Abstract
During the development of the central nervous system (CNS), neurons and glia are derived from multipotent neural stem cells (NSCs) undergoing self-renewal. NSC commitment and differentiation are tightly controlled by intrinsic and external regulatory mechanisms in space- and time-related fashions. SIRT1, a silent information regulator 2 (Sir2) ortholog, is expressed in several areas of the brain and has been reported to be involved in the self-renewal, multipotency, and fate determination of NSCs. Recent studies have highlighted the role of the deacetylase activity of SIRT1 in the determination of the final fate of NSCs. This review summarizes the roles of SIRT1 in the expansion and differentiation of NSCs, specification of neuronal subtypes and glial cells, and reprogramming of functional neurons from embryonic stem cells and fibroblasts. This review also discusses potential signaling pathways through which SIRT1 can exhibit versatile functions in NSCs to regulate the cell fate decisions of neurons and glia.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sirtuins (silent mating type information regulation 2 homolog, SIRTs) are members of the class-III histone/lysine deacetylase (HDAC) family and require β-nicotinamide adenine dinucleotide (NAD+) as an obligatory cofactor [1]. SIRTs have been highly conserved throughout evolution. They have been shown to be involved in calorie restriction and have been proposed to be involved in longevity [2, 3]. Seven SIRTs in mammals (SIRT1-SIRT7) are localized to different subcellular organelles and mediate different cellular functions depending on their typical substrate [4, 5]. SIRT1, SIRT6, and SIRT7 are nucleolar proteins that influence gene expression by deacetylating histones [6–8]. Although SIRT2 is generally considered a cytosolic protein, recent studies have found that it can be shuttled to the nucleus to modulate tumor progression and the cell cycle [9, 10]. Typical of sirtuins, SIRT3, SIRT4, and SIRT5 regulate oxidative stress by acting on metabolic enzymes in mitochondria [11, 12].
More recent studies have found that SIRTs have many important functions in the central nervous system (CNS) [13, 14]. SIRT6 is predominantly expressed in neuronal cells and exerts a protective effect on those cells by enhancing the release of high mobility group box-1 (HMGB1) from cell nuclei [15]. SIRT2 has been shown to be involved in the resistance to axonal degeneration in granule cells of slow Wallerian degeneration (WldS) mice through the acetylation of microtubules and to be protective in animal models of Parkinson’s and Huntington’s diseases [16, 17]. Mitochondrial SIRT3 in neurons promotes neuron survival under N-methyl-d-aspartate (NMDA)-induced excitotoxicity [18]. Noticeably, SIRT1 is widely expressed in the entire adult brain [19] and has been shown to trigger central metabolic actions in several areas of the brain to control feeding behavior, energy expenditure, glucose metabolism, and insulin sensitivity [20]. For example, enhanced SIRT1 activity in the dorsomedial and lateral hypothalamic nuclei (DMH and LH, respectively) delays aging through increased orexin type 2 receptor expression [21]. SIRT1 has emerged as a new therapeutic target for neurodegenerative diseases, such as Parkinson’s disease, Huntington’s disease, motor neuron diseases, and multiple sclerosis [1, 6, 17]. SIRT1 has drawn even more attention because it is considered to be one of the determining factors of the biology of stem cells, including neural stem cells (NSCs) and neural progenitor cells (NPCs) [14, 22]. Typically, SIRT1 has been observed in the developing mammalian brain, which indicates its influence on neural cell fate determination [19]. Thus, in this article, we summarize the emerging role of SIRT1 in the regulation of neural fate determination in embryonic and adult NSCs/NPCs, with a particular emphasis on NSCs/NPCs proliferation and differentiation.
Essential Characteristics of SIRT1
SIRT1 is most homologous to the founding member of the Sir2 family from yeast [23]. The human sirt1 gene is located on chromosome 10 and encodes a protein 747 amino acids in length. This protein comprises a conserved catalytic core domain that binds NAD+ between amino acids 254 and 489 and a NH2-terminal region that flanks the core domain. The core domain of SIRT1 possesses a low deacetylase activity, and the NH2 and COOH-terminal regions of SIRT1 intramolecularly potentiate the catalytic efficiency of the core domain and the deacetylase activity of SIRT1. The catalytic domain contains a large Rossmann-fold domain, which is characteristic of NAD+/NADH-binding proteins [4, 24–26]. Studies have shown that SIRT1 regulates a wide variety of biological processes and cellular functions through the deacetylation of several histone protein residues, including H3-K9, H4-K16, and H1-K26, and the deacetylation of a growing list of non-histone proteins, such as p53, FOXO, and PGC-1α [8, 27–29]. By deacetylating a variety of substrates, SIRT1 can mediate a broad range of vital signaling pathways, including pathways involved in the control of gene expression, DNA repair and apoptosis, neurogenesis, and aging; thus, SIRT1 promotes cellular longevity through a number of mechanisms [11, 23, 30]. Due to the important targets of pathways mediated by SIRT1, the activity and expression of SIRT1 seem to be regulated at many levels, including broader, more general regulatory mechanisms, such as substrate availability and tissue and subcellular localization, and gene-specific regulatory mechanisms, such as the modulation of transcription factors and the regulation of protein expression by microRNA (miRNA) [31–34]. In addition, SIRT1 activity can be controlled by posttranslational modifications including phosphorylation, methylation, nitrosylation, and SUMOylation [35–38]. More importantly, cross talk between the gene regulation pathways of multiple transcription factors and SIRT1 determines the expansion and differentiation of stem cells [39].
Because sirtuins have important effects on metabolism and physiological activities, they have attracted great attention as medicinal targets [40–42]. Several molecules have been found to be sirtuin-activating compounds (STACs), including plant-derived metabolites and synthetic chemicals. Resveratrol, a natural polyphenolic compound, was discovered to be a potent activator of SIRT1 [43]. There are many other synthetic pharmaceuticals, such as SRT1460, SRT1720, and SRT2183, that have been reported to activate SIRT1 with potencies of 1000-fold greater than resveratrol [44]. These STACs can activate sirtuins through two mechanisms: (i) direct allosteric activation of Sir2/SIRT1 by lowering the peptide substrate Km and (ii) coincidental, indirect activation through off-target effects [40]. Similarly, various types of natural SIRT1 inhibitors have also been successfully isolated from plants and sea organisms. Nicotinamide (NAM), commonly used as a sirtuin inhibitor, can physiologically inhibit the catalytic activity of sirtuins by binding to a conserved region of the sirtuin catalytic site [45]. Guttiferone G and amurensin G are potent SIRT1 inhibitors and are the most commonly used natural SIRT1 inhibitors [46, 47]. High-throughput screening and rational design studies have found a large number of potent sirtuin inhibitors. Tenovins, MC2141, and EX-527 are SIRT1-specific inhibitors with an IC50 in the submicromolar range [48–50]. In addition, the most advanced SIRT1 inhibitor compound is selisistat (also known as EX-527 or SEN196), and it is being studied in a phase II clinical trial in Huntington’s disease patients (see the Siena Biotech website) [50, 51]. The results of that study might reveal new methods of treating neurodegenerative disease.
SIRT1 Drives the Differentiation of NSCs from Pluripotent ESCs and iPSCs
Embryonic stem cells (ESCs) are derived from the inner cell mass of blastocysts and can self-renew indefinitely while maintaining pluripotency, the ability to differentiate into all cell types of the body [52]. SIRT1 is more highly expressed in mouse (m) and human (h) ESCs than in differentiated tissues, suggesting that SIRT1 is a key factor in maintaining the pluripotency of ESCs through the regulation of transcription factors, such as Nanog, OCT4, and Sox2, that maintain the self-renewal characteristic of stem cells [53–55]. Zhang and colleagues showed that the Oct4-mediated maintenance of hESC pluripotency is related to the inactivation of p53 through SIRT1-mediated deacetylation [54]. Similar to this study, SIRT1 has been found to inhibit the p53 signaling pathway during the differentiation of mESCs into mouse embryoid bodies (EB) [56]. Furthermore, SIRT1 has been found to block the nuclear translocation of p53 and repress the p53-mediated suppression of Nanog expression in cultured mESCs [53]. Sussman et al. showed that SIRT1 forms a functional complex with histone ubiquitin hydrolase ubiquitin-specific protease 22 (USP22), which is involved in the differentiation of mouse and human ESCs by facilitating the repression of Sox2 transcription [57]. Consistent with findings in ESCs, SIRT1-mediated deacetylation plays an important role in maintaining the self-renewal capability and multipotency of human bone marrow-derived mesenchymal stem cells by maintaining Sox2 protein expression in the nucleus [55].
In addition to the key role of SIRT1 in controlling the stemness of stem cells, profiling results have indicated its pivotal roles in fate decision during NSCs/NPCs differentiation of ESCs and induces pluripotent stem cells (iPSCs). ESCs are an excellent model for recapitulating early neuronal development in vitro [58]. Because SIRT1 can bind to and epigenetically repress a subset of developmental genes in pluripotent hESCs, it is reasonable to hypothesize that SIRT1 might be involved in determining specific differentiation programs during human and mouse ESC differentiation. The downregulation of SIRT1 mediated by coactivator-associated arginine methyltransferase 1 (CARM1)-dependent decrease in methyl-HuR/SIRT1 messenger RNA (mRNA) binding leads to the reactivation of key developmental genes, such as the neuroretinal morphogenesis effectors DLL4, TBX3, and PAX6, during hESC differentiation. Similarly, neuroectodermal markers have been shown to be overexpressed in Sirt1 KO EBs and to be downregulated in Super Sirt1 EBs (in which Sirt1 was increased) during mouse ESC differentiation [59]. This also strongly suggests that SIRT1 contributes to the establishment of specific developmental/differentiation programs that have particular relevance for neuroectodermal fates. It is well known that retinoic acid (RA) promotes the differentiation of ESCs to neuroectoderm and suppresses the differentiation of ESCs to mesoderm [60, 61]. One recent study found that SIRT1 contributes to homeostatic RA signaling and mouse ESC differentiation through the deacetylation of cellular retinoic acid-binding protein II (CRABPII) [62]. Furthermore, another study showed that RA-induced hESC-derived neuroectodermal cells retained the embryonic acetylation pattern on their chromatin, which is related to the inactivation of SIRT1 [63].
SIRT1 expression is closely correlated to the reprogramming of mouse embryonic fibroblasts (MEFs). SIRT1 activation promotes iPSC formation through deacetylation of p53, inhibition of p21, and enhancement of Nanog expression. On the other hand, SIRT1 has been shown to be downregulated during the differentiation process of mouse iPSCs into NSCs using serum-free medium combined with RA. Nicotinamide accelerated the formation of NSCs and mature nerve cells from iPSCs, suggesting that SIRT1 inhibition during the early stage of differentiation induction can facilitate the differentiation of iPSCs into NSCs. Furthermore, miR-34a, which targets SIRT1, has been shown to be required for the proper differentiation of mouse ESCs and NSCs. MiR-34a has been observed to inhibit iPSC formation from MEFs by suppressing SIRT1 expression. That study also demonstrated that miRNA-34a might be involved in the differentiation of iPSCs into NSCs through the repression of SIRT1 [64, 65]. The results of these studies indicate that the repression of SIRT1 at the early stage of differentiation induction may contribute to the initiation of NSCs/NPCs differentiation from ESCs and iPSCs. This may also explain developmental defects induced by SIRT1 deficiency observed in the mouse CNS.
SIRT1 Regulates the Expansion and Differentiation of NSCs
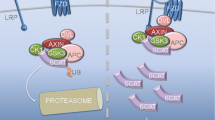
The features of NSCs are their ability to self-renew and generate different neural lineages, including neurons, astrocytes, and oligodendrocytes [66]. Neurogenesis is the basic process of neuron generation and includes the proliferation of NSCs/NPCs, the differentiation of neurons, and the integration of new neurons into the existing neural circuitry [67, 68]. An increasing number of studies have shown that SIRT1 is dynamically regulated in the embryonic brain. High levels of SIRT1 have been found in the brain, heart, spinal cord, and dorsal root ganglia of mouse embryos, with the highest SIRT1 mRNA levels detected at E4.5. A notable decrease in the expression of neural SIRT1 occurs between E13.5 and E14.5 and between E15.5 and E16.5, a timeframe that is coincident with the formation and maturation of neurons in the developing brain. A logarithmic decrease in SIRT1 expression in different areas of the brain during human embryonic development has also been confirmed by global transcriptome analysis [19]. Hisahara et al. have confirmed that SIRT1 is highly expressed in the cytoplasm of NPCs localized in the ventricular zone (VZ) and subventricular zone (SVZ) at E14.5, with a minor amount of nuclear staining [69]. Additionally, it has been demonstrated that the downregulation of SIRT1 expression in tumor neuroprogenitor cells (N2a cells) causes spontaneous neurite outgrowth coincident with a reduced growth rate, whereas SIRT1 overexpression inhibits induced neural differentiation of N2a cells and, subsequently, upregulates heat shock proteins (HSPs) through the deacetylation of heat shock factor 1 [70]. One recent study showed that human bone marrow-mesenchymal stem cells (hBM-MSCs) can be effectively differentiated into neurons by SIRT1 activator treatment combined with neuronal induction media [71]. It can be inferred from these reports that SIRT1 may be involved in embryonic neurogenesis. Emerging evidence has suggested that the Notch-Hes1 pathway, transcription factors, and miRNAs play important roles in the execution of SIRT1 functions in embryonic neurogenesis (Fig. 1).
Schematic representation of roles of SIRT1 in NSC fate determination. SIRT1 consumes NAD+ as a substrate and produces NAM and 2′-O-acetyl-ADP-ribose. NAM is recycled back to NAD+ through NAMPT, NMNAT, and NMN. This NAD recycling deacetylates a series of cell fate commitment factors of NSCs, such as TFs, TLX, Notch-Hes1, RA, and P53. These deacetylated proteins promote the self-renewing and neural commitment of NSCs. Otherwise, the NSCs may enter apoptosis. The deacetylase activity of SIRT1 is controlled by posttranslational modifications including phosphorylation, methylation, nitrosylation, and SUMOylation [35–38]. M methylation, N nitrosylation, NAMPT nicotinamide phosphorybosyltransferase, NMN nicotinamide mononucleotide, NMNAT nicotinamide mononucleotide adenylyltransferase, NSCs neural stem cells, P phosphorylation, RA retinoic acid, S SUMOylation, TFs transcription factors, TLX tailless
Notch-Hes1 Pathway
The Notch pathway is essential for maintaining NPCs in the developing brain through the activation of its downstream genes. Upon activation, the Notch intracellular domain (Notch-ICD) is released and translocates into the nucleus; there, it cooperates with the CBF-1–SuH–LAG-1(CSL) DNA-binding protein and its coactivator, mastermind-like (Maml), to induce the expression of downstream genes, such as Hes1 and Hes5 [72, 73]. Hes1 and Hes5 function as classical DNA-binding repressors that antagonize the expression of proneural genes, such as Mash, Math, and Neurogenin, to maintain NPCs in an undifferentiated state [69, 74, 75]. Studies have indicated that extensive chromatin remodeling at locations of genes in the Notch-dependent transcriptional program is involved in irreversible changes in gene expression during neurogenesis. It has been reported that mice lacking Hes1 exhibit exencephaly and retinal defects with rosette structures [76, 77]. The similarities between these phenotypes and those observed in sirt1 mutant mice suggest that SIRT1 interacts with the transcriptional repressor Hes1 [78]. Through its interaction with Hes1, SIRT1 may act as an attenuating influence on Notch effects, and therefore, SIRT1 could influence the role of the Notch pathway in asymmetric stem cell division and self-renewal division.
Hisahara et al. found that cytoplasmic SIRT1 in cultured embryonic NPCs was transiently translocated into the nucleus under differentiation conditions; this resulted in the suppression of Hes1 expression and an increased number of intermediate neural progenitors and Tuj1-positive neurons. Furthermore, SIRT1 has been shown to repress the expression of the Hes1 promoter by directly repressing RBP-J activity promoted by Notch1-ICD through its interaction with the nuclear receptor corepressor (NCoR). Both SIRT1 and NCoR can bind to the Hes1 promoter region in differentiating NPCs, and Hes1 transactivation by Notch1 was inhibited by SIRT1 and/or N-CoR [69]. In addition to the direct action of SIRT1 on the Notch1-Hes1 signaling pathway during NPCs differentiation, an indirect interaction of SIRT1 with the Notch1-Hes1 signaling pathway that is involved in embryonic neurogenesis has been confirmed. Acetylated Pax3, which directly regulates TGFβ2 transcription by binding to cis-regulatory elements within its promoter, is associated with SIRT1 during early embryonic development. Pax3 is involved in mouse embryonic caudal neural tube development at E9.5 by interacting with Hes1 and Neurog2 promoters. In the presence of SIRT1 at E9.5, deacetylated Pax3 causes the upregulation of Hes1 and promotes stem cell proliferation and maintenance. At E12.5, acetylated Pax3 downregulates Hes1 and upregulates Neurog2 promoter activities. These studies suggest that Pax3 acetylation results in decreased Hes1 and increased Neurog2 activity and thereby promotes neuronal differentiation [79]. BCL6 is required for proper neurogenesis of the mouse cerebral cortex. BCL6 acts by modifying the composition of transcriptional complexes and the chromatin structure at the promoter of the Notch target gene Hes5. By selective binding to the Hes5 promoter, BCL6 excludes Maml1 from the Notch transcriptional complex, preventing transcriptional activation, and recruits the NAD+-dependent deacetylase SIRT1, resulting in histone deacetylation and consequent epigenetic silencing of Hes5. Thus, BCL6 acts as an essential trigger for the neurogenic conversion of NPCs into neurons [80]. BCL6 has also been shown to induce neuronal differentiation in the granule neuron precursors of the cerebellum through the recruitment of BCOR corepressor and SIRT1 histone deacetylase, leading to epigenetic repression of Sonic Hedgehog signaling effectors Gli1 and Gli2 [81].
On the other hand, a study by Prozorovski et al. demonstrated that SIRT1 repressed the proliferation of NPCs and directed their differentiation toward the astroglial lineage at the expense of the neuronal lineage under oxidative stress conditions in vitro and in vivo. They found that oxidation results in a stronger association of the SIRT1-Hes1 complex to the Mash1 promoter, leading to a targeted and localized deacetylation of H3K9 and, subsequently, to Mash1 inhibition. However, in a reducing environment, Hes1 recruits transcription activators, such as CREB-binding protein (CBP), to the Mash1 promoter, and this drives NPCs toward a neuronal fate. The influence of the redox state on NPC fate decisions was eliminated by the blockage of SIRT1 activity either by RNAi or SIRT1 inhibitors [22]. The mitochondrial DNA (mtDNA) integrity of NSCs is vulnerable to oxidative damage and is related to NSC differentiation [82]. mtDNA damage induced by redox insult drives NSCs to astrogenesis at the expense of neurogenesis, and this is accompanied by an increase in SIRT1 [83]. Similarly, one study showed that antioxidants halted the shift in NSC differentiation potential from the neurogenic to gliogenic lineage while strongly reducing reactive oxygen species generation and the nuclear translocation of Nrf2 and SIRT1 in NSCs exposed to redox insult [84]. The opposing effects of SIRT1 on NSC lineage choice demonstrated by these independent studies raise the possibility that SIRT1 function in NSC differentiation depends heavily on the redox state of the cells, underscoring the significance of oxidative state on NSC fate.
Transcription Factors
Mounting evidence has shown that histone deacetylases interact with transcription factors and can be targeted to specific loci. The transcription factor tailless (TLX) has been suggested to be essential for NSC self-renewal and adult neurogenesis. It represses the transcriptional repression of cell cycle inhibitor p21Cip1/WAF1 and the Pten tumor suppressor in NSCs by interacting with HDAC3 and HDAC5. Furthermore, in NPCs, TLX has been shown to colocalize with SIRT1 and to increase SIRT1 expression by binding to the newly identified TLX-activating element in the SIRT1 gene promoter [85, 86]. Thus, TLX is an inducer of SIRT1 and may contribute to neurogenesis both as a transactivator and as a repressor. In addition, downstream of TLX, SIRT1 seems to have a role in retinal neurogenesis because eye phenotypes in TLX−/− mice are similar to that in SIRT1−/− mice [78, 87].
The retinoic acid receptor (RAR) contributes to RA-induced neuron differentiation [88]. Ski-interacting protein (SKIP) is a transcriptional coactivator of several nuclear receptors and other transcription factors [89]. SIRT1 competes with SKIP for RARα binding at the protein level and at the chromatin-associated RARβ2 promoter. Resveratrol treatment has been shown to inhibit RA-induced neuronal differentiation of ES-like P19 cells and to reduce the neuronal marker nestin and a RAR target gene, RARβ2. This inhibition was relieved by either knockdown of sirt1 or overexpression of SKIP. These results suggest that SIRT1 and SKIP play reciprocal roles in the regulation of RA-induced neuronal differentiation of P19 cells through RAR activity [90]. Furthermore, Yu et al. found that RA-induced neuronal differentiation of P19 stem cells is mediated by the CRABP-II/RAR pathway in the early stages and through the PPARβ/δ/FABP5 pathway in the late stages of the process. This switch in RA signaling is accomplished by a transient upregulation of RARβ with a concurrent inhibition of the FABP5/PPARβ/δ pathway by the RAR repressor SIRT1 in the early stages of differentiation [91].
miRNAs
miRNAs are short RNAs with an average length of 22 nucleotides. They cause gene silencing by binding to complementary sequences on target mRNAs, resulting in the degradation of the target mRNAs [92, 93]. To date, several miRNAs, such as miR-181a and b, miR-9, miR-204, miR-199b, and miR-135a, have been shown to reduce SIRT1 expression at the posttranscriptional level [34]. Aranha et al. have found that miR-34a negatively modulates SIRT1, contributing to neurite elongation of postmitotic neurons derived from mouse NSCs. In addition, acetylation of p53 (Lys 379) and p53-DNA binding activity was increased due to the repression of SIRT1 by miR-34a during neural differentiation [94]. Emerging evidence has indicated that p53 is involved in the regulation of the cell cycle and the transcription of neuronal genes. It is probable that the upregulation of p53 acetylation induced by miR-34a-mediated silencing of sirt1 contributes to the correct establishment of specific differentiation programs during NSC differentiation. It has also been confirmed that miR-9 acts early in the differentiation of mESCs to mEBs and during the directed differentiation of mESCs to neurons by downregulating SIRT1 expression. One study demonstrated that miR-9 has a key role in the coordination of the proliferation and migration of hNPCs. That study also found that miR-9 limits the migration of hESC-derived hNPCs at the early neurosphere stage, and this is accompanied by the downregulation of STMN1 (stathmin), CHMP2B, and SIRT1 [95]. This also suggests that SIRT1 is negatively correlated with miR-9 expression at the early neurosphere stage, although the true mechanism of the involvement of miR-9 in stem cell differentiation and migration requires further investigation.
Apoptosis Regulators
Surprisingly, recent evidence has suggested apoptosis-associated factors, such as p53 and caspases, participate in the differentiation process of mNSCs [34, 96]. Increased neurosphere-forming potential of precursor cells due to a lack of p53 is related to a reduction in apoptotic rates [97]. However, p53KO NSCs from embryonic olfactory bulbs (OBs) exhibit a differentiation bias toward the generation of neurons with a reduction in glial cells, particularly astrocytes, without an alteration in the rate of apoptosis [98]. This also suggests that p53 is directly involved in the differentiation of NSCs from embryonic OBs. SIRT1 has been shown to be critical for maintaining the stemness of stem cells, such as cancerous neural stem cells (F3.Ras.CNSCs) and glioma stem cells, through its suppression of p53-dependent tumor surveillance [99]. This suggests that SIRT1 deacetylation of p53 might be involved in proliferation and differentiation via an apoptotic or non-apoptotic pathway. Caspase-2, caspase-3, and caspase-9 have been shown to be selectively and transiently activated in an apoptosis-independent manner during the RA-induced neuronal differentiation of the human teratocarcinoma cell line Ntera2/cl.D1 (NT2 cells). Caspase-9 activation greatly reduces the cleavage of SIRT1 and facilitates the differentiation process due to a transiently increased expression of the proneural transcription factor MASH1 at early times of NT2-cell differentiation. In contrast, silencing of caspase-2 activation appears to hinder the differentiation process as shown by earlier and increased expression of MASH1, although SIRT1 cleavage is not significantly affected [100]. Thus, non-apoptotic functions of caspases are involved in the regulation of terminal neuronal differentiation.
Cell Fate Determination of Neuronal Subtypes by SIRT1
Directing ESCs to differentiate into large numbers of motoneurons is a key step for motor neuron replacement therapy [101, 102]. Nicotinamide treatment has been shown to produce a large amount of motoneurons from a cultured human ESC line (PKU1.1). As a specific inhibitor of SIRT1, nicotinamide activates Mash1 and Ngn2 by repressing the binding of SIRT1 to the Mash1 promoter and increases the expression of HB9 to drive motoneuron formation [103]. Additionally, Nurr1 has been shown to be essential for meso-diencephalic dopamine (mdDA) neuron development through its regulation of the transcription of the tyrosine hydroxylase (TH) gene, which encodes the rate-limiting enzyme in dopamine synthesis [104]. In the human neural stem cell (hNSC) line HB1.F3, Nurr1 strongly represses the transcription of hTH by increasing the SIRT1 occupancy of the NBRE-A hTH promoter region [105]. This effect can be reversed by mutating the NBRE-A element of the hTH promoter region and by nicotinamide treatment. However, Min-Ju Kim found that the deacetylation of FOXO3a by SIRT1 is involved in the all-trans retinoic acid (ATRA)-induced upregulation of TH and differentiation of neuroblastoma cells and is inhibited by SIRT1 and FOXO3a siRNA [106]. The different mechanisms induced by SIRT1 are still not clear, and there is still controversy as to whether TH is a transcriptional target of FOXO3a.
The orexin system plays a central role in the integration of sleep/wake and feeding behaviors in neural-metabolic physiology [107]. ManNAc has been shown to successfully generate functional orexin neurons from mouse ESCs, and the key step in the differentiation pathway is an epigenetic switch on histones from a hypoacetylated state with unidentified O-GlcNAcylated nuclear proteins to a hyperacetylated state at T-DMRs. ManNAc causes the dislocation of Ogt/Sirt1/Sin3A/Ezh2 from the T-DMRs of the Hcrt gene locus and the recruitment of p300, CBP, and Mgea5 [108].
Another study confirmed that SIRT1 overexpression in ND7 cells (fused neonatal rat dorsal root ganglion neurons with mouse neuroblastoma cells) increased Hes1 expression and decreased Neurog2 and Brn3a expression, thereby promoting sensory neuron differentiation [79]. Furthermore, SIRT1 seems to have important functions in eye morphogenesis and retinal development because abnormal retinal histology with rosette formation was observed in SIRT1-deficient embryos as early as E12.5. It has also confirmed that SIRT1 is ubiquitously expressed in human retinal progenitor cells (RPCs) and shows some nuclear localization and is involved in the genesis of specific retinal cell subtypes. In particular, the phenotype of abnormal eye morphogenesis in SIRT1 deficient mice is similar to that in mice with targeted deletion of the homeobox gene Vax2 [78, 109].
SIRT1 Regulates Survival of Differentiated Neurons
The survival of differentiated neurons has to be against various intrinsic and extrinsic stresses for the entire lifetime of the organism. Increasing evidence suggests that SIRT1 plays a prominent role in the survival of differentiated neurons under various cellular stresses. The acetylation status of p53 seemed to be crucial for neuron survival. This has been further supported by the finding that necdin, a p53-interacting protein expressed mainly in postmitotic neurons, downregulates the acetylation status of p53 via SIRT1 to protect neurons from DNA damage-induced apoptosis [110]. In addition, resveratrol activation of SIRT1 has been indicated to be related to the promotion of autophagic functions, thereby leading to prolong cell survival in differentiated PC12 cell [111]. It is widely accepted that CR can increase the expression of SIRT1 and prolong longevity. Similarly, Kotaro Fujino found that when glucose availability increased, total expression levels of SIRT1 in PC12 cells were reduced while increased by glucose deprivation. Therefore, glucose deprivation-induced SIRT1 upregulation may potentially play a pivotal role in PC12 cell survival [112]. Similarly, NeuroD6 has been demonstrated to initiate neuronal differentiation while promoting neuronal survival in cultured PC12-ND6 cell line (overexpresses NeuroD6) via the NeuroD6–PGC-1α–SIRT1 neuroprotective axis under oxidative stress [113].
Different from these reports, one study conducted by Sansone et al. suggested that SIRT1 silencing induced expression levels of insulin-like growth factor-1 (IGF-1) and IGF-1R protein levels in vitro differentiated NG108-15 cells which, in turn, prolonged neuron survival in the presence of an apoptotic insult [114]. All these reports regarding the regulation of SIRT1 in differentiated neurons reveal that SIRT1 can be neuroprotective or neurotoxic depending on conditions, cellular stress, and cellular type.
SIRT1 and Adult Stem Cells
NSCs/NPCs continue to produce neurons in two regions of the adult brain, the SVZ of the lateral ventricles, and the subgranular zone of the hippocampal dentate gyrus (DG) [115]. Histone acetylation is mainly associated with active gene transcription. SIRT1 is strongly expressed in the SVZ and DG of adult mice in Sox2+ stem cells [20, 116]. An in vivo study conducted by Sumiti Saharan indicated that SIRT1 signaling is a negative regulator of the neuronal differentiation of adult NPCs in the SVZ and hippocampal DG, although it does not affect the proliferation of precursor cells. They further demonstrated that cultured NPCs derived from the SVZ and DG of genetically ablated SIRT1 knockout mice exhibited intrinsically enhanced neurogenic potential without altered proliferation [14]. However, one recent study showed that the proliferation and self-renewal rates of adult NSCs and NPCs were elevated by the loss of SIRT1, and this may be mediated by the inhibition of Notch signaling [117]. These two findings suggest that SIRT1 probably has dual roles in the modulation of neurogenesis. In addition, constitutive activation of FOXO3, a target of SIRT1 in CaMKIIα-positive NPCs in adult mice, results in dysregulation of adult neurogenesis, which suggests that FOXO3 may be involved in the modulation of adult NPCs by SIRT1 [118].
Loss of SIRT1 deacetylase activity by conditional ablation of SIRT1 in nestin+ stem cells results in an increased number of oligodendrocyte transcription factor 2 (Olig2)+ OPCs being generated in the SVZ and migrating toward the septum and the striatum. These newly produced OPCs following SIRT1 inactivation differentiate into myelinating oligodendrocytes. Furthermore, the loss of SIRT1 promotes platelet-derived growth factor receptor alpha (PDGFRα) transcription through the upregulation of the acetylation of its promoter at histone 3 lysine 9 (H3K9) [119]. However, another study found that overexpression of SIRT1 or treatment with resveratrol and SRT1720 increased oligodendrocyte differentiation by upregulating PGC1α expression. It is likely that these conflicting results with regard to the effect of SIRT1 on oligodendrocyte differentiation are due to the modulation of downstream genes [120]. Similarly, resveratrol treatment enhanced myelination in cultured rat Schwann cells, with a slight reduction in the number of Schwann cells, suggesting that SIRT1 activation induced cell differentiation [121]. One recent study found that nicotinamide phosphoribosyl transferase (Nampt), the rate-limiting enzyme in mammalian NAD(+) biosynthesis, is the main source of NAD(+) in NSCs/NPCs and is critical in oligodendrocytic lineage fate decisions through a mechanism mediated redundantly by SIRT1 and SIRT2. Ablation of Nampt in adult NSCs/NPCs reduced oligodendrogenesis upon insult [122]. These results suggest that SIRT1 mediation of oligodendrocyte regeneration in adults may attenuate the development of demyelinating injuries and diseases, such as encephalomyelitis and multiple sclerosis.
Prospects and Challenges
Neural fate determination is a complex process that occurs in embryonic and adult brains and is tightly regulated at multiple levels in a time- and stage-dependent manner. Epigenetic mechanisms are involved in neural cell fate determination through the modulation of the expression and activity of a panel of transcription factors in a neurodevelopment-dependent fashion. SIRT1, a NAD+-dependent class III histone deacetylase, plays important roles in NSC derivation, proliferation, and differentiation. Notably, SIRT1 is a sensor of the redox/oxidative state of NSCs/NPCs that is involved in neurogenesis under oxidative insult; thus, it may be involved in ageing and neurodegeneration. Furthermore, SIRT1 not only determines the cell fate of NSCs undergoing differentiation but is also altered in stem cell therapy to generate adequate neurogenic potential. For example, in vitro neuroectoderm-derived Nurr1-positive hESC-I hNuPs in high purity with inactive SIRT1 yielded neurons efficiently and exclusively and retained an embryonic acetylated globally active chromatin state [123]. It is promising to determine hESC pluripotent fate through embedding lineage-specific genetic and epigenetic programs into the open epigenomic landscape of pluripotent hESCs [124]. Thereby, it is worth investigating the mechanism of SIRT1 involvement in the control of the acetylated globally active chromatin state in those cells.
Additionally, paradoxical effects of multiple transcription factors and SIRT1 exist to determine neural cell fate. Determining the interactions of these transcription factors and SIRT1 will help clarify the multifaceted mechanism of SIRT1 under different conditions. Lastly, given that SIRT1 can bind and regulate many regions in the genome of ESCs, unbiased identification of all SIRT1 genomic binding sites in NSCs would help reveal a network of genes by which SIRT1 exerts its effects on NSC fate choice.
References
Donmez G (2012) The neurobiology of sirtuins and their role in neurodegeneration. Trends Pharmacol Sci 33(9):494–501
Dali-Youcef N, Lagouge M, Froelich S, Koehl C, Schoonjans K, Auwerx J (2007) Sirtuins: the ‘magnificent seven’, function, metabolism and longevity. Ann Med 39(5):335–345
Morselli E, Maiuri MC, Markaki M, Megalou E, Pasparaki A, Palikaras K, Criollo A, Galluzzi L et al (2010) Caloric restriction and resveratrol promote longevity through the Sirtuin-1-dependent induction of autophagy. Cell Death Dis 1:e10
Frye RA (2000) Phylogenetic classification of prokaryotic and eukaryotic Sir2-like proteins. Biochem Biophys Res Commun 273(2):793–798
Blander G, Guarente L (2004) The Sir2 family of protein deacetylases. Annu Rev Biochem 73:417–435
Paraíso AF, Mendes KL, Santos SH (2013) Brain activation of SIRT1: role in neuropathology. Mol Neurobiol 48(3):681–689
Barber MF, Michishita-Kioi E, Xi Y, Tasselli L, Kioi M, Moqtaderi Z, Tennen RI, Paredes S et al (2012) SIRT7 links H3K18 deacetylation to maintenance of oncogenic transformation. Nature 487(7405):114–118
Vaquero A, Scher M, Lee D, Erdjument-Bromage H, Tempst P, Reinberg D (2004) Human SirT1 Interacts with Histone H1 and Promotes Formation of Facultative Heterochromatin. Mol Cell 16(1):93–105
Kwon HS, Ott M (2008) The ups and downs of SIRT1. Trends Biochem Sci 33(11):517–525
Harting K, Knöll B (2010) SIRT2-mediated protein deacetylation: an emerging key regulator in brain physiology and pathology. Eur J Cell Biol 89(2–3):262–269
Michan S, Sinclair D (2007) Sirtuins in mammals: insights into their biological function. Biochem J 404(1):1–13
Haigis MC, Sinclair DA (2010) Mammalian sirtuins: biological insights and disease relevance. Annu Rev Pathol 5:253–295
Guo W, Qian L, Zhang J, Zhang W, Morrison A, Hayes P, Wilson S, Chen T et al (2011) Sirt1 overexpression in neurons promotes neurite outgrowth and cell survival through inhibition of the mTOR signaling. J Neurosci Res 89(11):1723–1736
Saharan S, Jhaveri DJ, Bartlett PF (2013) SIRT1 regulates the neurogenic potential of neural precursors in the adult subventricular zone and hippocampus. J Neurosci Res 91(5):642–659
Lee OH, Kim J, Kim JM, Lee H, Kim EH, Bae SK, Choi Y, Nam HS et al (2013) Decreased expression of sirtuin 6 is associated with release of high mobility group box-1 after cerebral ischemia. Biochem Biophys Res Commun 438(2):388–394
Suzuki K, Koike T (2007) Resveratrol abolishes resistance to axonal degeneration in slow Wallerian degeneration (WldS) mice: activation of SIRT2, an NAD-dependent tubulin deacetylase. Biochem Biophys Res Commun 359(3):665–671
Donmez G, Outeiro TF (2013) SIRT1 and SIRT2: emerging targets in neurodegeneration. EMBO Mol Med 5(3):344–352
Kim SH, Lu HF, Alano CC (2011) Neuronal Sirt3 Protects against Excitotoxic Injury in Mouse Cortical Neuron Culture. PLoS One 6(3):e14731
Sakamoto J, Miura T, Shimamoto K, Horio Y (2004) Predominant expression of Sir2α, an NAD-dependent histone deacetylase, in the embryonic mouse heart and brain. FEBS Lett 556(1–3):281–286
Ramadori G, Lee CE, Bookout AL, Lee S, Williams KW, Anderson J, Elmquist JK, Coppari R (2008) Brain SIRT1: anatomical distribution and regulation by energy availability. J Neurosci 28(40):9989–9996
Satoh A, Brace CS, Ben-Josef G, West T, Wozniak DF, Holtzman DM, Herzog ED, Imai S (2010) SIRT1 promotes the central adaptive response to diet restriction through activation of the dorsomedial and lateralnuclei of the hypothalamus. J Neurosci 30(30):10220–10232
Prozorovski T, Schulze-Topphoff U, Glumm R, Baumgart J, Schröter F, Ninnemann O, Siegert E, Bendix I et al (2008) Sirt1 contributes critically to the redox-dependent fate of neural progenitors. Nat Cell Biol 10(4):385–394
Revollo JR, Li X (2013) The ways and means that fine tune Sirt1 activity. Trends Biochem Sci 38(3):160–167
Luo XY, Qu SL, Tang ZH, Zhang Y, Liu MH, Peng J, Tang H, Yu KL et al (2014) SIRT1 in cardiovascular aging. Clin Chim Acta 437:106–114
Wang Y, Xu C, Liang Y, Vanhoutte PM (2012) SIRT1 in metabolic syndrome: where to target matters. Pharmacol Ther 136(3):305–318
Peled T, Shoham H, Aschengrau D, Yackoubov D, Frei G, Rosenheimer GN, Lerrer B, Cohen HY et al (2012) Nicotinamide, a SIRT1 inhibitor, inhibits differentiation and facilitates expansion of hematopoietic progenitor cells with enhanced bone marrow homing and engraftment. Exp Hematol 40(4):342–355
Brunet A, Sweeney LB, Sturgill JF, Chua KF, Greer PL, Lin Y, Tran H, Ross SE et al (2004) Stress-dependent regulation of FOXO transcription factors by the SIRT1 deacetylase. Science 303(5666):2011–2015
Kim EJ, Kho JH, Kang MR, Um SJ (2007) Active regulator of SIRT1 cooperates with SIRT1 and facilitates suppression of p53 activity. Mol Cell 28(2):277–290
Yeung F, Hoberg JE, Ramsey CS, Keller MD, Jones DR, Frye RA, Mayo MW (2004) Modulation of NF-kappaB-dependent transcription and cell survival by the SIRT1 deacetylase. EMBO J 23(12):2369–2380
Seo JS, Moon MH, Jeong JK, Seol JW, Lee YJ, Park BH, Park SY (2012) SIRT1, a histone deacetylase, regulates prion protein-induced neuronal cell death. Neurobiol Aging 33(6):1110–1120
Motta MC, Divecha N, Lemieux M, Kamel C, Chen D, Gu W, Bultsma Y, McBurney M et al (2004) Mammalian SIRT1 represses forkhead transcription factors. Cell 116(4):551–563
Fusco S, Maulucci G, Pani G (2012) Sirt1: def-eating senescence? Cell Cycle 11(22):4135–4146
Yamakuchi M (2012) MicroRNA Regulation of SIRT1. Front Physiol 3:68
Saunders LR, Sharma AD, Tawney J, Nakagawa M, Okita K, Yamanaka S, Willenbring H, Verdin E (2010) miRNAs regulate SIRT1 expression during mouse embryonic stem cell differentiation and in adult mouse tissues. Aging (Albany NY) 2(7):415–431
Guo X, Williams JG, Schug TT, Li X (2010) DYRK1A and DYRK3 promote cell survival through phosphorylation and activation of SIRT1. J Biol Chem 285(17):13223–13232
Liu X, Wang D, Zhao Y, Tu B, Zheng Z, Wang L, Wang H, Gu W et al (2011) Methyltransferase Set7/9 regulates p53 activity by interacting with Sirtuin1 (SIRT1). Proc Natl Acad Sci U S A 108(5):1925–1930
Yang Y, Fu W, Chen J, Olashaw N, Zhang X, Nicosia SV, Bhalla K, Bai W (2007) SIRT1 sumoylation regulates its deacetylase activity and cellular response to genotoxic stress. Nat Cell Biol 9(11):1253–1262
Kornberg MD, Sen N, Hara MR, Juluri KR, Nguyen JV, Snowman AM, Law L, Hester LD et al (2010) GAPDH mediates nitrosylation of nuclear proteins. Nat Cell Biol 12(11):1094–1100
Lin Z, Yang H, Kong Q, Li J, Lee SM, Gao B, Dong H, Wei J et al (2012) USP22 antagonizes p53 transcriptional activation by deubiquitinating Sirt1 to suppress cell apoptosis and is required for mouse embryonic development. Mol Cell 46(4):484–494
Hubbard BP, Sinclair DA (2014) Small molecule SIRT1 activators for the treatment of aging and age-related diseases. Trends Pharmacol Sci 35(3):146–154
Jiang M, Wang J, Fu J, Du L, Jeong H, West T, Xiang L, Peng Q et al (2011) Neuroprotective role of Sirt1 in mammalian models of Huntington’s disease through activation of multiple Sirt1 targets. Nat Med 18(1):153–158
Milne JC, Denu JM (2008) The Sirtuin family: therapeutic targets to treat diseases of aging. Curr Opin Chem Biol 12(1):11–17
Borra MT, Smith BC, Denu JM (2005) Mechanism of human SIRT1 activation by resveratrol. J Biol Chem 280(17):17187–17195
Milne JC, Lambert PD, Schenk S, Carney DP, Smith JJ, Gagne DJ, Jin L, Boss O et al (2007) Small molecule activators of SIRT1 as therapeutics for the treatment of type 2 diabetes. Nature 450(7170):712–716
Avalos JL, Bever KM, Wolberger C (2005) Mechanism of Sirtuin Inhibition by Nicotinamide:Altering the NAD+Cosubstrate Specificity of a Sir2 Enzyme. Mol Cell 17(6):855–868
Gey C, Kyrylenko S, Hennig L, Nguyen LH, Buttner A, Pham HD, Giannis A (2007) Phloroglucinol derivatives guttiferone G, aristoforin, and hyperforin: inhibitors of human sirtuins SIRT1 and SIRT2. Angew Chem Int Ed Engl 46(27):5219–5222
Oh WK, Cho KB, Hien TT, Kim TH, Kim HS, Dao TT, Han HK, Kwon SM et al (2010) Amurensin G, a potent natural SIRT1 inhibitor, rescues doxorubicin responsiveness via down-regulation of multidrug resistance 1. Mol Pharmacol 78(5):855–864
Medda F, Russell RJ, Higgins M, McCarthy AR, Campbell J, Slawin AM, Lane DP, Lain S et al (2009) Novel cambinol analogs as sirtuin inhibitors: synthesis, biological evaluation, and rationalization of activity. J Med Chem 52(9):2673–2682
Rotili D, Tarantino D, Carafa V, Lara E, Meade S, Botta G, Nebbioso A, Schemies J et al (2010) Identification of Tri- and Tetracyclic Pyrimidinediones as Sirtuin Inhibitors. ChemMedChem 5(5):674–677
Solomon JM, Pasupuleti R, Xu L, McDonagh T, Curtis R, DiStefano PS, Huber LJ (2006) Inhibition of SIRT1 Catalytic Activity Increases p53 Acetylation but Does Not Alter Cell Survival following DNA Damage. Mol Cell Biol 26(1):28–38
Arrowsmith CH, Bountra C, Fish PV, Lee K, Schapira M (2012) Epigenetic protein families: a new frontier for drug discovery. Nat Rev Drug Discov 11(5):384–400
Bishop AE, Buttery LD, Polak JM (2002) Embryonic stem cells. J Pathol 197(4):424–429
Han MK, Song EK, Guo Y, Ou X, Mantel C, Broxmeyer HE (2008) SIRT1 regulates apoptosis and Nanog expression in mouse embryonic stem cells by controlling p53 subcellular localization. Cell Stem Cell 2(3):241–251
Zhang ZN, Chung SK, Xu Z, Xu Y (2014) Oct4 maintains the pluripotency of human embryonic stem cells by inactivating p53 through Sirt1-mediated deacetylation. Stem Cells 32(1):157–165
Yoon DS, Choi Y, Jang Y, Lee M, Choi WJ, Kim SH, Lee JW (2014) SIRT1 Directly Regulates SOX2 to Maintain Self-Renewal and Multipotency in Bone Marrow-Derived Mesenchymal Stem Cells. Stem Cells 32(12):3219–3231
Lhee SJ, Song EK, Kim YR, Han MK (2012) SIRT1 Inhibits p53 but not NF-κB Transcriptional Activity during Differentiation of Mouse Embryonic Stem Cells into Embryoid Bodies. Int J Stem Cells 5(2):125–129
Sussman RT, Stanek TJ, Esteso P, Gearhart JD, Knudsen KE, McMahon SB (2013) The epigenetic modifier ubiquitin-specific protease 22 (USP22) regulates embryonic stem cell differentiation via transcriptional repression of sex-determining region Y-box 2 (SOX2). J Biol Chem 288(33):24234–24246
Dvash T, Benvenisty N (2004) Human embryonic stem cells as a model for early human development. Best Pract Res Clin Obstet Gynaecol 18(6):929–940
Calvanese V, Lara E, Suárez-Alvarez B, Abu Dawud R, Vázquez-Chantada M, Martinez-Chantar ML, Embade N, López-Nieva P et al (2010) Sirtuin 1 regulation of developmental genes during differentiation of stem cells. Proc Natl Acad Sci U S A 107(31):13736–13741
Bain G, Kitchens D, Yao M, Huettner JE, Gottlieb DI (1995) Embryonic stem cells express neuronal properties in vitro. Dev Biol 168(2):342–357
Bain G, Ray WJ, Yao M, Gottlieb DI (1996) Retinoic Acid Promotes Neural and Represses Mesodermal Gene Expression in Mouse Embryonic Stem Cells in Culture. Biochem Biophys Res Commun 223(3):691–694
Tang S, Huang G, Fan W, Chen Y, Ward JM, Xu X, Xu Q, Kang A et al (2014) SIRT1-Mediated Deacetylation of CRABPII Regulates Cellular Retinoic Acid Signaling and Modulates Embryonic Stem Cell Differentiation. Mol Cell 55(6):843–855
Parsons XH (2012) MicroRNA Profiling Reveals Distinct Mechanisms Governing Cardiac and Neural Lineage-Specification of Pluripotent Human Embryonic Stem Cells. J Stem Cell Res Ther 2(3):124
Lee YL, Peng Q, Fong SW, Chen AC, Lee KF, Ng EH, Nagy A, Yeung WS (2012) Sirtuin 1 facilitates generation of induced pluripotent stem cells from mouse embryonic fibroblasts through the miR-34a and p53 pathways. PLoS One 7(9):e45633
Hu B, Guo Y, Chen C, Li Q, Niu X, Guo S, Zhang A, Wang Y et al (2014) Repression of SIRT1 promotes the differentiation of mouse induced pluripotent stem cells into neural stem cells. Cell Mol Neurobiol 34(6):905–912
Kennea NL, Mehmet H (2002) Neural stem cells. J Pathol 197(4):536–550
Kempermann G, Kuhn HG, Gage FH (1998) Experience-induced neurogenesis in the senescent dentate gyrus. J Neurosci 18(9):3206–3212
Kempermann G (2002) Regulation of adult hippocampal neurogenesis – implications for novel theories of major depression. Bipolar Disord 4(1):17–33
Hisahara S, Chiba S, Matsumoto H, Tanno M, Yagi H, Shimohama S, Sato M, Horio Y (2008) Histone deacetylase SIRT1 modulates neuronal differentiation by its nuclear translocation. Proc Natl Acad Sci U S A 105(40):15599–15604
Liu DJ, Hammer D, Komlos D, Chen KY, Firestein BL, Liu AY (2014) SIRT1 Knockdown Promotes Neural Differentiation and Attenuates the Heat Shock Response. J Cell Physiol 229(9):1224–1235
Joe IS, Jeong SG, Cho GW (2015) Resveratrol-induced SIRT1 activation promotes neuronal differentiation of human bone marrow mesenchymal stem cells. Neurosci Lett 584:97–102
Kageyama R, Shimojo H, Imayoshi I (2015) Dynamic expression and roles of Hes factors in neural development. Cell Tissue Res 359(1):125–133
Kopan R, Ilagan MX (2009) The canonical Notch signaling pathway: unfolding the activation mechanism. Cell 137(2):216–233
Louvi A, Artavanis-Tsakonas S (2006) Notch signalling in vertebrate neural development. Nat Rev Neurosci 7(2):93–102
Kageyama R, Ohtsuka T, Kobayashi T (2007) The Hes gene family: repressors and oscillators that orchestrate embryogenesis. Development 134(7):1243–1251
Ishibashi M, Ang SL, Shiota K, Nakanishi S, Kageyama R, Guillemot F (1995) Targeted disruption of mammalian hairy and Enhancer of split homolog-1 (HES-1) leads to up-regulation of neural helix-loop-helix factors, premature neurogenesis, and severe neural tube defects. Genes Dev 9(24):3136–3148
Tomita K, Ishibashi M, Nakahara K, Ang SL, Nakanishi S, Guillemot F, Kageyama R (1996) Mammalian hairy and Enhancer of Split Homolog 1 Regulates Differentiation of Retinal Neurons and Is Essential for Eye Morphogenesis. Neuron 16(4):723–734
Cheng HL, Mostoslavsky R, Saito S, Manis JP, Gu Y, Patel P, Bronson R, Appella E et al (2003) Developmental defects and p53 hyperacetylation in Sir2 homolog (SIRT1)-deficient mice. Proc Natl Acad Sci U S A 100(19):10794–10799
Ichi S, Boshnjaku V, Shen YW, Mania-Farnell B, Ahlgren S, Sapru S, Mansukhani N, McLone DG et al (2011) Role of Pax3 acetylation in the regulation of Hes1 and Neurog2. Mol Biol Cell 22(4):503–512
Tiberi L, van den Ameele J, Dimidschstein J, Piccirilli J, Gall D, Herpoel A, Bilheu A, Bonnefont J et al (2012) BCL6 controls neurogenesis through Sirt1-dependent epigenetic repression of selective Notch targets. Nat Neurosci 15(12):1627–1635
Tiberi L, Bonnefont J, van den Ameele J, Le Bon SD, Herpoel A, Bilheu A, Baron BW, Vanderhaeghen P (2014) A BCL6/BCOR/SIRT1 Complex Triggers Neurogenesis and Suppresses Medulloblastoma by Repressing Sonic Hedgehog Signaling. Cancer Cell 26(6):797–812
Wang W, Osenbroch P, Skinnes R, Esbensen Y, Bjørås M, Eide L (2010) Mitochondrial DNA integrity is essential for mitochondrial maturation during differentiation of neural stem cells. Stem Cells 28(12):2195–2204
Wang W, Esbensen Y, Kunke D, Suganthan R, Rachek L, Bjørås M, Eide L (2011) Mitochondrial DNA damage level determines neural stem cell differentiation fate. J Neurosci 31(26):9746–9751
Santos DM, Santos MM, Moreira R, Solá S, Rodrigues CM (2013) Synthetic condensed 1,4-naphthoquinone derivative shifts neural stem cell differentiation by regulating redox state. Mol Neurobiol 47(1):313–324
Sun G, Yu RT, Evans RM, Shi Y (2007) Orphan nuclear receptor TLX recruits histone deacetylases to repress transcription and regulate neural stem cell proliferation. Proc Natl Acad Sci U S A 104(39):15282–15287
Iwahara N, Hisahara S, Hayashi T, Horio Y (2009) Transcriptional activation of NAD+-dependent protein deacetylase SIRT1 by nuclear receptor TLX. Biochem Biophys Res Commun 386(4):671–675
Yu RT, Chiang MY, Tanabe T, Kobayashi M, Yasuda K, Evans RM, Umesono K (2000) The orphan nuclear receptor Tlx regulates Pax2 and is essential for vision. Proc Natl Acad Sci U S A 97(6):2621–2625
Horn V, Minucci S, Ogryzko VV, Adamson ED, Howard BH, Levin AA, Ozato K (1996) RAR and RXR selective ligands cooperatively induce apoptosis and neuronal differentiation in P19 embryonal carcinoma cells. FASEB J 10(9):1071–1077
Zhang C, Dowd DR, Staal A, Gu C, Lian JB, van Wijnen AJ, Stein GS, MacDonald PN (2003) Nuclear Coactivator-62 kDa/Ski-interacting Protein Is a Nuclear Matrix-associated Coactivator That May Couple Vitamin D Receptor-mediated Transcription and RNA Splicing. J Biol Chem 278(37):35325–35336
Kang MR, Lee SW, Um E, Kang HT, Hwang ES, Kim EJ, Um SJ (2010) Reciprocal roles of SIRT1 and SKIP in the regulation of RAR activity: implication in the retinoic acid-induced neuronal differentiation of P19 cells. Nucleic Acids Res 38(3):822–831
Yu S, Levi L, Siegel R, Noy N (2012) Retinoic acid induces neurogenesis by activating both retinoic acid receptors (RARs) and peroxisome proliferator-activated receptor beta/delta (PPARbeta/delta). J Biol Chem 287(50):42195–42205
Xie Z, Kasschau KD, Carrington JC (2003) Negative Feedback Regulation of Dicer-Like1in Arabidopsis by microRNA-Guided mRNA Degradation. Curr Biol 13(9):784–789
Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J, Bartel DP, Linsley PS et al (2005) Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433(7027):769–773
Aranha MM, Santos DM, Sola S, Steer CJ, Rodrigues CM (2011) miR-34a regulates mouse neural stem cell differentiation. PLoS One 6(8):e21396
Delaloy C, Liu L, Lee JA, Su H, Shen F, Yang GY, Young WL, Ivey KN et al (2010) MicroRNA-9 coordinates proliferation and migration of human embryonic stem cell-derived neural progenitors. Cell Stem Cell 6(4):323–335
Aranha MM, Solá S, Low WC, Steer CJ, Rodrigues CM (2009) Caspases and p53 Modulate FOXO3A/Id1 Signaling During Mouse Neural Stem Cell Differentiation. J Cell Biochem 107(4):748–758
Meletis K, Wirta V, Hede SM, Nistér M, Lundeberg J, Frisen J (2006) p53 suppresses the self-renewal of adult neural stem cells. Development 133(2):363–369
Armesilla-Diaz A, Bragado P, Del Valle I, Cuevas E, Lazaro I, Martin C, Cigudosa JC, Silva A (2009) p53 regulates the self-renewal and differentiation of neural precursors. Neuroscience 158(4):1378–1389
Lee JS, Park JR, Kwon OS, Lee TH, Nakano I, Miyoshi H, Chun KH, Park MJ et al (2015) SIRT1 is required for oncogenic transformation of neural stem cells and for the survival of “cancer cells with neural stemness” in a p53-dependent manner. Neuro Oncol 17(1):95–106
Pistritto G, Papaleo V, Sanchez P, Ceci C, Barbaccia ML (2012) Divergent modulation of neuronal differentiation by caspase-2 and -9. PLoS One 7(5):e36002
Shin SDS, Stice SL (2005) Human Motor Neuron Differentiation from Human Embryonic Stem Cells. Stem Cells Dev 14(3):266–269
Li XJ, Hu BY, Jones SA, Zhang YS, Lavaute T, Du ZW, Zhang SC (2008) Directed differentiation of ventral spinal progenitors and motor neurons from human embryonic stem cells by small molecules. Stem Cells 26(4):886–893
Zhang Y, Wang J, Chen G, Fan D, Deng M (2011) Inhibition of Sirt1 promotes neural progenitors toward motoneuron differentiation from human embryonic stem cells. Biochem Biophys Res Commun 404(2):610–614
Jacobs FM, van Erp S, van der Linden AJ, von Oerthel L, Burbach JP, Smidt MP (2009) Pitx3 potentiates Nurr1 in dopamine neuron terminal differentiation through release of SMRT-mediated repression. Development 136(4):531–540
Kim TE, Seo JS, Yang JW, Kim MW, Kausar R, Joe E, Kim BY, Lee MA (2013) Nurr1 Represses Tyrosine Hydroxylase Expression via SIRT1 in Human Neural Stem Cells. PLoS One 8(8):e71469
Kim MJ, Ahn K, Park SH, Kang HJ, Jang BG, Oh SJ, Oh SM, Jeong YJ et al (2009) SIRT1 regulates tyrosine hydroxylase expression and differentiation of neuroblastoma cells via FOXO3a. FEBS Lett 583(7):1183–1188
Beuckmann CT, Yanagisawa M (2002) Orexins: from neuropeptides to energy homeostasis and sleep/wake regulation. J Mol Med (Berl) 80(6):329–342
Hayakawa K, Hirosawa M, Tabei Y, Arai D, Tanaka S, Murakami N, Yagi S, Shiota K (2013) Epigenetic switching by the metabolism-sensing factors in the generation of orexin neurons from mouse embryonic stem cells. J Biol Chem 288(24):17099–17110
Barbieri AM, Broccoli V, Bovolenta P, Alfano G, Marchitiello A, Mocchetti C, Crippa L, Bulfone A et al (2002) Vax2 inactivation in mouse determines alteration of the eye dorsal-ventral axis, misrouting of the optic fibres and eye coloboma. Development 129(3):805–813
Hasegawa K, Yoshikawa K (2008) Necdin regulates p53 acetylation via Sirtuin1 to modulate DNA damage response in cortical neurons. J Neurosci 28(35):8772–8784
Hayakawa N, Shiozaki M, Shibata M, Koike M, Uchiyama Y, Matsuura N, Gotow T (2013) Resveratrol affects undifferentiated and differentiated PC12 cells differently, particularly with respect to possible differences in mitochondrial and autophagic functions. Eur J Cell Biol 92(1):30–43
Fujino K, Ogura Y, Sato K, Nedachi T (2013) Potential neuroprotective effects of SIRT1 induced by glucose deprivation in PC12 cells. Neurosci Lett 557:148–153
Uittenbogaard M, Baxter KK, Chiaramello A (2010) The neurogenic basic helix-loop-helix transcription factor NeuroD6 confers tolerance to oxidative stress by triggering an antioxidant response and sustaining the mitochondrial biomass. ASN Neuro 2(2):e00034
Sansone L, Reali V, Pellegrini L, Villanova L, Aventaggiato M, Marfe G, Rosa R, Nebbioso M et al (2013) SIRT1 silencing confers neuroprotection through IGF-1 pathway activation. J Cell Physiol 228(8):1754–1761
Taupin P, Gage FH (2002) Adult neurogenesis and neural stem cells of the central nervous system in mammals. J Neurosci Res 69(6):745–749
Zakhary SM, Ayubcha D, Dileo JN, Jose R, Leheste JR, Horowitz JM, Torres G (2010) Distribution analysis of deacetylase SIRT1 in rodent and human nervous systems. Anat Rec (Hoboken) 293(6):1024–1032
Ma CY, Yao MJ, Zhai QW, Jiao JW, Yuan XB, Poo MM (2014) SIRT1 suppresses self-renewal of adult hippocampal neural stem cells. Development 141(24):4697–4709
Schmidt-Strassburger U, Schips TG, Maier HJ, Kloiber K, Mannella F, Braunstein KE, Holzmann K, Ushmorov A et al (2012) Expression of constitutively active FoxO3 in murine forebrain leads to a loss of neural progenitors. FASEB J 26(12):4990–5001
Rafalski VA, Ho PP, Brett JO, Ucar D, Dugas JC, Pollina EA, Chow LM, Ibrahim A et al (2013) Expansion of oligodendrocyte progenitor cells following SIRT1 inactivation in the adult brain. Nat Cell Biol 15(6):614–624
Xiang Z, Krainc D (2013) Pharmacological upregulation of PGC1alpha in oligodendrocytes: implications for Huntington’s Disease. J Huntingtons Dis 2(1):101–105
Stettner M, Wolffram K, Mausberg AK, Albrecht P, Derksen A, Methner A, Dehmel T, Hartung HP et al (2013) Promoting myelination in an in vitro mouse model of the peripheral nervous system: the effect of wine ingredients. PLoS One 8(6):e66079
Stein LR, Imai S (2014) Specific ablation of Nampt in adult neural stem cells recapitulates their functional defects during aging. EMBO J 33(12):1321–1340
Parsons XH (2012) An Engraftable Human Embryonic Stem Cell Neuronal Lineage-Specific Derivative Retains Embryonic Chromatin Plasticity for Scale-Up CNS Regeneration. J Regen Med Tissue Eng 1(1):3
Parsons XH (2013) Embedding the Future of Regenerative Medicine into the Open Epigenomic Landscape of Pluripotent Human Embryonic Stem Cells. Annu Res Rev Biol 3(4):323–349
Acknowledgments
This study was supported by the National Nature Science Foundation of China (No. (81371197, 31271051), Natural Science Foudation Project of CQ CSTC 2013jjB10028.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Cai, Y., Xu, L., Xu, H. et al. SIRT1 and Neural Cell Fate Determination. Mol Neurobiol 53, 2815–2825 (2016). https://doi.org/10.1007/s12035-015-9158-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-015-9158-6