Abstract
Wilms tumor 1 (WT1), a tumor suppressor gene, was originally identified in the homonymous renal neoplasm but is also involved in other cancers. Its function is still unclear, since it acts both as a pro- and an anti-oncogene. At least 14 WT1 transcriptional variants have been described; yet most investigations have focused on a small number of isoforms. We describe their structural features and review the evidence of their involvement in cancer with emphasis on neuroblastoma. In future, full characterization of all WT1 isoforms is expected to identify new molecular tumor markers and/or therapeutic targets.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Wilms tumor 1 gene (WT1) maps on the short arm of human chromosome 11 and has been one of the first tumor suppressor genes to be cloned (Call et al. 1990; Gessler et al. 1990; Hung et al. 2016; Goyal et al. 2016; Naitoh et al. 2016). It takes its name from Wilms tumor, a renal tumor that largely affects children, but is also involved in several other cancers.
WT1 plays a crucial role in fetal life, due to its involvement in the development of the organs of mesodermal origin (kidney, spleen, gonads, and cardiovascular system) and in the proliferation of some nervous system progenitor cells. It promotes kidney differentiation by blocking cell cycle progression, whereas in other tissues it sustains cell proliferation (Moore et al. 1998; Moore et al. 1999). In healthy adults it is found only in podocytes, highly specialized cells forming the visceral layer of the Bowman’s capsule in the kidney (Hohenstein and Hastie 2006), whereas it is widely expressed in tissue from patients suffering from a variety of pathological conditions, including tumors.
Its function is the subject of extensive debate, since WT1 acts as an oncogene in some neoplasms and as an oncosuppressor in others, including Wilms tumor. Interestingly in neuroblastoma, a neuroepithelial tumor of embryonic origin that also largely affects children, it seems to act in both ways (Sebire et al. 2005; Wang et al. 2011; Kletzel et al. 2015; Maugeri et al. 2016; Masserot et al. 2016). Neuroblastoma is characterized by a highly heterogeneous clinical behavior, as it can either progress to more aggressive cancer or regress spontaneously to less malignant ganglioneuroblastoma or ganglioneuroma. WT1 is expressed in tumor tissue both in the nuclear and the cytoplasmic compartment (Niksic et al. 2004; Parenti et al. 2013; Magro et al. 2015) and its level of expression is related to tumor malignancy (Maugeri et al. 2016).
WT1 encodes a protein having an N-terminal region rich in lysine and glutamine residues and a C-terminal region composed of four consecutive zinc finger motifs that are highly homologous to those of early growth response-1 (Egr-1) transcription factor (Rauscher et al. 1990). These domains have led to the classification of WT1 as a transcription factor, where the N-terminal region interacts with transcriptional co-activators and the C-terminal tract binds to GC-rich DNA regions (Rauscher et al. 1990), WTE sites (Nakagama et al. 1995), or (TCC)n motifs (Wang et al. 1993). Through these regions, WT1 regulates the transcription of target genes such as Bcl-2, Bcl-xL, Bfl-1, and c-myc, which are involved in cell growth or apoptosis (Mayo et al. 1999; Han et al. 2004; Simpson et al. 2006; Bansal et al. 2012).
Despite the existence of multiple WT1 isoforms—generated through alternative splicing—only a few have been investigated (Oji et al. 2002; Koesters et al. 2004; Luna et al. 2013). The cloned variants deposited in the various databases (NCBI, Ensembl, UniProt) are listed in Table 1. The published evidence has suggested to us that their expression profile may be associated to a specific cellular phenotype. Immunolocalization studies performed in tissue samples from cancer patients have found a variable staining pattern— cytoplasmic, nuclear, or both—depending on whether the antibody employed is directed against an epitope on the C or the N-terminus (Carpentieri et al. 2002; Nakatsuka et al. 2006; Schittenhelm et al. 2010; Bisceglia et al. 2011; Salvatorelli et al. 2011; Magro et al. 2014a, b, 2015; Salvatorelli et al. 2015).
Furthermore, WT1 protein shows a discrete distribution in embryonic organs: exclusively in the nucleus of urogenital apparatus progenitor cells and only in the cytoplasm of cardiovascular and nervous system germ cells (Pritchard-Jones et al. 1990; Sharma et al. 1992; Armstrong et al. 1993; Mundlos et al. 1993; Ramani and Cowell 1996; Charles et al. 1997; Parenti et al. 2013, 2015; Salvatorelli et al. 2015). Based on these data, it may be hypothesized that WT1 regulates gene transcription when it is expressed in the nucleus, whereas it participates in post-transcriptional events when it is expressed in the cytoplasm. Such compartmentalization could well reflect the different actions and functions of the diverse WT1 isoforms.
Here we describe the features of the WT1 transcript variants that have already been cloned and review the literature regarding their involvement in tumors, especially neuroblastoma.
Alternative Splicing and the WT1 Gene Isoforms
WT1, a complex gene measuring approximately 50 kb in length, comprises 10 exons and 9 introns. Various mRNA transcripts are translated into different isoforms by alternative splicing (Haber et al. 1991; Tadokoro et al. 1992).
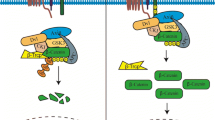
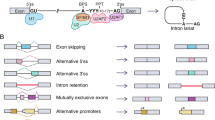
During precursor mRNA (pre-mRNA) splicing, a crucial stage of post-transcriptional regulation, introns are removed and exons joined to form mature mRNA, which is exported to the cytoplasm for translation into a protein (Crick 1979; D’Agata et al. 2000; Hastings and Krainer 2001; Jurica and Moore 2003; Hollander et al. 2016). Four highly conserved regions in the intronic sequences are involved in the splicing process: two splice sites (a donor and an acceptor) respectively at the 5′ (GU) and the 3′ (AG) end and a branching point in the intron followed by a polypyrimidine tract (Hertel and Graveley 2005). The spliceosome, a molecular complex consisting of small nuclear ribonucleoproteins (snRNP) and of non-snRNP factors, recognizes these sites and enables the synthesis of mature mRNA through a number of steps (Collins and Guthrie 2000; Das et al. 2000; Maroney et al. 2000; Hertel and Graveley 2005). Different transcript variants are produced by the exclusion of whole exons and/or introns during alternative splicing (D’Agata and Cavallaro 2004; Scuderi et al. 2014; La Cognata et al. 2014; D’Amico et al. 2015; Maugeri et al. 2015).
As regards WT1, an alternative splicing site is found on exon 5, which encodes 17 amino acids; another is located on the 3′ end of exon 9, which codes for the amino acids lysine, threonine, and serine; this tract is also referred to as KTS region (Haber et al. 1991). Notably, WT1 pre-mRNA has three different alternative start codons including a CUG and an AUG, respectively, upstream and downstream of the regular AUG (Bruening and Pelletier 1996; Scharnhorst et al. 1999). This combination of alternative splicing sites and start codons results in generation of at least 12 transcript variants, although in theory up to 24 isoforms can be formed through subsequent RNA editing events. Moreover, additional mRNA variants (AWT1) are generated by an alternative promoter found on the first intron of WT1, which starts transcription from exon 1a (Dallosso et al. 2004). A further variant, Ex4a(+), containing the additional exon 4a downstream of exon 4, has recently been cloned (Schnerwitzki et al. 2014; Tatsumi et al. 2015).
Fourteen variants are found in public repositories (Ensembl, NCBI, UniProt) and have lately been given a new ID code (Maugeri et al. 2016). Their list, accession number, nucleotide and amino acid composition, isoelectric point, and molecular weight as well as the articles describing any variants are reported in Table 1.
To identify the structural differences among the transcripts, we aligned their nucleotide sequence variants using the CLC Sequence viewer 7 software. WT1.6, the most widely studied isoform, was arbitrarily designated as the canonical sequence.
To date, most studies have examined exclusively the WT1 variants containing or lacking exon 5 and the KTS region, besides the canonical sequence (Loeb et al. 2001; Oji et al. 2002; Oji et al. 2003; Koesters et al. 2004; Oji et al. 2004a, b; Luna et al. 2013). Yet, transcripts differing in the initial part of the N-terminal domain and, in some cases, also in the C-terminal domain have also been cloned. These nucleotide regions found in the 14 splice variants are shown in Figs. 1, 2, 3, and 4.
The canonical sequence (WT1.6), containing both exon 5 and the KTS region, is the +/+ variant; WT1.7, which contains exon 5 but not the KTS region, is the +/− variant; WT1.8 contains the KTS region and lacks exon 5 and is the −/+ variant; and finally WT1.9, which lacks both regions, is the −/− transcript. WT1.1 and WT1.4 mRNAs are similar to WT1.6, except that they have a longer N-terminal domain, because the start codon is found upstream of the one of the canonical isoform. WT1.2 diverges from WT1.1 by a few initial nucleotides; WT1.3 is a “+/−” transcript, and WT1.5 a −/− splice variant (Maugeri et al. 2016).
WT1.10 and WT1.14 differ at the 5′ end (Fig. 4). As shown in Fig. 3, they share a 153 nucleotide insertion downstream of exon 4, corresponding to exon 4a. The latter exon bears a stop codon whose presence could expose such variants to nonsense-mediated mRNA decay, a process where mRNA degradation is triggered by nonsense mutations to prevent abnormal or truncated protein formation. However, the degradation has been excluded in these transcripts by demonstration that WT1.14 is translated into a truncated protein lacking four zinc finger domains (Tatsumi et al. 2015).
Variants WT1.11 and WT1.12, the former lacking the KTS region and the latter lacking exon 5, can be considered as AWT1 variants. Their transcription starts from exon 1a, following the splice out of exon 1. These transcripts encode a ∼33-kDa molecular weight protein having an N-terminal domain that is shorter than that of the other isoforms (Dallosso et al. 2004; Royer-Pokora 2013). Finally, WT1.13 is the smallest cloned variant. With regard to the translated protein, the Ensembl database reports the annotation: “5′ truncation in transcript evidence prevents annotation of the start of the CDS.” This suggests that further investigation is required to confirm its expression.
To summarize, the WT1.1, WT1.2, WT1.4, WT1.6, WT1.10, WT1.13, and WT1.14 transcripts contain exon 5 as well as the KTS region, both of which are absent in WT1.5 and WT1.9; variants WT1.3, WT1.7, and WT1.11 contain exon 5 but not the KTS region, and WT1.8 and WT1.12 contain the KTS tract but not exon 5.
Involvement of the WT1 Isoforms in Tumors, with Emphasis on Neuroblastoma
WT1 plays a role in a wide range of conditions, where it seems to exert conflicting effects. It is rarely detected in healthy tissue, whereas it is commonly expressed in several cancers including leukemia (Luna et al. 2013) and lung (Oji et al. 2002), colon (Koesters et al. 2004), pancreatic (Oji et al. 2004a), breast (Loeb et al. 2001; Oji et al. 2004b), and thyroid (Oji et al. 2003) tumors. Since its discovery, WT1 has been considered as a tumor suppressor gene with a mutation in Wilms tumor, WAGR (Wilms tumor, aniridia, genitourinary malformations, mental retardation) (Gessler et al. 1990), and Danys-Drash (Pelletier et al. 1991; Patek et al. 1999) and Frasier syndrome (Barbaux et al. 1997; Klamt et al. 1998). However, an oncogenic role has also been hypothesized in some solid tumors and leukemia (Oji et al. 2002; Bansal et al. 2012; Luna et al. 2013), leading it to be considered both as an oncosuppressor and an oncogene (Yang et al. 2007). Even though such conflicting actions may depend on changes in promoter methylation (Loeb et al. 2001; Brocato and Costa 2013; Guillaumet-Adkins et al. 2014; Zitzmann et al. 2014; Abete et al. 2015; Sahnane et al. 2015; Mžik et al. 2016), a correlation with the tissue- and cell-specific expression of the WT1 isoform cannot be ruled out. Indeed, the different roles played by different variants suggest a definite cellular phenotype.
Since most studies of WT1 isoforms have been confined to establishing the presence/absence of specific splicing regions, they have been unable to discriminate among the different variants. For instance, Oji et al. (2002) and Luna et al. (2013) assessed them respectively in lung cancer and acute myeloid leukemia (AML). Using probes for exon 5 and the KTS region, they identified the +/+, −/+, +/−, and −/− variants, which included several transcripts. All variants were found to be overexpressed in both tumors except −/− transcripts, which were underexpressed. These findings suggest that the scanty expression of the −/− isoforms (WT1.5 and WT1.9) might depend more on the small number of variants (only 2) than on downregulation. Furthermore, Luna et al. (2013) found that blood progenitor cells in AML patients also expressed WT1.11 and WT1.12 (AWT1 isoforms).
The Ex4a(+) variant, cloned by Tatsumi et al. (2015), is found in non-cancerous kidney cells and is downregulated in some tumors. By inhibiting the transcriptional activation of the anti-apoptotic gene Bcl-xL, it could regulate the oncogenic action of other WT1 isoforms which conversely induce this gene (Tatsumi et al. 2015).
Some studies have suggested that in neuroblastomaWT1 acts as an oncosuppressor as well as an oncogene (Haber et al. 1993; McMaster et al. 1995; Menke et al. 1997; Smith et al. 2000; Fraizer et al. 2004; Sebire et al. 2005; Wang et al. 2011; Kletzel et al. 2015; Masserot et al. 2016; Maugeri et al. 2016). Neuroblastoma is a childhood tumor occurring more often in boys (Spix et al. 2006; Gatta et al. 2012). It is a rare solid tumor characterized by a highly variable clinical behavior that may either regress spontaneously or progress to an aggressive form resistant to chemotherapy as well as radiation therapy. Outcome is related to conversion to a benign form, including ganglioneuroblastoma and ganglioneuroma, or to progression to a highly malignant form with metastases and a poor prognosis. The degree of malignancy is reflected in discrete histological features, undifferentiated cells with poor stroma being characteristic of aggressive cancer and differentiated cells with abundant stroma being found in benign forms (Maris et al. 2007; Salvatorelli et al. 2015).
Although the etiopathogenic mechanism of neuroblastoma development is unknown, epidemiological studies have identified some risk factors, which include maternal exposure to pesticides, volatile hydrocarbons, tobacco, and codeine-containing drugs during pregnancy or lactation (De Roos et al. 2001; Schüz et al. 2001; Cook et al. 2004; Heck et al. 2009).
The tumor originates from the cells of the neural crest, which are arranged as two cords on the sides of the neural tube. During development, these cells migrate to various sites in the thoracic and abdominal regions, differentiating into tissue-specific cells; this explains why the tumor rarely forms in the brain. The most common sites of origin are the adrenal glands and the paravertebral ganglia and involve formation of masses that induce abdominalcompression. Tumor development in the posterior mediastinal ganglia gives rise to respiratory symptoms. Occasionally, neuroblastoma forms in the neck, giving rise to Bernard-Horner syndrome, or in the pelvis, affecting the urinary and/or anorectal sphincter (Caron and Pearson 2005; Brodeur and Maris 2006; De Bernardi et al. 2008).
The prognosis is related to patient age and tumor stage: it is favorable in newborns and usually adverse in adolescents (Gatta et al. 2014). Surgical resection of localized tumor is associated to 85% survival, whereas metastasis and amplification of the MYCN gene, a biomarker of neuroblastoma, predicts a poor prognosis (Luksch et al. 2016).
The expression pattern of the WT1 isoforms in neuroblastoma has not been fully characterized. In a recent study, our group has identified 13 WT1 isoforms, although the analysis was limited to in vitro models mimicking a benign and a highly aggressive form (Maugeri et al. 2016). Although it is impossible to determine the expression profile of each isoform, upregulation was detected in less malignant cells, suggesting a tumor suppressor role. Variants WT1.1–9, whose molecular weight ranges from ∼56 to ∼47 kDa, were the most highly expressed. The study did not include the WT1.14 isoform, because it was the last to be cloned (Tatsumi et al. 2015). In line with our findings, Wang et al. (2011) have reported WT1 overexpression in ganglioneuroblastoma and suggested that it may act as a pro-differentiation and anti-proliferation factor involved in benign progression.
Other studies have hypothesized an oncogenic role for WT1 in neuroblastoma. In 2005, Sebire et al. demonstrated its cytoplasmic expression in some Bcl-2-positive tumors and suggested that WT1 may activate this endogenous anti-apoptotic gene through a transcriptional mechanism, thus providing resistance to chemotherapy (Mayo et al. 1999). In another study (Kletzel et al. 2015), investigation of WT1expression in the bone marrow, peripheral blood, or peripheral blood stem cells from young neuroblastoma patients sampled at the time of diagnosis, during treatment, and during clinical remission/relapse suggested a correlation with the level of tyrosine hydroxylase, a molecular marker of neuroblastoma (Naito et al. 1991; Parareda et al. 2005; Ootsuka et al. 2008; Avigad et al. 2009; Lee et al. 2010). During remission, WT1 levels were significantly reduced, albeit no data were provided regarding the isoform(s) involved. Amplification and PCR analysis of a 440 nucleotide tract corresponding to the 1350–1789 region of the canonical sequence excluded variant WT1.13. Then, amplification using the inner primers gave a 302 nucleotide product corresponding to the 1418–1719 region of WT1.6, which did not allow discriminating among the variants. However, the authors did not clarify whether the patients in clinical remission included both those who had been cured and those with benign tumor.
Masserot et al. (2016) have recently found a correlation between the levels of WT1 and MYCN, a member of myc family transcription factors, in tumor tissue and neoplastic cell lines. The MYCN gene is considered as a marker of malignant neuroblastoma progression, because its overexpression is often related to a fatal outcome, yet 70–80% of MYCN-negative tumors have a poor prognosis (Hiyama et al. 1991; Schwab et al. 2003). Masserot et al. (2016) found that WT1 overexpression was associated to a poor prognosis in samples lacking MYCN amplification. However, the amplification product, including the canonical sequence from 1218 to 1313, did not allow to discriminate among variants, since it is found in all isoforms.
As summarized in Table 2, the available data do not allow the establishment of the role played by each isoform or it cannot be ruled out that the conflicting results reported in different studies depend on differences in pathogenicity or analytical methods. The hypothesis may be advanced that the oncogenic and oncosuppressor action of the WT1 isoforms may correlate with tumor malignancy (Fig. 5).
Conclusions
Generation of splice variants with a tissue- and cell-specific expression profile enhances the functional diversity of the gene. Although the over- or underexpression of some isoforms could play a key role in tumor phenotype, their high homology currently hampers discrimination. Most studies have confined the analysis to overall transcript expression. In future, full characterization of all WT1 isoforms is expected to identify new molecular tumor markers and/or therapeutic targets.
References
Abete I, Gómez-Úriz AM, Mansego ML, De Arce A, Goyenechea E, Blázquez V, Martínez-Zabaleta MT, González-Muniesa P, López De Munain A, Martínez JA, Campión J, Milagro FI (2015) Epigenetic changes in the methylation patterns of KCNQ1 and WT1 after a weight loss intervention program in obese stroke patients. Curr Neurovasc Res 12:321–333
Armstrong JF, Pritchard-Jones K, Bickmore WA, Hastie ND, Bard JB (1993) The expression of the Wilms’ tumour gene, WT1, in the developing mammalian embryo. Mech Dev 40:85–97
Avigad S, Feinberg-Gorenshtein G, Luria D, Jeison M, Stein J, Grunshpan A, Sverdlov Y, Ash S, Yaniv I (2009) Minimal residual disease in peripheral blood stem cell harvests from high-risk neuroblastoma patients. J Pediatr Hematol Oncol 31:22–26
Bansal H, Seifert T, Bachier C, Rao M, Tomlinson G, Iyer SP, Bansal S (2012) The transcription factor Wilms tumor 1 confers resistance in myeloid leukemia cells against the proapoptotic therapeutic agent TRAIL (tumor necrosis factor α-related apoptosis-inducing ligand) by regulating the antiapoptotic protein Bcl-xL. J Biol Chem 287:32875–32880
Barbaux S, Niaudet P, Gubler MC, Grünfeld JP, Jaubert F, Kuttenn F, Fékété CN, Souleyreau-Therville N, Thibaud E, Fellous M, McElreavey K (1997) Donor splice-site mutations in WT1 are responsible for Frasier syndrome. Nat Genet 17:467–470
Bisceglia M, Vairo M, Galliani C, Lastilla G, Parafioriti A, De Maglio G, Rosai J (2011) Immunohistochemical investigation of WT1 expression in 117 embryonal tumors. Patologica 103:182–183
Brocato J, Costa M (2013) Basic mechanics of DNA methylation and the unique landscape of the DNA methylome in metal-induced carcinogenesis. Crit Rev Toxicol 43:493–514
Brodeur GM, Maris JM (2006) Neuroblastoma. In: Pizzo DA, Poplack DG (eds) Principles and practice of pediatric oncology, 5th edn. Williams&Wilkins, Philadelphia, pp 933–970
Bruening W, Pelletier J (1996) A non-AUG translational initiation event generates novel WT1 isoforms. J Biol Chem 271:8646–8654
Bruening W, Bardeesy N, Silverman BL, Cohn RA, Machin GA, Aronson AJ, Housman D, Pelletier J (1992) Germline intronic and exonic mutations in the Wilms’ tumour gene (WT1) affecting urogenital development. Nat Genet 1:144–148
Call KM, Glaser T, Ito CY, Buckler AJ, Pelletier J, Haber DA, Rose EA, Kral A, Yeger H, Lewis WH, Jones C, Housman DE (1990) Isolation and characterization of a zinc finger polypeptide gene at the human chromosome 11 Wilms’ tumor locus. Cell 60:509–520
Caron HN, Pearson AD (2005) Neuroblastoma. In: Voute PA, Barrett A, Stevens MCG, Caron HN (eds) Cancer in children, 5th edn. Oxford University Press, Oxford, pp 337–352
Carpentieri DF, Nichols K, Chou PM, Matthews M, Pawel B, Huff D (2002) The expression of WT1 in the differentiation of rhabdomyosarcoma from other pediatric small round blue cell tumors. Mod Pathol 15:1080–1086
Charles AK, Mall S, Watson J, Berry PJ (1997) Expression of the Wilms’ tumour gene WT1 in the developing human and in paediatric renal tumours: an immunohistochemical study. Mol Pathol 50:138–144
Collins CA, Guthrie C (2000) The question remains: is the spliceosome a ribozyme? Nat Struct Biol 7:850–854
Cook MN, Olshan AF, Guess HA, Savitz DA, Poole C, Blatt J, Bondy ML, Pollock BH (2004) Maternal medication use and neuroblastoma in offspring. Am J Epidemiol 159:721–731
Crick F (1979) Split genes and RNA splicing. Science 204:264–271
D’Agata V, Cavallaro S (2004) Parkin transcript variants in rat and human brain. Neurochem Res 29:1715–1724
D’Agata V, Zhao W, Cavallaro S (2000) Cloning and distribution of the rat parkin mRNA. Brain Res Mol Brain Res 75:345–349
Dallosso AR, Hancock AL, Brown KW, Williams AC, Jackson S, Malik K (2004) Genomic imprinting at the WT1 gene involves a novel coding transcript (AWT1) that shows deregulation in Wilms’ tumours. Hum Mol Genet 13:405–415
D’Amico AG, Maugeri G, Magro G, Salvatorelli L, Drago F, D’Agata V (2015) Expression pattern of parkin isoforms in lung adenocarcinomas. Tumour Biol 36:5133–5141
Das R, Zhou Z, Reed R (2000) Functional association of U2 snRNP with the ATP-independent spliceosomal complex E. Mol Cell 5:779–787
De Bernardi B, Pistoia V, Gambini C, Granata C (2008) Peripheral neuroblastic tumours. In: Sheaves R, Jenkins PJ, Wass JA (eds) Clinical endocrine oncology, 2nd edn. Blackwell Science, Oxford, pp 360–369
De Roos AJ, Teschke K, Savitz DA, Poole C, Grufferman S, Pollock BH, Olshan AF (2001) Parental occupational exposures to electromagnetic fields and radiation and the incidence of neuroblastoma in offspring. Epidemiology 12:508–517
Fraizer G, Leahy R, Priyadarshini S, Graham K, Delacerda J, Diaz M (2004) Suppression of prostate tumor cell growth in vivo by WT1, the Wilms’ tumor suppressor gene. Int J Oncol 24:461–471
Gatta G, Ferrari A, Stiller CA, Pastore G, Bisogno G, Trama A, Capocaccia R (2012) Embryonalcancers in Europe. Eur J Cancer 48:1425–1433
Gatta G, Botta L, Rossi S, Aareleid T, Bielska-Lasota M, Clavel J, Dimitrova N, Jakab Z, Kaatsch P, Lacour B, Mallone S, Marcos-Gragera R, Minicozzi P, Sánchez-Pérez MJ, Sant M, Santaquilani M, Stiller C, Tavilla A, Trama A, Visser O, Peris-Bonet R (2014) Childhood cancer survival in Europe 1999-2007: results of EUROCARE-5—a population-based study. Lancet Oncol 15:35–47
Gessler M, Poustka A, Cavenee W, Neve RL, Orkin SH, Bruns GA (1990) Homozygous deletion in Wilms tumours of a zinc-finger gene identified by chromosome jumping. Nature 343:774–778
Goyal S, Mishra K, Sarkar U, Sharma S, Kumari A (2016) Diagnostic utility of Wilms’ tumour-1 protein (WT-1) immunostaining in paediatric renal tumours. Indian J Med Res 143:S59–S67
Guillaumet-Adkins A, Richter J, Odero MD, Sandoval J, Agirre X, Catala A, Esteller M, Prósper F, Calasanz MJ, Buño I, Kwon M, Court F, Siebert R, Monk D (2014) Hypermethylation of the alternative AWT1 promoter in hematological malignancies is a highly specific marker for acute myeloid leukemias despite high expression levels. J Hematol Oncol 7:4
Haber DA, Sohn RL, Buckler AJ, Pelletier J, Call KM, Housman DE (1991) Alternative splicing and genomic structure of the Wilms tumor gene WT1. Proc Natl Acad Sci U S A 88:9618–9622
Haber DA, Park S, Maheswaran S, Englert C, Re GG, Hazen-Martin DJ, Sens DA, Garvin AJ (1993) WT1-mediated growth suppression of Wilms tumor cells expressing a WT1 splicing variant. Science 262:2057–2059
Han Y, San-Marina S, Liu J, Minden MD (2004) Transcriptional activation of c-myc proto-oncogene by WT1 protein. Oncogene 23:6933–6941
Hastings ML, Krainer AR (2001) Pre-mRNA splicing in the new millennium. CurrOpin Cell Biol 13:302–309
Heck JE, Ritz B, Hung RJ, Hashibe M, Boffetta P (2009) The epidemiology of neuroblastoma: a review. Paediatr Perinat Epidemiol 23:125–143
Hertel KJ, Graveley BR (2005) RS domains contact the pre-mRNA throughout spliceosome assembly. Trends Biochem Sci 30:115–118
Hiyama E, Hiyama K, Yokoyama T, Ishii T (1991) Immunohistochemical analysis of N-myc protein expression in neuroblastoma: correlation with prognosis of patients. J Pediatr Surg 26:838–843
Hohenstein P, Hastie ND (2006) The many facets of the Wilms’ tumour gene, WT1. Hum Mol Genet 15(2):R196–R201
Hollander D, Naftelberg S, Lev-Maor G, Kornblihtt AR, Ast G (2016) How are short exons flanked by long introns defined and committed to splicing? Trends Genet 32:596–606
Hung YP, Fletcher CD, Hornick JL (2016) Evaluation of ETV4 and WT1 expression in CIC-rearranged sarcomas and histologic mimics. Mod Pathol 29:1324–1334
Jinno Y, Yun K, Nishiwaki K, Kubota T, Ogawa O, Reeve AE, Niikawa N (1994) Mosaic and polymorphic imprinting of the WT1 gene in humans. Nat Genet 6:305–309
Jurica MS, Moore MJ (2003) Pre-mRNA splicing: awash in a sea of proteins. Mol Cell 12:5–14
Klamt B, Koziell A, Poulat F, Wieacker P, Scambler P, Berta P, Gessler M (1998) Frasier syndrome is caused by defective alternative splicing of WT1 leading to an altered ratio of WT1 +/-KTS splice isoforms. Hum Mol Genet 7:709–714
Kletzel M, Chou PM, Olszewski M, Rademaker AW, Khan S (2015) Expression of Wilms tumor gene in high risk neuroblastoma: complementary marker to tyrosine hydroxylase for detection of minimal residual disease. Transl Pediatr 4:219–225
Koesters R, Linnebacher M, Coy JF, Germann A, Schwitalle Y, Findeisen P, von KnebelDoeberitz M (2004) WT1 is a tumor-associated antigen in colon cancer that can be recognized by in vitro stimulated cytotoxic T cells. Int J Cancer 109:385–392
La Cognata V, Iemmolo R, D’Agata V, Scuderi S, Drago F, Zappia M, Cavallaro S (2014) Increasing the coding potential of genomes through alternative splicing: the case of PARK2 gene. Curr Genomics 15:203–216
Lee ST, Suh YL, Ko YH, Ki CS, Sung KW, Kim HJ, Kim JW, Kim SH, Chueh H, Lee SH, Yoo KH, Koo HH (2010) Measurement of tyrosine hydroxylase transcripts in bone marrow using biopsied tissue instead of aspirates for neuroblastoma. Pediatr Blood Cancer 55:273–278
Loeb DM, Evron E, Patel CB, Sharma PM, Niranjan B, Buluwela L, Weitzman SA, Korz D, Sukumar S (2001) Wilms’ tumor suppressor gene (WT1) is expressed in primary breast tumors despite tumor-specific promoter methylation. Cancer Res 61:921–925
Luksch R, Castellani MR, Collini P, De Bernardi B, Conte M, Gambini C, Gandola L, Garaventa A, Biasoni D, Podda M, Sementa AR, Gatta G, Tonini GP (2016) Neuroblastoma (peripheral neuroblastic tumours). Crit Rev Oncol Hematol 107:163–181
Luna I, Such E, Cervera J, Barragán E, Ibañez M, Gómez-Seguí I, López-Pavía M, Llop M, Fuster O, Dolz S, Oltra S, Alonso C, Vera B, Lorenzo I, Martínez-Cuadrón D, Montesinos P, Senent ML, Moscardó F, Bolufer P, Sanz MA (2013) WT1 isoform expression pattern in acute myeloid leukemia. Leuk Res 37:1744–1749
Magro G, Salvatorelli L, Vecchio GM, Musumeci G, Rita A, Parenti R (2014a) Cytoplasmic expression of Wilms tumor transcription factor-1 (WT1): a useful immunomarker for young-type fibromatoses and infantile fibrosarcoma. Acta Histochem 116:1134–1140
Magro G, Longo F, Salvatorelli L, Vecchio GM, Parenti R (2014b) Wilms’ tumor protein (WT1) in mammary myofibroblastoma: an immunohistochemical study. Acta Histochem 116:905–910
Magro G, Salvatorelli L, Puzzo L, Musumeci G, Bisceglia M, Parenti R (2015) Oncofetal expression of Wilms’ tumor 1 (WT1) protein in human fetal, adult and neoplastic skeletal muscle tissues. Acta Histochem 117:492–504
Maki T, Ikeda H, Kuroda A, Kyogoku N, Yamamura Y, Tabata Y, Abiko T, Tsuchikawa T, Hida Y, Shichinohe T, Tanaka E, Kaga K, Hatanaka K, Matsuno Y, Imai N, Hirano S (2017) Differential detection of cytoplasmic Wilms tumor 1 expression by immunohistochemistry, western blotting and mRNA quantification. Int J Oncol 50:129–140
Maris JM, Hogarty MD, Bagatell R, Cohn SL (2007) Neuroblastoma. Lacent 369:2106–2120
Maroney PA, Romfo CM, Nilsen TW (2000) Functional recognition of 5′ splice site by U4/U6.U5 tri-snRNP defines a novel ATP-dependent step in early spliceosome assembly. Mol Cell 6:317–328
Masserot C, Liu Q, Nguyen E, Gattolliat CH, Valteau-Couanet D, Bénard J, Huber C, Ségal-Bendirdjian E (2016) WT1 expression is inversely correlated with MYCN amplification or expression and associated with poor survival in non-MYCN-amplified neuroblastoma. Mol Oncol 10:240–252
Maugeri G, D’Amico AG, Magro G, Salvatorelli L, Barbagallo GM, Saccone S, Drago F, Cavallaro S, D’Agata V (2015) Expression profile of parkin isoforms in human gliomas. Int J Oncol 47:1282–1292
Maugeri G, D’Amico AG, Rasà DM, Reitano R, Saccone S, Federico C, Parenti R, Magro G, D’Agata V (2016) Expression profile of Wilms tumor 1 (WT1) isoforms in undifferentiated and all-trans retinoic acid differentiated neuroblastoma cells. Genes Cancer 7:47–58
Mayo MW, Wang CY, Drouin SS, Madrid LV, Marshall AF, Reed JC, Weissman BE, Baldwin AS (1999) WT1 modulates apoptosis by transcriptionally upregulating the bcl-2 proto-oncogene. EMBO J 18:3990–4003
McMaster ML, Gessler M, Stanbridge EJ, Weissman BE (1995) WT1 expression alters tumorigenicity of the G401 kidney-derived cell line. Cell Growth Differ 6:1609–1617
Menke AL, Shvarts A, Riteco N, van Ham RC, van der Eb AJ, Jochemsen AG (1997) Wilms’ tumor 1-KTS isoforms induce p53-independent apoptosis that can be partially rescued by expression of the epidermal growth factor receptor or the insulin receptor. Cancer Res 57:1353–1363
Mitsuya K, Sui H, Meguro M, Kugoh H, Jinno Y, Niikawa N, Oshimura M (1997) Paternal expression of WT1 in human fibroblasts and lymphocytes. Hum Mol Genet 6:2243–2246
Moore AW, Schedl A, McInnes L, Doyle M, Hecksher-Sorensen J, Hastie ND (1998) YAC transgenic analysis reveals Wilms’ tumour 1 gene activity in the proliferating coelomic epithelium, developing diaphragm and limb. Mech Dev 79:169–184
Moore AW, McInnes L, Kreidberg J, Hastie ND, Schedl A (1999) YAC complementation shows a requirement for WT1 in the development of epicardium, adrenal gland and throughout nephrogenesis. Development 126:1845–1857
Mundlos S, Pelletier J, Darveau A, Bachmann M, Winterpacht A, Zabel B (1993) Nuclear localization of the protein encoded by the Wilms’ tumor gene WT1 in embryonic and adult tissues. Development 119:1329–1341
Mžik M, Chmelařová M, John S, Laco J, Slabý O, Kiss I, Bohovicová L, Palička V, Nekvindová J (2016) Aberrant methylation of tumour suppressor genes WT1, GATA5 and PAX5 in hepatocellular carcinoma. Clin Chem Lab Med 54:1971–1980
Naito H, Kuzumaki N, Uchino J, Kobayashi R, Shikano T, Ishikawa Y, Matsumoto S (1991) Detection of tyrosine hydroxylase mRNA and minimal neuroblastoma cells by the reverse transcription-polymerase chain reaction. Eur J Cancer 27:762–765
Naitoh K, Kamigaki T, Matsuda E, Ibe H, Okada S, Oguma E, Kinoshita Y, Takimoto R, Makita K, Ogasawara S, Goto S (2016) Immunohistochemical analysis of WT1 antigen expression in various solid cancer cells. Anticancer Res 36:3715–3724
Nakagama H, Heinrich G, Pelletier J, Housman DE (1995) Sequence and structural requirements for high-affinity DNA binding by the WT1 gene product. Mol Cell Biol 15:1489–1498
Nakatsuka S, Oji Y, Horiuchi T, Kanda T, Kitagawa M, Takeuchi T, Kawano K, Kuwae Y, Yamauchi A, Okumura M, Kitamura Y, Oka Y, Kawase I, Sugiyama H, Aozasa K (2006) Immunohistochemical detection of WT1 protein in a variety of cancer cells. Mod Pathol 19:804–814
Niksic M, Slight J, Sanford JR, Caceres JF, Hastie ND (2004) The Wilms’ tumour protein (WT1) shuttles between nucleus and cytoplasm and is present in functional polysomes. Hum Mol Genet 13:463–471
Oji Y, Miyoshi S, Maeda H, Hayashi S, Tamaki H, Nakatsuka S, Yao M, Takahashi E, Nakano Y, Hirabayashi H, Shintani Y, Oka Y, Tsuboi A, Hosen N, Asada M, Fujioka T, Murakami M, Kanato K, Motomura M, Kim EH, Kawakami M, Ikegame K, Ogawa H, Aozasa K, Kawase I, Sugiyama H (2002) Overexpression of the Wilms’ tumor gene WT1 in de novo lung cancers. Int J Cancer 100:297–303
Oji Y, Miyoshi Y, Koga S, Nakano Y, Ando A, Nakatsuka S, Ikeba A, Takahashi E, Sakaguchi N, Yokota A, Hosen N, Ikegame K, Kawakami M, Tsuboi A, Oka Y, Ogawa H, Aozasa K, Noguchi S, Sugiyama H (2003) Overexpression of the Wilms’ tumor gene WT1 in primary thyroid cancer. Cancer Sci 94:606–611
Oji Y, Nakamori S, Fujikawa M, Nakatsuka S, Yokota A, Tatsumi N, Abeno S, Ikeba A, Takashima S, Tsujie M, Yamamoto H, Sakon M, Nezu R, Kawano K, Nishida S, Ikegame K, Kawakami M, Tsuboi A, Oka Y, Yoshikawa K, Aozasa K, Monden M, Sugiyama H (2004a) Overexpression of the Wilms’ tumor gene WT1 in pancreatic ductal adenocarcinoma. Cancer Sci 95:583–587
Oji Y, Miyoshi Y, Kiyotoh E, Koga S, Nakano Y, Ando A, Hosen N, Tsuboi A, Kawakami M, Ikegame K, Oka Y, Ogawa H, Noguchi S, Sugiyama H (2004b) Absence of mutations in the Wilms’ tumor gene WT1 in primary breast cancer. Jpn J Clin Oncol 34:74–77
Ootsuka S, Asami S, Sasaki T, Yoshida Y, Nemoto N, Shichino H, Chin M, Mugishima H, Suzuki T (2008) Useful markers for detecting minimal residual disease in cases of neuroblastoma. Biol Pharm Bull 31:1071–1074
Parareda A, Gallego S, Roma J, Llort A, Sábado C, Gros L, De Toledo JS (2005) Prognostic impact of the detection of microcirculating tumor cells by a real-time RT-PCR assay of tyrosine hydroxylase in patients with advanced neuroblastoma. Oncol Rep 14:1021–1027
Parenti R, Perris R, Vecchio GM, Salvatorelli L, Torrisi A, Gravina L, Magro G (2013) Immunohistochemical expression of Wilms’ tumor protein (WT1) in developing human epithelial and mesenchymal tissues. Acta Histochem 115:70–75
Parenti R, Salvatorelli L, Musumeci G, Parenti C, Giorlandino A, Motta F, Magro G (2015) Wilms’ tumor 1 (WT1) protein expression in human developing tissues. Acta Histochem 117:386–396
Patek CE, Little MH, Fleming S, Miles C, Charlieu JP, Clarke AR, Miyagawa K, Christie S, Doig J, Harrison DJ, Porteous DJ, Brookes AJ, Hooper ML, Hastie ND (1999) A zinc finger truncation of murine WT1 results in the characteristic urogenital abnormalities of Denys-Drash syndrome. Proc Natl Acad Sci U S A 96:2931–2936
Pelletier J, Bruening W, Kashtan CE, Mauer SM, Manivel JC, Striegel JE, Houghton DC, Junien C, Habib R, Fouser L, Fine RN, Silverman BL, Haber DA, Housman D (1991) Germline mutations in the Wilms’ tumor suppressor gene are associated with abnormal urogenital development in Denys-Drash syndrome. Cell 67:437–447
Pritchard-Jones K, Fleming S, Davidson D, Bickmore W, Porteous D, Gosden C, Bard J, Buckler A, Pelletier J, Housman D, Van Heyningen V, Hastie N (1990) The candidate Wilms’ tumour gene is involved in genitourinary development. Nature 346:194–197
Ramanathan AS, Vijayan M, Rajagopal S, Rajendiran P, Senguttuvan P (2017) WT1 and NPHS2 gene mutation analysis and clinical management of steroid-resistant nephrotic syndrome. Mol Cell Biochem 426:177–181
Ramani P, Cowell JK (1996) The expression pattern of Wilms’ tumour gene (WT1) product in normal tissues and paediatric renal tumours. J Pathol 179:162–168
Rauscher FJ, Morris JF, Tournay OE, Cook DM, Curran T (1990) Binding of the Wilms’ tumor locus zinc finger protein to the EGR-1 consensus sequence. Science 250:1259–1262
Royer-Pokora B (2013) Genetics of pediatric renal tumors. Pediatr Nephrol 28:13–23
Sahnane N, Magnoli F, Bernasconi B, Tibiletti MG, Romualdi C, Pedroni M, Ponz de Leon M, Magnani G, Reggiani-Bonetti L, Bertario L, Signoroni S, Capella C, Sessa F, Furlan D, AIFEG (2015) Aberrant DNA methylation profiles of inherited and sporadic colorectal cancer. Clin Epigenetics 7:131
Salvatorelli L, Bisceglia M, Vecchio G, Parenti R, Galliani C, Alaggio R, Gurrera A, Giurato E, Torrisi A, Magro G (2011) A comparative immunohistochemicalstudy of oncofetal cytoplasmic WT1 expression in human fetal, adult and neoplastic skeletal muscle. Pathologica 103:186
Salvatorelli L, Parenti R, Leone G, Musumeci G, Vasquez E, Magro G (2015) Wilms tumor 1 (WT1) protein: diagnostic utility in pediatrictumors. Acta Histochem 117:367–378
Scharnhorst V, Dekker P, van der Eb AJ, Jochemsen AG (1999) Internal translation initiation generates novel WT1 protein isoforms with distinct biological properties. J Biol Chem 274:23456–23462
Schittenhelm J, Thiericke J, Nagel C, Meyermann R, Beschorner R (2010) WT1 expression in normal and neoplastic cranial and peripheral nerves is independent of grade of malignancy. Cancer Biomark 7:73–77
Schnerwitzki D, Perner B, Hoppe B, Pietsch S, Mehringer R, Hänel F, Englert C (2014) Alternative splicing of Wilms tumor suppressor 1 (WT1) exon 4 results in protein isoforms with different functions. Dev Biol 393:24–32
Schüz J, Kaletsch U, Meinert R, Kaatsch P, Spix C, Michaelis J (2001) Risk factors for neuroblastoma at different stages of disease. Results from a population-based case-control study in Germany. J Clin Epidemiol 54:702–709
Schwab M, Westermann F, Hero B, Berthold F (2003) Neuroblastoma: biology and molecular and chromosomal pathology. Lancet Oncol 4:472–480
Scuderi S, La Cognata V, Drago F, Cavallaro S, D’Agata V (2014) Alternative splicing generates different parkin protein isoforms: evidences in human, rat, and mouse brain. Biomed Res Int 2014:690796
Sebire NJ, Gibson S, Rampling D, Williams S, Malone M, Ramsay AD (2005) Immunohistochemical findings in embryonal small round cell tumors with molecular diagnostic confirmation. Appl Immunohistochem Mol Morphol 13:1–5
Sharma PM, Yang X, Bowman M, Roberts V, Sukumar S (1992) Molecular cloning of rat Wilms’ tumor complementary DNA and a study of messenger RNA expression in the urogenital system and the brain. Cancer Res 52:6407–6412
Sharma PM, Bowman M, Madden SL, Rauscher FJ III, Sukumar S (1994) RNA editing in the Wilms’ tumor susceptibility gene, WT1. Genes Dev 8:720–731
Simpson LA, Burwell EA, Thompson KA, Shahnaz S, Chen AR, Loeb DM (2006) The antiapoptotic gene A1/BFL1 is a WT1 target gene that mediates granulocytic differentiation and resistance to chemotherapy. Blood 107:4695–4702
Smith SI, Down M, Boyd AW, Li CL (2000) Expression of the Wilms’ tumor suppressor gene, WT1, reduces the tumorigenicity of the leukemic cell line M1 in C.B-17 scid/scid mice. Cancer Res 60:808–814
Spix C, Pastore G, Sankila R, Stiller CA, Steliarova-Foucher E (2006) Neuroblastoma incidence and survival in European children (1978-1997): report from the Automated Childhood Cancer Information System project. Eur J Cancer 42:2081–2091
Tadokoro K, Oki N, Fujii H, Ohshima A, Inoue T, Yamada M (1992) Genomic organization of the human WT1 gene. Jpn J Cancer Res 83:1198–1203
Tatsumi N, Hojo N, Sakamoto H, Inaba R, Moriguchi N, Matsuno K, Fukuda M, Matsumura A, Hayashi S, Morimoto S, Nakata J, Fujiki F, Nishida, Nakajima H, Tsuboi A, Oka Y, Hosen N, Sugiyama H, Oji Y (2015) Identification of a novel C-terminal truncated WT1 isoform with antagonistic effects against major WT1 isoforms. PLoS One 10:e0130578
Wagner KD, Wagner N, Schedl A (2003) The complex life of WT1. J Cell Sci 116:1653–1658
Wang ZY, Qiu QQ, Enger KT, Deuel TF (1993) A second transcriptionally active DNA-binding site for the Wilms tumor gene product, WT1. Proc Natl Acad Sci U S A 90:8896–8900
Wang J, Oue T, Uehara S, Yamanaka H, Oji Y, Fukuzawa M (2011) The role of WT1 gene in neuroblastoma. J PediatrSurg 46:326–331
Yang L, Han Y, Suarez SF, Minden MD (2007) A tumor suppressor and oncogene: the WT1 story. Leukemia 21:868–876
Zitzmann F, Mayr D, Berger M, Stehr M, von Schweinitz D, Kappler R, Hubertus J (2014) Frequent hypermethylation of a CTCF binding site influences Wilms tumor 1 expression in Wilms tumors. Oncol Rep 31:1871–1876
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interests.
Rights and permissions
About this article
Cite this article
Rasà, D.M., D’Amico, A.G., Maugeri, G. et al. WT1 Alternative Splicing: Role of Its Isoforms in Neuroblastoma. J Mol Neurosci 62, 131–141 (2017). https://doi.org/10.1007/s12031-017-0930-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-017-0930-0