Abstract
Nerve growth factor (NGF) promotes pleiotropic gene transcription-dependent biological effects, in neuronal and non-neuronal cells, including survival, proliferation, differentiation, neuroprotection, pain, and angiogenesis. It is hypothesized that during odontogenesis, NGF may be implicated in morphogenetic and mineralization events by affecting proliferation and/or differentiation of dental cells. Tuftelin belongs to the enamel associated teeth proteins and is thought to play a role in enamel mineralization. We previously reported that tuftelin transcript and protein, which are ubiquitously expressed in various tissues of embryos, adults, and tumors, were significantly upregulated during NGF-induced PC12 differentiation. To further confirm the involvement of tuftelin in the differentiation process, we established a tuftelin-knockdown neuronal PC12 cell model, using a non-cytotoxic siRNA directed towards sequences at the 3′ UTR of the tuftelin gene. Using real-time PCR, we quantified tuftelin mRNA expression and found that tuftelin siRNA, but not scrambled siRNA or transfection reagents, efficiently depleted about 60% of NGF-induced tuftelin mRNA transcripts. The effect of tuftelin siRNA was quantified up to 6 days of NGF-induced differentiation. Using immunofluorescence and western blot analyses, we also found a direct correlation between reduction of 60–80% in tuftelin protein expression and inhibition of about 50–70% in NGF-induced differentiation of the cells, as was detected after 3–6 days of treatment. These results demonstrate an important role for tuftelin in NGF-induced differentiation of PC12 cells. Tuftelin could be a useful target for drug development in disease where neurotrophin therapy is required.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nerve growth factor (NGF) is widely recognized as a target tissue-derived growth factor, promoting a variety of cellular physiological processes including survival, proliferation, differentiation, neurotransmission, neuroprotection, pain, and angiogenesis during embryonic development and in the adult (Skaper 2017). These activities are not limited to certain subsets of peripheral sensory and sympathetic neurons and basal forebrain cholinergic neurons but also occur in non-neuronal cells such as fibroblasts, hematopoietic, epithelial, endothelial, and smooth muscle cells (Aloe et al. 2015; Skaper 2017). NGF promotes these activities by transcription-dependent and transcription-independent mechanisms in both neuronal (Cho et al. 1989; Chung et al. 2010; Greene and Angelastro 2005) and non-neuronal target cells (Arien-Zakay et al. 2009; Dutta et al. 2011; Lin et al. 2016). Therefore, understanding the mechanism of NGF physiological activities requires identification of the genes and their encoded proteins, which are subjected to regulation by this neurotrophin. Cellular models which can be tested before and after NGF treatment were established for detection of these genes. A majority of NGF-induced gene regulation studies have employed the PC12 pheochromocytoma cell line (Dijkmans et al. 2008; Levi et al. 1989; Vician et al. 1997). These cells, originated from a tumor of the adrenal gland, respond to NGF by changing their morphological phenotype from proliferating chromaffin-like cells into non-proliferative cells with neurite outgrowths, resembling sympathetic neurons (Fujita et al. 1989; Rudkin et al. 1989). NGF-induced morphological differentiation, expressed by neurite outgrowth, is a key cellular process involving a cascade of complex intracellular signaling during neuronal differentiation (Vaudry et al. 2002). This process includes initiation of neurite outgrowth (sprouting), elongation of neurites over long distances, and synapse formation followed by functional maturation (Fujita et al. 1989). Using different experimental approaches, hundreds of genes encoding proteins with established neuronal functions, which respond to NGF treatment, have been identified, including immediate early genes, intermediate, as well as those that are regulated relatively late in the NGF-induced differentiation process (Angelastro et al. 2000; D'Onofrio et al. 2011; Leonard et al. 1987). Despite this impressive progress, many additional NGF-responsive genes remain to be identified. It has been estimated that 5–10% of the genes expressed in PC12 cells may be NGF-regulated (Angelastro et al. 2000), which would suggest more than 1000 transcripts involved. Detecting and identifying these transcripts are crucial in order to understand how NGF exerts its multiple physiological activities and should provide valuable new information, in particular on how NGF and other neurotrophins induce neuronal differentiation.
The Tuftelin gene (TUFT1), a highly conserved gene (Delgado et al. 2017) mapped to chromosome 1q21-31, contains 13 exons, encoding an acidic glycoprotein of 390 amino acids, with a molecular mass of 44.3 kDa, belonging to the enamel associated teeth proteins (Deutsch et al. 1997; Mao et al. 2001). The timing of tuftelin protein expression in the developing tooth (just before enamel mineralization), its accumulation at the dentin-enamel-junction, and its acidic nature suggested that tuftelin might be involved in the initial stages of enamel mineralization, perhaps by nucleation (Deutsch et al. 1991; Deutsch et al. 1998). Several alternatively spliced tuftelin mRNA transcripts have been identified in human and rodent tissues. The 1.6 kb tuftelin promoter contains several transcription factor binding sites, including motifs for regulation of expression by activation proteins (activator protein 1 and stimulatory protein 1), as well as for hypoxia inducible factors (HIFs) (Leiser et al. 2011; Mao et al. 2001). Expression of tuftelin mRNA and protein was detected in early embryonic tissues as well as various adult normal and cancerous soft tissues (Mao et al. 2001; Deutsch et al. 2002; Leiser et al., 2007). Tuftelin gene expression was found to be elevated in kidney and testis compared to brain, liver, lung, and eye (Leiser et al. 2007). The kidney and testis are hovering closely to hypoxia during normal physiological conditions. Tuftelin expression was also upregulated in mesenchymal stem cells and in PC12 cells during hypoxia (Deutsch et al. 2011; Leiser et al. 2011). In a previous study, we found that during NGF-induced PC12 cell differentiation, tuftelin expression was significantly induced in correlation with the morphological differentiation expressed by neurite outgrowth elongation. This induction was selectively inhibited by an antagonist of the NGF receptor TrkA (Leiser et al. 2011). Therefore, in the present study, we sought to further establish the role of tuftelin in NGF-induced differentiation using a “loss of function” approach. Employing a selective siRNA directed towards the tuftelin gene, a causal, temporal, and significant direct relationship between tuftelin protein expression and NGF-induced PC12 cell neurite outgrowths was found, further supporting the hypothesis of tuftelin involvement in NGF-induced differentiation.
Materials and Methods
Cell Culture
PC12 cells were grown in Falcon T-200 culture flasks and 24-well culture plates or for microscopy in Nunc™ TC-Dish, 92 × 17 (Roshilde, Denmark) with 4.5 mg/l DMEM, supplemented with 7% fetal calf serum, 7% horse serum, 100 μg/ml streptomycin, and 100 U/ml penicillin as previously described (Leiser et al. 2011). Cultures were maintained at 37 °C in a humidified atmosphere of 6% CO2. Medium was changed twice weekly, and the cultures were split once a week at 1:6 ratio using gentle tapping to remove adherent cells. In the experiments involving the use of tuftelin siRNA, about 2 × 105 cells were used, according to the manufacturer’s instructions (Ambion, Austin, TX), to maximize the transfection efficiency. In experiments involving tuftelin induction by NGF, about 4 × 105 cells were used to allow enough space for neurite outgrowths elongation. Cells were suspended in DMEM and plated on tissue culture dishes coated with 0.1 mg/ml rat tail type-I collagen (Be’it Ha’emek, Israel). For immunofluorescence tracking of neurite extension, 5 × 103 cells were cultured on coverslips in a 24-well Nunc™ TC-Dish. Following 72 h of incubation, the medium was changed in the 96 h and 144 h groups and a new medium containing transfection reagent and the appropriate siRNA was added to the cells.
Neuronal Differentiation Analysis
In mouse βNGF, high analytical grade (50 ng/ml; Alomone Labs, Jerusalem, Israel) was applied daily to PC12 cell culture to induce differentiation. Cells were harvested after 24, 48, 72, 96, and 144 h of exposure to NGF and tuftelin siRNA. Neurite outgrowths were measured using the SigmaScanPro-5.0 (Systat Software, San Jose, CA, USA). By dividing the total length of neurites of a specific cell by its diameter, a parameter termed elongation factor (Ef) was generated. Percentage responsive cell (PRC) was calculated, reflecting the percentage of cells which exhibit neurites longer than twice the cell diameter. The experiments were performed three to five times in duplicates. In each well, three random areas were photographed. In each area, 15–25 cells were analyzed (n = 90–250 cells). Results of siRNA treated cultures were compared to PC12 cell cultures supplemented with NGF (positive control), PC12 cells treated with transfection reagents (negative control), or cells transfected with scrambled siRNA (second negative control).
RNA Isolation, Reverse Transcription and Real-Time PCR
RNA isolation was performed using the TRI-REAGENT standard protocol (TRI-REAGENT RNA/DNA/protein isolation reagent, Molecular Research Center, Cincinnati, OH, USA). Total RNA from PC12 cells was extracted using 1 ml of TRI-REAGENT per 10 cm2 culture dish, according to the manufacturer’s protocol. Total RNA concentration was determined using the NanoDrop ND-1000 spectrophotometer (Nano-Drop Technologies, Wilmington, DE, USA). Total RNA was subjected to reverse transcription according to the manufacturer’s protocol using the High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems®, Foster City, CA, USA).
Real-time quantitative PCR was performed using Applied Biosystems 7300 Real-Time-PCR System with Applied Biosystems TaqMan® AOD (Assays-On-Demand) and Master Mixes (TaqMan®, Gene Expression Assays). AOD used for PC12 cells (rat origin); GAPDH = RN9999996_s1, Tuftelin [TUFT1] = RN01762781_M1. The results were normalized to GAPDH.
Confocal Laser Scanning Microscopy
Cells were fixed with 4%-paraformaldehyde, permeabilized with Triton X-100, and blocked in goat-serum. The primary antibodies, anti-tuftelin polyclonal rabbit antibody (LF-74) as previously described (Deutsch et al. 1991), and anti-tubulin monoclonal mouse antibody (cytoskeleton-marker, anti-βIII-tubulin, MAB1637, Chemicon (Millipore), Billerica, MA, USA) diluted in PBS (1:200 and 1:1000 accordingly) were incubated overnight at 4 °C followed by incubation for 2 h at room temperature with secondary antibodies (diluted 1:200 in PBS). The secondary antibodies used were goat anti-rabbit conjugated to Cy5 (red) and goat anti-mouse conjugated to Alexa-Fluor-488 (green). Nuclear staining was performed with DAPI diluted 1:1000 in PBS for 3 min (blue staining). Cells were then rinsed, mounted, and examined by laser confocal microscopy at a magnification of × 600 (LSM 410; Zeiss, Germany).
Western Blotting and Protein Bands Densitometry
Proteins were extracted from PC12 cells subjected to siRNA transfection or controls, using TRI-REAGENT. Thirty micrograms of total protein, determined using the Bicinchoninic Acid Assay (Pierce Chemical Company, Rockford, IL, USA), was loaded onto 4–20% Tris-glycine precast gels (Novex, Invitrogen, Carlsbad, CA, USA). SDS–PAGE was performed using a mini-cell apparatus (Invitrogen) at 150 V for 50 min using Tris-running buffer (25 mM Tris pH 8.8, 1% (v/v) SDS, 0 .2M glycine). Following electrophoresis, the proteins were transferred onto Protran® nitrocellulose membranes (Schleicher & Schuell BioScience, Dassel, Germany) by electric blotting (Bio-Rad apparatus; Bio-Rad, Munich, Germany) at 19 mA for 60 min in ice. The nitrocellulose membranes were incubated in “superblotto” buffer (10% (v/v) PBS, 1 M glucose, 3% (w/v) BSA, 1% (w/v) milk powder, 10% (v/v) glycerol, 0.5% (v/v) Tween-20™) and shaken at 37 °C for 2 h. The membranes were then incubated under shaking overnight at room temperature with tuftelin polyclonal antibody LF-74 (Deutsch et al. 1991), diluted 1:200 in 1% (w/v) gelatine–Tris-buffered saline, followed by incubation in alkaline phosphatase-conjugated anti-rabbit IgG (Promega, Madison, WI, USA; diluted 1:7500) for 2 h at room temperature. Visualization was performed with BCIP/NBT reagents (Promega, Madison, WI, USA). Tuftelin protein band intensity was determined by densitometry using the Tina-2.0 software and calculated as net values of protein band optical density after subtraction of background optical density. β-Actin was used to normalize the quantity of the tuftelin protein bands.
Knockdown of Tuftelin Gene Expression Using siRNA
The commercial Amine Transfection Agent protocol was followed (Ambion, Austin, TX, USA). SiRNA SiPORT™ Amine–Polyamine–Based Transfection Agent is a propriety blend of polyamines formulated for transfection of small RNAs. The reagent functions by complexing with RNA and facilitating its transfer into the cells. In brief, 10 nM of Silencer® Select Pre-designed tuftelin siRNAs s172375 and s172376 cat#4390771 or Silencer® Select Negative Control (scrambled) siRNA cat#4390843 (Ambion), diluted in Gibco™ OPTI-MEM®I (Invitrogen, NY, USA), was combined with siPORT™ Amine Transfection Agent (cat#AM4502) diluted in Gibco™ OPTI-MEM®I. This mixture was added to a 6-well NUNC™ TC plate containing 2 × 105 PC12 cells per well. While studying NGF-induced differentiation, cells were subjected to daily treatment with mouse βNGF (50 ng/ml), starting 12 h after treatment with the siRNAs, and up to 6 days (144 h) post transfection, followed by RNA extraction and RT-PCR analyses. Following 72 h of incubation, a new transfection reagent was added, containing tuftelin or scrambled siRNA in the appropriate groups.
Cytotoxicity Measured by LDH Assay
Necrosis of PC12 cells following siRNA treatment was estimated by the rate of lactate dehydrogenase (LDH) leakage from the cells into the medium, as previously described (Tabakman et al. 2002). LDH activity was determined using a Power-WaveXTM (Bio-Tek Instruments®, USA) ELISA reader at 490 nm by following the rate of conversion of oxidized nicotinamide adenine dinucleotide (NAD+) to the reduced form (NADH). Total LDH (extracellular + intracellular) was obtained by freezing and thawing the cultures. The basal LDH (background) release was measured in cultures of untreated cells and was subtracted from all the experimental values.
Statistical Analysis
The experiments were independently repeated three to five times, in duplicates or triplicates (n = 6–15). The data, expressed as means ± SE, was analyzed by ANOVA. When only two groups were compared, the two-tailed Student’s t test for paired observations was used. Values are presented as mean ± SE; p values less than 0.05 were considered as statistically significant.
Results
Tuftelin siRNAs: Design, Safety, and Silencing of NGF-Induced Tuftelin mRNA Expression
In our previous studies, we have shown that NGF-induced differentiation is correlated with upregulation of tuftelin gene in PC12 cells (Leiser et al. 2011). NGF-induced tuftelin mRNA expression was silenced using two different siRNA probes to examine their efficacy. Both siRNAs were directed towards sequences in the 3′ UTR of the tuftelin gene (Fig. 1a), and efficacy of silencing was evaluated 72 h post transfection, using real-time PCR. The mean reduction in tuftelin mRNA expression using tuftelin siRNA1 as compared to the scrambled siRNA was over 60%, while tuftelin siRNA2 brought about less than 35% reduction in tuftelin expression (Fig. 1a). Hence, tuftelin siRNA1 was used in the following experiments. siRNAs were found effective for 12-72 h (data not shown). To evaluate the biological effect of tuftelin inhibition on NGF-induced PC12 differentiation, we had to extend the effect of tuftelin siRNAs. For this purpose, we established a treatment protocol in which the cultures received a second dose of siRNAs 72 h after the first transfection. Quantification of tuftelin expression in PC12 cells using real-time PCR was conducted 24, 48, 72, 96, and 144 h post-transfection with tuftelin siRNA1, scrambled siRNA (negative control), or transfection reagent alone (second negative control, termed “no treatment”), all followed by daily supplementation with 50 ng/ml NGF. Tuftelin siRNA significantly reduced tuftelin expression at 48, 72, 96, and 144 h post transfection and NGF application, as compared to the controls (Fig. 1b).
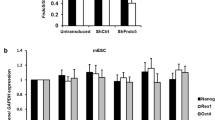
Tuftelin siRNA significantly reduced NGF-induced tuftelin mRNA expression in PC12 cells. a Inhibition of tuftelin mRNA expression by siRNA1 and siRNA2 as compared to scrambled siRNA, 72 h post infection. Upper right corner contains a schematic representation of the tuftelin gene and location of the siRNAs in the 3′ UTR. b Quantification of tuftelin mRNA levels, using real-time PCR, in PC12 cells subjected to siRNA prior to treatment with NGF for 24–144 h; untreated cells (black), scrambled siRNA (gray), and tuftelin siRNA1 (white). c Effect of tuftelin siRNA and NGF treatment on the viability of cultures. Cell death was calculated by measuring lactate dehydrogenase release in the medium of cell cultures treated with NGF and tuftelin siRNA. Transfection medium but no siRNA (black): scrambled siRNA (gray); tuftelin siRNA (white). Cytotoxicity was measured at 48, 72, and 144 h post treatment. *p value < 0.05 as compared to scrambled siRNA at each time point. mRNA levels were normalized to GAPDH
To verify the safety of siRNA transfection, LDH was measured at different time intervals from 48 to 144 h after treatment with transfection medium but no addition of siRNA or with siRNA transfections (with scrambled siRNA and with siRNA1) followed by daily NGF supplementation. No significant differences were detected between the different groups at each time point evaluated, indicating the safety of the siRNA and transfection medium (Fig. 1c). Thus, tuftelin siRNA1 significantly reduced NGF-induced tuftelin mRNA levels, in an efficient and safe manner, during 6 days of treatment with NGF in PC12 cells.
Tuftelin siRNA-Mediated Silencing of NGF-Induced Tuftelin Protein Expression
We assessed the effect of tuftelin siRNA1 on NGF-induced tuftelin protein expression by Western blot analysis. Quantification of tuftelin protein expression revealed 4.6 and 4.3-fold reduction in tuftelin protein expression, 72 and 96 h post transfection with tuftelin siRNA1 followed by daily NGF supplementation, as compared to transfection with scrambled siRNA, respectively. Treatment with scrambled siRNA and transfection medium without any siRNA (no treatment) yielded similar results (Fig. 2).
Tuftelin siRNA1 reduced tuftelin protein expression in PC12 cells treated with NGF; Western blot evaluation. a Western blot showing tuftelin protein expression compared to β-actin. Proteins were extracted 72 and 96 h post siRNA treatments or treatment only with transfection medium (no treatment) followed by NGF supplementation to PC12 cell cultures. b Quantification of tuftelin protein expression as compared to β-actin from the experiments described in a. Black—untreated PC12 cells (treated only with transfection medium) supplemented with NGF. Gray—PC12 cells treated with scrambled siRNA and supplemented with NGF compared to untreated cells. White—PC12 cells treated with tuftelin siRNA1 and supplemented with NGF relative to untreated cells. *p value < 0.05 as compared to scrambled siRNA
In a second approach, tuftelin protein expression was evaluated by immunofluorescence following treatment with tuftelin siRNA1 for 24, 48, 72, 96, and 144 h and daily supplementation of NGF (Fig. 3a, b). Quantification of intensity of immunofluorescence staining showed 5.9-, 2.9-, and 2.6-fold reduction in tuftelin protein expression after application of tuftelin siRNA for 72, 96, and 144 h, respectively (Fig. 3c). Thus, siRNA efficiently and safely silenced NGF-induced tuftelin protein levels during 6 days of treatment in PC12 cells.
Tuftelin siRNA1 reduced tuftelin protein expression in PC12 cells treated with NGF; immunofluorescence staining evaluation. a Tuftelin expression (red staining) was visualized, using anti-tuftelin antibody, in PC12 cells treated with tuftelin siRNA1 followed by NGF supplementation. b Same figures as in a with the additional staining of neuronal cytoskeleton biomarker—anti-tubulin antibody (green staining) and nuclei staining using DAPI (blue staining). c Quantification of tuftelin expression in PC12 cells treated with tuftelin siRNA followed by NGF supplementation, measured by fluorescence intensity normalized to the background. Bar represents 20 μm. *p value < 0.05 as compared to no treatment (0 h)
siRNA-Mediated Silencing of Tuftelin Expression Significantly Reduced NGF-Induced Neurite Outgrowth
Next, we evaluated the effect of siRNA-mediated silencing of tuftelin expression on NGF-induced differentiation. We focused on the morphological differentiation of PC12 cells, examined by neurite outgrowth. Figure 4 clearly indicates that NGF-induced neurite outgrowth, evaluated either by percentage responsive cells (PRC) (Fig. 5a) or elongation factor, Ef (Fig. 5b), was significantly reduced in cells transfected with tuftelin siRNA1, 72, 96, and 144 h after transfection followed by daily NGF supplementation. A significant decrease of 2.9-, 2-, and 1.9-fold in Ef at 72, 96, and 144 h after treatment, respectively, was obtained in cells treated with tuftelin siRNA compared to scrambled siRNA followed by daily NGF supplementation (Fig. 5b). A similar inhibitory effect of reduction in neurite outgrowth was observed by measuring PRC after treatment with tuftelin siRNA followed by NGF (Fig. 5a). A correlation coefficient of 0.83 (Fig. 5c) characterizes the relationship between NGF-induced neurite outgrowth (Fig. 5b) and tuftelin mRNA levels (Fig. 1b). These findings confirm that tuftelin is involved in initiating the NGF-induced neurite outgrowth and indicates its important role in NGF-induced differentiation.
Tuftelin siRNA1 inhibition of NGF-induced neurite outgrowth and of tuftelin protein expression in PC12 cells. a Light microscopy. b Immunofluorescence using anti-tuftelin antibody (red staining), antibody against a neuronal cytoskeleton biomarker—anti-tubulin antibody (green staining) and nuclei staining using DAPI (blue staining). I, untreated. II–VI, treatment with scrambled siRNA followed by daily NGF supplementation. VII–XI, treatment with tuftelin siRNA1 followed by daily NGF supplementation. I—0 h. II, VII—24 h. III, VIII—48 h. IV, IX—72 h. V, X—96 h. VI, XI—144 h post treatment. Bar 20 μm
Tuftelin siRNA1 reduced NGF-induced differentiation of PC12 cells. Neurite elongation was measured in cultures exposed to tuftelin siRNA or scrambled siRNA followed by daily NGF supplementation: black—untreated cells (treated only with transfection medium), gray—scrambled siRNA, and white—tuftelin siRNA. a Results are presented as percentage of cells which exhibit neurites longer than cell diameter (percentage responsive cells, PRC). b Results are presented as fold increase of neurite length compared to cell diameter (elongation factor, Ef). c Correlation between neurite elongation, taken from Fig. 5b, and tuftelin mRNA levels, taken from Fig. 1b—dotted line represents the elongation factor, and full line represents tuftelin mRNA levels. Results represent the relationship between cells treated with tuftelin siRNA1 and cells treated with scambled siRNA. Correlation coefficient = 0.83. *p value < 0.05
Discussion
Our previous attempts, made to understand the role of tuftelin in the PC12 neuronal cell model, found that this protein is upregulated during hypoxia (Leiser et al. 2011) and by NGF (Leiser et al. 2011). NGF is a well-known neurotrophin-inducing PC12 differentiation, and therefore, molecules such as tuftelin, which are upregulated by NGF, are likely candidates for the transient siRNA mediated silencing. To determine whether tuftelin plays a role in NGF-induced morphological differentiation, or its induction during NGF differentiation is a consequence of the differentiation process, we used siRNA to silence tuftelin expression in PC12 cells. The effect of siRNA on NGF-induced tuftelin mRNA expression was detected already 24 h after transfection while the effect on tuftelin protein levels was evident at an onset of 72 h after transfection. NGF-induced differentiation in wild-type PC12 cells is usually measured by neurite outgrowth 5–7 days after initiation of treatment (Katzir et al. 2002). Since the effect of transfection with siRNAs is maintained up to 72 h, it was required to prolong the effect of tuftelin siRNA by transfection with a second dose, 72 h after the first transfection. Our present data indicates a direct correlation between tuftelin expression and NGF-induced neurite outgrowth formation, initiation and elongation, providing convincing evidence that tuftelin is involved in NGF-induced commitment for differentiation of PC12 cells. Thus, our results contribute towards understanding of the physiological role of tuftelin and contribute a new gene target in the mechanism of NGF-induced differentiation.
NGF has a major role not only during neuronal embryonic development but also in the immune and cardiovascular systems, as well as in the craniofacial complex development and function (Amano et al. 1999; Lazarovici et al. 2006; Nosrat et al. 2002). NGF and its receptors are expressed in the developing rodent (Mitsiadis and Pagella 2016; Yan and Johnson 1988) and human teeth (Mitsiadis and Pagella 2016), suggesting that NGF has multiple roles in odontogenesis, dental cell function, as well as in outgrowth, maintenance, and regeneration of nerve fibers that surround the developing tooth pulp. Previously, we have also shown that tuftelin recruits embryonic mesenchymal cells (Shay et al. 2009) and induces embryonic mesenchymal stem cell proliferation (personal communication). Recent findings from our laboratories indicated that tuftelin expression exhibited significant spatio-temporal changes along neuronal differentiation in the developing brain, the trigeminal ganglion, and the eye (Shilo et al. submitted). It was also established by our group that tuftelin is present in the mouse brain specifically in the neurons, is differentially expressed in different brain regions, and its expression increases with age in the mouse brain, from prenatal to 7-month-old mice (Leiser et al. 2007). Therefore, it is tempting to propose that NGF-dependent induction of tuftelin expression, and tuftelin activity, is not only limited to teeth physiology but also important in neurodevelopment of NGF’s target neurons in the nervous system.
References
Aloe L, Luisa Rocco M, Omar Balzamino B, Micera A (2015) Nerve growth factor: a focus on neuroscience and therapy. Curr Neuropharmacol 13:294–303
Amano O, Bringas P, Takahashi I, Takahashi K, Yamane A, Chai Y, Nuckolls GH, Shum L, Slavkin HC (1999) Nerve growth factor (NGF) supports tooth morphogenesis in mouse first branchial arch explants. Dev Dyn 216:299–310
Angelastro JM, Klimaschewski L, Tang S, Vitolo OV, Weissman TA, Donlin LT, Shelanski ML, Greene LA (2000) Identification of diverse nerve growth factor-regulated genes by serial analysis of gene expression (SAGE) profiling. Proc Natl Acad Sci 97:10424–10429
Arien-Zakay H, Lecht S, Bercu MM, Amariglio N, Rechavi G, Galski H, Lazarovici P, Nagler A (2009) Interferon-γ-induced neuronal differentiation of human umbilical cord blood-derived progenitors. Leukemia 23:1790–1800
Cho K, Skarnes W, Minsk B, Palmieri S, Jackson-Grusby L, Wagner J (1989) Nerve growth factor regulates gene expression by several distinct mechanisms. Mol Cell Biol 9:135–143
Chung J, Kubota H, Y-i O, Uda S, Kuroda S (2010) Timing-dependent actions of NGF required for cell differentiation. PLoS One 5:e9011
D’Onofrio M, Paoletti F, Arisi I, Brandi R, Malerba F, Fasulo L, Cattaneo A (2011) NGF and proNGF regulate functionally distinct mRNAs in PC12 cells: an early gene expression profiling. PLoS One 6:e20839
Delgado S, Deutsch D, Sire J (2017) Evolutionary analysis of the mammalian tuftelin sequence reveals features of functional importance. J Mol Evol 84:214–224
Deutsch D, Palmon A, Fisher LW, Kolodny N, Termine JD, Young MF (1991) Sequencing of bovine enamelin (“tuftelin”) a novel acidic enamel protein. J Biol Chem 266:16021–16028
Deutsch D, Dafni L, Palmon A, Hekmati M, Young MF, Fisher LW (1997) Tuftelin: enamel mineralization and amelogenesis imperfecta. CIBA Found Symp 205:135–147 discussion 147-155
Deutsch D, Palmon A, Dafni L, Mao Z, Leytin V, Young M, Fisher LW (1998) Tuftelin--aspects of protein and gene structure. Eur J Oral Sci 106(1):315–323
Deutsch D et al (2002) The human tuftelin gene and the expression of tuftelin in mineralizing and nonmineralizing tissues. Connect Tissue Res 43:425–434
Deutsch D, Silverstein N, Shilo D, Lecht S, Lazarovici P, Blumenfeld A (2011) Biphasic influence of hypoxia on tuftelin expression in mouse mesenchymal C3H10T1/2 stem cells. Eur J Oral Sci 119(Suppl 1):55–61
Dijkmans TF, van Hooijdonk LWA, Schouten TG, Kamphorst JT, Vellinga ACA, Meerman JHN, Fitzsimons CP, de Kloet ER, Vreugdenhil E (2008) Temporal and functional dynamics of the transcriptome during nerve growth factor-induced differentiation. J Neurochem 105:2388–2403
Dutta P, Koch A, Breyer B, Schneider H, Dittrich-Breiholz O, Kracht M, Tamura T (2011) Identification of novel target genes of nerve growth factor (NGF) in human mastocytoma cell line (HMC-1 (V560G c-Kit)) by transcriptome analysis. BMC Genomics 12:196
Fujita K, Lazarovici P, Guroff G (1989) Regulation of the differentiation of PC12 pheochromocytoma cells. Environ Health Perspect 80:127–142
Greene LA, Angelastro JM (2005) You can’t go home again: transcriptionally driven alteration of cell signaling by NGF. Neurochem Res 30:1347–1352
Katzir I, Shani J, Regev K, Shabashov D, Lazarovici P (2002) A quantitative bioassay for nerve growth factor, using PC12 clones expressing different levels of trkA receptors. J Mol Neurosci 18:251–264
Lazarovici P, Marcinkiewicz C, Lelkes PI (2006) Cross talk between the cardiovascular and nervous systems: neurotrophic effects of vascular endothelial growth factor (VEGF) and angiogenic effects of nerve growth factor (NGF)-implications in drug development. Curr Pharm Des 12:2609–2622
Leiser Y, Blumenfeld A, Haze A, Dafni L, Taylor AL, Rosenfeld E, Fermon E, Gruenbaum-Cohen Y, Shay B, Deutsch D (2007) Localization, quantification, and characterization of tuftelin in soft tissues. Anat Rec 290:449–454
Leiser Y, Silverstein N, Blumenfeld A, Shilo D, Haze A, Rosenfeld E, Shay B, Tabakman R, Lecht S, Lazarovici P, Deutsch D (2011) The induction of tuftelin expression in PC12 cell line during hypoxia and NGF-induced differentiation. J Cell Physiol 226:165–172
Leonard D, Ziff E, Greene L (1987) Identification and characterization of mRNAs regulated by nerve growth factor in PC12 cells. Mol Cell Biol 7:3156–3167
Levi A, Biocca S, Cattaneo A, Calissano P (1989) The mode of action of nerve growth factor in PC12 cells. In: Molecular neurobiology 1988. Springer, pp 201-226
Lin JY-S, Wu CL, Liao CN, Higuchi A, Ling Q-D (2016) Chemogenomic analysis of neuronal differentiation with pathway changes in PC12 cells. Mol BioSyst 12:283–294
Mao Z, Shay B, Hekmati M, Fermon E, Taylor A, Dafni L, Heikinheimo K, Lustmann J, Fisher LW, Young MF, Deutsch D (2001) The human tuftelin gene: cloning and characterization. Gene 279:181–196
Mitsiadis TA, Pagella P (2016) Expression of nerve growth factor (NGF), TrkA, and p75(NTR) in developing human fetal teeth. Front Physiol 7:338
Nosrat I, Seiger A, Olson L, Nosrat CA (2002) Expression patterns of neurotrophic factor mRNAs in developing human teeth. Cell Tissue Res 310:177–187
Rudkin B, Lazarovici P, Levi B, Abe Y, Fujita K, Guroff G (1989) Cell cycle-specific action of nerve growth factor in PC12 cells: differentiation without proliferation. EMBO J 8:3319–3325
Shay B, Gruenbaum-Cohen Y, Tucker AS, Taylor AL, Rosenfeld E, Haze A, Dafni L, Leiser Y, Fermon E, Danieli T, Blumenfeld A, Deutsch D (2009) High yield expression of biologically active recombinant full length human tuftelin protein in baculovirus-infected insect cells. Protein Expr Purif 68:90–98
Skaper SD (2017) Nerve growth factor: a neuroimmune crosstalk mediator for all seasons. Immunology 151:1–15
Tabakman R, Lazarovici P, Kohen R (2002) Neuroprotective effects of carnosine and homocarnosine on pheochromocytoma PC12 cells exposed to ischemia. J Neurosci Res 68:463–469
Vaudry D, Stork P, Lazarovici P, Eiden L (2002) Signaling pathways for PC12 cell differentiation: making the right connections. Science 296:1648–1649
Vician L, Basconcillo R, Herschman HR (1997) Identification of genes preferentially induced by nerve growth factor versus epidermal growth factor in PC12 pheochromocytoma cells by means of representational difference analysis. J Neurosci Res 50:32–43
Yan Q, Johnson EM Jr (1988) An immunohistochemical study of the nerve growth factor receptor in developing rats. J Neurosci 8:3481–3498
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Shilo, D., Cohen, G., Blumenfeld, A. et al. Tuftelin Is Required for NGF-Induced Differentiation of PC12 Cells. J Mol Neurosci 68, 135–143 (2019). https://doi.org/10.1007/s12031-019-01292-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-019-01292-1