Abstract
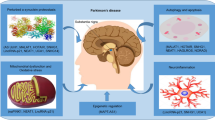
Parkinson’s disease (PD) is a slowly progressing neurodegenerative disorder that affects approximately seven million patients worldwide. Despite intensive research, the molecular mechanisms initiating and promoting PD are still unknown. However, it is assumed that environmental factors trigger PD. Recent research demonstrated that long noncoding RNAs (lncRNA) interfere in transcriptional and translational processes modulating gene expression reflecting environmental influences. Nevertheless, there is no systematic analysis available that investigates the impact of lncRNAs on PD. In the current study, we performed a comprehensive analysis on expression levels of 90 well-annotated lncRNAs in 30 brain specimens deriving from 20 PD patients and 10 controls as a preliminary report on the significance of lncRNAs in PD. Expression profiling of lncRNAs revealed that five lncRNAs are significantly differentially expressed in PD. While H19 upstream conserved 1 and 2 is significantly downregulated in PD, lincRNA-p21, Malat1, SNHG1, and TncRNA are significantly upregulated. An analysis on expression levels and PD stages revealed that the identified dysregulated lncRNA are altered already in early disease stage and that they precede the course of PD. In summary, this is the first comprehensive analysis on lncRNAs in PD revealing significantly altered lncRNAs. Additionally, we found that lncRNA dysregulations precede the course of the disease. Thus, the five newly identified lncRNAs may serve as potential new biomarkers appropriate even in early PD. They may be used in monitoring disease progression and they may serve as potential new targets for novel therapeutic approaches.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Parkinson’s disease (PD) is the most frequent neurodegenerative disorder altering the movement abilities [1]. This slowly progressing disease affects approximately seven million patients worldwide. In the population over 80 years, up to 4 % of persons get the disease [1]. It is estimated that there are 8 to 18 new PD cases per 100,000 persons per year [1].
Clinically, PD is characterized by progressive impairments of motoric abilities. Till date, the disease can only be treated inadequately: while the currently available drugs provide only a relief of the symptoms and a delay in progression, there are no causal therapies available. Despite intensive research, the molecular mechanisms initiating and promoting PD still remain unclear [2]. Recent research emphasizes that epigenomic factors modulating gene expression, transcription, and translation play a crucial role in the development of neurodegenerative diseases [2, 3]. Thus, it is assumed that the initiation is promoted by a combination of a genetic predisposition and environmental triggers [1--3].
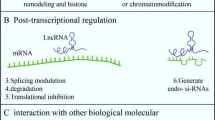
Recently, it has been revealed that the vast majority of genetic information coded within the DNA is transcribed into RNA. This “dark matter” of noncoding RNA (ncRNA) is not only “transcriptional noise” but interferes in transcription, translation, and numerous other cellular mechanisms [4--7]. Consisting of RNA molecules of 200 bp to >10 kbp in length, the group of long noncoding RNA (lncRNA) is assumed to be responsible for numerous regulatory processes [8--10].
The best studied events that are linked to lncRNAs are imprinting and X-chromosome inactivation [11--13]. They are regulated by cis-acting master control regions. In case of X-chromosome inactivation experiments have shown that a single X-inactivation center (Xic) is sufficient for X-chromosome inactivation [13, 14]. With regard to functions and mechanisms of lncRNAs, Xic contains the best characterized lncRNA: X-inactive specific transcript (Xist) [13, 14]. This 17–20-kbp-long RNA marks one X-chromosome in the course of inactivation [12]. Only the X-chromosome that is inactivated expresses Xist subsequently leading to the inactivation of the whole X-chromosome and the change of chromatin and transcription by binding Polycomb repressive complex 2 (PRC2) being responsible for H3K27 trimethylation [13, 15--17]. However, Xist itself is controls by other lncRNAs, e.g., Tsix and Jpx, leading to a complex regulatory mechanism [14, 18, 19]. It is obvious that abnormalities in these regulations can lead to severe diseases. Besides the importance of lncRNAs in the field of tumorigenesis [20, 21], lncRNAs are closely linked to the development of cognitive dysfunctions, autism, schizophrenia, and Alzheimer’s disease [22]. However, till date, there are no comprehensive studies available analyzing the significance of lncRNAs in Parkinson’s disease.
In the current study, we addressed this question and performed expression profiling of 90 well-annotated lncRNAs in neurons separated from anterior cingulate gyrus of 30 human postmortem brain specimens. Tissue was derived from healthy donors as well as PD patients. Finding severe dysregulations of lncRNAs, the data presented in this study is a preliminary report on lncRNAs in PD that will boost research on lncRNAs in PD and will contribute to better understand the molecular mechanisms underlying PD as well as to develop new therapeutic approaches.
Materials and Methods
Sample Collection and Preparation of Tissue Specimen
In this study, we selected 30 well-characterized human postmortem brain tissue samples that were stored at −80 °C. All samples were provided by the Neurobiobank Munich (NBM) and were clinically as well as neuropathologically classified according to the NBM standard protocols. The extent of Parkinson’s disease (PD)-associated changes was assessed according to the classification recommended by McKeith et al. [23]. Cases with hypoxia, inflammation, infection, infarction, and tumor were excluded. Written informed consent was obtained according to the guidelines of the local ethics committee. We included 10 control cases without PD and 20 cases with PD. Age distribution of controls ranged from 46 to 85 years with mean age of 66 years, and age distribution of PD cases ranged from 57 to 85 years with mean age of 74 years. Sex distribution showed 6 female and 4 male patients in the control cohort (female to male ratio of 1.5) and 11 female and 9 male patients in the PD cohort (female to male ratio of 1.2). Detailed information on patients can be found in Table 1. According to McKeith, PD can be assigned based on the pattern of Lewy-related pathology in brain stem, limbic, and neocortical regions [23]. Early PD cases show Lewy-related pathology only in brain stem regions. Subsequently, basal forebrain and limbic regions are affected (such as the cingulate gyrus). In late PD, Lewy-associated pathology can also be detected in neocortical regions. By selecting the cingulate gyrus as target region in this study, we were able to perform an analysis of lncRNA expression preceding the course of Parkinson’s disease as the anterior cingulate gyrus is affected in limbic and neocortical type PD but not in brain stem type PD. Additionally, whereas the substantia nigra shows a loss of neurons and reactive gliosis in very early PD with a raddled state during advancing of PD, the anterior cingulate gyrus shows an active state during a long time period. Thus, the anterior cingulate gyrus represents the ideal anatomical region to be studied in order to reveal molecular mechanisms preceding and promoting PD. As the cortex consists of a crude mixture of different cell types [24] and as molecular alterations are cell type specific [22], we performed an enrichment of neurons using a NeuN selective antibody as described previously [25] to further increase the validity of our approach.
Extraction of Long Noncoding RNAs, Reverse Transcription Reaction, and Quantitative Polymerase Chain Reaction
We performed enrichment of lncRNAs by using the miRNeasy Micro Kit (Qiagen) and the RNeasy MinElute Kit (Qiagen) according to the manufacturer’s protocols. Detecting the 260/280 nm absorbance ratio using a NanoDrop device (Thermo Fischer), we determined the quantity and quality of lncRNA. In all cases, the absorption ratio was between 1.90 and 2.20. To avoid amplification bias, no preamplification of lncRNA was performed and lncRNA was directly processed for subsequent analysis. Reverse transcription reaction (RT) was performed using the human LncProfiler qPCR Assay Kit (SBI) including polyadenylation reactions optimized for lncRNAs as described previously [26, 27]. We used equal amounts of 50 ng lncRNA with a concentration of 10 ng/μl. To boost cDNA yield, reverse transcription reactions were performed using Oligo dT primers and random primer. All procedures were performed in accordance with the manufacturer’s protocols. As a first approach to assess the importance of lncRNAs in PD, we selected the top 90 disease-associated lncRNAs that are annotated in the lncRNA database (lncRNAdb) [28]. These notable lncRNAs have already been described as key factors in a vast majority of human diseases such as cancers (ANRIL, HOTAIR, HOTAIRM1, MALAT1, MEG3, Tsix, Xist) [20, 29], neurdegeneration (BASE1AS, BC200, NEAT1) [30, 31], and neurodevelopmental/psychiatric diseases (Gomafu, NRON, Sox2ot) [32--35]. Analysis was performed using the predesigned and validated qPCR primer library set of the human LncProfiler qPCR Assay Kit (SBI). Predesigned assays have been validated by the supplier across numerous cell types for high specificity and robustness. Quantitative PCR (qPCT) was performed on a LightCycler 480 II device (Roche) using the SensiFAST SYBR No-ROX Kit (Bioline) in combination with standard protocols. The relative amount of cDNA was calculated using the comparative CT method (ΔΔCT) [36]. To enhance data quality, three valid normalizers were used according to the guidelines for real-time qPCR experiments [37].
Computational Data Analysis
Computational analysis of stably expressed lncRNAs was performed using NormFinder algorithm [38]. We determined the stability value (sv) for each lncRNA. High expression stability is reflected by a low sv, and low expression stability is reflected by a high sv. We assumed stable expression for stability values of ≤0.009. Only highly abundant lncRNAs with CT (cycle threshold) values of ≤ 32 were assumed as suitable references. Graphical data visualization by heat map generation was performed using the Gene-E software (http://www.broadinstitute.org/cancer/software/GENE-E/). Statistical analysis was performed applying unpaired t test. We using Prism 6 software suite (GraphPad) as statistical environment. Statistical significance was assumed for p values <0.05.
Results
Identification of Stably Expressed lncRNAs
In this study, we performed an analysis on 90 well-annotated lncRNAs that are annotated in the lncRNAdb [28]. As we explored neuronal enriched fractions of 30 human brain tissue samples of the anterior cingulate gyrus, we initially performed an analysis on lncRNA expression stability in order to identify stably expressed lncRNAs that serve as appropriate normalisers in the subsequent analysis.
Applying NormFinder algorithm [38] on 2700 individual expression data, the sv of each lncRNA was determined. Only lncRNAs with stability values of ≤0.009 were assumed as stably expressed. Additionally, we assumed only highly abundant lncRNAs with mean cycle threshold (CT) values of ≤32 as suitable references.
Applying these criteria, only 3 out of 90 explored lncRNAs fulfilled the requirements for suitable normalizers (Table 2 and Supplementary Table S1): Gas5-family (growth arrest specific 5) showed a sv of 0.007 and a mean CT of 31.05; highly accelerated region 1B (HAR1B) showed a sv of 0.008 and a mean CT of 31.23; and small nucleolar RNA host gene 4 (SNHG4) showed a sv of 0.009 and a mean CT of 31.75 (Table 2 and Supplementary Fig. S1). Thus, these three lncRNAs were regarded as appropriate references and used for normalization strategy in the subsequent analysis.
Detection of Significantly Dysregulated lncRNAs in Parkinson’s Disease
In order to compute the relative expression levels of the investigated lncRNAs, we applied the comparative CT method [36]. According to the guidelines for real-time qPCR experiments that were promoted by Abdel Nour et al., we used all three stably expressed lncRNAs (Gas5 family, HAR1B, and SNHG4) as normalizers to enhance data quality [37] in calculating expression levels (Fig. 1).
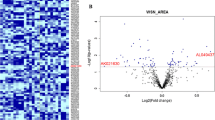
Heatmap showing expression levels of lncRNAs analyzed in this study. Data were normalized according to the comparative CT method using the three stably expressed references GAS5-family, HAR1B, and SNHG4. Low relative expression levels correspond with blue marks; high relative expression levels correspond with red marks. Donors 1 to 10 are controls; donors 11 to 30 are PD patients. Dnr donor (color figure online)
Statistical testing on expression data revealed that five lncRNAs were significantly differentially expressed in PD compared with controls (Fig. 2 and Supplementary Table S2): while only one lncRNA was significantly downregulated in PD, four lncRNAs were significantly upregulated in PD. The two ∼400-bp-long conserved elements H19 upstream conserved 1 and 2 (Huc 1 and 2) that can be found ∼10 kb of the 5′ end of the H19 gene showed a significant twofold decrease in PD compared with controls (p = 0.0142, unpaired t test, Fig. 2a). The lincRNA-p21 that is a transcription target of p53 and HIF1α showed a significant twofold increase in PD compared with controls (p = 0.0224, unpaired t test, Fig. 2b). The lncRNA metastasis-associated lung adenocarcinoma transcript 1 (Malat1) showed a significant threefold increase in PD compared with controls (p = 0.0475, unpaired t test, Fig. 2c). Small nucleolar RNA host gene 1 (SNHG1) showed a significant twofold increase in PD compared with controls (p = 0.0039, unpaired t test, Fig. 2d). Tiny noncoding RNA (TncRNA) showed a significant twofold increase in PD compared with controls (p = 0.0427, unpaired t test, Fig. 2e).
Normalized relative expression levels of differentially expressed lncRNAs. Indicated are the normalized relative expression levels of the five lncRNAs that were significantly differentially expressed in PD compared with controls. H19 upstream conserved 1 and 2 shows a significantly decreased expression in PD compared with controls (a). The lncRNAs lincRNA-p21 (b), Malat1 (c), SNHG1 (d), and TncRNA (e) show significantly increased expression levels in PD compared with controls. Indicated are mean and SEM (color figure online)
Thus, the five identified differentially expressed lncRNAs H19 upstream conserved, lincRNA-p21, Malat1, SNHG1, and TncRNA may be crucial for the understanding of the molecular mechanisms occurring in the disease and may serve as new targets for an advanced molecular therapy.
Altered Expression of lncRNAs Precedes the Course of Parkinson’s Disease
Parkinson’s disease is a neurodegenerative disorder that shows a distinct spread throughout the brain during the course of the disease. In order to perform analysis on the dynamics of lncRNAs during progression of PD, all cases analyzed in this study were classified according to the guidelines suggested by McKeith et al., which were revealed by the dementia with Lewy bodies (DLB) consortium [23]. The assignment of disease stage is based upon the pattern of Lewy-related pathology in brain stem, limbic, and neocortical regions [23]. Early PD cases show Lewy-related pathology only in brain stem regions (e.g., in the locus caeruleus and the substantia nigra). Subsequently, basal forebrain and limbic regions are affected (e.g., amygdala and the cingulate gyrus). In late PD, Lewy-associated pathology can also be detected in neocortical regions (e.g., frontal and occipital cortex). By selecting the cingulate gyrus as target region in this study, we were able to perform an analysis of lncRNA expression preceding the course of Parkinson’s disease.
Neuropathological staging of the 20 PD cases included in this study showed that 4 cases corresponded with brain stem type, 8 corresponded with limbic type, and 8 corresponded with neocortical type (Table 1). Performing stage-dependent analysis on expression profiles of the five lncRNAs that we identified as differentially expressed in PD (H19 upstream conserved, lincRNA-p21, Malat1, SNHG1, and TncRNA) revealed that alterations can already be detected in early, brain stem type PD (Fig. 3 and Supplementary Table S3). The significantly downregulated lncRNA H19 upstream conserved 1 and 2 was already downregulated in brain stem type PD and then remained at decreased levels in the progression of PD with a significantly decreased expression in neocortical PD compared with limbic PD (p = 0.0309, unpaired t test, Fig. 3a). In case of the significantly upregulated lincRNA-p21, we found that this lncRNA was already upregulated in early PD and remained at high levels during disease progression with a significantly increased expression in brain stem type PD (p = 0.0050, unpaired t test), limbic type PD (p = 0.0199, unpaired t test), and neocortical type PD compared with controls (p = 0.0399, unpaired t test, Fig. 3b). The significantly upregulated lncRNA Malat1 showed already increased expression in brain stem type PD, highest expression levels were reached in limbic PD cases with a significantly increased expression in limbic type PD compared with controls (p = 0.0027, unpaired t test), in neocortical type PD expression deceases compared with limbic type PD (p = 0.0291, unpaired t test, Fig. 3c). The significantly overexpressed SNHG1 lncRNA showed increased levels already in brain stem type PD cases, during the course of the disease, expression further increased with a significantly increased expression in brain stem type PD (p = 0.0262, unpaired t test), limbic type PD (p = 0.0072, unpaired t test), and neocortical type PD compared with controls (p = 0.0035, unpaired t test, Fig. 3d). The significantly overexpressed TncRNA showed already overexpression in early brain stem type PD cases, and the overexpression remained stable during the course of the disease a significantly increased expression in limbic type PD (p = 0.0247, unpaired t test) and neocortical type PD compared with controls (p = 0.0301, unpaired t test, Fig. 3e).
PD stage-dependent normalized relative expression levels of differentially expressed lncRNAs. Indicated are the PD stage-dependent normalized relative expression levels of the five lncRNAs that were significantly differentially expressed in PD compared with controls. PD stages were assigned to brain stem type, limbic type, and neocortical type according to the classification of McKeith et al. [23]. In early PD, pathological changes can be detected in brain stem regions only. During the course of the disease, basal forebrain and limbic regions, such as the cingulate gyrus, are affected. In late PD, there are also pathological changes in neocortical regions. The lncRNAs H19 upstream conserved 1 and 2 (a), lincRNA-p21 (b), Malat1 (c), SNHG1 (d), and TncRNA (e) show altered expression in the anterior cingulate gyrus during the course of PD. Indicated are mean and SEM
Summing up these findings, we revealed that the expression of differentially expressed lncRNAs was already altered in early PD, e.g. expression of TncRNA was already significantly increased in limbic type PD and remained at high levels during progression of PD, and expression of lincRNA-p21 and SNHG1 was already significantly increased in early brain stem type PD and then remained at overexpressed levels during the course of PD. As morphological changes in the investigated target brain region (anterior cingulate gyrus) can be detected earliest in limbic stage, altered expression of these identified lncRNAs precedes the course of Parkinson’s disease.
Discussion
Parkinson’s disease is the most frequent neurodegenerative disorder that shows altered movement abilities [1]. During the progression of PD, there is a spread of pathological synuclein deposits beginning in the brain stem and successively affecting limbic and neocortical regions in a distinct pattern [23]. However, the molecular mechanisms underlying and initiating PD still remain unknown. It is assumed that the initiation of neurodegenerative diseases is promoted by a combination of genetic predisposition and environmental influences [2, 3]. In this context, especially mechanisms modulating transcription and translation are supposed to be crucial in disease initiation and progression. The recent discovery of long noncoding RNAs opened up a new field of transcriptional and translational control [22].
Exploring this new field of lncRNA expression, the identification of suitable references that serve as normalizers is essential [37, 39, 40]. As there were no studies available analyzing neurons of the human anterior cingulate cortex, we explored 90 lncRNAs according to their expression stability using the NormFinder algorithm [38]. We identified three lncRNAs that are highly abundant and stably expressed in controls and PD samples and thus are appropriate as normalizers: GAS5-family, HAR1B, and SNHG4. Applying the comparative CT method and computational analysis, we found that five lncRNAs are significantly differentially expressed in PD. H19 upstream conserved 1 and 2 is significantly downregulated, and lincRNA-p21, Malat1, SNHG1, and TncRNA are significantly upregulated in PD. H19 upstream conserved 1 and 2 (Huc 1 and 2) are ∼400-bp-long conserved elements that are located ∼10 kb at the 5′ end of the H19 gene. It is proposed that these genomic regions interact with imprinting control regions and epigenetically regulated silencers and that the transcription products are overexpressed and stabilized in various human tissues [41, 42]. The identified lincRNA-p21 represents a transcription target of p53 and HIF1α, and recent research indicated that lincRNA-p21 regulates mRNA translation, gene expression, protein stability, as well as p53-dependent apoptosis and cell cycle arrest [43]. Hall et al. hypothesized that lincRNA-p21 represents a functional key role in cell cycle arrest and in apoptosis [43]. Furthermore, they found that loss of lincRNA-p21 resulted in the evasion of cellular apoptosis and cell cycle arrest [43]. The lncRNA Malat1 is reported to be highly abundant in neurons [31]. Experimental knockdown of Malat1 resulted in a decreased synaptic density while the overexpression showed a cell-autonomous increase in synaptogenesis [44]. In case of the lncRNA SNHG1, it was shown that SNHG1 promotes cellular proliferation and affects p53 stability as well as downstream p53 regulated pathways [45, 46]. TncRNA is hypothesized being a new target of TP53 that may play a role in mediating DNA damage response, but there is also evidence for associations with Malat1 lncRNA [47, 48]. Summing these findings up, the lncRNAs that we identified to be overexpressed in PD are reported being highly abundant in neurons and being closely linked with synaptogenesis (Malat1) as well as being linked with cellular proliferation, cell cycle control and apoptosis (lincRNA-p21, SNHG1, and TncRNA). Interestingly, analysis on lncRNA expression levels and disease stage revealed that altered expression of lncRNAs can already be detected in early PD, e.g. overexpression of TncRNA was already in limbic type PD statistically significant, and overexpression of lincRNA-p21 and SNHG1 was already in very early brain stem type statistically significant and remained at high levels during the course of the disease. As we analyzed the anterior cingulate gyrus, a brain region where morphological changes can be detected earliest in limbic stage PD, altered expression of these lncRNAs precedes the course of PD. Thus, these lncRNAs may serve as early biomarkers in PD.
In summary, we were able to identify five lncRNAs (H19 upstream conserved, lincRNA-p21, Malat1, SNHG1, and TncRNA) in this preliminary report that are differentially expressed in neurons of the anterior cingulate gyrus in PD brains. While H19 upstream conserved is significantly downregulated, lincRNA-p21, Malat1, SNHG1, and TncRNA are significantly upregulated in PD. It is worthwhile to mention that these lncRNAs have already been connected with synaptogenesis, cellular proliferation, and apoptosis. This is in good perception of the current notion of PD [2, 3, 31]. Interestingly, the expression of these lncRNAs precede the course of the disease: already in early brain stem type PD, altered lncRNA expression can be detected with lincRNA-p21 and SNHG1 showing significantly increased expression levels in brain stem type PD and remaining at high levels during the course of PD. It is noteworthy that the analyzed target region, the anterior cingulate gyrus, does not show morphological alterations in this early brain stem type PD according to the classification recommended by McKeith et al. [23]. Commonly, the cingulate gyrus is only affected in advancing PD, i.e., in limbic and neocortical type PD. Thus, it can be assumed that these newly identified differentially expressed lncRNAs may be appropriate as novel biomarkers. They may even serve as potential early indicators for PD. Thus, the data presented in this study will help to better understand the molecular mechanisms occurring during initiation and progression of PD. They will boost further research in the field of lncRNAs and neurodegeneration and will help to establish novel biomarkers and to develop new molecular approaches for molecularly targeted therapies.
References
de Lau LM, Breteler MM (2006) Epidemiology of Parkinson’s disease. Lancet Neurol 5:525–535. doi:10.1016/S1474-4422(06)70471-9
Noyce AJ et al (2012) Meta-analysis of early nonmotor features and risk factors for Parkinson disease. Ann Neurol 72:893–901. doi:10.1002/ana.23687
Van Maele-Fabry G, Hoet P, Vilain F, Lison D (2012) Occupational exposure to pesticides and Parkinson’s disease: a systematic review and meta-analysis of cohort studies. Environ Int 46:30–43. doi:10.1016/j.envint.2012.05.004
Carninci P et al (2005) The transcriptional landscape of the mammalian genome. Science 309:1559–1563. doi:10.1126/science.1112014
Derrien T et al (2012) The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res 22:1775–1789. doi:10.1101/gr.132159.111
Johnson JM, Edwards S, Shoemaker D, Schadt EE (2005) Dark matter in the genome: evidence of widespread transcription detected by microarray tiling experiments. Trends Genet 21:93–102. doi:10.1016/j.tig.2004.12.009
Mattick JS (2009) The genetic signatures of noncoding RNAs. PLoS Gen 5, e1000459. doi:10.1371/journal.pgen.1000459
Bu D et al (2012) NONCODE v3.0: integrative annotation of long noncoding RNAs. Nucleic Acids Res 40:D210–D215. doi:10.1093/nar/gkr1175
Ma L, Bajic VB, Zhang Z (2013) On the classification of long non-coding RNAs. RNA Biol 10
Mattick JS, Amaral PP, Dinger ME, Mercer TR, Mehler MF (2009) RNA regulation of epigenetic processes. BioEssays 31:51–59. doi:10.1002/bies.080099
Carrel L et al (1996) X inactivation analysis and DNA methylation studies of the ubiquitin activating enzyme E1 and PCTAIRE-1 genes in human and mouse. Hum Mol Genet 5:391–401
Clemson CM, McNeil JA, Willard HF, Lawrence JB (1996) XIST RNA paints the inactive X chromosome at interphase: evidence for a novel RNA involved in nuclear/chromosome structure. J Cell Biol 132:259–275
Lee JT, Bartolomei MS (2013) X-inactivation, imprinting, and long noncoding RNAs in health and disease. Cell 152:1308–1323. doi:10.1016/j.cell.2013.02.016
Lee JT, Davidow LS, Warshawsky D (1999) Tsix, a gene antisense to Xist at the X-inactivation centre. Nat Gen 21:400–404. doi:10.1038/7734
Penny GD, Kay GF, Sheardown SA, Rastan S, Brockdorff N (1996) Requirement for Xist in X chromosome inactivation. Nature 379:131–137. doi:10.1038/379131a0
Zhao J et al (2010) Genome-wide identification of polycomb-associated RNAs by RIP-seq. Mol Cell 40:939–953. doi:10.1016/j.molcel.2010.12.011
Zhao J, Sun BK, Erwin JA, Song JJ, Lee JT (2008) Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 322:750–756. doi:10.1126/science.1163045
Sun S, Del Rosario BC, Szanto A, Ogawa Y, Jeon Y, Lee JT (2013) Jpx RNA activates Xist by evicting CTCF. Cell 153:1537–1551. doi:10.1016/j.cell.2013.05.028
Tian D, Sun S, Lee JT (2010) The long noncoding RNA, Jpx, is a molecular switch for X chromosome inactivation. Cell 143:390–403. doi:10.1016/j.cell.2010.09.049
Nie L et al (2012) Long non-coding RNAs: versatile master regulators of gene expression and crucial players in cancer. Am J Transl Res 4:127–150
Prensner JR, Chinnaiyan AM (2011) The emergence of lncRNAs in cancer biology. Cancer Discovery 1:391–407. doi:10.1158/2159-8290.CD-11-0209
Schonrock N, Gotz J (2012) Decoding the non-coding RNAs in Alzheimer’s disease. Cell Mol Life Sci 69:3543–3559. doi:10.1007/s00018-012-1125-z
McKeith IG et al (2005) Diagnosis and management of dementia with Lewy bodies: third report of the DLB Consortium. Neurology 65:1863–1872. doi:10.1212/01.wnl.0000187889.17253.b1
Pelvig DP, Pakkenberg H, Stark AK, Pakkenberg B (2008) Neocortical glial cell numbers in human brains. Neurobiol Aging 29:1754–1762. doi:10.1016/j.neurobiolaging.2007.04.013
Wagner M et al (2015) Age-Dependent Levels of 5-Methyl-, 5-Hydroxymethyl-, and 5-Formylcytosine in Human and Mouse Brain Tissues. Angew Chem. doi:10.1002/anie.201502722
Kraus TF, Greiner A, Guibourt V, Kretzschmar HA (2014) Long non-coding RNA normalisers in human brain tissue. J Neural Transm. doi:10.1007/s00702-014-1352-6
Kraus TF, Greiner A, Guibourt V, Lisec K, Kretzschmar HA (2015) Identification of stably expressed lncRNAs as valid endogenous controls for profiling of human glioma. J Cancer 6:111–119. doi:10.7150/jca.10867
Amaral PP, Clark MB, Gascoigne DK, Dinger ME, Mattick JS (2011) lncRNAdb: a reference database for long noncoding RNAs. Nucleic Acids Res 39:D146–D151. doi:10.1093/nar/gkq1138
Tang L, Zhang W, Su B, Yu B (2013) Long noncoding RNA HOTAIR is associated with motility, invasion, and metastatic potential of metastatic melanoma. BioMed Res Intern 2013:251098. doi:10.1155/2013/251098
Tan L, Yu JT, Hu N, Tan L (2013) Non-coding RNAs in Alzheimer’s disease. Mol Neurobiol 47:382–393. doi:10.1007/s12035-012-8359-5
Wu P, Zuo X, Deng H, Liu X, Liu L, Ji A (2013) Roles of long noncoding RNAs in brain development, functional diversification and neurodegenerative diseases. Brain Res Bull 97:69–80. doi:10.1016/j.brainresbull.2013.06.001
Barry G (2014) Integrating the roles of long and small non-coding RNA in brain function and disease. Mol Psychiatry 19:410–416. doi:10.1038/mp.2013.196
Barry G et al (2014) The long non-coding RNA Gomafu is acutely regulated in response to neuronal activation and involved in schizophrenia-associated alternative splicing. Mol Psychiatry 19:486–494. doi:10.1038/mp.2013.45
van de Vondervoort II et al (2013) Long non-coding RNAs in neurodevelopmental disorders. Front Mol Neurosci 6:53. doi:10.3389/fnmol.2013.00053
Ziats MN, Rennert OM (2013) Aberrant expression of long noncoding RNAs in autistic brain. J Mol Neurosci 49:589–593. doi:10.1007/s12031-012-9880-8
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 3:1101–1108
Abdel Nour AM, Azhar E, Damanhouri G, Bustin SA (2014) Five years MIQE guidelines: the case of the Arabian countries. PLoS One 9, e88266. doi:10.1371/journal.pone.0088266
Andersen CL, Jensen JL, Orntoft TF (2004) Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res 64:5245–5250. doi:10.1158/0008-5472.CAN-04-0496
Gao Q et al (2008) Selection of reference genes for real-time PCR in human hepatocellular carcinoma tissues. J Cancer Res Clin Oncol 134:979–986. doi:10.1007/s00432-008-0369-3
Langnaese K, John R, Schweizer H, Ebmeyer U, Keilhoff G (2008) Selection of reference genes for quantitative real-time PCR in a rat asphyxial cardiac arrest model. BMC Mol Biol 9:53. doi:10.1186/1471-2199-9-53
Berteaux N et al (2008) A novel H19 antisense RNA overexpressed in breast cancer contributes to paternal IGF2 expression. Mol Cell Biol 28:6731–6745. doi:10.1128/MCB.02103-07
Drewell RA, Arney KL, Arima T, Barton SC, Brenton JD, Surani MA (2002) Novel conserved elements upstream of the H19 gene are transcribed and act as mesodermal enhancers. Development 129:1205–1213
Hall JR, Messenger ZJ, Tam HW, Phillips SL, Recio L, Smart RC (2015) Long noncoding RNA lincRNA-p21 is the major mediator of UVB-induced and p53-dependent apoptosis in keratinocytes. Cell Death Dis 6, e1700. doi:10.1038/cddis.2015.67
Bernard D et al (2010) A long nuclear-retained non-coding RNA regulates synaptogenesis by modulating gene expression. EMBO J 29:3082–3093. doi:10.1038/emboj.2010.199
You J et al (2014) Noncoding RNA small nucleolar RNA host gene 1 promote cell proliferation in nonsmall cell lung cancer. Indian J Cancer 51(Suppl 3):e99–e102. doi:10.4103/0019-509X.154092
Yu F et al (2015) p53 Represses the Oncogenic Sno-MiR-28 Derived from a SnoRNA. PLoS One 10, e0129190. doi:10.1371/journal.pone.0129190
Adamsen BL, Kravik KL, Clausen OP, De Angelis PM (2007) Apoptosis, cell cycle progression and gene expression in TP53-depleted HCT116 colon cancer cells in response to short-term 5-fluorouracil treatment. Int J Oncol 31:1491–1500
Hutchinson JN, Ensminger AW, Clemson CM, Lynch CR, Lawrence JB, Chess A (2007) A screen for nuclear transcripts identifies two linked noncoding RNAs associated with SC35 splicing domains. BMC Genomics 8:39. doi:10.1186/1471-2164-8-39
Acknowledgments
We thank the Neurobiobank Munich (Thomas Arzberger) for providing human brain tissue.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This work was supported by the German Federal Ministry of Education and Research (BMBF) through the EpiPD (Epigenomics of Parkinson’s disease) project, under the auspices of the bilateral BMBF/ANR (French National Research Agency) Epigenomics of Common and Age-related Diseases Programme (grant no. 01KU1403B to TFJK and HAK) and by the BMBF through the Integrated Network IntegraMent (Integrated Understanding of Causes and Mechanisms in Mental Disorders), under the auspices of the e:Med Programme (grant no. 01ZX1314I to TFJK and HAK).
Conflict of Interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
Overview on the expression stability of all 90 analysed lncRNAs. Indicated are stability values, intragroup and intergroup variations as calculated using the NormFinder algorithm. Stability values of ≤ 0.009 and CT values of ≤ 32 are indicated in grey. Only the 3 lncRNAs GAS5-family, HAR1B and SNHG4 that are indicated in grey fulfill the requirements of valid normalisers. (XLSX 18 kb)
Table S2
Relative expression levels of lncRNAs. Indicated are the relative expression levels of 87 unstably expressed lncRNAs as calculated using the comparative CT method. As references, we used the three stably expressed lncRNAs GAS5-family, HAR1B and SNHG4. Only 5 lncRNAs showed significant expression differences with p-values of < 0.05 (indicated in grey) in PD compared with controls: H19 upstream conserved 1 and 2, lincRNA-p21, Malat1, SNHG1, and TncRNA (indicated in grey). (XLSX 17 kb)
Table S3
Expression levels of candidate lncRNAs in PD stages. Classifying PD cases according to McKeith enables us to perform PD stage dependent analysis. Indicated are mean relative expression levels of the 5 identified candidate lncRNAs H19 upstream conserved 1 and 2, lincRNA-p21, Malat1, SNHG1, and TncRNA in controls, brain stem type PD, limbic type PD and neocortical type PD. (XLSX 9 kb)
Figure S1
Stably expressed lncRNAs. Indicated are the expression levels of the 3 identified valid lncRNA normalisers GAS5-family, HAR1B and SNHG4. Displayed are the CT values. Indicated are mean and SEM. (GIF 719 kb)
Rights and permissions
About this article
Cite this article
Kraus, T.F.J., Haider, M., Spanner, J. et al. Altered Long Noncoding RNA Expression Precedes the Course of Parkinson’s Disease—a Preliminary Report. Mol Neurobiol 54, 2869–2877 (2017). https://doi.org/10.1007/s12035-016-9854-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-016-9854-x