta Y, Reese BE, Behringer RR (2007) Lim1 is essential for the correct laminar positioning of retinal horizontal cells. J Neurosci 27(51):14099–14107. doi:10.1523/JNEUROSCI.4046-07.2007
Inoue T, Hojo M, Bessho Y, Tano Y, Lee JE, Kageyama R (2002) Math3 and NeuroD regulate amacrine cell fate specification in the retina. Development 129(4):831–842
Morrow EM, Furukawa T, Lee JE, Cepko CL (1999) NeuroD regulates multiple functions in the developing neural retina in rodent. Development 126(1):23–36
Ahmad I, Acharya HR, Rogers JA, Shibata A, Smithgall TE, Dooley CM (1998) The role of NeuroD as a differentiation factor in the mammalian retina. J Mol Neurosci 11(2):165–178
Tomita K, Moriyoshi K, Nakanishi S, Guillemot F, Kageyama R (2000) Mammalian achaete-scute and atonal homologs regulate neuronal versus glial fate determination in the central nervous system. EMBO J 19(20):5460–5472
Watanabe S, Sanuki R, Sugita Y, Imai W, Yamazaki R, Kozuka T, Ohsuga M, Furukawa T (2015) Prdm13 regulates subtype specification of retinal amacrine interneurons and modulates visual sensitivity. J Neurosci 35(20):8004–8020. doi:10.1523/JNEUROSCI.0089-15.2015
Jiang H, Xiang M (2009) Subtype specification of GABAergic amacrine cells by the orphan nuclear receptor Nr4a2/Nurr1. J Neurosci 29(33):10449–10459. doi:10.1523/JNEUROSCI.3048-09.2009
Elshatory Y, Everhart D, Deng M, Xie X, Barlow RB, Gan L (2007) Islet-1 controls the differentiation of retinal bipolar and cholinergic amacrine cells. J Neurosci 27(46):12707–12720. doi:10.1523/JNEUROSCI.3951-07.2007
Mizeracka K, DeMaso CR, Cepko CL (2013) Notch1 is required in newly postmitotic cells to inhibit the rod photoreceptor fate. Development 140(15):3188–3197. doi:10.1242/dev.090696
Dvoriantchikova G, Perea-Martinez I, Pappas S, Barry AF, Danek D, Dvoriantchikova X, Pelaez D, Ivanov D (2015) Molecular characterization of notch1 positive progenitor cells in the developing retina. PLoS One 10(6):e0131054. doi:10.1371/journal.pone.0131054
Burmeister M, Novak J, Liang MY, Basu S, Ploder L, Hawes NL, Vidgen D, Hoover F et al (1996) Ocular retardation mouse caused by Chx10 homeobox null allele: impaired retinal progenitor proliferation and bipolar cell differentiation. Nat Genet 12(4):376–384. doi:10.1038/ng0496-376
Chen CM, Cepko CL (2000) Expression of Chx10 and Chx10-1 in the developing chicken retina. Mech Dev 90(2):293–297
Livne-Bar I, Pacal M, Cheung MC, Hankin M, Trogadis J, Chen D, Dorval KM, Bremner R (2006) Chx10 is required to block photoreceptor differentiation but is dispensable for progenitor proliferation in the postnatal retina. Proc Natl Acad Sci U S A 103(13):4988–4993. doi:10.1073/pnas.0600083103
Wang S, Sengel C, Emerson MM, Cepko CL (2014) A gene regulatory network controls the binary fate decision of rod and bipolar cells in the vertebrate retina. Dev Cell 30(5):513–527. doi:10.1016/j.devcel.2014.07.018
Nishida A, Furukawa A, Koike C, Tano Y, Aizawa S, Matsuo I, Furukawa T (2003) Otx2 homeobox gene controls retinal photoreceptor cell fate and pineal gland development. Nat Neurosci 6(12):1255–1263. doi:10.1038/nn1155
Ng L, Hurley JB, Dierks B, Srinivas M, Salto C, Vennstrom B, Reh TA, Forrest D (2001) A thyroid hormone receptor that is required for the development of green cone photoreceptors. Nat Genet 27(1):94–98
Haider NB, Jacobson SG, Cideciyan AV, Swiderski R, Streb LM, Searby C, Beck G, Hockey R et al (2000) Mutation of a nuclear receptor gene, NR2E3, causes enhanced S cone syndrome, a disorder of retinal cell fate. Nat Genet 24(2):127–131. doi:10.1038/72777
Sato S, Inoue T, Terada K, Matsuo I, Aizawa S, Tano Y, Fujikado T, Furukawa T (2007) Dkk3-Cre BAC transgenic mouse line: a tool for highly efficient gene deletion in retinal progenitor cells. Genesis 45(8):502–507. doi:10.1002/dvg.20318
Furukawa T, Morrow EM, Li T, Davis FC, Cepko CL (1999) Retinopathy and attenuated circadian entrainment in Crx-deficient mice. Nat Genet 23(4):466–470. doi:10.1038/70591
Chen S, Wang QL, Nie Z, Sun H, Lennon G, Copeland NG, Gilbert DJ, Jenkins NA et al (1997) Crx, a novel Otx-like paired-homeodomain protein, binds to and transactivates photoreceptor cell-specific genes. Neuron 19(5):1017–1030
Andzelm MM, Cherry TJ, Harmin DA, Boeke AC, Lee C, Hemberg M, Pawlyk B, Malik AN et al (2015) MEF2D drives photoreceptor development through a genome-wide competition for tissue-specific enhancers. Neuron 86(1):247–263. doi:10.1016/j.neuron.2015.02.038
Omori Y, Kitamura T, Yoshida S, Kuwahara R, Chaya T, Irie S, Furukawa T (2015) Mef2d is essential for the maturation and integrity of retinal photoreceptor and bipolar cells. Genes Cells 20(5):408–426. doi:10.1111/gtc.12233
Furukawa T, Mukherjee S, Bao ZZ, Morrow EM, Cepko CL (2000) rax, Hes1, and notch1 promote the formation of Muller glia by postnatal retinal progenitor cells. Neuron 26(2):383–394
Satow T, Bae SK, Inoue T, Inoue C, Miyoshi G, Tomita K, Bessho Y, Hashimoto N et al (2001) The basic helix-loop-helix gene hesr2 promotes gliogenesis in mouse retina. J Neurosci 21(4):1265–1273
Jadhav AP, Mason HA, Cepko CL (2006) Notch 1 inhibits photoreceptor production in the developing mammalian retina. Development 133(5):913–923. doi:10.1242/dev.02245
Poche RA, Furuta Y, Chaboissier MC, Schedl A, Behringer RR (2008) Sox9 is expressed in mouse multipotent retinal progenitor cells and functions in Muller glial cell development. J Comp Neurol 510(3):237–250. doi:10.1002/cne.21746
Taranova OV, Magness ST, Fagan BM, Wu Y, Surzenko N, Hutton SR, Pevny LH (2006) SOX2 is a dose-dependent regulator of retinal neural progenitor competence. Genes Dev 20(9):1187–1202. doi:10.1101/gad.1407906
Lin YP, Ouchi Y, Satoh S, Watanabe S (2009) Sox2 plays a role in the induction of amacrine and Muller glial cells in mouse retinal progenitor cells. Invest Ophthalmol Vis Sci 50(1):68–74. doi:10.1167/iovs.07-1619
Muto A, Iida A, Satoh S, Watanabe S (2009) The group E Sox genes Sox8 and Sox9 are regulated by notch signaling and are required for Muller glial cell development in mouse retina. Exp Eye Res 89(4):549–558. doi:10.1016/j.exer.2009.05.006
Chow RL, Volgyi B, Szilard RK, Ng D, McKerlie C, Bloomfield SA, Birch DG, McInnes RR (2004) Control of late off-center cone bipolar cell differentiation and visual signaling by the homeobox gene Vsx1. Proc Natl Acad Sci U S A 101(6):1754–1759. doi:10.1073/pnas.0306520101
Cheng CW, Chow RL, Lebel M, Sakuma R, Cheung HO, Thanabalasingham V, Zhang X, Bruneau BG et al (2005) The Iroquois homeobox gene, Irx5, is required for retinal cone bipolar cell development. Dev Biol 287(1):48–60. doi:10.1016/j.ydbio.2005.08.029
Huang L, Hu F, Feng L, Luo XJ, Liang G, Zeng XY, Yi JL, Gan L (2014) Bhlhb5 is required for the subtype development of retinal amacrine and bipolar cells in mice. Dev Dyn 243(2):279–289. doi:10.1002/dvdy.24067
Jo HS, Kang KH, Joe CO, Kim JW (2012) Pten coordinates retinal neurogenesis by regulating notch signalling. EMBO J 31(4):817–828. doi:10.1038/emboj.2011.443
Rocha SF, Lopes SS, Gossler A, Henrique D (2009) Dll1 and Dll4 function sequentially in the retina and pV2 domain of the spinal cord to regulate neurogenesis and create cell diversity. Dev Biol 328(1):54–65. doi:10.1016/j.ydbio.2009.01.011
Jadhav AP, Cho SH, Cepko CL (2006) Notch activity permits retinal cells to progress through multiple progenitor states and acquire a stem cell property. Proc Natl Acad Sci U S A 103(50):18998–19003. doi:10.1073/pnas.0608155103
Yaron O, Farhy C, Marquardt T, Applebury M, Ashery-Padan R (2006) Notch1 functions to suppress cone-photoreceptor fate specification in the developing mouse retina. Development 133(7):1367–1378. doi:10.1242/dev.02311
Belecky-Adams T, Adler R (2001) Developmental expression patterns of bone morphogenetic proteins, receptors, and binding proteins in the chick retina. J Comp Neurol 430(4):562–572
Sakuta H, Takahashi H, Shintani T, Etani K, Aoshima A, Noda M (2006) Role of bone morphogenic protein 2 in retinal patterning and retinotectal projection. J Neurosci 26(42):10868–10878. doi:10.1523/JNEUROSCI.3027-06.2006
Liu J, Wilson S, Reh T (2003) BMP receptor 1b is required for axon guidance and cell survival in the developing retina. Dev Biol 256(1):34–48
Sakuta H, Suzuki R, Takahashi H, Kato A, Shintani T, Iemura S, Yamamoto TS, Ueno N et al (2001) Ventroptin: a BMP-4 antagonist expressed in a double-gradient pattern in the retina. Science 293(5527):111–115. doi:10.1126/science.1058379
Harpavat S, Cepko CL (2003) Thyroid hormone and retinal development: an emerging field. Thyroid 13(11):1013–1019. doi:10.1089/105072503770867183
Martinez-Morales JR, Del Bene F, Nica G, Hammerschmidt M, Bovolenta P, Wittbrodt J (2005) Differentiation of the vertebrate retina is coordinated by an FGF signaling center. Dev Cell 8(4):565–574. doi:10.1016/j.devcel.2005.01.022
Chen S, Li H, Gaudenz K, Paulson A, Guo F, Trimble R, Peak A, Seidel C et al (2013) Defective FGF signaling causes coloboma formation and disrupts retinal neurogenesis. Cell Res 23(2):254–273. doi:10.1038/cr.2012.150
Fang Y, Cho KS, Tchedre K, Lee SW, Guo C, Kinouchi H, Fried S, Sun X et al (2013) Ephrin-A3 suppresses Wnt signaling to control retinal stem cell potency. Stem Cells 31(2):349–359. doi:10.1002/stem.1283
Wilkinson DG (2014) Regulation of cell differentiation by Eph receptor and ephrin signaling. Cell Adhes Migr 8(4):339–348. doi:10.4161/19336918.2014.970007
Young RW (1985) Cell differentiation in the retina of the mouse. Anat Rec 212(2):199–205. doi:10.1002/ar.1092120215
Dyer MA, Cepko CL (2000) Control of Muller glial cell proliferation and activation following retinal injury. Nat Neurosci 3(9):873–880. doi:10.1038/78774
Fischer AJ, Reh TA (2001) Muller glia are a potential source of neural regeneration in the postnatal chicken retina. Nat Neurosci 4(3):247–252. doi:10.1038/85090
Ooto S, Akagi T, Kageyama R, Akita J, Mandai M, Honda Y, Takahashi M (2004) Potential for neural regeneration after neurotoxic injury in the adult mammalian retina. Proc Natl Acad Sci U S A 101(37):13654–13659. doi:10.1073/pnas.0402129101
Karl MO, Hayes S, Nelson BR, Tan K, Buckingham B, Reh TA (2008) Stimulation of neural regeneration in the mouse retina. Proc Natl Acad Sci U S A 105(49):19508–19513. doi:10.1073/pnas.0807453105
Chen M, Tian S, Glasgow NG, Gibson G, Yang X, Shiber CE, Funderburgh J, Watkins S et al (2015) Lgr5 amacrine cells possess regenerative potential in the retina of adult mice. Aging Cell. doi:10.1111/acel.12346
Reh TA, Levine EM (1998) Multipotential stem cells and progenitors in the vertebrate retina. J Neurobiol 36(2):206–220. doi:10.1002/(SICI)1097-4695(199808)36:2<206::AID-NEU8>3.0.CO;2-5
Perron M, Harris WA (2000) Retinal stem cells in vertebrates. Bioessays 22(8):685–688. doi:10.1002/1521-1878(200008)22:8<685::AID-BIES1>3.0.CO;2-C
Cicero SA, Johnson D, Reyntjens S, Frase S, Connell S, Chow LM, Baker SJ, Sorrentino BP et al (2009) Cells previously identified as retinal stem cells are pigmented ciliary epithelial cells. Proc Natl Acad Sci U S A 106(16):6685–6690. doi:10.1073/pnas.0901596106
Gualdoni S, Baron M, Lakowski J, Decembrini S, Smith AJ, Pearson RA, Ali RR, Sowden JC (2010) Adult ciliary epithelial cells, previously identified as retinal stem cells with potential for retinal repair, fail to differentiate into new rod photoreceptors. Stem Cells 28(6):1048–1059. doi:10.1002/stem.423
Krol J, Krol I, Alvarez CP, Fiscella M, Hierlemann A, Roska B, Filipowicz W (2015) A network comprising short and long noncoding RNAs and RNA helicase controls mouse retina architecture. Nat Commun 6:7305. doi:10.1038/ncomms8305
Karali M, Persico M, Mutarelli M, Carissimo A, Pizzo M, Singh Marwah V, Ambrosio C, Pinelli M et al (2016) High-resolution analysis of the human retina miRNome reveals isomiR variations and novel microRNAs. Nucleic Acids Res. doi:10.1093/nar/gkw039
Wride MA, Geatrell J, Guggenheim JA (2006) Proteases in eye development and disease. Birth Defects Res C Embryo Today 78(1):90–105. doi:10.1002/bdrc.20063
Gong L, Li DW (2010) SUMOylation in ocular development and pathology. Curr Mol Med 10(9):794–801
Iida A, Iwagawa T, Kuribayashi H, Satoh S, Mochizuki Y, Baba Y, Nakauchi H, Furukawa T et al (2014) Histone demethylase Jmjd3 is required for the development of subsets of retinal bipolar cells. Proc Natl Acad Sci U S A 111(10):3751–3756. doi:10.1073/pnas.1311480111
Iida A, Iwagawa T, Baba Y, Satoh S, Mochizuki Y, Nakauchi H, Furukawa T, Koseki H et al (2015) Roles of histone H3K27 trimethylase Ezh2 in retinal proliferation and differentiation. Dev Neurobiol 75(9):947–960. doi:10.1002/dneu.22261
Arya R, White K (2015) Cell death in development: signaling pathways and core mechanisms. Semin Cell Dev Biol 39:12–19. doi:10.1016/j.semcdb.2015.02.001
Fan Y, Bergmann A (2014) Multiple mechanisms modulate distinct cellular susceptibilities toward apoptosis in the developing Drosophila eye. Dev Cell 30(1):48–60. doi:10.1016/j.devcel.2014.05.007
Al-Shamekh S, Goldberg JL (2014) Retinal repair with induced pluripotent stem cells. Transl Res 163(4):377–386. doi:10.1016/j.trsl.2013.11.002
Gill KP, Hewitt AW, Davidson KC, Pebay A, Wong RC (2014) Methods of retinal ganglion cell differentiation from pluripotent stem cells. Translat Vis Sci Technol 3(4):7. doi:10.1167/tvst.3.3.7
Wu F, Kaczynski TJ, Sethuramanujam S, Li R, Jain V, Slaughter M, Mu X (2015) Two transcription factors, Pou4f2 and Isl1, are sufficient to specify the retinal ganglion cell fate. Proc Natl Acad Sci U S A 112(13):E1559–E1568. doi:10.1073/pnas.1421535112
Acknowledgments
The author thanks Drs. Courtni Newsome, Min Zou, Shengguo Li, and Rashade A. H. Haynes II for critical readings and helpful comments on the manuscript. The author apologizes that so many great works in the field are not cited due to space limit and accessibility of full articles. This work is partially supported by funding from the Zhongshan Ophthalmic Center.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares that he has no competing interests.
Additional information
An erratum to this article is available at http://dx.doi.org/10.1007/s12035-016-0037-6.
Rights and permissions
About this article
Cite this article
Jin, K. Transitional Progenitors during Vertebrate Retinogenesis. Mol Neurobiol 54, 3565–3576 (2017). https://doi.org/10.1007/s12035-016-9899-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-016-9899-x