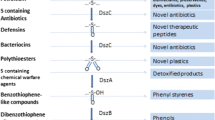
Abstract
Metabolic pathways of aerobic bacteria able to assimilate sulfur can provide biocatalysts for biodesulfurization of petroleum and of other sulfur-containing pollutants. Of major interest is the so-called “4S pathway,” in that C–S bonds are specifically cleaved leaving the carbon skeleton of substrates intact. This pathway is carried out by four enzymes, named Dsz A, B, C, and D. In view of a possible application of recombinant Dsz enzymes in biodesulfurization treatments, we have investigated the structural features of enzymes cloned from a Rhodococcus strain isolated from polluted environmental samples and their resistance to temperature (20–95 °C) and to organic solvents (5, 10, and 20 % v/v methanol, acetonitrile, hexane, and toluene). Changes in protein structures were assessed by circular dichroism and intrinsic fluorescence spectroscopy. We found that all Dsz proteins are unfolded by temperatures in the range 45–60 °C and by all solvents tested, with the most dramatic effect being produced by toluene. These results suggest that stabilization of the biocatalysts by protein engineering will be necessary for developing biodesulfurization technologies based on Dsz enzymes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biotechnology may take advantage of the metabolic abilities of living beings that offer an astonishing repertoire of enzymatic catalysts. Enzymes can be applied in reactions of commercial and/or environmental relevance, either directly or upon improvement by protein engineering techniques. In this context, enzymes evolved by bacterial strains able to assimilate sulfur are of interest for application in the transformation of sulfur-containing chemical compounds, environmental decontamination, and fossil fuels desulfurization [1]. Major organosulfur contaminants of oils are aromatic compounds, such as benzothiophene, dibenzothiophene (DBT), and their alkylated derivates. Desulfurization of crude and refined petroleum products is conventionally carried out by high-pressure and high-temperature hydrodesulfurization (HDS) [2, 3]. Increasingly strict environmental regulations about sulfur emission from fossil fuels and the fact that some organosulfur compounds are recalcitrant to HDS are driving the development of alternate technologies, in particular microbial (or bio) desulfurization (BDS). Sulfur-metabolizing bacteria such as Rhodococcus, Nocardia, Paenibacillus, and others are able to desulfurize aromatic heterocycles, including DBT, via the aerobic “4S pathway,” in which carbon–sulfur bonds are cleaved leaving the carbon skeleton of substrates intact [1, 4]. Reactions of this four-step pathway convert DBT into 2-hydroxybiphenyl (2-HBP) and sulfate. Four enzymes are involved: two flavin-dependent monooxygenases (DszA and DszC), a desulfinase (DszB) and a FMN:NADH oxidoreductase (DszD), necessary to recycle the monooxygenase coenzyme [4, 5]. Desulfurization occurs through sequential oxidation of the sulfur moiety and cleavage of C–S bonds. DszC monooxygenase converts DBT first to DBT sulfoxide (DBTO) and then to DBT sulfone (DBTO2). DszA attacks the C-S bond of DBTO2 and produces 2′-hydroxybiphenyl-2-sulfinate (HBPS), which is then converted to 2-HBP and sulfate by the DszB desulfinase. Finally, DszD recycles reduced flavins through the oxidation of NADH. Genetic studies demonstrated that dszA, dszC, and dszB genes are plasmid-encoded and belong to the same operon [6–8], while dszD is located on the bacterial chromosome [9]. Tough it was already demonstrated that whole bacterial cells can desulfurize diesel fuels [10, 11], replacement of HDS by microbial desulfurization is still hindered by major limitations [1], notably the inhibition of the expression of dsz genes by end products and sulfate [12], the costs of cultivation of desulfurizing bacteria [13], and the sensitivity of bacterial cells to temperature or to components of the reaction mixtures. Several strategies are under study to alleviate these hurdles, for example, the use of solvent-tolerant [14] or thermostable [15] bacterial strains. A promising approach is the production of recombinant enzymes to be used alone or as additives in the biological treatments. Following this strategy, Dsz genes have been overexpressed in heterologous hosts [13], at high copy number or under the control of strong promoters [16, 17]. Still poorly investigated is the issue of maintaining biocatalysts activity over time and in the presence of challenging reaction conditions, a major prerequisite to their industrial exploitation. As a first step towards this goal, we have investigated the conformational stability of three recombinant Dsz proteins, cloned from a novel Rhodococcus strain. Enzymes were assayed towards solvents commonly employed in biocatalysis and towards temperature, another key factor in biotransformations. We report that Dsz proteins are rather unstable to organic solvents and heating and require to be stabilized before their use in biotechnological processes.
Materials and Methods
Strains, Growth Media, and Materials
Escherichia coli strains DH5α ™ (Thermo Fisher Scientific) and BL21 (DE3) (Novagen, Madison, USA) were used as the host for DNA amplification and for heterologous expression, respectively. E. coli cells were grown in low-salt Luria–Bertani (ls-LB) medium (10 g peptone, 5 g yeast extract, 5 g NaCl in 1 L water) or in auto-induction medium ZYM-5052 [18] containing glucose as the carbon source and lactose as the inducer and secondary carbon source. Transformants of both strains were selected on agarized plates of ls-LB supplemented with 100 mg/L ampicillin and, when appropriate, with 40 mg/L 5-Bromo-4-chloro-3-Indolyl-β-D-Galactopyranoside (X-Gal). Oligonucleotides were from Eurofin MWG (Alabama, USA), and restriction enzymes and DNA ligase from New England Biolabs (NEB, Ipswich, USA). Dibenzothiophene (DBT), 2-hydroxybiphenyl (2-HBP), and organic solvents were from Sigma Aldrich (Germany). The Rhodococcus sp. AF21875 strain was isolated from environmental samples and was a kind gift of G. Bestetti and A. Tatangelo, University of Milano-Bicocca. Rhodococcus cells were grown in minimal medium (10 g glucose, 4.5 g K2HPO4, 1.5 g NaH2PO4, 2 g NH4Cl, 0.2 g MgCl2, 0.02 g CaCl2, 5 mg MnCl2, 0.5 mg H3BO3, 0.5 mg ZnCl2, 0.003 mg NaSeO3, 0.008 mg Na2WO, 1.49 mg FeCl2, 0.5 mg CoCl2, 0.46 mg NiCl2, 0.3 mg CuCl2, 1 mg NaMoO4, and 1 mL of vitamin solution in 1 L water) [19].
Amplification and Cloning of dsz Genes
Standard recombinant DNA techniques were applied according to Sambrook et al. [20]. Genes dszA, dszB, dszC, and dszD were amplified by PCR using 100 ng of total DNA extracted from Rhodococcus sp. AF21875 as the template and each primer 0.5 μM. Oligonucleotide primers (Table SI) were designed based on available sequences of the Rhodococcus dsz operon (gene bank accession number L37363.1). Each oligonucleotide contained restriction sites useful for cloning amplified genes: NdeI/HindIII for dszA and dszB, NdeI/XhoI for dszC and dszD. Before their use, oligonucleotides were phosphorylated at 37 °C for 1 h by polynucleotide kinase A (NEB, Ipswich, USA). PCR was carried out in a volume of 50 μL with 1 μL of Q5 ®high-fidelity PfuII Ultra DNA polymerase (NEB, Ipswich, USA), according to the manufacturer’s instructions and applying the following temperature program: 30 s denaturation at 98 °C, 30 cycles of 10 s at 98 °C, 30 s at 61 °C, 30 s at 72 °C, and a final extension step of 2 min at 72 °C. Amplified blunt-end DNA was cloned into a SmaI-linearized pUC18 (Thermo Fisher Scientific) plasmid that, in the presence of X-Gal (5-Bromo-4-chloro-3-Indolyl-β-D-Galactopyranoside), allows for white-blue screening of transformed colonies. Each gene was separately back-cloned into pET22 (Novagen, Madison, USA) linearized with the appropriate restriction enzymes (NdeI/HindIII for dszA and dszB, NdeI/XhoI for dszC and dszD). Constructs were checked by restriction pattern analysis and by DNA sequencing (Primm, Milan, Italy).
Expression and Purification of Dsz Proteins
Transformed BL21 cells were grown over night in 50 mL of auto-induction ZYM-5052 medium [18] at either 25 °C or 30 °C by shaking (220 rpm) and harvested by centrifugation at 1600 g at 4 °C. Cells were re-suspended in Purification Buffer (PB—50 mM sodium phosphate pH 8.0, 300 mM NaCl, and 10 mM imidazole) and lysed using a cell disruptor (Constant Systems Ltd) at 25,000 psi. Insoluble proteins were pelleted by centrifugation (15 min at 15,000×g, 4 °C). Recombinant, his-tagged proteins were purified from the supernatant by immobilized-metal affinity chromatography (IMAC) on Ni2+/NTA beads. The clear lysate was loaded on a hand-packed column containing 1.5 mL of HIS-Select Nickel Affinity Gel (Sigma Aldrich, Germany) equilibrated with 3 mL of PB. The column was washed with 2 mL PB and 2 mL PB containing 20 mM imidazole and proteins eluted with PB containing 250 mM imidazole. Protein-containing fractions were exchanged to the final buffer (10 mM sodium phosphate, pH 7) by two consecutive gel filtrations on PD-10 columns (GE Healthcare, Little Chalfont, United Kingdom) according to the manufacturer’s instructions. Protein concentration was determined by the Bradford protein assay (Bio-Rad, California, USA), using bovine serum albumin as a standard. SDS-PAGE was carried out on 14 % acrylamide Laemmli gels [21] stained with GelCode Blue (Pierce, Rockford, USA) after electrophoresis. Broad-range, pre-stained molecular-weight markers (GeneSpin, Milan, Italy) were used as standards.
Circular Dichroism Spectroscopy
CD spectra were recorded at room temperature by a spectropolarimeter J-815 (JASCO Corporation, Easton, USA) in a 1-mm path-length cuvette. Samples were in 10 mM sodium phosphate buffer, pH 7. Measurements were performed at variable wavelength (190–260 nm) with scanning velocity 20 nm/min and data pitch 0.2 nm. Spectra were averaged over three acquisitions, and smoothed by the Means-Movement algorithm [22]. Unfolding of 0.150 mg/mL protein solutions was monitored recording consecutive spectra in the temperature range 20–95 °C. Thermal ramps were acquired with the same parameters as above, in the range 20–95 °C with increments of 1 °C/min. The decrease in CD signal was monitored at 195 nm. Deconvolution of CD spectra was carried out by the CDPro software package [22] using the reference protein sets 1, 3, 4, and 10. Secondary-structure contents were expressed as percent content average over the results obtained with the algorithms SELCON3, CDSSTR, and CONTIN [22]. Buffer solutions without protein were used as blank.
Fluorescence Spectroscopy
Protein samples were re-suspended in 10 mM sodium phosphate buffer, pH 7, and fluorescence emission spectra were measured on a Cary Eclipse Fluorescence Spectrophotometer (Varian), with excitation at 290 nm and emission range 300–450 nm, employing a 1-cm path-length quartz cuvette. To detect structural changes induced by organic solvents, DszA, DszC, and DszD were incubated 3 h under continuous agitation at room temperature with 5, 10, or 20 % methanol, acetonitrile, hexane, and toluene. Fluorescence emission was measured at different times. All measurements were made in triplicate.
Sequence Analysis and Structural Modeling
DNA sequence analysis was performed using BLAST-P [23] and Psi-BLAST [24]. 3D models of DszA and DszD were built using SWISS-MODEL [25, 26]. BLAST [23] and HHBlits [27] template search were performed against the SWISS-MODEL template library.
Results
Sequence Analysis
The Rhodococcus strain used in this work was isolated from polluted environmental samples and classified as Rhodococcus sp. AF21875 (G. Bestetti and V. Tatangelo, personal communication). Genes encoding DszA, DszB, DszC, and DszD were amplified by PCR from total DNA using primers designed on the basis of corresponding Rhodococcus sp. sequences available in databases. When PSI-BLAST [24] was used to iteratively search profile similarity for DszA-D, even at the convergence of the search, no sequence was found outside the Phylum of Eubacteria. The four deduced amino acid sequences were analyzed by BLAST-P [23] against non-redundant protein sequence databases. The most significant hits for each sequence are reported in supplementary materials (Tables S2–S5), while results are summarized in the following. Sequences closest to DszA were identified within the superfamily of flavin-utilizing monooxygenases, a distinct clade of the larger luciferase-like monooxygenases family. Most retrieved sequences are annotated as dibenzothiophene desulfurizing proteins, DBT sulfone monooxygenase, and nitriloacetate monooxygenase. DszB search retrieved closest hits within the superfamily of periplasmic binding protein type 2 (PBP2). This includes the subfamily of “domains binding the substrates of 2′-hydroxybiphenyl-2-sulfinate desulfinase” to which also DszB belongs. Most sequences are annotated as dibenzothiophene desulfurization (enzyme) B, 2′-hydroxybiphenyl-2-sulfinate desulfinase, or simply as desulfurase or desulfinase. Analysis of the putative DszC confirmed its similarity to proteins of the superfamily of acyl-CoA dehydrogenase (ACAD), domain 2 type. Most sequences retrieved are annotated as dibenzothiophene desulfurization (enzyme) C or as (dibenzothiophene) monooxygenases. Analysis of putative DszD identified highly similar sequences within the pyridoxine 5′-phosphate (PNP) oxidase-like protein superfamily that includes flavin mononucleotide (FMN) binding proteins, which catalyze FMN-mediated redox reactions.
Comparison of the trees of sequence distances (SD) provided by BLAST-P (Fig. S1) highlights at a glance phylogenetic differences among Dsz enzymes. Overall, closest sequences belong to Gram-positive bacteria, notably Rhodococcus species. Besides, DszA-C are closely related to sequences from other Mycolata, for example, Gordonia, Nocardia, and Mycobacterium, while DszD is the most variable, even within the genus Rhodococcus (Fig. S1).
Homologous sequences at given SDs (i.e., at SD 0.1; 0.2; 0.3, …, 0.6) were selected within each SD tree and used to build, for each Dsz enzyme, a “reference set” containing seven sequences related by comparable degrees of similarity/diversity (BLAST-P outputs and SDs values are reported in supplementary Tables S2–S5). The high frequency of Rhodococcus sp. in the reference set of DszD (5 out of 7 reference species) is worth of note, and confirms that intra-genus variability is higher than for other Dsz proteins. The different degree of conservation of Dsz sequences may be accounted for by different evolutionary constrains. Differently than dszD, dszA-C genes are located on plasmid DNA [28, 29] that can be more easily exchanged among species. For this reason, it is reasonable to hypothesize that they spread only recently among a restricted group of Gram-positive microorganisms. It is also worth to mention that DszD is the less specific out of the four enzymes and that in microorganisms other than Rhodococcus its activity could be provided by other oxidoreductases.
Structural Features of Dsz Two-Component Enzymes
The conversion of DBT to 2-hydroxybiphenyl-2-sulfinate requires the activity of two monooxygenases, DszC and DszA. Both of them rely on the same FMN reductase, DszD, to regenerate FMNH2 that is essential to activate molecular oxygen. The two FMN-dependent monooxygenases (DszA and DszC) and the DszD FMN reductase build therefore a two-component system in which the reductive and oxidative half-reactions of FMN are carried out by separate enzymes [30]. In such systems, flavoenzymes utilize flavin as a reaction substrate rather than binding it tightly as other flavoproteins do [30]. Indeed, the strength of interactions with the cofactor and the mechanism of its intermolecular transfer between the reductase and the cognate monooxygenase are very relevant, and still debated issues for the functional characterization of these enzymes [31–35]. We appreciate that structural knowledge about the members of this enzyme system may be helpful for designing tailored processes that take advantage from the combination of catalytic functions from different sources and for implementing robust and productive catalytic systems [36–39]. In this perspective, we built by homology a 3D model of DszA, whose crystallographic structure has not been solved yet and integrated this information with data available on the other enzymes of the two-component system.
BLAST search against the Protein Data Bank (PDB) revealed that DszA shares the highest identity with a nitrilotriacetate monooxygenase (PDB code 3SDO.1.A, 41 % identity) and with the alkane monooxygenase LadA from Geobacillus thermodenitrificans Ng80-2 (PDB entry 3B9O.1.A, 37 % identity). The prototypical structure of these monooxygenases is that of bacterial luciferases (EC 1.14.14.3), able to emit light upon oxidation of long-chain aliphatic aldehydes. The homology model of DszA obtained by SWISS-MODEL [25, 26] was based on apo-LadA and on the holo-enzyme in complex with the FMN cofactor. As expected for class C flavoprotein monooxygenases [40], the structural core of DszA model conforms to a TIM-barrel scaffold from which different extensions protrude. The most remarkable is an unstructured region (loop E, residues 176–218) that contains a short α-helix and is probably involved in DszA dimerization, as it is in LadA [41], and a large pocket (P) formed by α-helices α11–α14 (DszA numbering). This pocket is assumed to build the substrate-binding region and the catalytic core of the enzyme (Fig. 1a).
Three-dimensional models of DszA and DszD. a Homology model of DszA based on the LadA structure. In the ribbon diagram, α-helices are cyan, β-sheets pink, FMN-binding residues are highlighted in red. The E loop, represented in green, and P pocket, in blue, are marked by “e” and “p” letters. b Alignment of amino acid sequences of DszA from Rhodococcus sp. AF21875 (DszA) and LadA from Geobacillus thermodenitrificans Ng80-2 (LadA). Symbols below the alignment indicate the extent of residue conservation (asterisks for absolute, colons for strong, and periods for weak conservation). Red stars above the alignment mark residues involved in FMN binding. c Structural alignment of the homology model of DszD from Rhodococcus AF12875 (green) and the structure of M. goodii DszD (PDB code: 3PFT; pink). d Alignment of amino acid sequences of DszD from Rhodococcus sp. AF21875 (DszD) and from M. goodii (3PFT) (Color figure online)
The structural alignment (Fig. 1b) highlighted a large insertion in the region corresponding to the LadA substrate-binding site (residues 320–360). This feature is consistent with the activity of DszA on aromatic substrates [42], bulkier than the long-chain, linear alkanes accepted by LadA [43]. Analysis of sequence and structural alignments also showed that most LadA residues involved in FMN binding are conserved in DszA (Asp59, Ser 138, Tyr160, Ala 228, Ser 230, Phe 246- DszA numbering), and suggested a similar binding mechanism. BLAST search against PDB database allowed to retrieve DszC enzymes from Rhodococcus sp. with known 3D structures (PDB entries: 4DOY 4JEK, 4NXL, 3X0X) [44–47]. As the DszC sequence determined in this work shares 100 % sequence identity with them, its structural and biochemical features were inferred from available information. A first outcome of this analysis is the observation that although both DszA and DszC are FMN-dependent monooxygenases, they conform to different folds. DszC forms tetramers in solution [46, 48]. Each subunit contains a β-barrel domain (β1–β8, residues 125–246) and two α-helical domains (α1–α6, residues 19–104; α7–α12, residues 257–417), with helices α8–α11 nearly parallel, forming a bundle also parallel to the longitudinal axis of the β-barrel [46]. Between the α-helix bundle and the β-barrel is located the substrate-binding pocket that is composed of an inner and an outer chamber [46]. The latter has been observed to host the flavin cofactor, [47], suggesting that the inner chamber accommodates DBT.
Finally, we focussed on DszD oxidoreductase. Irrespective of the enzymatic system they belong to, only a few 3D structures of FMN reductases are reported, and the one referred to as a DszD enzyme is from Mycobacterium goodii (PDB entry: 3PFT, 33 % identity). This structure, however, is different from the template selected by SWISS-MODEL [25, 26] to build up the homology model (Fig. 1c) that is instead an oxidoreductase from Sinorhizobium meliloti (PDB entry: 3RH7, 35 % identity) [49, 50]. Hence, the model of Rhodococcus sp. AF21875 DszD generated by SWISS-MODEL was compared with the experimental structure of M. goodii DszD by performing a pairwise 3D structural alignment with the Dali server [51]. The spatial difference between Cα atoms is 1.9 Å, indicating a degree of structural conservation higher than expected for two proteins sharing only 33 % sequence similarity (Fig. 1c, d).
Production of Recombinant Dsz Proteins and Stability Analysis
Amplified gene sequences were cloned in the expression vector pET22 and used to transform E. coli BL21 cells. Production was in auto-induction ZYM-5052 medium [18] at different temperatures. SDS-PAGE analysis revealed the presence of proteins with the expected molecular mass of 50.7 kDa (DszA), 46,0 kDa (DszC); 40.1 kDa (DszB), and 21.6 kDa (DszD). Mr were calculated including the histidine tag. At 30 °C, DszA, DszC, and DszD were produced as soluble proteins, and were easily purified by IMAC affinity chromatography (Fig. 2). Obtaining recombinant DszB was more difficult because of its low production, aggregation propensity and for the presence of contaminants in the purified samples. Similar results were obtained with all production protocols applied, including LB medium supplemented with IPTG, auto-induction broth containing glucose and lactose [18], and cultivation at different temperatures from 22 to 37 °C (data not shown). For this reason, structural analysis was not performed. As detailed in materials and methods, cells producing Dsz enzymes were harvested by centrifugation, re-suspended in PB, and lysed using a cell disruptor. Raw cell extracts thus obtained were mixed together and their activity tested on DBT, revealing that the four enzymes together produced low but detectable amounts of 2-HBP (data not shown).
SDS-PAGE analysis of recombinant DszA-D proteins. CE crude extract, SOL soluble protein fraction, INS insoluble protein fraction, PUR proteins purified by IMAC, MWM molecular-weight markers. DszA and DszC correspond to the bands at ~50 and 46 kDa, respectively; DszB corresponds to the band at ~40 kDa highlighted with an arrow; DszD, to the band at ~22 kDa. Production was at 30 °C for Dsz A, C, and D and at 25 °C for DszB. Proteins were purified from soluble protein fractions
One of the major issues in biodesulfurization is the biocatalysts stability over time. For this reason, we set up to investigate the conformational stability of the enzymes in our hands by exposing them to temperature and organic solvents.
As a first step, purified enzymes were investigated by circular dichroism spectroscopy. The model of DszA described in this work and available 3D structures of DszC [45] indicate in both cases a major contribution of alpha–beta secondary structures. Accordingly, the far-UV CD profiles of DszA and DszC show negative minima at 209 and 220 nm and a positive maximum at 195 nm, typical of alpha–beta structures (Fig. 3a, b). Spectra deconvolution with the CDPro software [22] confirms that DszA contains 20 % α-helix and 31 % β-sheets, while DszC is mainly composed by α-helix structures (40 %) and only 10 % by β-sheets. Deconvolution data further show that DszD mainly contains beta structures (38 %) with a few α-helices (11 %), as also indicated by the overall lower intensity of the ellipticity signal (Fig. 3c). To cope with the variability that can affect CD spectra deconvolution, we used four different data sets (1, 3, 4, and 10) that provided comparable results, in good agreement with the secondary structure determined for DszC (PDB code: 4JEK) and with our homology model of DszD. On the other hand, CD deconvolution data poorly agree with the secondary-structure content derived from the homology model of DszA, where α-helices seem to be markedly over-estimated (+23 %) and β-sheets under-estimated (−16.5 %). This discrepancy might depend on the lower content of proline residues in LadA (3.6 %) than in DszA (5.1 %), where the role of proline as a α-helix breaker could have been not accurately predicted by our model.
Heat denaturation of DszA, C, D proteins. a–c far-UV spectra at 20 °C (bold lines), 30, 40, 50, 60, 70, 80, 95 °C (continuous lines), and 20 °C after denaturation (dashed lines) of DszA (a), DszC (b), and DszD (c). Arrows point to changes of signal intensity at increasing temperatures. d Kinetics of thermal denaturation of DszA (continuous line), DszC (dotted line), and DszD (dashed line) heated at temperatures ranging from 20 to 95 °C measured at 195 nm
Thermal unfolding of DszA, DszC, and DszD was induced by heating protein solutions from 20 to 95 °C and monitored by circular dichroism in the far-UV (Fig. 3a–c), while unfolding transitions were recorded at 195 nm fixed wavelength (Fig. 3d). All enzymes unfolded with growing temperature, as highlighted by the loss of signal at 195 nm. In the case of DszA and DszC, we also observed a decrease of the peak at 220 nm (Fig. 3a, b). In DszC spectra, the isodichroic point at ~207 nm (Fig. 3b) was consistent with a two-state transition of α/β structures (Fig. 3d, dotted line). This protein unfolded through a cooperative process, with a melting temperature of 48 °C. DszA unfolding was a multi-stage process characterized by two different melting temperatures, at ~45 °C and ~87 °C (Fig. 3d, continuous line). The denaturation profile of DszA (Fig. 3d, continuous line) hints at the existence of at least one conformational intermediate induced by heating at 40–50 °C and stable at 50–75 °C. This intermediate has still a very high content of secondary structure (Fig. 3a, continuous line) and is highly resistant to heating (Tm = 87 °C). This particular behavior may suggest a two-step unfolding, the first implying the loss of quaternary structure (DszA is a dimer) [52, 53] and the second, the unfolding of monomers. The reversibility of thermally induced denaturation proceeds apparently until the restoration of this unfolding intermediate, as suggested by the good overlap of spectra obtained at 50–60 °C and after refolding. The denaturation of DszD did not show any isodichroic point and resulted in a final spectrum with low ellipticity at 195 nm and intense negative ellipticity at 202 nm, overall suggesting that at 95 °C the protein was almost fully unfolded (Fig. 3c). DszD showed a low cooperative and barely detectable multi-stage denaturation process (Fig. 3d, dashed lines). At 60 °C, 50 % of the protein was unfolded. For all enzymes, unfolding was (though at different extent) reversible, since samples were able to partially recover secondary structure when re-cooled to 20 °C (Fig. 3a–c, dashed lines).
Not only stability to temperature but also the enzymes robustness towards solvents is of relevance in processes where the biocatalysts can be challenged by harsh reaction conditions. Therefore, we investigated the structural stability of Dsz proteins towards solvents of common use. Protein unfolding was monitored by fluorescence spectroscopy since, in this technique, the impact of the solvent on the signal is lower than in CD spectroscopy. To this end, proteins were incubated in the presence of methanol, acetonitrile, hexane, and toluene at 5, 10, and 20 % v/v concentration and analyzed at different times through the emission spectra of intrinsic tryptophan fluorescence. Spectra were recorded after 5, 30, 60, and 180 min of incubation for each concentration. In Fig. 4, we report spectra obtained in the presence of 20 % v/v solvents, after 3 h of incubation. Lower concentrations of solvents and shorter incubation times induced similar though milder effects. Data are therefore not reported. Overall, solvents caused a red shift of the maximum peak of tryptophan fluorescence, reflecting the increased solvent accessibility of these residues. Dsz proteins were differently affected by each assayed solvent, e.g., the red shift was more evident for DszD, more sensitive to all solvents, and almost undetectable for DszC. Data on red shifts are summarized in Table 1.
Chemical denaturation of DszA, C, D proteins. Spectra of intrinsic fluorescence emission of Dsz proteins were recorded in the presence of 20 % v/v of organic solvents, after 3 h of incubation: acetonitrile (purple), toluene (green), methanol (cyan), and hexane (red). Control spectra, in the absence of solvents (dark blue), show a maximum of fluorescence emission at 338 nm for DszA, at 343 nm for DszC, and at 332 nm for DszD. Samples were excited at 290 nm and emissions were collected from 300 to 450 nm (Color figure online)
The intensity of tryptophan fluorescence increases or decreases, depending on solvent features, concentration, and duration of exposition. Overall, fluorescence intensity increases when a protein expands in a more loose structure and tryptophan residues become more exposed to the solvent [54, 55], while loss of signal intensity is often due to protein aggregation. We observed that hexane and toluene induce an increase of fluorescence intensity in the case of DszA, even upon short-time incubation (5 min, spectra not shown). Prolonged expositions to acetonitrile and methanol induced an intense increase of the tryptophan fluorescence of DszD. Toluene caused a marked decrease of the signal upon long incubation times, at whatever concentration. Both intrinsic fluorescence and thermal stability profiles support the conclusion that DszC is the most stable among Dsz enzymes.
Discussion
Biodesulfurization is gaining interest because it is milder than the HDS technology and acts on heterocyclic sulfur compounds (i.e., DBT) that are recalcitrant to HDS. However, major technical problems have still to be faced, in particular, the issue of maintaining biocatalyst activity for long time and in non-physiological conditions. Most processes described to date use whole bacterial cells, and therefore, most efforts have focused on the search for strains resistant to temperature and to inhibition of their growth and metabolism by chemical compounds present in the oil [56–58]. An alternate and still poorly explored strategy is the use of recombinant enzymes. This approach does not require the use of living cells whose growth and metabolism might be inhibited during the process. Moreover, individual enzymes from different sources and with different properties might be mixed in enzyme cocktails to be used alone or as additives to whole-cell biocatalysis. This work aims at contributing knowledge about the conformational robustness of desulfurizing enzymes. At the best of our knowledge, this issue was not tackled yet. We found out that the two monooxygenases (DszA and DszC) and the oxidoreductase (DszD) under study are only moderately resistant to temperature and to organic solvents. In view of a rational mutagenesis approach to stabilize these proteins, we collected information on their sequences and structures. Very often, homologous stable proteins offer information useful for planning stabilizing changes in a protein structure. While a few studies report about the isolation of bacterial cells active in desulfurization and tolerant to temperature, for example, Paenibacillus and Klebsiella species [56, 57], information about properties of specific enzymes is still very poor. At the present, protein engineering may rely on the structures solved or obtained by modeling. DszC is the best studied among desulfurizing enzymes, with 3 crystallographic structures available and information on both the apo- and FMN-bound enzyme [44–47]. Typical points of weakness in a protein are loops, cofactor binding sites, and surface of interaction among subunits. DszC possesses flexible loops adjacent to the flavin binding site and the substrate-binding site (between strands β1 and β2, b4 and b5 and helices α9 and α10). The loop between α-helices (residues 280–295) acts as a lid at the entrance of the active site. The lid, being by definition flexible, can initiate protein unfolding as it was shown for other proteins classes [59], and would be worth of stabilization. Obviously, planning of mutants should take into consideration that the lid participates in substrate recognition and binding. Accordingly, Guan and colleagues [47] suggested that the position of this loop influences the binding affinity of FMN. Residues involved in FMN binding and in the binding of DBT were also identified in the crystallographic structure [44] and might be targeted by mutagenesis. The same Authors report that mutations introduced to test the binding site hypothesized by molecular docking abolished enzyme activity. Though mutagenesis was performed with other purposes, this information is important and indicates that caution is necessary because the position of flavin and substrate is crucial for activity.
The DszA model localizes the position of the substrate-binding pocket P, of the FMN-binding site and of an unstructured region (loop E) that could be involved in protein dimerization. As in the previous case, all these regions might be targeted by site-directed mutagenesis.
Worth to be mentioned is the different pathway of denaturation of the two monooxygenases DszC and DszA. Both bind a flavin mononucleotide coenzyme and are organized in a quaternary structure. Differences in heat-induced denaturation pathway may reflect differences in the tightness of quaternary interactions. Indeed, DszC is a homotetramer formed by two dimers that tightly interact by hydrophobic bonds and by exchanging their C-terminal regions [46]. Its cooperative two-state transition during thermal unfolding might affect the tertiary structure, without causing a preliminary loss of quaternary structure. On the contrary, thermal unfolding of the dimeric DszA [41] seems to proceed through multiple stages, the first of them could be an early loss of quaternary structure. This information may be of some relevance in selecting the strategies of stabilization, in particular, for the choice of focussing either on the robustness of the tertiary structure or on the strength of monomers association.
An accurate study of the denaturation of the two monooxygenases to follow the first steps of unfolding (subunits dissociation? Flavin release?) might be useful to this end as well as molecular dynamics simulations to evidence other regions of weakness.
Abbreviations
- DBT:
-
Dibenzothiophene
- DBTO:
-
DBT sulfoxide
- DBTO2 :
-
DBT sulfone
- HDS:
-
Hydrodesulfurization
- 2-HBP:
-
2-Hydroxybiphenyl
- FMN:
-
Flavin mononucleotide
References
Mohebali, G., & Ball, A. S. (2008). Biocatalytic desulfurization (BDS) of petrodiesel fuels. Microbiology, 154, 2169–2183.
Babich, I. V., & Moulijn, A. C. (2003). Science and technology of novel processes for deep desulfurization of oil refinery streams: a review. Fuel, 82, 607–631.
Furimsky, E., & Massoth, F. E. (1999). Deactivation of hydroprocessing catalysts. Catalysis Today, 52, 381–495.
McFarland, B. L. (1999). Biodesulfurization. Current Opinion in Microbiology, 2, 257–264.
Kilbane, J. J, 2nd. (2006). Microbial biocatalyst developments to upgrade fossil fuels. Current Opinion in Biotechnology, 17, 305–314.
Li, M. Z., Squires, C. H., Monticello, D. J., & Childs, J. D. (1996). Genetic analysis of the dsz promoter and associated regulatory regions of Rhodococcus erythropolis IGTS8. Journal of Bacteriology, 178, 6409–6418.
Shavandi, M., Soheili, M., Zareian, S., Akbari, N., & Khajeh, K. (2013). The gene cloning, overexpression, purification, and characterization of dibenzothiophenemonooxygenase and desulfinase from Gordonia alkanivorans RIPI90A. Journal of Petroleum Science and Technology, 3, 57–64.
Ma, T., Li, S., Li, G., Wang, R., Liang, F., et al. (2006). Cloning and expressing DBT (dibenzothiophene) monooxygenase gene (dszC) from Rhodococcus sp. DS-3 in Escherichia coli. Frontiers of Biology in China, 4, 375–380.
Matsubara, T., Ohshiro, T., Nishina, Y., & Izumi, Y. (2001). Purification, characterization, and overexpression of flavin reductase involved in dibenzothiophene desulfurization by Rhodococcus erythropolis D-1. Applied and Environment Microbiology, 67, 1179–1184.
Zhang, Q., Tong, M. Y., Li, Y. S., Gao, H. J., & Fang, X. C. (2007). Extensive desulfurization of diesel by Rhodococcus erythropolis. Biotechnology Letters, 29, 123–127.
Yu, B., Ma, C., Zhou, W., Wang, Y., Cai, X., et al. (2006). Microbial desulfurization of gasoline by free whole-cells of Rhodococcus erythropolis XP. FEMS Microbiology Letters, 258, 284–289.
Alves, L., Salgueiro, R., Rodrigues, C., Mesquita, E., Matos, J., et al. (2005). Desulfurization of dibenzothiophene, benzothiophene, and other thiophene analogs by a newly isolated bacterium, Gordonia alkanivorans strain 1B. Applied Biochemistry and Biotechnology, 120, 199–208.
Alves, L., Melo, M., Mendonça, D., Simões, F., Matos, J., Tenreiro, R., & Gírio, F. M. (2007). Sequencing, cloning and expression of the dsz genes required for dibenzothiophene sulfone desulfurization from Gordonia alkanivorans strain 1B. Enzyme and Microbial Technol, 40, 1598–1603.
Tao, F., Yu, B., Xu, P., & Ma, C. Q. (2006). Biodesulfurization in biphasic systems containing organic solvents. Applied and Environment Microbiology, 72, 4604–4609.
Konishi, J., Onaka, T., Ishii, Y., & Suzuki, M. (2000). a) Demonstration of the carbon-sulfur bond targeted desulfurization of benzothiophene by thermophilic Paenibacillus sp. strain A11-2 capable of desulfurizing dibenzothiophene. FEMS Microbiology Letters, 187, 151–154.
Li, G. Q., Ma, T., Li, S. S., Li, H., Liang, F. L., et al. (2007). Improvement of dibenzothiophene desulfurization activity by removing the gene overlap in the dsz operon. Bioscience, Biotechnology, and Biochemistry, 71, 849–854.
Hirasawa, K., Ishii, Y., Kobayashi, M., Koizumi, K., & Maruhashi, K. (2001). Improvement of desulfurization activity in Rhodococcus erythropolis KA2-5-1 by genetic engineering. Bioscience, Biotechnology, and Biochemistry, 65, 239–246.
Studier, F. W. (2005). Protein production by auto-induction in high density shaking cultures. Protein Expression and Purification, 41, 207–234.
Chang, J. H., Rhee, S. K., Chang, Y. K., & Chang, H. N. (1998). Desulfurization of diesel oils by a newly isolated dibenzothiophene-degrading Nocardia sp. strain CYKS2. Biotechnology Progress, 14, 851–855.
Sambrook, J., Fritsch, E. F., & Maniatis, T. (1989). Molecular cloning. A laboratory manual. Cold Spring Harbor: Cold Spring Harbor Laboratory Press.
Laemmli, U. K. (1970). Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature, 227, 680–685.
Sreerama, N., & Woody, R. W. (2000). Estimation of protein secondary structure from circular dichroism spectra: comparison of CONTIN, SELCON, and CDSSTR methods with an expanded reference set. Analytical Biochemistry, 287, 252–260.
Altschul, S. F., Gish, W., Miller, W., Myers, E. W., & Lipman, D. J. (1990). Basic local alignment search tool. Journal of Molecular Biology, 215, 403–410.
Altschul, S. F., Madden, T. L., Schaffer, A. A., Zhang, J., Zhang, Z., et al. (1997). Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Research, 25, 3389–3402.
Arnold, K., Bordoli, L., Kopp, J., & Schwede, T. (2006). The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics, 22, 195–201.
Biasini, M., Bienert, S., Waterhouse, A., Arnold, K., Studer, G., et al. (2014). SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Research, 42, W252–W258.
Remmert, M., Biegert, A., Hauser, A., & Soding, J. (2012). HHblits: Lightning-fast iterative protein sequence searching by HMM-HMM alignment. Nature Methods, 9, 173–175.
Piddington, C. S., Kovacevich, B. R., & Rambosek, J. (1995). Sequence and molecular characterization of a DNA region encoding the dibenzothiophene desulfurization operon of Rhodococcus sp. strain IGTS8. Applied and Environment Microbiology, 61, 468–475.
Denome, S. A., Oldfield, C., Nash, L. J., & Young, K. D. (1994). Characterization of the desulfurization genes from Rhodococcus sp. strain IGTS8. Journal of Bacteriology, 176, 6707–6716.
Ellis, H. R. (2010). The FMN-dependent two-component monooxygenase systems. Archives of Biochemistry and Biophysics, 497, 1–12.
Tu, S. C., Lei, B., Liu, M., Tang, C. K., & Jeffers, C. (2000). Probing the mechanisms of the biological intermolecular transfer of reduced flavin. Journal of Nutrition, 130, 331S–332S.
Tu, S. C. (2001). Reduced flavin: donor and acceptor enzymes and mechanisms of channeling. Antioxidants & Redox Signaling, 3, 881–897.
Tu, S. C. (2008). Activity coupling and complex formation between bacterial luciferase and flavin reductases. Photochemical & Photobiological Sciences, 7, 183–188.
Lei, B., & Tu, S. C. (1998). Mechanism of reduced flavin transfer from Vibrio harveyi NADPH-FMN oxidoreductase to luciferase. Biochemistry, 37, 14623–14629.
Jeffers, C. E., & Tu, S. C. (2001). Differential transfers of reduced flavin cofactor and product by bacterial flavin reductase to luciferase. Biochemistry, 40, 1749–1754.
Ohshiro, T., Aoi, Y., Torii, K., & Izumi, Y. (2002). Flavin reductase coupling with two monooxygenases involved in dibenzothiophene desulfurization: purification and characterization from a non-desulfurizing bacterium, Paenibacillus polymyxa A-1. Applied Microbiology and Biotechnology, 59, 649–657.
Louie, T. M., Xie, X. S., & Xun, L. (2003). Coordinated production and utilization of FADH2 by NAD(P)H-flavin oxidoreductase and 4-hydroxyphenylacetate 3-monooxygenase. Biochemistry, 42, 7509–7517.
Gisi, M. R., & Xun, L. (2003). Characterization of chlorophenol 4-monooxygenase (TftD) and NADH:flavin adenine dinucleotide oxidoreductase (TftC) of Burkholderia cepacia AC1100. Journal of Bacteriology, 185, 2786–2792.
Ishii, Y., Konishi, J., Suzuki, M., & Maruhashi, K. (2000). Cloning and expression of the gene encoding the thermophilic NAD(P)H-FMN oxidoreductase coupling with the desulfurization enzymes from Paenibacillus sp. A11-2. Journal of Bioscience and Bioengineering, 90, 591–599.
van Berkel, W. J., Kamerbeek, N. M., & Fraaije, M. W. (2006). Flavoprotein monooxygenases, a diverse class of oxidative biocatalysts. Journal of Biotechnology, 124, 670–689.
Li, L., Liu, X., Yang, W., Xu, F., Wang, W., et al. (2008). Crystal structure of long-chain alkane monooxygenase (LadA) in complex with coenzyme FMN: unveiling the long-chain alkane hydroxylase. Journal of Molecular Biology, 376, 453–465.
Ohshiro, T., Kojima, T., Torii, K., Kawasoe, H., & Izumi, Y. (1999). Purification and characterization of dibenzothiophene (DBT) sulfone monooxygenase, an enzyme involved in DBT desulfurization, from Rhodococcus erythropolis D-1. Journal of Bioscience and Bioengineering, 88, 610–616.
Feng, L., Wang, W., Cheng, J., Ren, Y., Zhao, G., et al. (2007). Genome and proteome of long-chain alkane degrading Geobacillus thermodenitrificans NG80-2 isolated from a deep-subsurface oil reservoir. Proc Natl Acad Sci U S A, 104, 5602–5607.
Liu, S., Zhang, C., Su, T., Wei, T., Zhu, D., et al. (2014). Crystal structure of DszC from Rhodococcus sp. XP at 1.79 A. Proteins, 82, 1708–1720.
Duan, X., Zhang, L., Zhou, D., Ji, K., Ma, T., et al. (2013). Crystallization and preliminary structural analysis of dibenzothiophene monooxygenase (DszC) from Rhodococcus erythropolis. Acta Crystallographica, Section F: Structural Biology and Crystallization Communications, 69, 597–601.
Zhang, L., Duan, X., Zhou, D., Dong, Z., Ji, K., et al. (2014). Structural insights into the stabilization of active, tetrameric DszC by its C-terminus. Proteins, 82, 2733–2743.
Guan, L. J., Lee, W. C., Wang, S., Ohshiro, T., Izumi, Y. et al. (2015). Crystal structures of apo-DszC and FMN-bound DszC from Rhodococcus erythropolis D-1. FEBS Journal, 282, 3126–3135.
Ohshiro, T., Ohkita, R., Takikawa, T., Manabe, M., Lee, W. C., et al. (2007). Improvement of 2′-hydroxybiphenyl-2-sulfinate desulfinase, an enzyme involved in the dibenzothiophene desulfurization pathway, from Rhodococcus erythropolis KA2-5-1 by site-directed mutagenesis. Bioscience, Biotechnology, and Biochemistry, 71, 2815–2821.
Zhang, Y., Edwards, T. E., Begley, D. W., Abramov, A., Thompkins, K. B., et al. (2011). Structure of nitrilotriacetate monooxygenase component B from Mycobacterium thermoresistibile. Acta Crystallographica, Section F: Structural Biology and Crystallization Communications, 67, 1100–1105.
Knobel, H. R., Egli, T., & van der Meer, J. R. (1996). Cloning and characterization of the genes encoding nitrilotriacetate monooxygenase of Chelatobacter heintzii ATCC 29600. Journal of Bacteriology, 178, 6123–6132.
Holm, L., & Rosenstrom, P. (2010). Dali server: Conservation mapping in 3D. Nucleic Acids Research, 38, W545–W549.
Doble, M., & Kruthiventi, A. K. (2005). Biotreatment of Industrial Effluents. Burlington: Elsevier Butterworth-Heinemann.
Stapleton, R. D. J., & Singh, V. P. E. (2002). Biotransformations: Bioremediation technology for health and environmental protection. In: Progress in industrial microbiology. Amsterdam, The Netherlands: Elsevier B.V.
Kamatari, Y. O., Konno, T., Kataoka, M., & Akasaka, K. (1996). The methanol-induced globular and expanded denatured states of cytochrome c: A study by CD fluorescence, NMR and small-angle X-ray scattering. Journal of Molecular Biology, 259, 512–523.
Uversky, V. N., Narizhneva, N. V., Kirschstein, S. O., Winter, S., & Lober, G. (1997). Conformational transitions provoked by organic solvents in beta-lactoglobulin: can a molten globule like intermediate be induced by the decrease in dielectric constant? Folding and Design, 2, 163–172.
Konishi, J., Ishii, Y., Onaka, T., Ohta, Y., Suzuki, M., et al. (2000). b) Purification and characterization of dibenzothiophene sulfone monooxygenase and FMN-dependent NADH oxidoreductase from the thermophilic bacterium Paenibacillus sp. strain A11-2. Journal of Bioscience and Bioengineering, 90, 607–613.
Bhatia, S., & Sharma, D. K. (2012). Thermophilic desulfurization of dibenzothiophene and different petroleum oils by Klebsiella sp. 13T. Environmental Science and Pollution Research International, 19, 3491–3497.
Aggarwal, S., Karimi, I. A., Kilbane Ii, J. J., & Lee, D. Y. (2012). Roles of sulfite oxidoreductase and sulfite reductase in improving desulfurization by Rhodococcus erythropolis. Molecular BioSystems, 8, 2724–2732.
Santarossa, G., Gatti Lafranconi, P., Alquati, C., DeGioia, L., Alberghina, L., Fantucci, P., & Lotti, M. (2005). Mutations in the ‘‘lid’’ region affect chain length specificity and thermostability of a Pseudomonas fragi lipase. FEBS Letters, 579, 2383–2386.
Acknowledgments
This work was supported by CORIMAV, a consortium between Pirelli and University of Milano-Bicocca, through a doctoral fellowship to F.P. The authors are grateful to C. Santambrogio for fruitful discussion and to J. Pleiss, University of Stuttgart, for help in the bioinformatic analysis and for hosting F.P. for a stage.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interests
The authors declare that they have no conflict of interests and that all authors read and approved this manuscript.
Ethical approval
This research did not imply human participants or animals.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Parravicini, F., Brocca, S. & Lotti, M. Evaluation of the Conformational Stability of Recombinant Desulfurizing Enzymes from a Newly Isolated Rhodococcus sp.. Mol Biotechnol 58, 1–11 (2016). https://doi.org/10.1007/s12033-015-9897-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-015-9897-7