Abstract
Despite recent advances in clinical treatments, identifying high-risk osteosarcoma (OS) patients remains an unresolved clinical challenge. Mitophagy, a specialized form of cellular autophagy, selectively reduces the number of mitochondria or repairs their abnormal functions in response to external stress, thereby ensuring mitochondrial quality and maintaining mitochondrial function. Mitophagy plays a crucial role in cancer development, including processes such as mitochondrial repair, homeostasis maintenance, and tumor metabolism. However, its impact on OS has not yet been reported. In this study, we collected 58 mitophagy-related genes (MPRGs) from the TARGET and GEO databases and bioinformatically screened for those associated with OS prognosis. By LASSO-multivariable Cox regression algorithm, we subsequently developed a novel scoring system, the MPRG score, and validated its significance in predicting OS prognosis. Immune landscape analysis showed patients in the low MPRG group had a higher immune infiltration level than those in the high MPRG group. Drug sensitivity differences highlighted the potential need for alternative therapeutic strategies based on MPRG scoring system. The distribution characteristics of the MPRG signature in different cell subtypes of OS were explored by single-cell sequencing analyses. In vitro experiments further confirmed the abnormal expression of screened targets in OS. Our findings highlight the role of mitophagy in OS and its potential as a therapeutic target.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Osteosarcoma (OS) is the most common primary bone malignancy, with its bimodal incidence peaked at 18 and 60 years of age [1]. Despite advancements in chemotherapy and surgical techniques, the 5-year survival rate for OS patients remains unsatisfactory, and there is a critical need for reliable biomarkers to predict patient outcomes [2, 3]. How to identify high-risk OS patients is still an unsolved clinical issue. Specific histologic features assessed on resection has been considered an indicator of therapeutic response in OS patients receiving neoadjuvant chemotherapy, but this protocol has not been widely accepted [4]. Strategies that can predict treatment sensitivity and risk stratification of outcomes in patients with OS are still urgently needed to further identify patients with OS with poor prognosis.
Autophagy is a catabolic pathway that recycles intracellular components by encapsulating them in bilayer vesicles called autophagosomes [5]. These vesicles can then fuse with lysosomes for degradation purposes [6, 7]. As a special form of cellular autophagy, the process of mitophagy allows mitochondria to respond to external pressures by selectively reducing the number of mitochondria or repairing abnormal mitochondrial function, thereby ensuring mitochondrial quality and maintaining mitochondrial function [8, 9]. Current studies have divided the mechanisms of mitophagy into ubiquitin-dependent and non-ubiquitin-dependent pathways [10]. The classical ubiquitin-dependent pathway refers to the ubiquitination of proteins on mitochondria by mitophagy-related proteins such as PINK1 and leads to the aggregation of microtubule-associated protein 1 light chain 3 (LC3). Finally, the lysosome is polymerized into autophagy lysosome [11]. In addition, NIP3-like protein X (NIX) receptor, the bcl2-interacting protein 3 (BNIP3) receptor and other proteins could directly aggregate with LC3. This pathway is known as the non-ubiquitin-dependent pathway [12].
Mitophagy has been reported to play an important role in the development of cancer [13]. According to different tumor types and different stages, mitophagy can play two opposite roles in cancer development: cancer inhibition and cancer promotion [14]. In the early stages of cancer, mitophagy can maintain cell metabolism while reducing mitochondrial damage, thereby avoiding further cancer development by maintaining homeostasis [15]. However, in advanced cancer cells, mitophagy provides nutrients that sustain cancer cell survival, invasion, and metastasis, thereby increasing tolerance to hypoxia and hypo-nutrition and supporting the tumor's high bioenergy requirements [16]. As critical regulator of mitophagy, the PINK1/Parkin pathway is significantly upregulated in OS cells, leading to enhanced removal of damaged mitochondria and improved cellular resilience [17]. Mitophagy promotion by BNIP3 in OS can attenuates cisplatin resistance [18]. Based on these evidences, we hypothesize that mitophagy may contribute to OS progression by promoting mitochondrial quality control, thereby enabling cancer cells to survive under metabolic stress conditions commonly found in the tumor microenvironment.
In this study, we aimed to identify mitophagy-related genes (MPRGs) associated with OS prognosis and develop and validate a prognostic scoring system based on MPRGs. The relationship between MPRG expression, immune infiltration, and drug resistance in OS was also explored. Our results suggest the role of mitophagy in OS and its potential value as a therapeutic target.
Materials and Methods
Data Acquisition and Download of Mitophagy Gene Signature
The osteosarcoma (OS) transcriptome data and baseline clinical characteristics used in this study were obtained from the TARGET and GEO databases. From the TARGET database, we collected a total of 87 OS samples that included clinical features such as survival time and status. Additionally, we acquired two gene chip datasets containing OS sample information (GSE21257 and GSE39058) from the GEO database. To eliminate batch effects between the TARGET, GSE21257, and GSE39058 data, we standardized the transcriptome data using the "sva" script. This process resulted in a final dataset of 140 OS samples for subsequent analysis. The mitophagy-related genes (MPRG) were obtained from the MSigDB database. Supplementary Table 1 lists the 58 mitophagy-related genes used in this study.
Identification of Prognosis-Related MPRGs and Consensus Clustering Molecular Subtype Analysis
To identify mitophagy-related genes (MPRGs) associated with OS prognosis, we extracted the expression of MPRGs from the gene expression matrix of OS samples using the “limma” package. By combining survival time and clinical outcomes of OS samples from the TARGET, GSE21257, and GSE39058 datasets, we employed the LASSO univariate Cox regression analysis algorithm to identify MPRGs associated with clinical survival outcomes of OS. To explore the molecular subtypes associated with MPRGs in OS, we used the “ConsensusClusterPlus” package to cluster OS patients into 2–9 subgroups based on the expression matrix of MPRGs related to clinical survival prognosis, with standards of pItem = 0.8 and pFeature = 1. The optimal K value was determined from the clustering results. We then used the “survival” package to analyze the survival differences among the different OS subgroups by integrating patients' survival time and clinical outcomes. The “limma” package was employed to identify differentially expressed genes among the subgroups. Furthermore, pathway enrichment analysis of these differential genes was conducted using the “GSVA” package to discover signaling pathways associated with the prognostic differences between subgroups.
Assessment of Immune Cell Infiltration Characteristics and Immune Regulatory Function
Based on the OS transcriptome matrix, we applied the “estimate” script to assess the immune infiltration status of each OS sample. This assessment included calculating the immune, stromal, ESTIMATE scores, and tumor purity for each OS sample. Using marker gene signatures of 23 immune cell types, we evaluated the relative proportions of immune cells and immune regulatory function scores in OS samples through the ssGSEA algorithm. To explore the potential association between MPRGs and the OS immune microenvironment, we used the Pearson correlation algorithm to assess the correlations between MPRGs and immune cells. The results were visualized using the "pheatmap" package.
Construction and Validation of the MPRG Scoring System
To accurately predict the clinical survival outcomes of OS, we constructed a new scoring system, the MPRG scoring system, based on the results of multivariate Cox regression. The scoring algorithm is defined as MPRG score = Σ (Prognosis-related MPRGi * Coefi). Using this algorithm, all osteosarcoma patients were scored, and patients were divided into high MPRG score and low MPRG score groups based on the median score. To evaluate the accuracy of the MPRG scoring system, patients were randomly divided into training (70%) and testing (30%) groups using the create Data Partition function from the “caret” R package. Similar to the whole dataset, patients in the training and testing groups were categorized into high and low MPRG score groups based on their scores. The distribution of MPRG scores in the whole group, training group, and testing group was displayed using scatter plots to study the relationship between MPRG scores and patients' survival time and clinical outcomes. The "survival" package was used to analyze the survival differences between different MPRG score groups, thereby clarifying the relationship between MPRG scores and patient prognosis.
Construction and Validation of the Prognostic Nomogram
Clinical characteristics can significantly influence patient prognosis. To provide a more accurate prognostic assessment, we incorporated patients' clinical features as covariates in our modeling process. By combining clinical characteristics with the MPRG score, we constructed a nomogram for predicting patient prognosis, thereby minimizing potential biases from other clinical factors. In this study, we integrated the MPRG score with clinical information (including age, gender, and metastasis) to construct the nomogram using the “rms” package. Calibration curves were employed to evaluate the predictive accuracy of the nomogram. These curves visualize the agreement between the actual and predicted incidence rates, with closer alignment indicating a more accurate prediction model.
Single-Cell Sequencing Data Acquisition and Analysis
The single-cell RNA sequencing dataset GSE152048, comprising transcriptomic profiles from osteosarcoma samples, was obtained from the Gene Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/geo/). Raw sequencing data were subjected to quality control using FastQC. Low-quality reads and adapters were removed using Trimmomatic, cleaned reads were aligned to the reference genome using STAR, and gene expression quantification was performed with feature Counts. The Seurat R package was utilized for downstream analysis: Cells with low library sizes and high mitochondrial gene content were filtered out. Data normalization was performed to adjust for differences in sequencing depth and cell size. Cells with fewer than 200 detected genes or more than 5% mitochondrial gene content were excluded from the analysis. Clustering was performed using the Louvain algorithm with a resolution parameter of 0.8. Principal component analysis (PCA) was applied to reduce data dimensionality. UMAP was used for visualization. Cells were clustered based on significant principal components using a graph-based clustering approach. Known marker genes and differential expression analysis were used to annotate cell types within the SingleR database. Differential gene expression between identified cell clusters was assessed using the FindMarkers function in Seurat. Statistical significance was determined using the Wilcoxon rank-sum test, with adjustments for multiple testing (Benjamini–Hochberg correction). Results were visualized using t-SNE and UMAP plots to explore cell clustering and differential gene expression patterns. Heatmaps and violin plots were generated to visualize expression profiles of marker genes across cell clusters.
Drug Sensitivity Analysis
Drug sensitivity analysis was performed using the “pRRophetic” package, which allows us to understand the sensitivity of different patient groups to anti-tumor drugs. This understanding can facilitate personalized treatment for different patients, enhancing the clinical applicability of our predictive model. The results of drug sensitivity analysis were visualized using the "ggplot2" package, incorporating osteosarcoma patient subgroup and score group information to reveal differences in drug sensitivity among various subgroups or scoring groups.
Cell Culture
U2OS human osteosarcoma cells were obtained from ATCC and cultured in DMEM supplemented with 10% fetal bovine serum (FBS) and 1% penicillin–streptomycin. U2OS cells were maintained in a humidified incubator at 37 °C with 5% CO2. Normal human (Human Fibroblast Cells) cells were cultured in appropriate medium DMEM supplemented with 10% FBS and 1% penicillin–streptomycin under similar conditions as U2OS cells. U2OS cells were used at passage numbers between 5 and 20 and were authenticated using short tandem repeat (STR) profiling before experiments.
Western Blot
Total protein from U2OS cells and control cells was extracted using RIPA lysis buffer (R0010, Solarbio, China) supplemented with 1 mM protease inhibitor mixture and 1 mM phosphatase inhibitor mixture. After adding an appropriate amount of lysis buffer on ice, the mixture was shaken for 30 min and then centrifuged at 12,000 g for 10 min at 4 °C. The supernatant was collected, and total protein quantification was performed using the BCA protein assay kit (PC0020, Solarbio, China). For quantification, 20 µl of protein sample was mixed with 200 µl of BCA working solution and incubated at 37 °C for 30 min. Absorbance at 562 nm was measured using a microplate reader (CMax Plus, USA), and protein concentration was inferred from the standard curve of BSA protein standards. Protein supernatant was mixed with 5 × protein loading buffer (P1040, Solarbio, China) at a 4:1 (V/V) ratio and boiled for 10 min. Proteins were separated by SDS-PAGE and transferred to a polyvinylidene fluoride (PVDF) membrane. The PVDF membrane was blocked with 5% skim milk at room temperature for 1 h. The membrane was then incubated overnight at 4 °C with primary antibodies: β-actin (E-AB-48018, Elabscience, China), ATG4D (ab237751, Abcam), RPS27A (ab172293, Abcam), and TOMM20 (ab78547, Abcam). The next day, the membrane was incubated with the corresponding HRP-conjugated secondary antibody (Elabscience, China) for 1 h. ECL reagent (34,577, Thermo Fisher, USA) was used for detection. The grayscale values of the bands were quantified using ImageJ software. Primary antibodies were diluted 1:1000 for β-actin and 1:500 for ATG4D, RPS27A, and TOMM20. Secondary antibody incubation was performed at a 1:2000 dilution, and ECL exposure times ranged from 1 to 5 min, depending on the signal strength.
Real-Time Quantitative PCR Analysis
RNA from tumor tissues was isolated using the TRIzol™ Plus reagent (Thermo Fisher, USA) according to the manufacturer's instructions. The isolated RNA was then reverse-transcribed into cDNA using a reverse transcription kit (AG11728, Accurate Biology, China). The cDNA was mixed with primers and SYBR Green Pro Taq HS premix (AG11701, Accurate Biology, China) and detected using a Bio-Rad real-time quantitative PCR system (Bio-Rad, USA). The relative expression levels of target genes ATG4D, RPS27A, and TOMM20 were normalized to glyceraldehyde-3-phosphate dehydrogenase (GAPDH) as the reference gene. The relative gene expression was calculated using the 2−∆∆CT method. The primer sequences used are listed in Table 1.
Statistical Analysis
All analyses were conducted using R software (version 4.4.1) with Bioconductor packages and custom scripts, along with the Perl programming environment. Survival curves between different subgroups were analyzed using the log-rank algorithm. Pearson correlation analysis was employed to assess relationships between variables. Statistical differences between two groups were assessed using the Wilcoxon rank-sum test, while one-way ANOVA was utilized for comparisons among multiple groups. Cell experiments in this study were performed in triplicate, and statistical analyses of differences were conducted using SPSS and GraphPad software (Student's t-test) for both statistical testing and visualization. Statistical significance was set at corrected p < 0.05. Data are presented as mean ± SD, with significance levels indicated as *p < 0.05, **p < 0.01, ***p < 0.001.
Results
Identification of Mitophagy-Associated Prognostic Genes and Unsupervised Consensus Clustering Analysis
We retrieved 58 genes associated with mitophagy from gene set databases. Univariate Cox regression analysis of these 58 mitophagy-associated genes revealed 6 genes significantly associated with prognosis in osteosarcoma patients. Further analysis using LASSO-multivariable Cox regression identified three mitophagy-associated prognostic genes: ATG4D, RPS27A, and TOMM20 (Fig. 1a, b). Additionally, based on the expression levels of these three mitophagy-associated genes, patients were stratified into 2–9 distinct subtypes. Notably, when K = 3, osteosarcoma patients were effectively classified into three subtypes: MPRG cluster A, MPRG cluster B, and MPRG cluster C (Fig. 1c). Survival and clinical outcome curves among the three MPRG subtypes demonstrated statistically significant differences in prognosis. Patients in MPRG cluster C exhibited favorable outcomes, whereas those in MPRG cluster B showed the poorest prognosis (p < 0.05, Fig. 1d). In summary, the expression of mitophagy-associated prognostic genes in osteosarcoma correlates closely with patient prognosis. Alterations in the expression of mitophagy-associated prognostic genes may significantly impact patient outcomes.
Identification of mitophagy-associated prognostic genes and unsupervised consensus clustering analysis results. a, b LASSO regression analysis results of mitophagy-associated genes. c Results of unsupervised consensus clustering analysis based on mitophagy-associated prognostic genes. d Kaplan–Meier (KM) curve results of MPRG subtypes
Immune Landscape Analysis Based on MPRG Clusters
To assess differences in immune landscapes among MPRG clusters, we employed multiple analytical algorithms to evaluate immune profiles across samples. ESTIMATE was used to assess immune status in tumor patients based on expression matrices. Results revealed that MPRG cluster C exhibited the highest stromal score, immune score, and ESTIMATE score, and the lowest Tumor Purity, whereas MPRG cluster B exhibited the opposite trends (Fig. 2a–d). This suggests that MPRG cluster C is characterized by greater immune cell infiltration compared to the other two subtypes. Combining these findings with prognostic outcomes indicates a positive correlation between immune scores, immune cell infiltration, and patient prognosis. Further analysis using single-sample Gene Set Enrichment Analysis (ssGSEA) was conducted to delineate immune cell infiltration and functional differences among different subgroups. Results showed that MPRG cluster C patients exhibited significantly higher infiltration levels of most immune cells compared to MPRG cluster B, particularly Activated B cells, Activated CD8 T cells, Activated dendritic cells, and Immature B cells (Fig. 2e). In addition, immune functional analysis indicated superior immune function in MPRG cluster C across multiple immune parameters compared to the other groups (Fig. 2f). These findings underscore significant immune landscape disparities among different MPRG clusters, particularly the heightened infiltration of key immune cells, which may impact patient prognosis. Lastly, pathway analysis using Gene Set Variation Analysis (GSVA) identified pathways contributing to these differences. Results indicated distinctions in pathways such as KEGG focal adhesion, KEGG adherens junction, KEGG FC gamma R mediated phagocytosis, KEGG glycosaminoglycan biosynthesis chondroitin sulfate, KEGG sulfur metabolism, KEGg nicotinate and nicotinamide metabolism, and KEGG steroid biosynthesis among MPRG clusters (Fig. 2g, h).
Immune Landscape of MPRG Clusters. a–d Analysis of ESTIMATE scores among MPRG clusters using the ESTIMATE algorithm. e Analysis depicting variations in immune cell infiltration among different MPRG clusters. f Comparison of immune function differences among different MPRG clusters. g KEGG Pathway Enrichment Analysis in MPRG Clusters A and C. h KEGG Pathway Enrichment Analysis in MPRG Clusters B and C
Construction and Validation of the MPRG Scoring Model
Based on LASSO-Cox regression analysis results, we developed a novel scoring system termed MPRG score. The formula for calculating MPRG score is: MPRG score = ATG4D * 3.20 + RPS27A * 5.940 + TOMM20 * 2.00. Furthermore, to validate the stability of the scoring model, osteosarcoma patients were randomly divided into training and testing groups at a ratio of 7:3. As shown in Fig. 3a–c, osteosarcoma patients were stratified into low (blue) and high (red) MPRG score groups based on the median MPRG score. Scatter plots indicated that patients with clinical outcomes of death tended to have higher MPRG scores, consistent across the entire cohort, training, and testing groups (Fig. 3d–f). Heatmap results demonstrated that expression levels of mitochondria autophagy-related prognostic genes increased with higher MPRG scores, indicating their role as risk factors (Fig. 3g–i). Kaplan–Meier survival curves showed that high MPRG score groups were associated with shorter survival times, while low MPRG score groups had longer survival times, consistently observed across the entire cohort (p < 0.001), training group (p < 0.001), and testing group (p = 0.006) (Fig. 4a–c). ROC analysis indicated AUCs of 0.797 at 1 year, 0.716 at 3 years, and 0.744 at 5 years for the entire cohort (Fig. 4d). For the training group, AUCs were 0.852 at 1 year, 0.630 at 3 years, and 0.676 at 5 years (Fig. 4e), while for the testing group, AUCs were 0.793 at 1 year, 0.757 at 3 years, and 0.778 at 5 years (Fig. 4f). In conclusion, these results demonstrate that MPRG score correlates with patient survival time and clinical outcomes in osteosarcoma, showing promising accuracy in predicting prognosis using MPRG score.
Construction and Validation of the MPRG Scoring Model. a–c Distribution of MPRG scores in entire cohort, training cohort and testing cohort. d–f Scatter plot of MPRG scores, survival time, and clinical outcomes in the entire cohort, training cohort and testing cohort. g–i Heatmap of prognostic MPRGs expression in the entire cohort, training cohort and testing cohort
Construction and Evaluation Of Prognostic Model Based on Clinical Features and MPRG Score
Clinical features of patients can also impact prognosis. To accurately predict prognosis in osteosarcoma patients, we integrated clinical features and MPRG scores to construct a prognostic model. Patient age, gender, tumor metastasis status, and MPRG score are shown in Fig. 5a. Univariate Cox regression analysis results indicated that metastases (p < 0.001) and MPRG score (p < 0.001) were associated with patient prognosis (Fig. 5b). Multivariate Cox regression analysis showed that metastases (p < 0.001) and MPRG score (p < 0.001) independently assessed prognosis (Fig. 5c). Finally, we constructed a nomogram based on age, gender, metastases, and MPRG score, accurately predicting prognosis in osteosarcoma patients based on the sum of scores for each indicator (Fig. 5d). Calibration curve results demonstrated close alignment between predicted and actual curves for prognosis at 1, 3, and 5 years (Fig. 5e), indicating high accuracy, minimal prediction bias, and strong predictive capability of the model. Figure 5f, g depicts differences in MPRG scores among different MPRG clusters. Consistent with previous findings, MPRG cluster C, associated with the best prognosis, had the lowest MPRG scores, whereas MPRG cluster B, associated with the poorest prognosis, had the highest MPRG scores (Fig. 5h). The relationship between MPRG clusters, MPRG scores, and clinical outcomes is illustrated in Fig. 5i. In conclusion, considering the diversity of clinical features among patients, integrating these with MPRG scores into prognostic assessment systems enables more accurate evaluation and prognosis prediction for osteosarcoma patients, demonstrating good predictive accuracy.
Construction and evaluation of prognostic model based on clinical features and MPRG Score. a Heatmap of patient clinical features and MPRG scores. b Univariate Cox regression analysis results of clinical features and MPRG scores. c Multivariate Cox regression analysis results of clinical features and MPRG scores. d Nomogram construction based on clinical features and MPRG scores. e Calibration curve results of the nomogram. f Concordance index results of the nomogram. g Decision curve analysis results of the nomogram. h MPRG score results across MPRG clusters. i Sankey diagram illustrating the association between MPRG clusters, MPRG scores, and clinical outcomes
Immune Landscape and Correlation Analysis in Different MPRG Score Groups
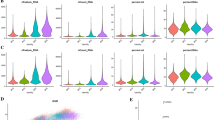
Previous results have shown that varying degrees of immune infiltration can impact patient prognosis. Therefore, focusing on MPRG scores, we conducted immune landscape analysis among patients grouped by MPRG scores. ESTIMATE results indicated that the low MPRG score group generally exhibited higher Immune Score, ESTIMATE Score, Stromal Score, and lower Tumor Score (Fig. 6a–d), suggesting greater immune cell infiltration in these patients. ssGSEA results further highlighted increased infiltration levels of Activated B cells, activated dendritic cells, Immature B cells, myeloid-derived suppressor cells (MDSCs), and various T helper cells in the low MPRG score group, indicating a close association of these immune cells with prognosis in osteosarcoma patients (Fig. 6e). Additionally, compared to the high MPRG score group, most immune functions were more activated in the low MPRG score group (Fig. 6f). As Fig. 6g shows that, apart from Activated CD4 T cells, Type 2 T helper cells, Gamma delta T cells, and Immature dendritic cells, most immune cells were negatively correlated with MPRG scores in patients. Correlation analysis revealed that mitochondrial autophagy-related prognostic genes ATG4D and RPS27A were positively correlated with most immune cells, while TOMM20 gene showed an inverse correlation with most immune cells (Fig. 6h). In conclusion, there are differences in immune landscapes among patients with different MPRG scores, and MPRG scores to some extent can indicate the extent of immune infiltration in patients.
Immune landscape and correlation analysis in different MPRG score groups. a–d Immune score, ESTIMATE score, tumor purity and stromal score based on ESTIMATE algorithm. e Differences in immune cell infiltration among patients with different MPRG scores. f Differences in immune functions among patients with different MPRG scores. g Correlation analysis between immune cells and MPRG scores. h Correlation analysis between immune cells and prognostic MPRGs
Drug sensitivity Results and Correlation Analysis in Different MPRG Score Groups
In order to tailor precise treatment plans for patients with different MPRG scores and enhance the clinical utility of drugs, we conducted drug sensitivity analysis and correlation analysis to clarify the relationship between drug sensitivity and MPRG scores. The results are shown in Fig. 7. Patients in different MPRG score groups exhibited varying sensitivity to the same drugs. For instance, drugs like TGX211, Midostaurin, and Bleomycin showed lower IC50 values in patients with low MPRG scores, whereas Pyrimethamine, Sorafenib, and Imatinib showed the opposite trend. Furthermore, IC50 values of TGX211 (R = 0.66, p = 1.9e − 07), Midostaurin (R = 0.47, p = 0.00045), and Bleomycin (R = 0.49, p = 0.00025) were positively correlated with MPRG scores, while Pyrimethamine (R = − 0.43, p = 0.0015), Sorafenib (R = − 0.44, p = 0.0033), and Imatinib (R = − 0.47, p = 0.00046) were negatively correlated with MPRG scores. This indicates that MPRG scores can be used for individualized assessment of drug sensitivity, providing high clinical utility for personalized treatment of patients.
Single-Cell Sequencing Analysis Reveals Expression Characteristics of MPRG Prognostic Signature
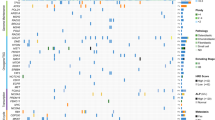
We further evaluated the immune cell landscape and the distribution characteristics of the MPRG signature at the single-cell level in OS. Based on the GSE152048 dataset, we extracted single-cell sequencing data from four OS samples for subsequent analysis. After batch effect correction and normalization for each sample, we used PCA for dimensionality reduction and identified 21 cell clusters in OS samples, as shown in Fig. 8a. Expression level results indicated that among these 21 cell clusters, RPS27A and TOMM20 were highly expressed (Fig. 8b). Using “SingleR” method, we annotated the 21 cell clusters and identified six cell subtypes: Macrophages, chondrocytes, tissue stem cells, monocytes, MSCs, and endothelial cells (Fig. 8c). Notably, RPS27A was significantly highly expressed among these six cell subtypes, while ATG4D showed low expression, and TOMM20 was highly expressed in Macrophages and Tissue Stem Cells (Fig. 8d–f). The prominent expression of these genes in macrophages and tissue stem cells underscores their potential role in OS progression [19]. Additionally, we assessed the expression distribution of the MPRG signature in the six cell subtypes, and the results indicated that the MPRG signature was prominently expressed in all six cell subtypes, especially in Macrophages (Fig. 8g).
Single-cell sequencing analysis reveals the expression distribution characteristics of the MPRG prognostic signature. a Identification of OS cell clusters based on PCA analysis. b Expression levels of the MPRG signature in different cell clusters. c Annotation of cell subtypes based on SingleR. d–f UMAP dimensionality reduction analysis reveals the expression of the MPRG signature in different cell subtypes. g Expression distribution of the MPRG signature in different cell subtypes
Experimental Validation of the MPRG Prognostic Signature In Vitro
To validate the bioinformatics results, we performed in vitro experiments to assess the expression of mitochondria autophagy-related prognostic genes in OS cells. Compared to control cells, the OS cell line exhibited higher protein levels of ATG4D, RPS27A, and TOMM20 (Fig. 9a–d). The qRT-PCR results also confirmed higher mRNA levels of ATG4D, RPS27A, and TOMM20 in OS cells (Fig. 9e–g).
Abnormal expression of mitochondrial autophagy related prognostic genes in OS verification in vitro. a Western blot analysis of ATG4D, RPS27A and TOMM20 in OS cells and control cells. b–d The statistical results of western blot (n = 3). e–g qRT-PCR result of ATG4D, RPS27A and TOMM20 in OS cells and control cells
Discussion
Risk factor screening and therapeutic responsiveness evaluation of OS have become a major unresolved issue. In recent years, the establishment of OS risk assessment models has helped to analyze multiple factors affecting OS prognosis, and several preliminary verifies have been carried out [20]. Our study on MPRGs effectively stratified the prognostic risk of OS patients by ATG4D, RPS27A and TOMM20, aligning with existing research and further enriching the assessment content [21].
Our results provide new evidence for the important role of mitophagy in carcinogenesis and tumor progression [22]. Cancer cells are able to avoid programmed apoptosis that occurs after separation from neighboring cells or from the extracellular matrix [23]. There is evidence that mitophagy enhances the survival of cancer cells after separation from the original site by avoiding programmed apoptosis [23]. For example, in hepatocellular cancer cells, hypoxia can up-regulate the expression of mitophagy receptor BNIP3 and inhibit the mTOR/S6K1 pathway to promote mitophagy, thereby increasing the survival of hepatocellular cancer cells [24]. In OS cell, mitophagy promotion by BNIP3 can attenuate cisplatin resistance, showing drug sensitivity adjustment might be a key mechanism of how mitophagy affects tumor progression [18, 25]. In addition, the mTOR/AKT pathway and beta-catenin pathway have also been shown to be related to the mitophagy process in OS and influence tumor progression [26,27,28]. Therefore, further research on mitophagy is of clinical value.
The risk stratification associated with MPRG was related with resistance to chemotherapy drugs. There is evidence for the effect of mitophagy on drug sensitivity. Mitophagy has been observed in acute myeloid leukemia tumor cells treated with chemotherapy-targeted drugs such as FLT3-ITD and may be the cause of drug resistance [29]. In pancreatic cancer, the E3 ubiquitin ligase ARIH1 can promote cancer cell growth by increasing the level of mitophagy in tumor cells and increasing cancer resistance to chemotherapy drugs through an ubiquitination-dependent pathway [30]. It has also been reported that enhanced mitophagy can increase the effect of drugs. Chemotherapy drugs and tumor targeting drugs directly attack mitochondria or induce ROS production, resulting in the production of a large number of damaged mitochondria. The subsequent significant enhancement of mitophagy can lead to cell death [15]. Enhanced mitophagy after Ceramide also made tumor cells more sensitive to the treatment [31]. There is also evidence that the anti-tumor mechanism of rapamycin is also realized by inhibiting mTORC1 and thus promoting mitophagy [32]. Therefore, these evidences, together with our results, demonstrate the important value of targeting mitophagy as an intervention in cancer therapy. Our results suggested the impact of mitophagy on drug sensitivity, for example bleomycin sensitivity is associated with MPRG score. A regimen containing bleomycin is effective in patients with OS and can be used for preoperative chemotherapy in patients with OS [33, 34]. MPRG scores are promising as a basis for determining whether bleomycin should be used in patients with OS.
Our results showed that macrophages had the highest correlation coefficient with the established MPRG score and were associated with better OS prognosis. Macrophage genetic signatures in OS have been shown to predict prognosis and effectiveness of therapeutic response [35]. Tumor-associated macrophages (TAMs) play an important role in OS carcinogenesis progress. TAM can promote tumor growth by protecting cancer stem cells [36]. Moreover, TAM secretes immunosuppressive molecules to suppress the host immune response, thereby promoting tumor immune escape and promoting cancer cell metastasis [37]. In addition, matrix metalloproteinases released by TAM can activate the NF-kB signaling pathway to enhance epithelial–mesenchymal transformation (EMT) to promote OS metastasis [38]. Based on the above evidences, it has been redefined as a potential therapeutic strategy to treat patients with OS by reducing the proportion of TAM or M2-type macrophages to activate the anti-cancer effects of macrophages [19]. For example, All-trans retinoic acid (ATRA) can exert anti-tumor and anti-tumor metastasis activity by inhibiting the polarization of macrophages to M2, which demonstrates the importance of the immune landscape in the treatment of OS [39]. In OS-related in vitro experiments, ATRA prevented invasion and migration potency of OS cancer cells as well as stem cell properties [39, 40]. OS patients treated with ATRA also showed promising results [41]. Multiple OS-targeting strategies by interfering with macrophage polarization are being developed [19].
Together, our results provide evidence that mitophagy plays a role in the development of OS. At present, the research on mitophagy is not enough, and its clinical potential still needs to be further explored. This study provides insight for further in vitro/in vivo experiments of mitophagy in OS. Further validating the MPRG score in larger, independent cohorts, exploring the mechanistic role of identified genes in mitophagy, and testing the efficacy of mitophagy-targeting drugs in preclinical OS models would be of great significance.
Data Availability
The original contributions presented in the study are included in the article/supplementary material. Further inquiries can be directed to the corresponding authors.
References
Beird, H. C., Bielack, S. S., Flanagan, A. M., Gill, J., Heymann, D., Janeway, K. A., Livingston, J. A., Roberts, R. D., Strauss, S. J., & Gorlick, R. (2022). Osteosarcoma. Nature Reviews Disease Primers, 8, 77.
Ferrari, S., Smeland, S., Mercuri, M., Bertoni, F., Longhi, A., Ruggieri, P., Alvegard, T. A., Picci, P., Capanna, R., Bernini, G., et al. (2005). Neoadjuvant chemotherapy with high-dose Ifosfamide, high-dose methotrexate, cisplatin, and doxorubicin for patients with localized osteosarcoma of the extremity: A joint study by the Italian and Scandinavian Sarcoma Groups. The Journal of Clinical Oncology, 23, 8845–8852.
Fernandes, J.S., Blattmann, C., Hecker-Nolting, S., Kager, L., Kevric, M., Mettmann, V., Sorg, B., Fernandes, M. and Bielack, S.S. (2024) Beyond 5-year survival. A report from the Cooperative Osteosarcoma Study Group (COSS). Cancer Medicine, 13, e6893.
Chui, M. H., Kandel, R. A., Wong, M., Griffin, A. M., Bell, R. S., Blackstein, M. E., Wunder, J. S., & Dickson, B. C. (2016). Histopathologic features of prognostic significance in high-grade osteosarcoma. The Archives of Pathology & Laboratory Medicine, 140, 1231–1242.
Smith, A. G., & Macleod, K. F. (2019). Autophagy, cancer stem cells and drug resistance. Journal of Pathology, 247, 708–718.
Kroemer, G., Marino, G., & Levine, B. (2010). Autophagy and the integrated stress response. Molecular Cell, 40, 280–293.
Galluzzi, L., Baehrecke, E. H., Ballabio, A., Boya, P., Bravo-San Pedro, J. M., Cecconi, F., Choi, A. M., Chu, C. T., Codogno, P., Colombo, M. I., et al. (2017). Molecular definitions of autophagy and related processes. EMBO Journal, 36, 1811–1836.
Lu, Y., Li, Z., Zhang, S., Zhang, T., Liu, Y., & Zhang, L. (2023). Cellular mitophagy: Mechanism, roles in diseases and small molecule pharmacological regulation. Theranostics, 13, 736–766.
Pickles, S., Vigie, P., & Youle, R. J. (2018). Mitophagy and quality control mechanisms in mitochondrial maintenance. Current Biology, 28, R170–R185.
Palikaras, K., Lionaki, E., & Tavernarakis, N. (2018). Mechanisms of mitophagy in cellular homeostasis, physiology and pathology. Nature Cell Biology, 20, 1013–1022.
Igarashi, R., Yamashita, S. I., Yamashita, T., Inoue, K., Fukuda, T., Fukuchi, T., & Kanki, T. (2020). Gemcitabine induces Parkin-independent mitophagy through mitochondrial-resident E3 ligase MUL1-mediated stabilization of PINK1. Scientific Report, 10, 1465.
Gatica, D., Lahiri, V., & Klionsky, D. J. (2018). Cargo recognition and degradation by selective autophagy. Nature Cell Biology, 20, 233–242.
Soengas, M. S. (2012). Mitophagy or how to control the Jekyll and Hyde embedded in mitochondrial metabolism: Implications for melanoma progression and drug resistance. Pigment Cell Melanoma Research, 25, 721–731.
Wang, H., Luo, W., Chen, H., Cai, Z., & Xu, G. (2024). Mitochondrial dynamics and mitochondrial autophagy: Molecular structure, orchestrating mechanism and related disorders. Mitochondrion, 75, 101847.
Mazure, N. M., Brahimi-Horn, M. C., & Pouyssegur, J. (2011). Hypoxic mitochondria: Accomplices in resistance. Bulletin du Cancer, 98, 40–46.
Vara-Perez, M., Felipe-Abrio, B. and Agostinis, P. (2019) Mitophagy in cancer: A tale of adaptation. Cells, 8.
Si, Z., Shen, Z., Luan, F., & Yan, J. (2023). PINK1 regulates apoptosis of osteosarcoma as the target gene of cisplatin. Journal of Orthopaedic Surgery and Research, 18, 132.
Pan, B., Li, Y., Han, H., Zhang, L., Hu, X., Pan, Y., & Peng, Z. (2024). FoxG1/BNIP3 axis promotes mitophagy and blunts cisplatin resistance in osteosarcoma. Cancer Science, 115, 2565–2577.
Anand, N., Peh, K.H. and Kolesar, J.M. (2023). Macrophage repolarization as a therapeutic strategy for osteosarcoma. The International Journal of Molecular Sciences, 24.
Yao, Y., Wang, D., Zheng, L., Zhao, J., & Tan, M. (2024). Advances in prognostic models for osteosarcoma risk. Heliyon, 10, e28493.
Zeng, Z., Zhou, X., Wang, Y., Cao, H., Guo, J., Wang, P., Yang, Y., & Wang, Y. (2022) Mitophagy-a new target of bone disease. Biomolecules, 12.
Panigrahi, D. P., Praharaj, P. P., Bhol, C. S., Mahapatra, K. K., Patra, S., Behera, B. P., Mishra, S. R., & Bhutia, S. K. (2020). The emerging, multifaceted role of mitophagy in cancer and cancer therapeutics. Seminars in Cancer Biology, 66, 45–58.
Nguyen, T. T., Wei, S., Nguyen, T. H., Jo, Y., Zhang, Y., Park, W., Gariani, K., Oh, C. M., Kim, H. H., Ha, K. T., et al. (2023). Mitochondria-associated programmed cell death as a therapeutic target for age-related disease. Experimental & Molecular Medicine, 55, 1595–1619.
Sun, L., Li, T., Wei, Q., Zhang, Y., Jia, X., Wan, Z., & Han, L. (2014). Upregulation of BNIP3 mediated by ERK/HIF-1alpha pathway induces autophagy and contributes to anoikis resistance of hepatocellular carcinoma cells. Future Oncology, 10, 1387–1398.
Vianello, C., Cocetta, V., Catanzaro, D., Dorn, G. W., 2nd., De Milito, A., Rizzolio, F., Canzonieri, V., Cecchin, E., Roncato, R., Toffoli, G., et al. (2022). Cisplatin resistance can be curtailed by blunting Bnip3-mediated mitochondrial autophagy. Cell Death and Disease, 13, 398.
Shen, W., Zeng, X., Zeng, X., Hu, B., Ren, C., Lin, Z., Zhang, L., Rui, G., Yasen, M., & Chen, X. (2024). Trifluoperazine activates AMPK/mTOR/ULK1 signaling pathway to induce mitophagy in osteosarcoma cells. Chemico-Biological Interactions, 392, 110904.
Zheng, Z., Zhao, X., Yuan, B., Jiang, S., Yan, R., Dong, X., Yao, Q., & Liang, H. (2024). Soy isoflavones induces mitophagy to inhibit the progression of osteosarcoma by blocking the AKT/mTOR signaling pathway. Molecular Medicine, 30, 5.
He, G., Nie, J. J., Liu, X., Ding, Z., Luo, P., Liu, Y., Zhang, B. W., Wang, R., Liu, X., Hai, Y., et al. (2023). Zinc oxide nanoparticles inhibit osteosarcoma metastasis by downregulating beta-catenin via HIF-1alpha/BNIP3/LC3B-mediated mitophagy pathway. Bioactive Materials, 19, 690–702.
Dany, M., Gencer, S., Nganga, R., Thomas, R. J., Oleinik, N., Baron, K. D., Szulc, Z. M., Ruvolo, P., Kornblau, S., Andreeff, M., et al. (2016). Targeting FLT3-ITD signaling mediates ceramide-dependent mitophagy and attenuates drug resistance in AML. Blood, 128, 1944–1958.
Villa, E., Proics, E., Rubio-Patino, C., Obba, S., Zunino, B., Bossowski, J. P., Rozier, R. M., Chiche, J., Mondragon, L., Riley, J. S., et al. (2017). Parkin-independent mitophagy controls chemotherapeutic response in cancer cells. Cell Reports, 20, 2846–2859.
Dorn, G. W. (2010). Mitochondria*l pruning by Nix and BNip3: An essential function for cardiac-expressed death factors. Journal of Cardiovascular Translational Research, 3, 374–383.
Zou, Z., Tao, T., Li, H., & Zhu, X. (2020). mTOR signaling pathway and mTOR inhibitors in cancer: Progress and challenges. Cell & Bioscience, 10, 31.
Meyers, P. A., Heller, G., Healey, J., Huvos, A., Lane, J., Marcove, R., Applewhite, A., Vlamis, V., & Rosen, G. (1992). Chemotherapy for nonmetastatic osteogenic sarcoma: The Memorial Sloan-Kettering experience. Journal of Clinical Oncology, 10, 5–15.
Goorin, A. M., Schwartzentruber, D. J., Devidas, M., Gebhardt, M. C., Ayala, A. G., Harris, M. B., Helman, L. J., Grier, H. E., Link, M. P., & Pediatric Oncology, G. (2003). Presurgical chemotherapy compared with immediate surgery and adjuvant chemotherapy for nonmetastatic osteosarcoma: Pediatric Oncology Group Study POG-8651. Journal of Clinical Oncology, 21, 1574–1580.
Mao, X., Song, F., Jin, J., Zou, B., Dai, P., Sun, M., Xu, W., Wang, L., & Kang, Y. (2023). Prognostic and immunological significance of an M1 macrophage-related gene signature in osteosarcoma. Front Immunology, 14, 1202725.
Yuan, R., & Li, J. (2024). Role of macrophages and their exosomes in orthopedic diseases. PeerJ, 12, e17146.
Terabe, M., Matsui, S., Park, J. M., Mamura, M., Noben-Trauth, N., Donaldson, D. D., Chen, W., Wahl, S. M., Ledbetter, S., Pratt, B., et al. (2003). Transforming growth factor-beta production and myeloid cells are an effector mechanism through which CD1d-restricted T cells block cytotoxic T lymphocyte-mediated tumor immunosurveillance: Abrogation prevents tumor recurrence. Journal of Experimental Medicine, 198, 1741–1752.
Lee, C. H., Liu, S. Y., Chou, K. C., Yeh, C. T., Shiah, S. G., Huang, R. Y., Cheng, J. C., Yen, C. Y., & Shieh, Y. S. (2014). Tumor-associated macrophages promote oral cancer progression through activation of the Axl signaling pathway. Annals of Surgical Oncology, 21, 1031–1037.
Zhou, Q., Xian, M., Xiang, S., Xiang, D., Shao, X., Wang, J., Cao, J., Yang, X., Yang, B., Ying, M., et al. (2017). All-trans retinoic acid prevents osteosarcoma metastasis by inhibiting M2 polarization of tumor-associated macrophages. Cancer Immunology Research, 5, 547–559.
Shao, X. J., Xiang, S. F., Chen, Y. Q., Zhang, N., Cao, J., Zhu, H., Yang, B., Zhou, Q., Ying, M. D., & He, Q. J. (2019). Inhibition of M2-like macrophages by all-trans retinoic acid prevents cancer initiation and stemness in osteosarcoma cells. Acta Pharmacologica Sinica, 40, 1343–1350.
Todesco, A., Carli, M., Iacona, I., Frascella, E., Ninfo, V., & Rosolen, A. (2000). All-trans retinoic acid and interferon-alpha in the treatment of a patient with resistant metastatic osteosarcoma. Cancer, 89, 2661–2666.
Acknowledgements
Not Applicable.
Funding
This study was supported by Youth Research Fund of Affiliated Hospital of Qingdao University in 2022 (QDFYQN202201012).
Author information
Authors and Affiliations
Contributions
S.R and Z.W conceived and designed the study. R.P performed the expression verification in cell lines. All authors participated in the acquisition, analysis, interpretation of data, drafting of the manuscript, and approval of the submitted version.
Corresponding author
Ethics declarations
Conflict of interests
The authors declare no conflicts of interest related to this study.
Ethical Approval
Not Applicable.
Consent for Publication
All listed authors have actively participated in the study and have read and approved the submitted manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary Table1. The gene symbol of mitophagy signature.
Supplementary file1 (XLSX 9 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ren, S., Pan, R. & Wang, Z. Multi-omics and Single Cell Sequencing Analyses Reveal Associations of Mitophagy-Related Genes Predicting Clinical Prognosis and Immune Infiltration Characteristics in Osteosarcoma. Mol Biotechnol (2024). https://doi.org/10.1007/s12033-024-01280-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12033-024-01280-w