Abstract
Since the discovery of microRNAs (miRNAs) as a class of important regulatory molecules, miRNAs are involved in the occurrence and development of tumors. In this paper, we aimed to identify the role of miR-1274a in non-small cell lung cancer (NSCLC). The miR-1274a expression levels in four NSCLC cells and tissues from 125 patients were determined by qRT-PCR assays. Kaplan–Meier survival curves and Cox regression analysis were used to examine the prognostic significance of miR-1274a in NSCLC patients. The CCK-8 and Transwell assays were performed to evaluate the cell proliferation, invasion, and migration ability of NSCLC cells. The miR-1274a expression levels were significantly higher in NSCLC tissues than in adjacent normal tissues, and overexpression of miR-1274a had a poor prognosis in NSCLC patients. Functional studies in two NSCLC cell lines have shown that overexpression of miR-1274a could promote cell proliferation, migration, and invasion. miR-1274a expression levels are upregulated in NSCLC tissues, and a high expression is associated with a poor prognosis in patients with NSCLC. Moreover, miR-1274a promotes cell proliferation, migration, and invasion. Based on our findings, miR-1274a may act as a tumor miRNA in the occurrence and development of NSCLC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lung cancer has become the most common type of cancer in China, ranking first in cancer incidence and mortality for many years. The mortality of lung cancer is much higher than that of other malignant tumors worldwide [1]. Due to the lack of specific biomarkers for early screening and diagnosis, most lung cancer patients are in an advanced stage. Many scientists are committed to the application of imaging and sputum cytology for early screening of lung cancer at present, but the situation has not improved and the five-year overall survival rate of lung cancer is still less than 15% [2, 3]. Lung cancer is divided into small cell lung cancer (SCLC) and non-small cell lung cancer (NSCLC), among which NSCLC accounts for 80%–85% of the total lung cancer. The research in molecular and cellular biology allowed the development of specific drugs against some molecular targets in the process of oncology resulting in targeted therapies, which is the second fundamental breakthrough in the field of medical oncology [4]. At present, targeted therapies and immunotherapies have proved to be of great clinical significance to many patients with different types of cancers, including patients with NSCLC [5, 6]. Therefore, effective prognostic markers and combination therapies have become the focus of tumor research [7].
MicroRNAs (miRNAs) are a series of single-stranded RNA molecules with 20–25 nucleotides in length that are evolutionarily conserved and belong to endogenous non-coding RNA [8, 9]. miRNAs inhibit the process of protein translation or promote the degradation of mRNA through, respectively, the incomplete or complete complementary pairing with target gene mRNA [10]. It is now generally believed that the abnormal expression of miRNAs is closely related to the occurrence, development, diagnosis, treatment, and prognosis of tumors. At present, many studies have shown the relationship between abnormal expression of miRNAs and lung cancer, such as miR-155 [11], miR-203 [2], miR-451a [12], and miR-7-5p [13]. The significant changes in miRNAs expression in normal lung tissues and lung cancer tissues make miRNAs a promising new biomarker for lung cancer. This not only is helpful for the early diagnosis of lung cancer but also may be used as a new drug and target for the treatment of lung cancer and will certainly open up a new horizon for us to deeply understand the occurrence and development of lung cancer at the molecular level [14, 15]. A previous study by Lu C used the method of cluster analysis to prove that there are many miRNAs expression differences between NSCLC tissues and adjacent normal tissues, including miR-1274a [16]. The above study focused on the important role of miR-92a in NSCLC, while the role of abnormally expressed miR-1274a in lung cancer remains unclear.
In this paper, the expression levels of miR-1274a in NSCLC cells and tissues were firstly assessed and then the role of miR-1274a in the proliferation, migration, and invasion of NSCLC was explored. This study aimed to investigate the feasibility of miR-1274a as an effective target and prognostic factor for the treatment of NSCLC.
Materials and Methods
Patients and Tissue Samples
A total of 125 pairs of NSCLC tissues and para-cancerous tissue samples were collected from NSCLC patients (range from 28 to 72 with mean ± SD of 49.655 ± 10.087) undergoing surgical excision from March 2012 to March 2015 at the affiliated Tumor Hospital of Harbin Medical University. The obtained specimens were immediately placed in liquid nitrogen and stored at − 80 °C until use. The experiment and all specimens were treated anonymously. The sample inclusion criteria were as follows: (1) NSCLC was confirmed by pathological examination after tumor resection; (2) There was no radiotherapy, chemotherapy, or other immunotherapy before operation; (3) There was no distant tumor metastasis or other diseases that might seriously affect the survival time. A 5-year follow-up was conducted to estimate the prognostic outcomes of the patients.
Cell Culture and Transfection
Four NSCLC cell lines (A549, H1650, 95D, and H1299) and a normal human bronchial epithelial cell line (16HBE) were purchased from the Institute of Cell Research, Chinese Academy of Sciences (Shanghai, China). Cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM; Thermo Fisher Scientific, Waltham, USA) supplemented with 10% FBS (Thermo Fisher Scientific, Waltham, USA) in humidified air at 37 °C with 5% CO2.
A549 and H1299 cells (2 × 105) were selected for cell transfection because both cells exhibited relatively higher miR-1274a expression levels among the four cell lines. Both cells were cultured overnight in a 6-well plate for transfection. Then A549 and H1299 cells were transfected with miR-1274a mimic, mimic negative control (mimic NC), miR-1274a inhibitor, or inhibitor NC using the Lipofectamine 3000 transfection reagent (Thermo Fisher Scientific, Waltham, MA, USA). The mimics, inhibitors, and NCs for transfection were purchased from GenePharma (Shanghai, China). Non-transfected cells served as a blank control group.
Primer sequences are as follows: 5′-GUCCCUGUUCAGGCGCCA-3′ (miR-1274a mimic), 5′-UGGCGCCUGAACAGGGAC-3′ (miR-1274a inhibitor), 5′-UCACAACCUCCUAGAAAGAGUAGA-3′ (mimic NC), and 5′-ACGUGACACGUUCGGAGAATT-3′ (inhibitor NC).
RNA Extraction and Quantitative Real-Time PCR
Total RNA from NSCLC tissues or cells was isolated using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s protocol, and the RNA quality and quantity were verified with a NanoDrop 2000 (Thermo Fisher Scientific). Reverse transcription was carried out to obtain cDNA using the TaqMan miRNA reverse transcription kit (Thermo Fisher Scientific). Real-time PCR was performed using a TaqMan miRNA quantitative PCR kit (Thermo Fisher Scientific) on an ABI7500 real-time PCR detection system. The thermocycling conditions for PCR consisted of an initial denaturation at 95 °C for 3 min, followed by 35 cycles of amplification (94 °C for 30 s, 55 °C for 30 s, 72 °C for 30 s), and a final extension at 72 °C for 10 min. The forward primer sequence of miR-1274a was 5′-TCGGCAGGGTCCCTGTTCAGG-3′. The reverse primer sequence was 5′-CTCAACTGGTGTCGTGGA-3′. U6 was used as an internal reference for miRNA quantification, forward primer: 5′-CTCGCTTCGGCAGCACA-3′, and reverse primer: 5′-AACGCTTCACGAATTTGCGT-3′. The relative expression levels of miR-1274a were calculated using the 2−ΔΔCT method.
Cell Proliferation Assay
Cell proliferation was determined by Cell Counting Kit-8 (CCK-8; Dojindo, Kumamoto, Japan) assay. Briefly, after transfection, 5 × 103 cells were plated in 96-well plates, 10 μL of CCK-8 solution was added to each well, and the cells were incubated for another 2 h at 37 °C in 5% CO2. The absorbance (OD) of each well at 450 nm was detected, and the experiment was repeated 3 times.
Cell Migration and Invasion Assays
Cell migration and invasion were assessed by Transwell assay using 24-well Transwell plates (8 μm; BD Biosciences, San Jose, CA). For the migration assay, A549 and H1299 cells were suspended in serum-free DMEM medium and the cells (1 × 105 cells/well) were placed in the upper chamber and were incubated at 37 °C. The invasion assay was similar to that of the migration assay except that the upper chambers were first covered with 1 mg/mL Matrigel. A normal cell culture medium containing 10% FBS was added to the lower layer of the chamber. Following a 24 h of incubation, the migrating and invading cells were fixed and stained with 0.1% crystal violet. Five random fields were randomly selected for counting and statistical analysis in each well.
Statistical Analysis
Statistical analyses were carried out using Statistical Product and Service Solutions (SPSS) 19.0 software (IBM, Armonk, NY, USA) and GraphPad 5.0 (GraphPad Software, Inc., La Jolla, CA, USA). Data are expressed as the mean ± standard deviation (SD). The χ2 test was carried out to analyze the relationship between miR-1274a expression and the clinical characteristics of the patients. A two-tailed paired Student’s t test was used to compare the means between two sets, and a one-way analysis of variance (ANOVA) followed by Tukey’s post hoc test was used to compare the means among three groups. Patient overall survival curves were estimated by the Kaplan–Meier method. The joint effect of the covariables was determined by Cox regression. The significance level was set at P < 0.05.
Results
Expression of miR-1274a in NSCLC Tissues and Cell Lines
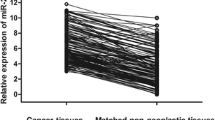
In the present study, the expression levels of miR-1274a in NSCLC tissues and para-carcinoma tissues were assessed by qRT-PCR. As shown in Fig. 1A, the results showed that the expression levels of miR-1274a were significantly higher in NSCLC tissues than that in adjacent tissues (P < 0.001). Moreover, the expression of miR-1274a in four NSCLC cell lines (A549, H1650, 95D, H1299) was also higher than that in the normal cell line (16HBE, P < 0.001, Fig. 1B). Among the four NSCLC cell lines, A549 and H1299 showed higher miR-1274a expression levels.
Correlation Between miR-1274a Expression and Clinical Factors in NSCLC Patients
To assess the relationship between clinicopathological characteristics and miR-1274a expression in patients with NSCLC, 125 patients with NSCLC were divided into a high expression group (n = 63) and a low expression group (n = 62) according to the median expression of miR-1274a in tissues. As shown in Table 1, the expression of miR-1274a in NSCLC tissues was related to TNM staging (P = 0.001) and lymph node metastasis (P = 0.001), While the expression of miR-1274a is not statistically significant with age, gender, tumor size, or tumor number (P > 0.05).
Upregulation of miR-1274a is Associated with Poor Prognosis in NSCLC Patients
We further evaluated whether miR-1274a expression was associated with the prognosis of patients with NSCLC. Using the Kaplan–Meier method and log-rank test, we found that the overall survival of patients in the high expression group was higher than that of the low expression group (P = 0.028, Fig. 2). In addition, multivariate Cox regression analysis assessing the abovementioned significant parameters revealed that miR-1274a (HR = 2.803, P = 0.001), TNM stage (HR = 0.515, P = 0.032), and lymph node metastasis (HR = 0.488, P = 0.024) were independent prognostic factors for overall survival (shown in Table 2).
Upregulation of miR-1274a Promotes NSCLC Cell Proliferation, Migration, and Invasion
Additionally, the biological role of miR-1274a in NSCLC cell lines was detected. The miR-1274a mimic, mimic NC, miR-1274a inhibitor, or inhibitor NC were transfected into A549 and H1299 cells. As shown in Fig. 3A, the expression levels of miR-1274a in miR-1274a mimic transfected cells were significantly higher than that in NC-transfected cells, while the miR-1274a inhibitor downregulated miR-1274a expression, compared with control (P < 0.001). CCK-8 assay showed that cell proliferation was significantly increased after miR-1274a mimics transfection, compared with the control group, while downregulation of miR-1274a decreased cell growth (P < 0.05, Fig. 3B).
Effects of miR-1274a expression levels on proliferation in A549 and H1299 cells. A The expression level of miR-1274a was studied by qRT-PCR after transient transfection with miR-1274a mimic/inhibitor (or mimic/inhibitor NC). B The CCK-8 assay was performed to examine cell proliferation. *P < 0.05, ***P < 0.001
Transwell invasion and migration assays were performed to detect the migration and invasion of A549 and H1299 cells. The results showed that upregulation of miR-1274a promoted the invasion and migration ability of NSCLC cells compared with the blank control group and NC group (P < 0.01, Fig. 4A, B).
Discussion
With increasing morbidity and mortality, lung cancer has become the leading cause of death caused by tumors worldwide [17]. Although the occurrence of molecular networks in lung cancer at the gene and protein levels has been partially clarified, personalized treatment based on genetic changes in the past few years has also made certain progress, but the high lung cancer case fatality rate over five years has not improved [18,19,20].
miRNAs can act as tumor suppressors and carcinogenic promoters to regulate cell proliferation, apoptosis, invasion, metastasis, and angiogenesis [21, 22]. The same miRNA can affect multiple protein-coding genes, while the same gene can be affected by multiple miRNAs. Understanding the role of miRNAs in the development of tumors and their use as tumor diagnostic markers, prognostic markers, and therapeutic targets is still in the early stages [23, 24]. In studies of lung cancer and miRNAs, it was found that the expression pattern of miRNAs in the lung changed from fetal to adult and from normal to lung cancer, which revealed specific miRNAs may play multiple functions in the formation of the lung, while the abnormal expression of miRNAs may also induce the occurrence of lung cancer [25, 26].
miR-1274a exerts regulatory functions in a number of human cancers. In the present study, the clinical significance and biological function of miR-1274a in lung cancer were investigated. A high expression of miR-1274a tended to be associated with positive lymph node metastasis and advanced TNM stage of NSCLC patients. In addition, Kaplan–Meier and multivariate Cox analyses results showed that patients with a high expression of miR-1274a might have a poor prognosis. These results suggest that the high expression of miR-1274 may be closely related to the development and progression of NSCLC. The above results were in line with previous studies in which the significance of miR-1274a overexpression in prostate cancer and the role of miR-1274a combined with PSA as a potential diagnostic biomarker in prostate cancer were studied [27]. In another study, upregulation of miR-1274a was also observed in clinical specimens and cell lines in clear cell renal cell carcinoma [28].
Then, we investigated the biological function of miR-1274a in NSCLC. Cell transfection experiments were conducted with miR-1274a using A549 and H1299 cells. The results showed that overexpression of miR-1274a could significantly promote cell proliferation, migration, and invasion, while the low expression of miR-1274a inhibits these cellular behaviors of NSCLC, compared with untreated cells. These results indicate that miR-1274a may play an oncogenic role in the pathological progression of NSCLC. Actually, the role of miR-1274a as an oncogene in tumorigenesis has recently been described in various types of cancer, such as colon cancer, and increased expression of miR-1274a promoted cell growth, migration, and invasion, while inhibition of miR-1274a suppressed these cellular activities by targeting forkhead box protein O4 (FOXO4) [29]. In a study in the tumorigenesis of gastric cancer, miR-1274a was confirmed to accelerate the proliferation and migration of gastric cancer cells by targeting FOXO4 directly [30]. miR-1274a was found to negatively regulate several target genes, such as bone morphogenetic protein receptor type 1B (BMPR1B) [28], a disintegrin and metalloproteinase 9 (ADAM9) [31, 32], and FOXQ4 [30]. When we studied the important role of miR-1274a in other cancers, we were surprised to find that the target gene of miR-1274a was FOXO4 when it was abnormally overexpressed and the corresponding FOXO4 showed low expression in tumors. The FOXO protein family mainly includes FOXO1, FOXO3a, FOXO4, and FOXO6. FOXO protein family is a key regulator of a variety of cellular processes, particularly in cancer and disease [33]. FOXO4 has been suggested to be a tumor metastasis suppressor [34]. In NSCLC, FOXO4 is involved in miR-150-induced metastasis of NSCLC cells, with downregulated FOXO4 expression promoting metastasis of NSCLC cells [35]. Here, miR-1274a can promote the proliferation, migration, and invasion of NSCLC in vitro. Therefore, we have reason to suspect that the target gene of miR-1274a for NSCLC is FOXO4.
This study has some limitations. For example, the limited sample size may lead to the deviation of the results, and miR-1274a was only extracted from the tissue sample, but not from the liquid. Liquid biopsy is a new analytical approach based on the analysis of a peripheral blood sample for the identification of tumor-specific alterations directly in the bloodstream [36]. We will continue to collect sample sizes and extract miR-1274a from liquid to enrich the study in future work.
In summary, this study showed that miR-1274a was significantly upregulated in NSCLC cell lines and tissues, and overexpression of miR-1274a was related to the poor prognosis and shorter survival time for NSCLC patients. In addition, miR-1274a was associated with cell proliferation, migration, and invasion of NSCLC. These data suggest that miR-1274a may serve as a prognostic molecular marker and may provide a novel therapeutic strategy for combating NSCLC.
Data Availability
All data generated or analyzed during this study are included in this published article.
References
Herbst, R. S., Morgensztern, D., & Boshoff, C. (2018). The biology and management of non-small cell lung cancer. Nature, 553(7689), 446–454.
Ruan, L., Chen, J., Ruan, L., Yang, T., & Wang, P. (2018). MicroRNA-186 suppresses lung cancer progression by targeting SIRT6. Cancer Biomarkers: Section A of Disease Markers, 21(2), 415–423.
Sun, Y., Li, L., Xing, S., Pan, Y., Shi, Y., Zhang, L., et al. (2017). miR-503-3p induces apoptosis of lung cancer cells by regulating p21 and CDK4 expression. Cancer Biomarkers: Section A of Disease Markers, 20(4), 597–608.
Falzone, L., Salomone, S., & Libra, M. (2018). Evolution of cancer pharmacological treatments at the turn of the third millennium. Frontiers in Pharmacology, 9, 1300.
Yuan, M., Huang, L. L., Chen, J. H., Wu, J., & Xu, Q. (2019). The emerging treatment landscape of targeted therapy in non-small-cell lung cancer. Signal Transduction and Targeted Therapy, 4, 61.
Christofi, T., Baritaki, S., Falzone, L., Libra, M., & Zaravinos, A. (2019). Current perspectives in cancer immunotherapy. Cancers, 11(10), 1472.
Yang, Z. Q., Wu, C. A., & Cheng, Y. X. (2018). Prognostic value of microRNA-133a expression and its clinicopathologic significance in non-small cell lung cancer: a comprehensive study based on meta-analysis and the TCGA database. Oncology Research and Treatment, 41(12), 762–768.
Bulgakova, O., Zhabayeva, D., Kussainova, A., Pulliero, A., Izzotti, A., & Bersimbaev, R. (2018). miR-19 in blood plasma reflects lung cancer occurrence but is not specifically associated with radon exposure. Oncology Letters, 15(6), 8816–8824.
Li, H., Feng, C., & Shi, S. (2018). miR-196b promotes lung cancer cell migration and invasion through the targeting of GATA6. Oncology Letters, 16(1), 247–252.
Chu, G. C. W., Lazare, K., & Sullivan, F. (2018). Serum and blood based biomarkers for lung cancer screening: A systematic review. BMC Cancer, 18(1), 181.
Shao, C., Yang, F., Qin, Z., Jing, X., Shu, Y., & Shen, H. (2019). The value of miR-155 as a biomarker for the diagnosis and prognosis of lung cancer: A systematic review with meta-analysis. BMC Cancer, 19(1), 1103.
Shen, Y. Y., Cui, J. Y., Yuan, J., & Wang, X. (2018). MiR-451a suppressed cell migration and invasion in non-small cell lung cancer through targeting ATF2. European Review for Medical and Pharmacological Sciences, 22(17), 5554–5561.
Xiao, H. (2019). MiR-7-5p suppresses tumor metastasis of non-small cell lung cancer by targeting NOVA2. Cellular & Molecular Biology Letters, 24, 60.
Zhang, L., & Yu, S. (2018). Role of miR-520b in non-small cell lung cancer. Experimental and Therapeutic Medicine, 16(5), 3987–3995.
Wang, Q., & Zhang, L. (2019). Possible molecular mechanisms for the roles of MicroRNA-21 played in lung cancer. Technology in Cancer Research & Treatment, 18, 1533033819875130.
Lu, C., Shan, Z., Hong, J., & Yang, L. (2017). MicroRNA-92a promotes epithelial-mesenchymal transition through activation of PTEN/PI3K/AKT signaling pathway in non-small cell lung cancer metastasis. International Journal of Oncology, 51(1), 235–244.
Munagala, R., Aqil, F., & Gupta, R. C. (2016). Exosomal miRNAs as biomarkers of recurrent lung cancer. Tumour Biology: The Journal of the International Society for Oncodevelopmental Biology and Medicine., 37(8), 10703–10714.
Hashemi, Z. S., Khalili, S., Forouzandeh Moghadam, M., & Sadroddiny, E. (2017). Lung cancer and miRNAs: A possible remedy for anti-metastatic, therapeutic and diagnostic applications. Expert Review of Respiratory Medicine, 11(2), 147–157.
Mao, Y., Yang, D., He, J., & Krasna, M. J. (2016). Epidemiology of lung cancer. Surgical Oncology Clinics of North America, 25(3), 439–445.
Rodriguez-Canales, J., Parra-Cuentas, E., & Wistuba, I. I. (2016). Diagnosis and molecular classification of lung cancer. Cancer Treatment and Research, 170, 25–46.
Sheervalilou, R., Lotfi, H., Shirvaliloo, M., Sharifi, A., Nazemiyeh, M., & Zarghami, N. (2019). Circulating MiR-10b, MiR-1 and MiR-30a expression profiles in lung cancer: possible correlation with clinico-pathologic characteristics and lung cancer detection. International Journal of Molecular and Cellular Medicine, 8(2), 118–129.
Zhang, H., Wang, R., & Wang, M. (2019). miR-331-3p suppresses cell invasion and migration in colorectal carcinoma by directly targeting NRP2. Oncology Letters, 18(6), 6501–6508.
Alizadeh, M., Safarzadeh, A., Beyranvand, F., Ahmadpour, F., Hajiasgharzadeh, K., Baghbanzadeh, A., et al. (2019). The potential role of miR-29 in health and cancer diagnosis, prognosis, and therapy. Journal of Cellular Physiology, 234(11), 19280–19297.
Zhu, L., Deng, H., Hu, J., Huang, S., Xiong, J., & Deng, J. (2018). The promising role of miR-296 in human cancer. Pathology, Research and Practice, 214(12), 1915–1922.
Zhang, X., Li, Y., Qi, P., & Ma, Z. (2018). Biology of MiR-17-92 cluster and its progress in lung cancer. International Journal of Medical Sciences, 15(13), 1443–1448.
Zhou, Y., & Shen, S. (2019). MiR-520f acts as a biomarker for the diagnosis of lung cancer. Medicine, 98(30), e16546.
Said, R., Garcia-Mayea, Y., Trabelsi, N., Setti Boubaker, N., Mir, C., Blel, A., et al. (2018). Expression patterns and bioinformatic analysis of miR-1260a and miR-1274a in Prostate Cancer Tunisian patients. Molecular Biology Reports, 45(6), 2345–2358.
Yoshino, H., Yonezawa, T., Yonemori, M., Miyamoto, K., Sakaguchi, T., Sugita, S., et al. (2018). Downregulation of microRNA-1274a induces cell apoptosis through regulation of BMPR1B in clear cell renal cell carcinoma. Oncology Reports, 39(1), 173–181.
Ren, B., Yang, B., Li, P., & Ge, L. (2020). Upregulation of MiR-1274a is correlated with survival outcomes and promotes cell proliferation, migration, and invasion of colon cancer. OncoTargets and Therapy, 13, 6957–6966.
Wang, G. J., Liu, G. H., Ye, Y. W., Fu, Y., & Zhang, X. F. (2015). The role of microRNA-1274a in the tumorigenesis of gastric cancer: Accelerating cancer cell proliferation and migration via directly targeting FOXO4. Biochemical and Biophysical Research Communications, 459(4), 629–635.
Feng, X. T., Wang, C., Zhang, F. J., Wu, X. Q., & Zhang, Z. (2021). MicroRNA-1274a serves as a prognostic biomarker in patients with osteosarcoma and is involved in tumor progression via targeting ADAM9. Journal of Biological Regulators and Homeostatic Agents, 35(1), 151–160.
Zhou, C., Liu, J., Li, Y., Liu, L., Zhang, X., Ma, C. Y., et al. (2011). microRNA-1274a, a modulator of sorafenib induced a disintegrin and metalloproteinase 9 (ADAM9) down-regulation in hepatocellular carcinoma. FEBS Letters, 585(12), 1828–1834.
Farhan, M., Wang, H., Gaur, U., Little, P. J., Xu, J., & Zheng, W. (2017). FOXO signaling pathways as therapeutic targets in cancer. International Journal of Biological Sciences, 13(7), 815–827.
Coomans de Brachène, A., & Demoulin, J. B. (2016). FOXO transcription factors in cancer development and therapy. Cellular and Molecular Life Sciences (CMLS), 73(6), 1159–1172.
Li, H., Ouyang, R., Wang, Z., Zhou, W., Chen, H., Jiang, Y., et al. (2016). MiR-150 promotes cellular metastasis in non-small cell lung cancer by targeting FOXO4. Scientific Reports, 6, 39001.
Tuaeva, N. O., Falzone, L., Porozov, Y. B., Nosyrev, A. E., Trukhan, V. M., Kovatsi, L., et al. (2019). Translational application of circulating DNA in oncology: Review of the last decades achievements. Cells, 8(10), 1251.
Funding
This work was supported by scientific research fund of Petrel (Grant No. JJQN2016-11]; the financial assistance of Heilongjiang Postdoctoral Fund (Grant No. LBH-Z16227).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical Approval
This study was approved by the hospital ethics committee of the affiliated Tumor Hospital of Harbin Medical University.
Consent to Participate
All patients signed a written informed consent form.
Consent to Publish
Patients signed informed consent regarding publishing their data.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhu, Sj., Wang, X., Hu, Sl. et al. Clinical Significance and Biological Function of miR-1274a in Non-small Cell Lung Cancer. Mol Biotechnol 64, 9–16 (2022). https://doi.org/10.1007/s12033-021-00385-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-021-00385-w