Abstract
This study aimed to develop and validate a nomogram based on immune checkpoint genes (ICGs) for predicting prognosis and immune checkpoint blockade (ICB) efficacy in lung adenocarcinoma (LUAD) patients. A total of 385 LUAD patients from the TCGA database and 269 LUAD patients in the combined dataset (GSE41272 + GSE50081) were divided into training and validation cohorts, respectively. Three different machine learning algorithms including random forest (RF), least absolute shrinkage and selection operator (LASSO) logistic regression analysis, and support vector machine (SVM) were employed to select the predictive markers from 82 ICGs to construct the prognostic nomogram. The X-tile software was used to stratify patients into high- and low-risk subgroups based on the nomogram-derived risk scores. Differences in functional enrichment and immune infiltration between the two subgroups were assessed using gene set variation analysis (GSVA) and various algorithms. Additionally, three lung cancer cohorts receiving ICB therapy were utilized to evaluate the ability of the model to predict ICB efficacy in the real world. Five ICGs were identified as predictive markers across all three machine learning algorithms, leading to the construction of a nomogram with strong potential for prognosis prediction in both the training and validation cohorts (all AUC values close to 0.800). The patients were divided into high- (risk score ≥ 185.0) and low-risk subgroups (risk score < 185.0). Compared to the high-risk subgroup, the low-risk subgroup exhibited enrichment in immune activation pathways and increased infiltration of activated immune cells, such as CD8 + T cells and M1 macrophages (P < 0.05). Furthermore, the low-risk subgroup had a greater likelihood of benefiting from ICB therapy and longer progression-free survival (PFS) than did the high-risk subgroup (P < 0.05) in the two cohorts receiving ICB therapy. A nomogram based on ICGs was constructed and validated to aid in predicting prognosis and ICB treatment efficacy in LUAD patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Non-small cell lung cancer (NSCLC) is a highly prevalent and deadly cancer worldwide, with approximately 2.2 million new cases and 1.8 million deaths reported annually [1]. The 5-year overall survival (OS) for NSCLC patients remains below 30% despite the continuous introduction of advanced diagnostic and therapeutic methods [2, 3]. The emerging immune checkpoint blockade (ICB) therapy has revolutionized the treatment landscape of NSCLC, particularly for patients in advanced stages or who are resistant to standard therapies [4,5,6]. However, a significant proportion of NSCLC patients still do not respond to ICB therapy effectively. Statistics have shown that approximately 40% of patients with high levels of PD-L1 show a positive response to treatment. Despite efforts to identify novel biomarkers such as PD-1/PD-L1, tumor mutational burden (TMB), and microsatellite instability (MSI) to predict the efficacy of ICB therapy [7,8,9], establishing new methods for predicting ICB treatment response in these patients is still needed.
Extensive research has been conducted on immune checkpoint genes (ICGs) to identify the most effective target genes involved in immune activation and evasion processes. While anti-PD-1/PD-L1 and CTLA-4 have been well-established as mechanisms for ICB therapies, emerging targets such as TIGIT, LAG3, and TIM-3 have garnered an increasing amount of attention and are currently being developed for clinical use [10,11,12,13]. This highlighted the importance of investigating the significance of ICGs in predicting prognosis and the efficacy of ICB therapy for cancer patients. High-throughput sequencing technique has been widely accepted as the optimal tool for the comprehensive interrogation of gene expression changes in cancer patients [14]. With the vast amount of data generated by high-throughput sequencing, machine learning algorithms have been introduced to analyze these datasets and have demonstrated a superior ability to identify biologically significant alteration profiles in high-dimensional datasets [15, 16]. Few previous studies have made some attempts to investigate the significance of ICGs in LUAD patients, but there is still room for improvement in this area. As such, Ling et al. first explored the prognostic role of ICGs in LUAD patients, but only half of the documented ICGs were investigated in their study [17]. In addition, the study from Jia et al. focused on the ability of ICGs to predict lymph node metastases in LUAD patients instead of assessing prognosis or ICB treatment efficacy [18]. Recent studies exploring the prognostic significance of ICGs failed to introduce multiple machine learning methods in the gene selection process, resulting in relatively low predictive power and limited ability to guide ICB treatment decisions [19].
Given the importance of ICGs in ICB therapy, our current study aimed to comprehensively investigate the significance of ICG expression profiles in predicting the prognosis and efficacy of ICB therapy for LUAD patients using machine learning methods.
Materials and methods
Data source
LUAD patients from The Cancer Genome Atlas (TCGA) (https://portal.gdc.cancer.gov/) (sequencing technique: IlluminaHiSeq_RNASeqV2), where data on patient survival, clinical features, and gene expression profiles were acquired, composed the training cohort. In contrast, validation was performed using the cohort that combines GSE41271 (sequencing technique: Illumina HumanWG-6 v3.0 expression beadchip) and GSE50081 (sequencing technique: Affymetrix Human Genome U133 Plus 2.0 Array) from the Gene Expression Omnibus database (https://www.ncbi.nlm.nih.gov/geo/). Patients who met the following criteria were excluded: (I) had a non-LUAD diagnostic pathology, (II) had a normal status or tissue, (III) patients who died within 30 days of the follow-up, (IV) had synchronous malignancies, or (V) had incomplete follow-up information regarding the survival time or status. Subsequently, the collected RNA sequencing data were normalized and corrected in batches with the “SVA package” in both the training and validation cohorts [20]. Information on the lung cancer patients who received ICB therapy was obtained from three independent cohorts, namely, the GSE126044 (sequencing technique: Illumina HiSeq 2500), GSE135222 (sequencing technique: Illumina HiSeq 2500) (https://www.ncbi.nlm.nih.gov/geo/), and Nanfang Hospital cohort (https://figshare.com/articles/dataset/Nanfang_hospital_NSCLC_immunotherapy_cohort). An ICG list containing 95 genes was obtained from previous literature exploring the mechanisms of ICGs in cancers [13, 21,22,23] (Table S1).
Selection of prognostic ICGs using machine learning algorithms
The selection process for ICGs in the training cohort involved three independent machine learning algorithms, including least absolute shrinkage and selection operator (LASSO) logistic regression [24], random forest (RF) [25], and the support vector machine (SVM) [26]. The “glmnet” R package was utilized to conduct the LASSO regression, which was useful for eliminating the nonsignificant variables by shrinking the size of the parameters. In this study, a tenfold cross-validation was performed to select the optimal lambda that met the minimum criteria. For RF selection, the “randomForest” package was utilized with 200 Monte Carlo iterations. ICGs with a relative importance greater than 0 were considered significant. Although the SVM was originally built for classification, it has recently been used for feature selection through the wrapper method with linear kernels. In this study, SVM was performed using the “e1071” R package where the prediction accuracy did not increase with the number of variables incorporated into the prediction model when the optimal number of variables was reached.
Construction and validation of the prognostic nomogram
The Venn diagram was used to identify the ICGs in the intersection part of the three machine learning algorithms. These selected ICGs were then incorporated into the development of the prognostic nomogram. The ability of the model to predict 1-year, 3-year, and 5-year OS for patients was evaluated using the area under the receiver operating characteristic curve (AUC). Calibration curves were also plotted to visualize the predictive accuracy of the model, where the closer the predictive curve was to the observed curve, the better the predictive accuracy of the model was [27]. The validation cohort was utilized to externally validate the results and the area under the curve (AUC) and calibration curves were also generated for the validation cohort. X-tile software (version 3.6.1; Yale University, New Haven, CT, USA) was used to determine the optimal cutoff value for the risk scores derived from the nomogram, which divided the patients into high- and low-risk subgroups [28]. Kaplan–Meier curves of different risk subgroups were plotted accordingly for both the training and validation cohorts. Moreover, the survival curves of each risk subgroup within different clinical subsets were also plotted. Principal component analysis (PCA) was conducted to visualize the clustering pattern of each risk subgroup.
Differences in functional enrichment between two risk subgroups
Gene set variation analysis (GSVA) was the principle method used to interrogate the biological and molecular differences between two risk subgroups [29]. We downloaded the h.all.v2023.1.Hs.symbols.gmt, c5.go.v2023.1.Hs.symbols.gmt, and c2.cp.kegg.v2023.1.Hs.
symbols.gmt from the Molecular Signatures Database (MSigDB) (https://www.gsea-msigdb.org/gsea/msigdb/index.jsp) for the broad hallmark genes, Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis in the GSVA, respectively [30], where a normal P < 0.05 and a false discovery rate (FDR) < 0.1 were set as the thresholds for the recognition of the significantly enriched gene sets. Additionally, gene set variation analysis (GSEA) was also performed to validate the results from GSVA, for which the “enrichplot” and “clusterProfier” R package were needed [31].
Comparison of the immune infiltration landscape between two risk subgroups
We performed the most comprehensive comparison of the immune infiltration landscape between the two risk subgroups using various algorithms, including CIBERSORT, ESTIMATE, EPIC, MCPCOUNTER, QUANTISEQ, ssGSEA, TIMER, and X_CELL, which provided us with a comprehensive understanding of the distinct microimmune landscape between the two subgroups [32,33,34,35,36,37,38,39].
Predicting the response to ICB therapy
The Tumor Immune Dysfunction and Exclusion (TIDE) score for each patient in the training cohort was calculated using the website (http://tide.dfci.harvard.edu/), which was helpful for identifying the patients who were more likely to benefit from ICB therapy [40]. Patients with higher TIDE scores tended to suffer from immune evasion and therefore be less responsive to ICB therapy. Subsequently, the TIDE scores between different risk subgroups were compared using the Wilcoxon rank sum test, and the correlation between TIDE scores and the risk scores derived from the nomogram was also investigated. Furthermore, the nomogram-derived risk scores were compared between responders and non-responders to ICB in the GSE126044 cohort. In addition, we plotted Kaplan–Meier curves to illustrate the differences in progression-free survival (PFS) among the different risk subgroups after receiving ICB therapy in the GSE135222 cohort and Nanfang Hospital cohort.
Statistical analysis
Categorical variables were presented as percentages or frequencies and were compared using the Pearson χ2 test or Fisher’s exact test, while continuous variables were described as medians with interquartile ranges (IQRs) and were compared with t-tests or Wilcoxon rank sum tests. The correlation between two continuous variables was estimated using the Spearman test. OS was defined as the time from the date of diagnosis to the date of all-cause death or censoring. Similarly, PFS represented the time interval between diagnosis and disease progression, recurrence, or death. Moreover, the survival outcomes were estimated and compared using the Kaplan–Meier method and log-rank test, respectively. All analyses were performed using R software (version.4.3.0; http://www.r-project.org). A two-tailed P < 0.05 was considered statistically significant.
Results
Identification of ICGs used for model construction
The workflow of the study is presented in detail in Fig. 1. After screening, a total of 385 LUAD patients in the TCGA database and 269 LUAD patients in the combined dataset (GSE41272 + GSE50081) were assigned to the training and validation cohorts, respectively. The clinical characteristics of these patients are summarized in Table 1.
The flow chart comprehensively depicted the procedure of the current study. ICGs, immune checkpoint genes; LASSO, least absolute shrinkage and selection operator; RF, random forest; SVM, support vector machine; PCA, principal component analysis; GSVA, gene set variation analysis; GSEA, gene set enrichment analysis
Among the 95 documented immune checkpoint genes (ICGs), 82 were sequenced in the TCGA-LUAD cohort. The specific names of the 82 ICGs are listed in Table.S2. The prognostic significance of the 82 ICGs was assessed using three different machine learning algorithms. Through LASSO logistic regression analysis, nine prognostic ICGs were identified when the optimal lambda value was reached (Fig. 2 a, b). The 82 ICGs were also subjected to the random forest algorithm, which showed that the error rate stabilized between 0.42 and 0.46 as the number of trees increased. Based on the importance ranking, 34 ICGs that positively contributed to the algorithm were selected (Fig. 2c). Furthermore, SVM was used to identify 12 prognostic ICGs, as the prediction accuracy did not improve when the number of included variables reached 12 (Fig. 2d). Ultimately, five ICGs were found in the overlapping region of the three machine learning algorithms, as shown in the Venn diagram (Fig. 2e).
The selection process of the prognostic ICGs. a, b the LASSO-based method to screen the ICGs and different colors means different genes; c the selection process via RF and the genes with blue bar would be selected for the model (variable importance > 0); d the SVM-based selection method; e Venn diagram marked the intersection of prognostic ICGs obtained by three different algorithms. ICGs, immune checkpoint genes; LASSO, least absolute shrinkage and selection operator; RF, random forest; SVM, support vector machine
Construction and validation of the ICGs-based nomogram
The five ICGs incorporated in the nomogram were VTCN1 (B7-H4), KIR3DL3, KIR2DL1, HLA-DRB5, and CD209 (DC-SIGN) (Fig. 3a). By drawing a line straight up from each independent risk factor to the point axis, we assigned each risk factor with one specific score, and the total score for each patient was obtained by adding each score together. Additionally, the risk score could be calculated through the following equation: risk score = ∑i expi × coefi (derived from the nomogram). The receiver operating characteristic (ROC) curves for 1-year, 3-year, and 5-year overall survival (OS) prediction demonstrated the excellent predictive power of the model in both the training and validation cohorts, with all AUC values closing to 0.800 (Fig. 3 b, c). Exceedingly, the calibration curves of the nomogram showed optimal consistency between the predicted and observed survival probabilities in both the training and validation cohorts (Fig. 3 d–i).
a Prognostic nomogram combining the selected 5 ICGs to predict the 1 year, 3 years, and 5 year OS for patients in the training cohort; The areas under the curves of the nomogram to predict 1 year, 3 years, and 5 years OS for patients in the training cohort (b) and validation cohort (c); Calibration curves of 1 year (d), 3 years (e), and 5 years OS (f) for patients in the training cohort and the calibration curves of 1 year (g), 3 years (h), and 5 years OS (i) in the validation cohort
Using X-tile software, we stratified the patients in the training cohort into low-risk (total score < 185.0) and high-risk (total score ≥ 185.0) subgroups based on the scores derived from the nomogram when the maximal chi-square value was reached, which represented the greatest differences in prognosis prediction among the subgroups (Fig. S1) [28].
Survival analysis of different risk subgroups
Kaplan–Meier curves were plotted to assess the differences in survival between the low-risk and high-risk subgroups in both the training and validation cohorts, and significant differences were observed (all P < 0.001) (Fig. 4 a, b). Furthermore, we plotted Kaplan–Meier curves based on risk subgroups within different subsets according to various clinical characteristics such as gender, age, clinical stage, and race, which demonstrated the promising ability of risk subgroups to differentiate the prognosis of LUAD patients regardless of the clinical characteristics (all P < 0.05) (Fig.S2a-g). However, for nonwhite patients, the difference in survival between risk subgroups was not statistically significant (P = 0.27) (Fig.S2h), which could be attributed to the extremely small population of nonwhite patients (84 total nonwhite patients were included, and the number of the high-risk group was 16).
Moreover, the PCA depicted the distinguishable distribution patterns of the five ICGs incorporated in the nomogram between the high- and low-risk subgroups in the training (Fig. 4c) and validation cohorts (Fig. 4d). PC1 clearly separated from PC2 in both cohorts.
Differences in the biological pathway enrichment between risk subgroups
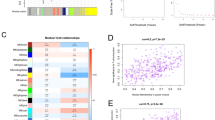
GSVA was performed to identify functional pathway differences between the low-risk and high-risk subgroups. In the high-risk subgroup, several pathways were found to be enriched including glutamine catabolic process, megakaryocyte differentiation regulation in biological process (BP), nucleosome, catenin complex in cellular component (CC), and glycerophospholipid activity and polypeptide transferase activity in molecular function (MF) (Fig. 5 a–c). In addition, hallmark gene sets such as pancreas beta cells and KRAS signaling were observed to be enriched in the high-risk subgroup, while interferon-gamma response and PI3K-AKT-mTOR signaling were enriched in the low-risk subgroup (Fig. 5d). Additionally, glycan biosynthesis and linoleic acid metabolism in the KEGG pathway were enriched in the high-risk subgroup (Fig. 5e). Notably, the correlation between GSVA scores (KEGG pathways) and risk scores was assessed and displayed by a lollipop plot, which revealed a negative correlation between the risk score and a significant proportion of immune-related pathways (Fig. 5f). Moreover, we performed GSEA to verify the results of GSVA, which showed good consistency between these two methods (Fig.S3). This further confirmed the significant differences in functional enrichment between the low-risk and high-risk subgroups.
Gene set variation analysis (GSVA) was conducted to show the differences between the high- and low-risk groups in terms of Gene ontology (GO) biological process (a), GO cellular component (b), GO molecular function gene sets (c), hallmark gene sets (d), and Kyoto Encyclopedia of Genes and Genomes (KEGG) gene sets (e); lollipop plot demonstrated the correlation between GSVA scores (KEGG) and nomogram-derived risk scores (f)
Comparison of immune infiltration profiles between risk subgroups
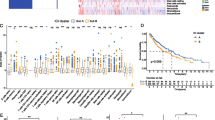
A comprehensive comparison of immune infiltration profiles between the two risk subgroups was performed using various algorithms including CIBERSORT, ESTIMATE, EPIC, MCPCOUNTER, QUANTISEQ, ssGSEA, TIMER, and X_CELL. Based on CIBERSORT, 22 infiltrating immune cells were counted. It was observed that M1 macrophages were significantly elevated in the low-risk subgroup (P < 0.001), while immune-resistant cells such as T-cell regulatory cells (Tregs) were more abundant in the high-risk subgroup (P < 0.01). Additionally, CD8 + T cells and activated NK cells were more strongly infiltrated in the low-risk subgroup than in the high-risk subgroup (P < 0.05) (Fig. 6a). Furthermore, the ESTIMATE algorithm was used to calculate the abundance of infiltrating immune cells in the two risk subgroups, which revealed that the low-risk subgroup had higher immune scores (P < 0.01) and ESTIMATE scores (P < 0.05) than did the high-risk subgroup (Fig. 6b). In brief, there were more immune-activated patients in the low-risk subgroup than in the high-risk subgroup, which could be further confirmed through other algorithms (Fig. 6 c–h).
Clinical utility of the model in predicting the efficacy of ICB therapy
The TIDE algorithm was utilized to estimate the TIDE score for each patient, and the scores were compared between the two risk subgroups. The TIDE scores in the high-risk subgroup were significantly greater than those in the low-risk subgroup (P = 4.1 × 10−5) (Fig. 7a), which indicated that the patients in the high-risk subgroup were more likely to suffer from immune evasion and, therefore, were less likely to respond to ICB therapy. A positive correlation between TIDE scores and nomogram-derived risk scores was observed despite the correlation was not that strong (r = 0.37, P = 9.7 × 10−14) (Fig. 7b). In terms of the predictive performance of the nomogram for immunotherapy response, patients who did not respond to ICB therapy had higher risk scores than those who respond to ICB therapy in the GSE126044 cohort (Fig. 7c). Similarly, a greater proportion in the low-risk subgroup than in the high-risk subgroup responded to ICB therapy (40% vs 16.7%) (Fig. 7d). Further validation was carried out using the GSE135222 cohort and Nanfang Hospital cohort, in which patients received similar ICB treatment, and the clinical characteristics of the included patients are presented in Table.S3. As a result, patients in the low-risk subgroup had longer PFS than those in the high-risk subgroup (Fig. 7 e–g), which further indicated that the low-risk subgroup was more likely to benefit from ICB therapy than the high-risk subgroup.
The implementation of the nomogram-derived risk subgroups to predict the efficacy of ICB therapy for lung cancer patients. a Box plot exhibited the difference in TIDE score between the two risk subgroups in the training cohort; b correlation between TIDE score and nomogram-derived score in the training cohort; c comparison of nomogram-derived score between different outcome subgroups in GSE126044 cohort; d comparison of the responding proportion to ICB therapy in different nomogram-derived risk subgroups in GSE126044 cohort; Kaplan–Meier progression-free survival curves to display the differences in the efficacy of ICB therapy in nomogram-derived risk subgroups in GSE135222 cohort (e), Nanfang hospital cohort (f) and GSE135222 + Nanfang hospital cohort (g). ICB, immune checkpoint blockade
Discussion
The PACIFIC trial has revolutionized the treatment landscape for advanced NSCLC patients by demonstrating the importance of adding immune checkpoint blockade (ICB) therapy to standard treatments [41]. Subsequently, studies have investigated the efficacy of ICB therapy in early-stage NSCLC patients, suggesting that ICB therapy is potentially applicable in any stage of NSCLC [42, 43]. However, a significant proportion of patients still do not respond effectively to ICB treatment, despite advancements in predicting responsiveness using biomarkers. Therefore, further research aimed at identifying promising predictive biomarkers or predictive models remains crucial for NSCLC patients, especially those with LUAD, which accounts for more than half of all NSCLC patients.
The expression of ICGs has long been recognized for its role in modulating the tumor microenvironment and impacting the efficacy of ICB therapy. Herein, we conducted a comprehensive analysis of the ability of ICGs to predict ICB response in patients with LUAD by introducing three different machine learning algorithms to identify the most significant ICGs for predicting prognosis and ICB efficacy. A nonparametric nonlinear technique was used in the RF model, which combines the effects of the expression of ICGs to reach the goal of classification [25]. Similarly, the LASSO regression and SVM were employed to reduce the number of ICGs incorporated into the model, aiming to minimize classification errors and maximize prediction accuracy [24, 26]. As a result, our predictive nomogram possessed significantly improved predictive ability compared to that of previous studies [19]. The AUC values in our study ranged from 0.776 to 0.801, whereas in the previous study, the AUC values ranged from 0.669 to 0.736. This further underscores the importance of incorporating various machine learning methods in the variable selection process, as it enhances the accuracy and robustness of the predictive model.
In this study, a total of five ICGs were identified through machine learning algorithms. Among them, VTCN1 (B7-H4) showed a positive correlation with risk score and a negative correlation with the survival outcome of LUAD patients (Fig. 3a), which was consistent with previous studies that have described VTCN1 as an immune-suppressive factor and reported a negative association between VTCN1 expression and immune cell infiltration [44, 45]. The kill-cell immunoglobulin-like receptor (KIR) family, chiefly expressed by nature killer (NK) cells, plays a role in controlling inflammatory and anti-inflammatory regulation. The KIR3DL3, expressed by CD56dim NK cells, mediates the differentiation of CD8 + T cells and suppresses their immune activation ability [46, 47]. However, KIR2DL1, an activating KIR, acts as a proinflammatory factor, and high expression of KIR2DL1 has been associated with longer survival time [48]. Furthermore, the expression of HLA-DRB5 has been reported to be associated with better prognosis in LUAD patients [49], which was further validated in our study where HLA-DRB5 was negatively correlated with OS (Fig. 3a). Moreover, CD209 (DC-SIGN) has long been recognized as a target of dendritic cells (DCs), and anti-CD209 antibodies can activate microenvironment immunity in vivo, leading to a persistent and strong CD8 + T-cell response [50].
As a result, we constructed a nomogram based on the five selected ICGs that possessed good ability to predict the prognosis of LUAD patients. According to the model, patients were divided into high- and low-risk subgroups. We observed significant differences in the microimmune environment between the two subgroups, with the low-risk subgroup showing a more pronounced immune activation environment, which was supported by the enrichment of the DNA damage repair hallmark in the low-risk subgroup (Fig. 5d), as DNA damage repair plays a crucial role in triggering innate immunity [51, 52]. Additionally, we calculated the infiltration of immune cells in the two risk subgroups using various methods, and the results confirmed that the low-risk subgroup had a greater abundance of immune activation cells, such as CD8 + T cells, NK cells, and M1 macrophages, further underpinning the immune activation status of the low-risk subgroup. Furthermore, our model was able to differentiate lung cancer patients who could benefit from ICB treatment from those who could not, which highlights the clinical utility of our model in identifying patients who are likely to respond to ICB therapy, thereby increasing its applicability in the real world.
In this study, we utilized sophisticated machine learning algorithms to assess the ability of ICGs to predict prognosis and ICB efficacy in LUAD patients, resulting in a robust nomogram. Additionally, the most comprehensive algorithms including CIBERSORT, ESTIMATE, EPIC, MCPCOUNTER, QUANTISEQ, ssGSEA, TIMER, and X_CELL were used to calculate and compare the immune infiltration between the high- and low-risk subgroups derived from the model. However, several limitations in our study also need to be noted. First, as this was a retrospective study spanning a long period, selection bias in terms of diagnostic methods, sequencing technique, and follow-up was difficult to avoid. Second, the validation datasets used to assess the predictive performance of the model for immunotherapy response included both LUAD and lung squamous cell carcinoma (LUSC) patients, which might slightly impair the predictive accuracy of the model. Third, the information on the common genetic mutational status of LUAD, such as that of EGFR, was not addressed in the database, and the prognostic significance of these genetic mutations could be further explored in the future. Last, due to the bioinformatics nature of this study, the results have not yet been confirmed and validated in real-world studies. Therefore, well-designed clinical trials to thoroughly explore and verify the prognostic significance and ability of this model to screen LUAD patients for ICB therapy are needed in the future. For example, the model could be assessed using only LUAD patients receiving ICB therapy, while a new model based on LUSC gene data could be developed for LUSC patients. In addition, RNA sequencing could be performed in the real-world cohort to validate the predictive ability of the model.
Conclusion
Three different machine learning algorithms were used in this study to select the most predictive ICGs for LUAD patients, and a nomogram was constructed and validated accordingly, which exhibited good accuracy in predicting the prognosis of LUAD patients (with an AUC value close to 0.8). Furthermore, we divided the patients into two risk subgroups according to the scores derived from the model. The comparison between these two subgroups revealed that the low-risk subgroup had a greater potential to benefit from ICB therapy than did the high-risk subgroup.
Data availability
No datasets were generated or analysed during the current study.
References
Thai AA, Solomon BJ, Sequist LV, Gainor JF, Heist RS. Lung cancer. Lancet. 2021;398(10299):535–54. https://doi.org/10.1016/S0140-6736(21)00312-3.
Alexander M, Kim SY, Cheng H. Update 2020: management of non-small cell lung cancer. Lung. 2020;198(6):897–907. https://doi.org/10.1007/s00408-020-00407-5.
Guerreiro T, Mayer A, Aguiar P, Araújo A, Nunes C. The effect of timeliness of care on lung cancer survival - a population-based approach. Ann Glob Health. 2023;89(1):39. https://doi.org/10.5334/aogh.3845.
Duma N, Santana-Davila R, Molina JR. Non-small cell lung cancer: epidemiology, screening, diagnosis, and treatment. Mayo Clin Proc. 2019;94(8):1623–40. https://doi.org/10.1016/j.mayocp.2019.01.013.
Hellmann MD, Paz-Ares L, Bernabe Caro R, et al. Nivolumab plus ipilimumab in advanced non-small-cell lung cancer. N Engl J Med. 2019;381(21):2020–31. https://doi.org/10.1056/NEJMoa191023.
Herbst RS, Giaccone G, de Marinis F, et al. Atezolizumab for first-line treatment of PD-L1-selected patients with NSCLC. N Engl J Med. 2020;383(14):1328–39. https://doi.org/10.1056/NEJMoa1917346.
Reck M, Rodríguez-Abreu D, Robinson AG, et al. Five-year outcomes with pembrolizumab versus chemotherapy for metastatic non-small-cell lung cancer with PD-L1 tumor proportion score ≥ 50. J Clin Oncol. 2021;39(21):2339–49. https://doi.org/10.1200/JCO.21.00174.
Havel JJ, Chowell D, Chan TA. The evolving landscape of biomarkers for checkpoint inhibitor immunotherapy. Nat Rev Cancer. 2019;19(3):133–50. https://doi.org/10.1038/s41568-019-0116-x.
Wu X, Gu Z, Chen Y, et al. Application of PD-1 blockade in cancer immunotherapy. Comput Struct Biotechnol J. 2019;17:661–74. https://doi.org/10.1016/j.csbj.2019.03.006.
Jiang Y, Chen M, Nie H, Yuan Y. PD-1 and PD-L1 in cancer immunotherapy: clinical implications and future considerations. Hum Vaccin Immunother. 2019;15(5):1111–22. https://doi.org/10.1080/21645515.2019.1571892.
Qin S, Xu L, Yi M, Yu S, Wu K, Luo S. Novel immune checkpoint targets: moving beyond PD-1 and CTLA-4. Mol Cancer. 2019;18(1):155. https://doi.org/10.1186/s12943-019-1091-2.
Andrews LP, Marciscano AE, Drake CG, Vignali DA. LAG3 (CD223) as a cancer immunotherapy target. Immunol Rev. 2017;276(1):80–96. https://doi.org/10.1111/imr.12519.
Pardoll DM. The blockade of immune checkpoints in cancer immunotherapy. Nat Rev Cancer. 2012;12(4):252–64. https://doi.org/10.1038/nrc3239.
Bacchelli C, Williams HJ. Opportunities and technical challenges in next-generation sequencing for diagnosis of rare pediatric diseases. Expert Rev Mol Diagn. 2016;16(10):1073–82. https://doi.org/10.1080/14737159.2016.1222906.
Choi RY, Coyner AS, Kalpathy-Cramer J, Chiang MF, Campbell JP. Introduction to machine learning, neural networks, and deep learning. Transl Vis Sci Technol. 2020;9(2):14. https://doi.org/10.1167/tvst.9.2.14.
Greener JG, Kandathil SM, Moffat L, Jones DT. A guide to machine learning for biologists. Nat Rev Mol Cell Biol. 2022;23(1):40–55. https://doi.org/10.1038/s41580-021-00407-0.
Ling B, Ye G, Zhao Q, Jiang Y, Liang L, Tang Q. Identification of an immunologic signature of lung adenocarcinomas based on genome-wide immune expression profiles. Front Mol Biosci. 2021;7:603701. https://doi.org/10.3389/fmolb.2020.603701.
Jia R, Sui Z, Zhang H, Yu Z. Identification and validation of immune-related gene signature for predicting lymph node metastasis and prognosis in lung adenocarcinoma. Front Mol Biosci. 2021;8:679031. https://doi.org/10.3389/fmolb.2021.679031.
Hua L, Wu J, Ge J, et al. Identification of lung adenocarcinoma subtypes and predictive signature for prognosis, immune features, and immunotherapy based on immune checkpoint genes. Front Cell Dev Biol. 2023;11:1060086. https://doi.org/10.3389/fcell.2023.1060086.
Leek JT, Johnson WE, Parker HS, Jaffe AE, Storey JD. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics. 2012;28(6):882–3. https://doi.org/10.1093/bioinformatics/bts034.
Xu D, Liu X, Wang Y, et al. Identification of immune subtypes and prognosis of hepatocellular carcinoma based on immune checkpoint gene expression profile. Biomed Pharmacother. 2020;126:109903. https://doi.org/10.1016/j.biopha.2020.109903.
Campbell KS, Purdy AK. Structure/function of human killer cell immunoglobulin-like receptors: lessons from polymorphisms, evolution, crystal structures and mutations. Immunology. 2011;132(3):315–25. https://doi.org/10.1111/j.1365-2567.2010.03398.x.
Afrache H, Gouret P, Ainouche S, Pontarotti P, Olive D. The butyrophilin (BTN) gene family: from milk fat to the regulation of the immune response. Immunogenetics. 2012;64(11):781–94. https://doi.org/10.1007/s00251-012-0619-z.
Cheung-Lee WL, Link AJ. Genome mining for lasso peptides: past, present, and future. J Ind Microbiol Biotechnol. 2019;46(9–10):1371–9. https://doi.org/10.1007/s10295-019-02197-z.
Wang H, Yang F, Luo Z. An experimental study of the intrinsic stability of random forest variable importance measures. BMC Bioinformatics. 2016;17:60. https://doi.org/10.1186/s12859-016-0900-5.
Huang S, Cai N, Pacheco PP, Narrandes S, Wang Y, Xu W. Applications of support vector machine (SVM) learning in cancer genomics. Cancer Genomics Proteomics. 2018;15(1):41–51. https://doi.org/10.21873/cgp.20063.
Zuo Z, Zhang G, Song P, et al. Survival nomogram for stage IB non-small-cell lung cancer patients, based on the SEER database and an external validation cohort. Ann Surg Oncol. 2021;28(7):3941–50. https://doi.org/10.1245/s10434-020-09362-0.
Camp RL, Dolled-Filhart M, Rimm DL. X-tile: a new bio-informatics tool for biomarker assessment and outcome-based cut-point optimization. Clin Cancer Res. 2004;10(21):7252–9. https://doi.org/10.1158/1078-0432.CCR-04-0713.
Hänzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics. 2013;14:7. https://doi.org/10.1186/1471-2105-14-7.
Liberzon A, Birger C, Thorvaldsdóttir H, Ghandi M, Mesirov JP, Tamayo P. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 2015;1(6):417–25. https://doi.org/10.1016/j.cels.2015.12.004.
Subramanian A, Tamayo P, Mootha VK, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102(43):15545–50. https://doi.org/10.1073/pnas.0506580102.
Racle J, de Jonge K, Baumgaertner P, Speiser DE, Gfeller D. Simultaneous enumeration of cancer and immune cell types from bulk tumor gene expression data. Elife. 2017;6:e26476. https://doi.org/10.7554/eLife.26476.
Aran D, Hu Z, Butte AJ. xCell: digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 2017;18(1):220. https://doi.org/10.1186/s13059-017-1349-1.
Becht E, Giraldo NA, Lacroix L, et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression [published correction appears in Genome Biol. 2016 Dec 1;17 (1):249]. Genome Biol. 2016;17(1):218. https://doi.org/10.1186/s13059-016-1070-5.
Yoshihara K, Shahmoradgoli M, Martínez E, et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun. 2013;4:2612. https://doi.org/10.1038/ncomms3612.
Newman AM, Liu CL, Green MR, et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015;12(5):453–7. https://doi.org/10.1038/nmeth.3337.
Yi M, Nissley DV, McCormick F, Stephens RM. ssGSEA score-based Ras dependency indexes derived from gene expression data reveal potential Ras addiction mechanisms with possible clinical implications. Sci Rep. 2020;10(1):10258. https://doi.org/10.1038/s41598-020-66986-8.
Li T, Fu J, Zeng Z, et al. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res. 2020;48(W1):W509–14. https://doi.org/10.1093/nar/gkaa407.
Finotello F, Mayer C, Plattner C, et al. Molecular and pharmacological modulators of the tumor immune contexture revealed by deconvolution of RNA-seq data. Genome Med. 2019;11(1):34. https://doi.org/10.1186/s13073-019-0638-6.
Jiang P, Gu S, Pan D, et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med. 2018;24(10):1550–8. https://doi.org/10.1038/s41591-018-0136-1.
Antonia SJ, Villegas A, Daniel D, et al. Durvalumab after chemoradiotherapy in stage III non-small-cell lung cancer. N Engl J Med. 2017;377(20):1919–29. https://doi.org/10.1056/NEJMoa1709937.
Forde PM, Spicer J, Lu S, et al. Neoadjuvant nivolumab plus chemotherapy in resectable lung cancer. N Engl J Med. 2022;386(21):1973–85. https://doi.org/10.1056/NEJMoa2202170.
Wakelee H, Liberman M, Kato T, et al. Perioperative pembrolizumab for early-stage non-small-cell lung cancer. N Engl J Med. 2023;389(6):491–503. https://doi.org/10.1056/NEJMoa2302983.
Sica GL, Choi IH, Zhu G, et al. B7–H4, a molecule of the B7 family, negatively regulates T cell immunity. Immunity. 2003;18(6):849–61. https://doi.org/10.1016/s1074-7613(03)00152-3.
Wei J, Loke P, Zang X, Allison JP. Tissue-specific expression of B7x protects from CD4 T cell-mediated autoimmunity. J Exp Med. 2011;208(8):1683–94. https://doi.org/10.1084/jem.20100639.
Bhatt RS, Berjis A, Konge JC, et al. KIR3DL3 is an inhibitory receptor for HHLA2 that mediates an alternative immunoinhibitory pathway to PD1. Cancer Immunol Res. 2021;9(2):156–69. https://doi.org/10.1158/2326-6066.CIR-20-0315.
Leaton LA, Shortt J, Kichula KM, et al. Conservation, extensive heterozygosity, and convergence of signaling potential all indicate a critical role for KIR3DL3 in higher primates. Front Immunol. 2019;10:24. https://doi.org/10.3389/fimmu.2019.00024.
Guillamón CF, Martínez-Sánchez MV, Gimeno L, et al. Activating KIRs on educated NK cells support downregulation of CD226 and inefficient tumor immunosurveillance. Cancer Immunol Res. 2019;7(8):1307–17. https://doi.org/10.1158/2326-6066.CIR-18-0847.
Schaafsma E, Fugle CM, Wang X, Cheng C. Pan-cancer association of HLA gene expression with cancer prognosis and immunotherapy efficacy. Br J Cancer. 2021;125(3):422–32. https://doi.org/10.1038/s41416-021-01400-2.
Hesse C, Ginter W, Förg T, et al. In vivo targeting of human DC-SIGN drastically enhances CD8+ T-cell-mediated protective immunity. Eur J Immunol. 2013;43(10):2543–53. https://doi.org/10.1002/eji.201343429.
Paludan SR. Activation and regulation of DNA-driven immune responses. Microbiol Mol Biol Rev. 2015;79(2):225–41. https://doi.org/10.1128/MMBR.00061-14.
Bednarski JJ, Sleckman BP. At the intersection of DNA damage and immune responses. Nat Rev Immunol. 2019;19(4):231–42. https://doi.org/10.1038/s41577-019-0135-6.
Acknowledgements
Sincere gratitude would be sent to the academic workers who were responsible for the creation, update, and maintenance of the TCGA and GEO databases. Also, we would like to thank the team from Nanfang Hospital for uploading their data to the website.
Author information
Authors and Affiliations
Contributions
Xi-Lin Yang: Conceptualization; Data curation; Formal analysis; Methodology; Resources; Software; Validation; Visualization; Roles/Writing – original draft;Chen Wang: Data curation; Formal analysis; Methodology; Resources; Software; Validation; Visualization;Zheng Zeng: Methodology; Resources; Software; Validation; Visualization; Guang-Yu Wang: Methodology; Resources; Software; Validation; Visualization;Fu-Quan Zhang: Conceptualization; Funding acquisition; Investigation; Project administration; Resources; Writing – review & editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Xi-Lin Yang and Zheng Zeng are co-first authors.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, XL., Zeng, Z., Wang, C. et al. Prognostic model incorporating immune checkpoint genes to predict the immunotherapy efficacy for lung adenocarcinoma: a cohort study integrating machine learning algorithms. Immunol Res 72, 851–863 (2024). https://doi.org/10.1007/s12026-024-09492-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12026-024-09492-7