Abstract
Autism is a severe neurodevelopmental condition with unknown pathobiology. Nevertheless, multiple pieces of evidence suggest long non-coding RNA (lncRNA) dysregulation may be a contributing factor to this disorder. We investigated the association between the expression of five specific lncRNAs and autism. Peripheral blood was collected from 30 children with autism and 41 healthy children. The expression levels of PCAT-29, lincRNA-ROR, LINC-PINT, lincRNA-p21, and PCAT-1 were calculated. Then, their significance as biomarkers was also evaluated. The expression of LincRNA-ROR (27 times), LINC-PINT (5.26 times), LincRNA-p21 (4.54 times), PCAT-29 (16.66 times), and PCAT-1 (25 times) genes was significantly decreased in patients compared to the control group (p values < 0.05). According to the ROC curve analysis for each lncRNA, LincRNA-ROR, LINC-PINT, LincRNA-p21, PCAT-29, and PCAT-1 lncRNAs with diagnostic power of 0.85, 0.67, 0.64, 0.74, and 0.84, respectively, could be used as diagnostic biomarkers for autism. Additionally, significant positive correlations were reported between expression levels of PCAT-1 and PCAT-29 genes. Moreover, a positive correlation was detected between expression levels of lincRNA-ROR and patients’ age. The current study shows further pieces of evidence for deregulation of lncRNAs in autistic patients that show these lncRNAs may play an important part in the pathogenesis of ASD. However, the role of lncRNA in the neurobiology of autism needs to be investigated further.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Autism is a neurodevelopmental disorder characterized by social communication defects, repetitive behaviors, and varying cognitive disabilities (Lord et al. 2020; Eyring and Geschwind 2021). It affects all societies, regardless of culture, race, ethnicity, or socioeconomic status (Lord et al. 2020). By 2020, the estimated prevalence of autism have risen to 1 in 36 children, with boys being 3.8 times more likely to be affected (Maenner et al. 2023). Autism is highly inheritable (74–93%), with many genetic loci and risk variants impacting early brain development (Lord et al. 2020).
The precise biological processes responsible for autism are still not fully understood (Ghafouri-Fard et al. 2023a). However, researchers suggested that the malfunction of various signaling pathways may play a role in its development (Jiang et al. 2022; Ghafouri-Fard et al. 2022). Long noncoding RNAs (lncRNAs) are crucial in modulating signaling pathways, serving as functional factors with multiple significant functions (Ghafouri-Fard et al. 2022). Furthermore, the involvement of various lncRNAs in the pathogenesis of autism has been extensively documented (Akbari et al. 2022; Pourtavakoli et al. 2024). These transcripts play crucial roles in gene expression regulation and epigenetic control and also function as scaffolds for protein complexes (Mattick et al. 2023). Extensive expression of lncRNAs in the human brain indicates their role in the brain development and potential involvement in the molecular pathogenesis of neurodevelopmental disorders (Srinivas et al. 2023). Many psychiatric disorders such as schizophrenia, autism, and anxiety are associated with the deregulation of lncRNA expression (Baruah et al. 2022; Dini et al. 2024). Here, we evaluated the expression levels of five lncRNAs, namely, LincRNA-ROR, LINC-PINT, LincRNA-p21, PCAT-29, and PCAT-1, in children with autism and healthy children. The lncRNAs chosen were based on their established involvement in the progression of neurological conditions and their impact on the pathways and cellular processes associated with the development of autism.
Long intergenic non-protein coding RNA, a regulator of reprogramming (LincRNA-ROR), regulates key pluripotency factors such as Oct4, Sox2, and Nanog (Wang et al. 2013). Research has shown that the absence of LincRNA-ROR results in the induction of apoptosis, indicating its crucial involvement in the p53 network (Grossi et al. 2016). The findings of another study showed that both LincRNA-ROR and p53 exhibited higher levels of expression in glioblastoma. Upregulation of p53 could potentially contribute to the elevation of LincRNA-ROR in glioblastoma (Toraih et al. 2019). The expression levels of LincRNA-ROR were found to be altered in individuals suffering from neurodegenerative and neurodevelopmental disorders (Ghafouri-Fard et al. 2023b). Furthermore, LincRNA-ROR was elevated in patients diagnosed with schizophrenia when compared to control groups. However, a significant difference in expression was observed exclusively among female subgroups when analyzing sex-based variations. Additionally, the expression of this lncRNA was linked to the age of patients with schizophrenia (Fallah et al. 2019).
Long intergenic non-protein coding, P53-induced transcript (LINC-PINT) has a direct interaction with the polycomb repressive complex 2 (PRC2) to modulate the expression of its target genes (Xu et al. 2019). This lncRNA is expressed in neurons and is up-regulated in several regions of the brain in individuals affected by neurodegenerative diseases (Simchovitz et al. 2020).
Long intergenic non-coding RNA p21 (LincRNA-p21) regulates a varied range of biological processes, such as apoptosis and cell cycle transition via affecting Notch signaling, JAK/STAT3, and AKT/mTOR pathways (Winkler et al. 2022; Amirinejad et al. 2020). Cross-talk with different miRNAs is one of the mechanisms by which lincRNA-p21 can alter these processes (Amirinejad et al. 2020). The activation of microglia cells and the increase in the expression levels of inflammatory mediators, as well as the induction of apoptosis in dopaminergic neurons, are observed upon the expression of lincRNA-p21 (Ye et al. 2018). Furthermore, lincRNA-p21 has been found to be upregulated in the early stages of Parkinson’s disease and continues to increase during disease progression, particularly in cases of brain stem type Parkinson’s disease (Kraus et al. 2017).
Moreover, the reduction in the blood levels of lincRNA-p21, lincRNA-ROR, and LINC-PINT could be potentially associated with a higher susceptibility to bipolar disorder (Maloum et al. 2022).
Prostate cancer-associated transcript 29 (PCAT-29) is considered to be a tumor suppressor inhibiting cell proliferation, migration, tumor growth, and metastasis (Al Aameri et al. 2017). An increase in PCAT-29 lncRNA expression has been documented in individuals diagnosed with major depressive disorder (Seki et al. 2019). PCAT-1 regulates cell response to genotoxic stress. It may also act as a sponge for miRNAs that regulate cell growth (Xiong et al. 2019).
These lncRNAs were demonstrated to have crucial involvement in the modulation of neurological responses and the differentiation of neuronal cells. Thus, in the current study, we compared the expression levels of these lncRNAs in the peripheral blood of children with autism and healthy individuals. We aimed to investigate any association between changes in their expression levels and autism and assess their potential as valuable biomarkers for diagnostic and prognostic purposes. The objective was to gain insights into the role of these lncRNAs in the development of autism and explore their potential as biomarkers for the disorder.
Methods and Material
Subjects
We performed this study on peripheral blood samples from 30 patients and 41 healthy controls. Samples were collected from the hospitals of Shahid Beheshti and Hamadan Universities of Medical Sciences. The research excluded children with neuropsychiatric conditions other than autism, metabolic disorders, or immune-related issues, as well as those with comorbid attention deficit hyperactivity disorder. Healthy children included in the study had no prior history of neuropsychiatric or developmental disorders. Table 1 presents the subjects’ demographic data, and detailed information of all samples are presented in Supplementary file 1. The patients were diagnosed using the DSM-5 criteria. Informed consent was obtained from all participants’ parents. The Ethical Committee of Shahid Beheshti University of Medical Sciences permitted this study, and it was conducted under the institution’s ethical standards (IR.SBMU.MSP.REC.1399.290).
Experiments
A total of 5 ml of peripheral blood samples were collected from both the patient and control groups in the EDTA tubes. Total RNA was then extracted using the RNXplus (Cinagen, Tehran, Iran) RNA extraction kit, following the guidelines provided by the manufacturer. The quality and quantity of the extracted RNA were evaluated using gel electrophoresis and nanodrop spectrophotometer, respectively. Any DNA contamination was removed using DNase I.
To produce cDNA from 3 μg of purified total RNA, the RT-PCR Pre-Mix Kit (BIOFACT™, South Korea) was used. Quantitative real-time PCR was performed using 7.5 μl of BIOFACT™ PCR Master Mix (High Rox including SYBR Green I), 10 ng cDNA, and 200 nM of each primer. Primer sequences provided in Table 2 were designed using GeneRunner v3.5 and Oligo7 (Rychlik 2007). The relative gene expression levels were determined by the housekeeping gene B2M, serving as an internal control. The average ΔCT for both the case and control groups was calculated, and then by using the Livak method (ratio = 2−∆∆Ct), fold changes of each gene expression were computed (Livak and Schmittgen 2001). Quality control checks were implemented in several steps, including assessment of abnormal amplification, evaluation of melting curves, assessment of variations between PCR replicates, inclusion of positive and negative control samples in each run, and evaluation of expression stability of the selected reference gene (B2M) among samples.
Statistical Analyses
Statistical analyses were performed using GraphPad Prism version 8.0.2 (GraphPad Software, Inc., San Diego, CA, USA), and a P value < 0.05 was deemed significant. The Kolmogorov–Smirnov test was employed to assess the normal distribution of the dataset. Because the samples had a normal distribution, the t-test was utilized to compare the expression levels of lncRNAs and identify any significant differences between patients and controls. The Pearson correlation coefficient was employed to assess the correlations between gene expression and participants’ clinical features, such as age. The ROC curve was used to determine the specificity and sensitivity of genes mentioned as possible biomarkers.
Results
Gene Expression Levels
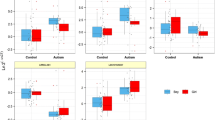
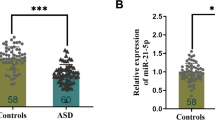
We observed a significant decrease in the expression levels of lncRNAs in patients with autism compared to healthy children. In detail, lincRNA-ROR was 27 times lower (p value < 0.0001) (Fig. 1 A and F), LINC-PINT was 5.26 times lower (p value = 0.013) (Fig. 1 B and F), lincRNA-p21 was 4.54 times lower (p value = 0.029) (Fig. 1 C and F), PCAT-29 was 16.66 times lower (p value = 0.0004) (Fig. 1 D and F), and PCAT-1 was 25 times lower (p value < 0.0001) (Fig. 1 E and F) in the patient group (Table 3). In the gender-specific comparison, we evaluated differences in the expression levels of lncRNAs in female and male patients compared to their respective healthy counterparts. Specifically, both female and male patients showed significantly decreased expression levels for lincRNA-ROR (p value = 0.0041 and p value < 0.0001, respectively), PCAT-29 (p value = 0.029 and p value = 0.004, respectively), and PCAT-1 (p value = 0.0014 and p value < 0.0001, respectively) (Fig. 2 A, D, and E, respectively). LINC-PINT demonstrated a significant decrease in female patients (p value = 0.0387), but this trend was not statistically significant in male patients (p value = 0.0989) (Fig. 2 B). Lastly, while lincRNA-p21 showed decreased expression levels in both genders, the difference was insignificant (female p value = 0.3909, male p value = 0.0535) (Fig. 2 C). We detected no significant decline when comparing expression levels between the female and male populations (Table 3).
The expression of lncRNAs in blood samples of autistic patients and controls. Expression analyses were normalized (∆Ct = Ct target gene –Ct reference gene) by comparing them to B2M internal control. ∆Cts of lincRNA-ROR (A), LINC-PINT (B), lincRNA-p21 (C), PCAT-29 (D), and PCAT-1 (E). F The relative expression (fold change = 2.−ΔΔCt) of lncRNAs shows downregulation of these genes in autistic patients compared with controls. *P < 0.05, ***P < 0.001, ****P < 0.0001
Expression analysis of lncRNAs in the blood samples of autistic patients in comparison with controls based on gender. Expression analyses were normalized by comparing them to B2M internal control. ∆Cts of lincRNA-ROR (A), LINC-PINT (B), lincRNA-p21 (C), PCAT-29 (D), and PCAT-1 (E). Expression analyses were normalized (∆Ct = Ct target gene –Ct reference gene) by comparing them to B2M internal control. *P < 0.05, ** P < 0.01, ****P < 0.0001
Correlation Analysis
Figure 3A shows the pairwise correlations between expression levels of lncRNAs in this study. A significant positive correlation was identified between PCAT-29 and PCAT-1 (r = 0.36, p value = 0.0004) and between LINC-PINT and PCAT-1 (r = 0.46 and p value = 0.011). No other two genes showed such a correlation. Correlations between clinical features and expression levels were also analyzed. Figure 3B illustrates a significant positive correlation between the expression of lincRNA-ROR and patient age (r = 0.46, p value = 0.0092), indicating an increase in its expression as the patient’s age increases. However, the correlations of age with the other four lncRNAs were not significant, as their p values were greater than 0.05 (Fig. 3C–F).
Correlation analysis. A A heat map illustrating the Pearson correlation among gene expression levels across various samples. The red color indicates a high degree of correlation between samples, while blue signifies a low correlation. B–F scatter plot of correlation analysis between age and expression levels of lncRNAs in patients with autism disorder. **P < 0.01
ROC Curve Analysis
To evaluate the diagnostic power of these lncRNAs for autism, ROC curve analysis was performed. LincRNA-ROR demonstrated the highest area under the ROC curve (AUC) of 0.85 (p value < 0.0001) with a sensitivity of 86.67% and a specificity of 65.85%, indicating its superiority in differentiation between groups. Closely following was PCAT-1 with an AUC of 0.84 (p value < 0.0001), a sensitivity of 80%, and a specificity of 70.73%. The AUC of PCAT-29, LINC-PINT, and lincRNA-p21 were 0.74 (p value = 0.0005), 0.67 (p value = 0.0138), and 0.64 (p value = 0.0394), respectively. These findings suggest that all tested lncRNAs could be used as potential biomarkers for autism to differentiate between patient and healthy conditions (Fig. 4).
Then, we conducted ROC curve analyses in males and females, separately (Table 4). While p values remained significant in both groups for LincRNA-ROR and PCAT-1, p values for LINC-PINT and PCAT-29 were significant only among females and males, respectively.
Discussion
Identification and characterization of lncRNAs that are involved in the neurodevelopmental processes and mental diseases are necessary for revealing the complex transcriptional processes in brain development. Many of the genes that are controlled by our investigated lncRNAs in this study are involved in brain development or are associated with neurodevelopmental and mental disorders. Therefore, we suggest their involvement in autism pathogenesis based on previous studies showing their function and association in similar conditions.
Previous studies have proven that PCAT-29 is significantly associated with glioma, the most common malignant brain tumor (Al Aameri et al. 2017). Besides, its expression is increased in major depressive disorder (Seki et al. 2019). Moreover, PCAT-29 downregulates miR-494, which regulates PTEN (Lu et al. 2021). Since mutations in PTEN are one of the key factors in microcephaly, cognitive impairment, seizure, and autism (Dhaliwal et al. 2021), the observed decline of PCAT-29 expression in our study suggests that this gene is involved in autism pathogenesis. In other words, a decrease in PCAT-29 might lead to a decline in PTEN, affecting neural cell growth and synaptic functions in the central nervous system, and causing many autism symptoms.
Decreased LincRNA-ROR expression in glioma affects cell proliferation and stem cell renewal by inhibiting KLF4 (Toraih et al. 2019). In addition, LincRNA-ROR can act as a molecular sponge and disrupt the usual function of miR-145 (as it does in triple-negative breast cancer) (Eades et al. 2015), which also plays a vital role in regulating neurogenesis and neural stem cell differentiation (Morgado et al. 2016). A rise in the expression of LincRNA-ROR in schizophrenia has been reported, which is one of the co-occurring disorders with autism and has similar symptoms (Jovčevska and Videtič Paska 2021). These suggest that decreased LincRNA-ROR expression in autism patients, observed in this study, could increase neuron apoptosis and impair stress responses, a key process in the development of this disorder (Kim et al. 2017).
Raised LINC-PINT expression has been detected in Parkinson’s disease (Bhattacharyya et al. 2021), and its accumulation has been observed in some regions of the brain in Alzheimer’s and Huntington’s diseases. This suggests a strong correlation between the up-regulation of LINC-PINT and the onset of neurodegeneration (Simchovitz et al. 2020; Reddy et al. 2021). Additionally, the reduction of LINC-PINT disrupts PRC2 function (Marín-Béjar et al. 2017). Based on the observed high expression of LINC-PINT in neural and microglial brain cells (Reddy et al. 2021), and significant role of PRC2 in the regulation of neurogenesis in the developing neocortex, responsible for higher-order functions such as sensory perception, cognition, and language (Elsen et al. 2018), we suggest the reduction of LINC-PINT expression in autism patients could account for some of the main autism symptoms.
Decreases in lincRNA-p21 expression in glioma stem cells (GSCs) have been linked to brain malignancies. These cells are directly associated with malignancy and angiogenesis in brain cells (Gimple et al. 2019). In addition, downregulation of miR-146b-5p in GSC cells (Wang et al. 2020) increases the Hu-antigen receptor (HuR) expression, leading to a decrease in lincRNA-p21 and an increase in β-catenin levels (Yang et al. 2016). Dysregulation of β-catenin disrupts the β-catenin/TGF pathway, which regulates expression of various genes, including NLGN3. Previous studies showed that NLGN3 overexpression leads to an increase in the number of synapses, altering the excitatory/inhibitory (E/I) ratio, which results in difficulties in higher cognitive function and social skills, which are important symptoms of autism (Guang et al. 2018). As the Wnt/β-catenin pathway is crucial for the formation of neurons, glial cells, and hippocampus, gain and loss of function mutations in this gene cause neurodevelopmental diseases such as Alzheimer’s, Parkinson’s, Huntington’s, and autism (Caracci et al. 2021). Therefore, we suggest that this decrease in lincRNA-p21 expression level in patients plays a significant role in autism pathogenesis.
Interestingly, both PCAT-1 and PCAT-29 expression levels rise in MDD patients (Seki et al. 2019). We also found that these two genes have a positive pairwise correlation. Additionally, previous research has reported increased expression of PCAT-1 upon treatment with a histone deacetylase (HDAC) inhibitor (Prensner et al. 2011). Alterations in the histone acetylation in various genes have been identified as a cause of autism due to their role in synaptic activities, neuronal excitability, and immune responses (Tseng et al. 2022). Moreover, studies have shown that parental or before-birth suppression of HDAC can delay brain neuron maturation and lead to changes in the synaptic molecules, ultimately contributing to autism symptoms (Kawanai et al. 2016). These findings underscore the interplay between gene expression, epigenetics, and neurodevelopmental disorders. In the context of autism, we propose that the observed reduction in PCAT-1 expression is associated with autism symptoms. However, the exact interactions or relationships that cause the effect of PCAT-1 on HDAC are not yet clear and need further research.
Conclusion
The study revealed that the expression levels of five lncRNAs examined—PCAT-29, lincRNA-ROR, LINC-PINT, lincRNA-p21, and PCAT-1—were significantly lower in individuals with autism compared to the healthy control group. ROC analysis indicated that these lncRNAs may serve as potential biomarkers for autism. Notably, the significant decrease in expression was observed in both genders for PCAT-29, lincRNA-ROR, and PCAT-1, while a significant reduction in LINC-PINT levels was specifically noted among girls with autism. Additionally, a positive correlation was identified between PCAT-29 and PCAT-1, as well as between the expression level of lincRNA-ROR and the age of the patients. To further substantiate these findings and to explore the underlying mechanisms of the observed associations, additional research with a larger sample size is warranted.
Data Availability
No datasets were generated or analysed during the current study.
References
AKBARI, M., EGHTEDARIAN, R., HUSSEN, B. M., ESLAMI, S., TAHERI, M., NEISHABOURI, S. M. & GHAFOURI-FARD, S. 2022. Assessment of expression of regulatory T cell differentiation genes in autism spectrum disorder. Frontiers in Molecular Neuroscience, 15.
Al Aameri RFH, Sheth S, Alanisi EMA, Borse V, Mukherjea D, Rybak LP, Ramkumar V (2017) Tonic suppression of PCAT29 by the IL-6 signaling pathway in prostate cancer: Reversal by resveratrol. PLoS One 12:e0177198
Amirinejad R, Rezaei M, Shirvani-Farsani Z (2020) An update on long intergenic noncoding RNA p21: a regulatory molecule with various significant functions in cancer. Cell Biosci 10:82
Baruah C, Nath P, Barah P (2022) LncRNAs in neuropsychiatric disorders and computational insights for their prediction. Mol Biol Rep 49:11515–11534
Bhattacharyya N, Pandey V, Bhattacharyya M, Dey A (2021) Regulatory role of long non coding RNAs (lncRNAs) in neurological disorders: from novel biomarkers to promising therapeutic strategies. Asian J Pharm Sci 16:533–550
Caracci MO, Avila ME, Espinoza-Cavieres FA, López HR, Ugarte GD, de Ferrari GV (2021) Wnt/β-catenin-dependent transcription in autism spectrum disorders. Front Mol Neurosci 14:764756
Dhaliwal N, Choi WWY, Muffat J, Li Y (2021) Modeling PTEN overexpression-induced microcephaly in human brain organoids. Mol Brain 14:131
Dini N, Taheri M, Shirvani-Farsani Z (2024) The expression analysis of long noncoding RNAs PCAT-1, PCAT-29, and MER11C in bipolar disorder. BMC Psychiatry 24:524
Eades G, Wolfson B, Zhang Y, Li Q, Yao Y, Zhou Q (2015) lincRNA-RoR and miR-145 regulate invasion in triple-negative breast cancer via targeting ARF6. Mol Cancer Res 13:330–338
Elsen GE, Bedogni F, Hodge RD, Bammler TK, Macdonald JW, Lindtner S, Rubenstein JLR, Hevner RF (2018) The epigenetic factor landscape of developing neocortex is regulated by transcription factors Pax6→ Tbr2→ Tbr1. Front Neurosci 12:571
Eyring KW, Geschwind DH (2021) Three decades of ASD genetics: building a foundation for neurobiological understanding and treatment. Hum Mol Genet 30:R236-r244
Fallah H, Azari I, Neishabouri SM, Oskooei VK, Taheri M, Ghafouri-Fard S (2019) Sex-specific up-regulation of lncRNAs in peripheral blood of patients with schizophrenia. Sci Rep 9:12737
Ghafouri-Fard S, Noroozi R, Brand S, Hussen BM, Eghtedarian R, Taheri M, Ebrahimzadeh K (2022) Emerging role of non-coding RNAs in autism spectrum disorder. J Mol Neurosci 72:201–216
Ghafouri-Fard S, Pourtavakoli A, Hussen BM, Taheri M, Ayatollahi SA (2023a) A review on the role of genetic mutations in the autism spectrum disorder. Mol Neurobiol 60:5256–5272
Ghafouri-Fard S, Pourtavakoli A, Hussen BM, Taheri M, Kiani A (2023b) A review on the importance of LINC-ROR in human disorders. Pathology - Research and Practice 244:154420
Gimple RC, Bhargava S, Dixit D, Rich JN (2019) Glioblastoma stem cells: lessons from the tumor hierarchy in a lethal cancer. Genes Dev 33:591–609
Grossi E, Sánchez Y, Huarte M (2016) Expanding the p53 regulatory network: LncRNAs take up the challenge. Biochim Biophys Acta 1859:200–208
Guang S, Pang N, Deng X, Yang L, He F, Wu L, Chen C, Yin F, Peng J (2018) Synaptopathology involved in autism spectrum disorder. Front Cell Neurosci 12:470
Jiang CC, Lin LS, Long S, Ke XY, Fukunaga K, Lu YM, Han F (2022) Signalling pathways in autism spectrum disorder: mechanisms and therapeutic implications. Signal Transduct Target Ther 7:229
JovčevskaVidetič Paska IA (2021) Neuroepigenetics of psychiatric disorders: Focus on lncRNA. Neurochem Int 149:105140
Kawanai T, Ago Y, Watanabe R, Inoue A, Taruta A, Onaka Y, Hasebe S, Hashimoto H, Matsuda T, Takuma K (2016) Prenatal exposure to histone deacetylase inhibitors affects gene expression of autism-related molecules and delays neuronal maturation. Neurochem Res 41:2574–2584
Kim C, Kang D, Lee EK, Lee JS (2017) Long noncoding RNAs and RNA-binding proteins in oxidative stress, cellular senescence, and age-related diseases. Oxid Med Cell Longev 2017:2062384
Kraus TFJ, Haider M, Spanner J, Steinmaurer M, Dietinger V, Kretzschmar HA (2017) Altered long noncoding RNA expression precedes the course of Parkinson’s disease-a preliminary report. Mol Neurobiol 54:2869–2877
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25:402–408
Lord C, Brugha TS, Charman T, Cusack J, Dumas G, Frazier T, Jones EJH, Jones RM, Pickles A, State MW, Taylor JL, Veenstra-Vanderweele J (2020) Autism spectrum disorder. Nat Rev Dis Primers 6:5
LU, B., LV, H., YANG, Z., SHU, J. & ZHANG, H. 2021. LncRNA PCAT29 up-regulates the expression of PTEN by down-regulating miR-494 in non-small-cell lung cancer to suppress tumor progression. 31, 9–15.
Maenner MJ, Warren Z, Williams AR, Amoakohene E, Bakian AV, Bilder DA, Durkin MS, Fitzgerald RT, Furnier SM, Hughes MM, Ladd-Acosta CM, McArthur D, Pas ET, Salinas A, Vehorn A, Williams S, Esler A, Grzybowski A, Hall-Lande J, Nguyen RHN, Pierce K, Zahorodny W, Hudson A, Hallas L, Mancilla KC, Patrick M, Shenouda J, Sidwell K, Dirienzo M, Gutierrez J, Spivey MH, Lopez M, Pettygrove S, Schwenk YD, Washington A, Shaw KA (2023) Prevalence and characteristics of autism spectrum disorder among children aged 8 years - autism and developmental disabilities monitoring network, 11 Sites, United States, 2020. MMWR Surveill Summ 72:1–14
Maloum Z, Ramezani S, Taheri M, Ghafouri-Fard S, Shirvani-Farsani Z (2022) Downregulation of long non-coding RNAs in patients with bipolar disorder. Sci Rep 12:7479
Marín-Béjar O, Mas AM, González J, Martinez D, Athie A, Morales X, Galduroz M, Raimondi I, Grossi E, Guo S, Rouzaut A, Ulitsky I, Huarte M (2017) The human lncRNA LINC-PINT inhibits tumor cell invasion through a highly conserved sequence element. Genome Biol 18:202
Mattick JS, Amaral PP, Carninci P, Carpenter S, Chang HY, Chen L-L, Chen R, Dean C, Dinger ME, Fitzgerald KA, Gingeras TR, Guttman M, Hirose T, Huarte M, Johnson R, Kanduri C, Kapranov P, Lawrence JB, Lee JT, Mendell JT, Mercer TR, Moore KJ, Nakagawa S, Rinn JL, Spector DL, Ulitsky I, Wan Y, Wilusz JE, Wu M (2023) Long non-coding RNAs: definitions, functions, challenges and recommendations. Nat Rev Mol Cell Biol 24:430–447
Morgado AL, Rodrigues CM, Solá S (2016) MicroRNA-145 Regulates Neural Stem Cell Differentiation Through the Sox2-Lin28/let-7 Signaling Pathway. Stem Cells 34:1386–1395
Pourtavakoli A, Ghafouri-Fard S, Eslami S, Brand S, Taheri M (2024) Expression assay of calcium signaling related lncRNAs in autism. Mol Biol Rep 51:185
Prensner JR, Iyer MK, Balbin OA, Dhanasekaran SM, Cao Q, Brenner JC, Laxman B, Asangani IA, Grasso CS, Kominsky HD, Cao X, Jing X, Wang X, Siddiqui J, Wei JT, Robinson D, Iyer HK, Palanisamy N, Maher CA, Chinnaiyan AM (2011) Transcriptome sequencing across a prostate cancer cohort identifies PCAT-1, an unannotated lincRNA implicated in disease progression. Nat Biotechnol 29:742–749
Reddy JS, Allen M, Ho CCG, Oatman SR, İş Ö, Quicksall ZS, Wang X, Jin J, Patel TA, Carnwath TP, Nguyen TT, Malphrus KG, Lincoln SJ, Carrasquillo MM, Crook JE, Kanekiyo T, Murray ME, Bu G, Dickson DW, Ertekin-taner N (2021) Genome-wide analysis identifies a novel LINC-PINT splice variant associated with vascular amyloid pathology in Alzheimer’s disease. Acta Neuropathol Commun 9:93
Rychlik W (2007) OLIGO 7 primer analysis software. Methods Mol Biol 402:35–60. https://doi.org/10.1007/978-1-59745-528-2_2
Seki T, Yamagata H, Uchida S, Chen C, Kobayashi A, Kobayashi M, Harada K, Matsuo K, Watanabe Y, Nakagawa S (2019) Altered expression of long noncoding RNAs in patients with major depressive disorder. J Psychiatr Res 117:92–99
Simchovitz A, Hanan M, Yayon N, Lee S, Bennett ER, Greenberg DS, Kadener S, Soreq H (2020) A lncRNA survey finds increases in neuroprotective LINC-PINT in Parkinson’s disease substantia nigra. Aging Cell 19:e13115
Srinivas T, Mathias C, Oliveira-Mateos C, Guil S (2023) Roles of lncRNAs in brain development and pathogenesis: emerging therapeutic opportunities. Mol Ther 31:1550–1561
Toraih EA, El-Wazir A, Hussein MH, Khashana MS, Matter A, Fawzy MS, Hosny S (2019) Expression of long intergenic non-coding RNA, regulator of reprogramming, and its prognostic value in patients with glioblastoma. Int J Biol Markers 34:69–79
Tseng CJ, McDougle CJ, Hooker JM, Zürcher NR (2022) Epigenetics of autism spectrum disorder: histone deacetylases. Biol Psychiatry 91:922–933
Wang Y, Xu Z, Jiang J, Xu C, Kang J, Xiao L, Wu M, Xiong J, Guo X, Liu H (2013) Endogenous miRNA sponge lincRNA-RoR regulates Oct4, Nanog, and Sox2 in human embryonic stem cell self-renewal. Dev Cell 25:69–80
Wang H, Tan L, Dong X, Liu L, Jiang Q, Li H, Shi J, Yang X, Dai X, Qian Z, Dong J (2020) MiR-146b-5p suppresses the malignancy of GSC/MSC fusion cells by targeting SMARCA5. Aging (albany NY) 12:13647–13667
Winkler L, Jimenez M, Zimmer JT, Williams A, Simon MD, Dimitrova N (2022) Functional elements of the cis-regulatory lincRNA-p21. Cell Rep 39:110687
Xiong T, Li J, Chen F, Zhang F (2019) PCAT-1: a novel oncogenic long non-coding RNA in human cancers. Int J Biol Sci 15:847–856
Xu Y, Wang H, Li F, Heindl LM, He X, Yu J, Yang J, Ge S, Ruan J, Jia R, Fan X (2019) Long non-coding RNA LINC-PINT suppresses cell proliferation and migration of melanoma via recruiting EZH2. Front Cell Dev Biol 7:350
Yang W, Yu H, Shen Y, Liu Y, Yang Z, Sun T (2016) MiR-146b-5p overexpression attenuates stemness and radioresistance of glioma stem cells by targeting HuR/lincRNA-p21/β-catenin pathway. Oncotarget 7:41505–41526
Ye Y, He X, Lu F, Mao H, Zhu Z, Yao L, Luo W, Sun X, Wang B, Qian C, Zhang Y, Lu G, Zhang S (2018) A lincRNA-p21/miR-181 family feedback loop regulates microglial activation during systemic LPS- and MPTP- induced neuroinflammation. Cell Death Dis 9:803
Author information
Authors and Affiliations
Contributions
SS and VE wrote the manuscript, analyzed the data, and performed the experiment. ZSF and SGF designed the study and revised the manuscript. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics Approval and Consent to Participate
All procedures were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration. Informed consent forms were obtained from all parents. This study was approved by the ethical committee of Shahid Beheshti University of Medical Science (IR.SBMU.MSP.REC.1399.290).
Consent of Publication
Not applicable.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sane, S., Ebrahimi, V., Shirvani Farsani, Z. et al. Assessment of Expression of lncRNAs in Autistic Patients. J Mol Neurosci 74, 81 (2024). https://doi.org/10.1007/s12031-024-02258-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12031-024-02258-8