Abstract
Breast cancer resistance protein (ABCG2) is a human ATP-binding cassette (ABC) that plays a paramount role in multidrug resistance (MDR) in cancer therapy. The discovery of ABCG2 inhibitors could assist in designing unprecedented therapeutic strategies for cancer treatment. There is as yet no approved drug targeting ABCG2, although a large number of drug candidates have been clinically investigated. In this work, binding affinities of 181 drug candidates in clinical-trial or investigational stages as ABCG2 inhibitors were inspected using in silico techniques. Based on available experimental data, the performance of AutoDock4.2.6 software was first validated to predict the inhibitor-ABCG2 binding mode and affinity. Combined molecular docking calculations and molecular dynamics (MD) simulations, followed by molecular mechanics-generalized Born surface area (MM-GBSA) binding energy calculations, were then performed to filter out the studied drug candidates. From the estimated docking scores and MM-GBSA binding energies, six auspicious drug candidates—namely, pibrentasvir, venetoclax, ledipasvir, avatrombopag, cobicistat, and revefenacin—exhibited auspicious binding energies with value < −70.0 kcal/mol. Interestingly, pibrentasvir, venetoclax, and ledipasvir were observed to show even higher binding affinities with the ABCG2 transporter with binding energies of < −80.0 kcal/mol over long MD simulations of 100 ns. The stabilities of these three promising candidates in complex with ABCG2 transporter were demonstrated by their energetics and structural analyses throughout the 100 ns MD simulations. The current study throws new light on pibrentasvir, venetoclax, and ledipasvir as curative options for multidrug resistant cancers by inhibiting ABCG2 transporter.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The emergence of multidrug resistance (MDR) in cancer cells, overwhelmingly due to the overexpression of ATP-binding cassette (ABC) transporters, is still one of the most significant challenges in cancer chemotherapy [1,2,3,4]. The multidrug resistance transporter ABCG2 (or breast cancer resistance protein 1, BCRP1) is a well-characterized member of the ATP-binding cassette (ABC) family [5, 6]. ABCG2 transporter is physiologically expressed in the blood-brain-barrier, the intestine, the liver, and the placenta and plays crucial role in eliminating drugs and other toxic materials such as steroid metabolites, uric acid, and porphyrin products [7]. Furthermore, ABCG2 modulates drug absorption, distribution, metabolism, excretion, and toxicity (ADMET) properties in pharmacological cures of different diseases [8]. Clinically, overexpression of ABCG2 has been related to the occurrence of resistance to various anticancer drugs such as anthracyclines, camptothecin derivatives, and mitoxantrone [9,10,11,12]. ABCG2 shapes a homodimer to act marvelously functional transporter, as well higher orders of oligomerization have been identified—particularly tetrameric and dodecameric forms [13,14,15].
Two exemplary chemotherapy drugs with extremely divergent chemical scaffolds, namely imatinib and mitoxantrone, were picked out to grasp how ABCG2 interacts with chemotherapy drugs [16]. Imatinib, a landmark drug utilized to treat chronic myelogenous leukemia (CML) [17], and mitoxantrone, a topoisomerase inhibitor, have been recognized as ABCG2 inhibitors [18,19,20,21].
Until now, no therapeutic agent has been permitted by the U.S. Food and Drug Administration (FDA) for the modulation of ABCG2 transporter [22]. A great number of potent synthetic inhibitors towards ABCG2 transporter have been tested over the last years [23]. However, the shortage of selectivity and factors linked with untoward drug-drug interactions has obstructed the further development of some of these inhibitors [2, 22]. At present, ongoing research focuses on estimating the efficacy of several repurposed drugs to contend with multidrug resistance in cancer cells [23, 24].
This study set out to evaluate binding affinities of clinical-trial and investigational ABCG2 drug candidates (counted, 181 drugs) and identify most potent ABCG2 inhibitors using in silico drug discovery approaches. Such methods, including flexible molecular docking and molecular dynamics (MD) simulations, were applied to gain structural and energetics insights into ABCG2 inhibition. First, the performance of the employed docking technique in predicting binding affinity and mode of drug-ABCG2 complex was evaluated based on available experimental data. The studied clinical-trial and investigational drugs were then filtered using molecular docking calculations. The most promising drugs were then subjected to MD simulations over 100 ns, and their binding affinities and stabilities were evaluated. This work sheds new light on drug candidates with high binding affinities towards ABCG2 transporter, which, in turn, have a high ability to fight multidrug resistance in cancer cells.
Computational Methodology
ABCG2 Preparation
For all molecular docking calculations in addition to molecular dynamics simulations, the experimentally resolved three dimensional (3D) structure of ABCG2 transporter bound with tert-butyl3-((3 S,6 S,12aS)-9-(cyclopentyloxy)-6-isobutyl-1,4-dioxo-1,2,3,4,6,7,12,12a-octahydro-pyrazino[1’,2’:1,6]pyrido[3,4-b]indol-3-yl) propanoate (MZ29) (PDB code: 6FFC [25], chains: A and B) was retrieved from the RCSB PDB database and used as template. All missing residues in the template were constructed with the assistance of Modeller software [26]. Water molecules, heteroatoms, the bound ligand, and ions were stripped out. H++ server was utilized to examine the protonation states of the amino acid residues of the ABCG2 transporter [27]. Then, all missing hydrogen atoms were inserted, while the salinity, external and internal dielectrics, and pH values were set to 0.15, 80, 10, and 6.5, respectively.
Validation Set
Validation of the performance of the utilized molecular docking protocol in predicting the inhibitor-ABCG2 binding mode was carried out on a set containing three experimentally resolved structures of ABCG2 transporter in complex with their inhibitors. The three complexes were MZ29, 4-(4-methyl-piperazin-1-YLmethyl)-N-[4-methyl-3-(4-pyridin-3-YL-pyrimidin-2-YLamino)-phenyl]-benzamide (imatinib/STI), and 1,4-dihydroxy-5,8-bis({2-[(2-hydro-xyethyl)amino]ethyl}amino)-9,10-anthracenedione (mitoxantrone/MIX) bound with ABCG2 transporter (PDB codes: 6FFC [25], 6VXH [16] and 6VXI [16], respectively).
Inhibitors Preparation
Geometrical structures of the studied 181 ABCG2 drug candidates were obtained from the DrugBank database in SDF format [28, 29]. The 3D chemical structures of the candidates were generated using Omega2 software [30, 31]. Besides, the 3D molecular structures of MZ29, imatinib, and mitoxantrone were extracted from experimentally resolved structures of human ABCG2 complexes (PDB codes: 6FFC [25], 6VXH [16], and 6VXI [16], respectively). Energy minimization was then performed for the generated 3D structures of the drug candidates with SZYBKI using Merck Molecular Force Field94 (MMFF94S) [32]. The Marsilli-Gasteiger partial charges were then assigned to the candidates [33].
Molecular Docking
AutoDock4.2.6 was utilized to perform all molecular docking calculations [34]. The pdbqt file for ABCG2 transporter in the homodimer structure was prepared [35]. THR435 and ASN436, essential four residues inside the binding pocket of chains A and B, were introduced as conformationally flexible residues, while the rest of the residues were rigidified. For the docking calculation, number of genetic algorithms (GA) run was set to 250, while maximum number of energy evaluations (eval) was set to 25,000,000. For all other docking parameters, default values were utilized. Center of grid was placed at 130.869, 126.675, 145.206 (XYZ coordinates), with dimensions of 80 Å × 80 Å × 80 Å to envelop the ABCG2 active site. Grid spacing was set to 0.375 Å. For each studied drug cadidate, predicted binding modes were handled by built-in clustering analysis (1.0 Å RMSD tolerance). Binding mode of lowest energy within the largest cluster was selected as representative binding pose.
Molecular Dynamics Simulations
Molecular dynamics (MD) simulations for the studied drug candidates in complex with ABCG2 transporter were performed using AMBER16 software [36]. In these simulations, general AMBER force field (GAFF2) and AMBER force field 14SB were utilized to describe the drug candidates and ABCG2 transporter, respectively [37, 38]. Due to a large number of candidates investigated, implicit- and explicit-solvent MD simulations were utilized to reduce computational cost and time.
In implicit-solvent MD, AM1-BCC partial charges were assigned for the drug candidates using Antechamber tool [39]. No periodic boundary conditions, as well as a non-bonded cutoff of 999 Å, were utilized [40]. Solvation impact was estimated using the igb = 1 implicit-solvent model. Energy minimization was initially applied on the docked drug-ABCG2 complexes for 500 steps, and the minimized complexes were then smoothly heated from 0 K to 300 K over 50 ps using Langevin thermostat. Ultimately, a production stage of 5 ns was executed, and snapshots were collected every 1 ps, giving 5000 snapshots. The implicit MD simulations were conducted using pmemd.MPI implemented inside AMBER16 software.
For the most promising ABCG2 drug candidates, explicit-solvent MD simulations were conducted. Atomic partial charges of the studied candidates were assigned using the restrained electrostatic potential (RESP) approach at the HF/6-31 G* level with the assistance of Gaussian09 software [41, 42]. The drug-ABCG2 complexes were neutralized by adding appropriate counterions. The complexes were then solvated in a truncated octahedron box of TIP3P water molecules with an average distance of 12 Å. Steepest descent and conjugate-gradient methods were utilized to perform energy minimization for the solvated complexes over 5000 steps. The systems were gradually heated from 0 K to 300 K over 50 ps. A weak restraint of 10 kcal mol−1 Å−1 was employed on the solute over the heating stages. The drug-ABCG2 complexes were equilibrated for 1 ns. The production stage of 100 ns for each drug candidate was then performed. Long-range Coulombic interactions were treated utilizing the particle mesh Ewald (PME) method [43]. Besides, a cutoff distance for the Lennard-Jones interactions was set at 12.0 Å. Periodic boundary conditions were used. Langevin thermostat with a collision frequency gamma_ln set to 1.0 was employed to keep temperature at 298 K. The Berendsen barostat was used for pressure control with a relaxation time of 2 ps [44]. All bonds involving hydrogen atoms were restrained using SHAKE algorithm. The trajectories were assembled every 10 ps throughout the production stage for binding energy calculations and post-dynamics analyses. The GPU version of pmemd (pmemd.cuda) in AMBER16 software was utilized to conduct all explicit-solvent MD simulations. All molecular docking and molecular dynamics calculations were performed on the hybrid CPU/GPU CompChem cluster (hpc.compchem.net). BIOVIA DS Visualize 2020 was utilized to execute all molecular graphics [45].
MM-GBSA Binding Energy
The binding free energies (ΔGbinding) of the studied drug candidates complexed with ABCG2 transporter were calculated using molecular mechanics-generalized Born surface area (MM-GBSA) approach [46]. The modulated GB paradigm proposed by Onufriev [47] (igb = 2) was employed to specify polar solvation energy. Based on the collected uncorrelated snapshots over MD simulations, MM-GBSA binding energies were calculated. The MM-GBSA binding energy can be conceptually summarized with the following equation:
where, the energy term (G) is evaluated as:
Eele and Evdw are electrostatic and van der Waals energies, respectively. GSA is the nonpolar contribution to the solvation-free energy from the solvent-accessible surface area (SASA). GGB is the electrostatic solvation-free energy evaluated from the generalized Born equation. A single-trajectory approach was applied in which the coordinates of each of the drugs, ABCG2, and drug-ABCG2 were obtained from a single trajectory. For all studied drug candidates, entropy contributions were ignored.
Results and Discussion
Cancer is ranked as a leading cause of death and an essential barrier to increased life expectancy worldwide in the 21st century [48]. The development of resistance to cancer therapy thus poses the major bottleneck in cancer treatment and patient mortality [1,2,3,4]. These situations clearly highlight a dire need for potent and selective inhibitors against multidrug resistance targets.
The ABCG2 is indeed a key transporter of the ATP-binding-cassette (ABC) family that affords protection to tissue against xenobiotics, exerts influence on the pharmacokinetics of drugs, and participates in multidrug resistance [49]. In this study, the potentialities of 181 clinical-trial and investigational drugs were evaluated as ABCG2 inhibitors using in silico approaches.
Molecular Docking Validation
Prior to docking calculations, AutoDock4.2.6 parameters and protocol were first validated based on available experimental data. The co-crystallized MZ29, imatinib, and mitoxantrone ligands in complex with ABCG2 transporter (PDB codes: 6FFC [25], 6VXH [16], and 6VXI [16], respectively) were re-docked, and the predicted binding modes were compared to experimental structures (Fig. 1).
The comparison between predicted docked structures and experimental structures unveiled that AutoDock4.2.6 precisely portended the right binding modes of MZ29, mitoxantrone, and imatinib inside the active site of the ABCG2 transporter (Fig. 1). The calculated root-mean-square deviation (RMSD) between the predicted docked and experimental structures were 0.25, 0.30, and 0.35 Å for MZ29, mitoxantrone and imatinib, respectively. According to the predicted docking scores, MZ29 showed the lowest docking score of −12.5 kcal/mol, followed by mitoxantrone and imatinib with docking scores of −9.5 and −8.2 kcal/mol, respectively. The highest potency of MZ29 towards ABCG2 transporter may be justified by its capability of exhibiting two fundamental hydrogen bonds with ASN436:A and THR435:A with bond lengths of 2.16 and 2.14 Å, respectively. Mitoxantrone forms two hydrogen bonds with ASN436 in chains A and B with bond lengths 2.31 and 3.12 Å, respectively, with a docking score of −9.5 kcal/mol. However, imatinib was not able to form any hydrogen bond inside the active site, but its good binding affinity with a docking score of −8.2 kcal/mol may be imputed to other interactions like hydrophobic and van der Waals interactions (Fig. 1). Due to the surpassed potentiality of MZ29 over mitoxantrone and imatinib towards ABCG2 transporter, MZ29 was considered as a reference inhibitor in latter sections.
In conclusion, current results confirmed the outperformance of the employed docking technique to predict correct binding modes of drug candidates inside the binding pocket of the ABCG2 transporter.
ABCG2 Drug Candidates
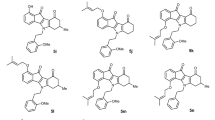
To identify potent inhibitors towards ABCG2 transporter, a total of 181 clinical-trial and investigational drug candidates were retrieved from DrugBank and prepared (see computational methodology section for details). All drug candidates were docked inside the active site of the ABCG2 transporter utilizing docking parameters of GA = 250 and eval = 25,000,000. The calculated docking scores for the investigated candidates towards ABCG2 transporter are summarized in Table S1. Calculated docking scores, 2D chemical structures, and binding features for the six potent drug candidates complexed with ABCG2 transporter are listed in Table 1, and compared to data on MZ29 as reference inhibitor.
As shown in Table 1, most of the investigated drug candidates demonstrated auspicious binding affinities towards ABCG2 homodimer structure with docking scores in the range of −10.5 to −15.1 kcal/mol. The high docking scores of the investigated drug candidates are attributed to their abilities to exhibit multiple hydrogen bonds, hydrophobic, pi-based, and van der Waals interactions with the proximal amino acid residues inside the active site of the ABCG2 transporter. Ledipasvir demonstrated the preeminent binding affinity against ABCG2 transporter with a docking score of −15.1 kcal/mol. Inspecting the binding mode of the ledipasvir inside the ABCG2 transporter revealed that the C=O of N-methylacetamide and the C=O of aminobutone form two hydrogen bonds with NH2 of ASN436:A with bond lengths of 2.04 and 1.72 Å, respectively (Fig. 2 and Table 1). At the same time, the NH of the carbamate moiety forms a hydrogen bond with ASN436:B with a bond length of 2.18 Å (Fig. 2 and Table 1). Besides, the imidazole ring interacts with the C=O of PHE439:B with a bond length of 2.06 Å (Fig. 2 and Table 1).
Compared to ledipasvir, MZ29 exhibited satisfactory binding affinity towards ABCG2 transporter, forming only two hydrogen bonds. The NH of indoline performs a hydrogen bond with the C=O of ASN436:A with a bond length of 2.16 Å. As well, the oxygen of cyclopentyloxy benzene makes a hydrogen bond with the hydroxy group of THR435:A with a bond length of 2.14 Å (Fig. 2). A most striking observation from the data comparison was the high potency of ledipasvir as a promising ABCG2 transporter inhibitor.
Molecular Dynamics (MD) Simulations
To understand conformational flexibilities, solvent influences, and stability of the drug-ABCG2 complexes, and to achieve reliable drug-ABCG2 binding affinities, molecular dynamics (MD) simulations should be executed [50, 51]. Therefore, MD simulations followed by MM-GBSA binding energy calculations for all studied drug candidates in complex with ABCG2 transporter were conducted over 5 ns in an implicit water solvent. The estimated MM-GBSA binding energies are presented in Table S1.
From Table S1, about 5% of the screened drug candidates showed appreciable binding energies (ΔGbinding) lower than −70.0 kcal/mol. To obtain more reliable binding affinities of drug-ABCG2 complexes, those potent 5% drugs (i.e., six drug candidates) were further submitted to 25 ns MD simulations in an explicit water solvent. The corresponding 25 ns MD//MM-GBSA binding energies calculations were also estimated (Fig. 3).
It is apparent from Fig. 3 that all investigated drug candidates displayed similar binding energies over both the 5 ns implicit-solvent and the 25 ns explicit-solvent MD simulations. However, three out of the investigated drugs—namely, pibrentasvir, venetoclax, and ledipasvir—exhibited the highest binding affinities over the 25 ns MD courses with ΔGbinding lower than −80.0 kcal/mol. Precisely, the estimated MM-GBSA binding energies for pibrentasvir, venetoclax, and ledipasvir in complex with ABCG2 transporter were −90.6, −89.8, and −88.0 kcal/mol over 25 ns MD simulations, respectively. The high binding affinities of pibrentasvir, venetoclax, and ledipasvir with ABCG2 transporter are explained by their ability to exhibit high stable hydrogen bond with ASN436 amino acid residue inside the active site of ABCG2 transporter (Fig. 3). It is noteworthy that the docked poses of pibrentasvir and venetoclax inside the active site of the ABCG2 transporter showed no hydrogen bond between these two drug candidates and ASN436 (Fig. S1). This observation highlights the crucial role of conformational flexibility of the drug inside the binding site of the target. Therefore, MD simulations for pibrentasvir, venetoclax, and ledipasvir complexed with ABCG2 transporter were prolonged to 100 ns and the corresponding MM-GBSA binding energies were estimated (Fig. 3).
From data in Fig. 3, there were no remarkable differences between the calculated MM-GBSA binding energies for pibrentasvir, venetoclax, and ledipasvir in complex with ABCG2 transporter over the 25 ns and 100 ns explicit-solvent MD simulations. From the estimated MM-GBSA binding energies over 100 ns explicit-solvent MD simulations, pibrentasvir, venetoclax, and ledipasvir demonstrated superior binding affinities towards ABCG2 transporter with ΔGbinding of −97.5, −85.1, and −83.3 kcal/mol, respectively, compared to ΔGbinding of −61.0 kcal/mol for MZ29.
To puzzle out the contribution of the prime driving forces in the binding of the investigated drug candidates with ABCG2 transporter, the MM-GBSA binding free energies were decomposed and presented in Table 2. The data in Table 2 interestingly showed that the van der Waals (Evdw) had predominant role in the total MM-GBSA binding free energies of pibrentasvir, venetoclax, ledipasvir, and MZ29 with average values of −115.8, −108.6, −102.5, and −74.6 kcal/mol, respectively. Most surprisingly, data in Table 2 also showed that Evdw of the three drug candidates was approximately one and a half times lower than for MZ29. The electrostatic interactions (Eele) were also appropriate with average values of −75.3, −15.3, −31.7, and −21.7 kcal/mol for pibrentasvir-, venetoclax-, ledipasvir-, and MZ29-ABCG2 complexes, respectively.
To better recognize the most significant residues that played a pivotal role in the active site of ABCG2 transporter, MM-GBSA decomposition per residue was performed for pibrentasvir-, venetoclax-, and ledipasvir-ABCG2 complexes over 100 ns MD simulations (Fig. 4). As displayed in Fig. 4, PHE439, ASN436, SER440, and THR542 amino acid residues remarkably contributed to the interactions of pibrentasvir, venetoclax, and ledipasvir with the ABCG2 transporter.
A significant contribution of the PHE439 residue in chains A and B to the total binding free energy was observed with values of −1.7 and −3.1, −3.7 and −3.7, and −1.7 and −2.9 kcal/mol for pibrentasvir-, venetoclax- and, ledipasvir-ABCG2 complexes, respectively.
Post-dynamics Analyses
To further verify the stability of pibrentasvir-, venetoclax-, and ledipasvir-ABCG2 complexes, structural and energetic analyses were carried out over 100 ns MD simulations. Four characteristics were estimated from respective simulation trajectories, including binding energy per frame, center-of-mass (CoM) distance, hydrogen bond length, and root-mean-square deviation (RMSD).
Binding Energy Per Frame
To investigate the stability of the three drugs inside the active site of ABCG2 transporter, the correlation between the binding energy per frame and time was scrutinized and plotted in Fig. 5. Most interesting was the overall stabilities observed for the three drug candidates and MZ29 inside the ABCG2 active site over 100 ns MD simulations with average binding energy values of −97.5, −87.7, −83.3, and −61.0 kcal/mol, respectively.
Hydrogen Bond Length
Hydrogen bonds play a pivotal role in biological systems and the conservation of the protein flexibility, and the binding of a drug with a target. Thus hydrogen bond distance and occupancy were inspected for pibrentasvir-, venetoclax-, and ledipasvir-ABCG2 complexes and compared to MZ29-ABCG2 complexes throughout the 100 ns MD simulations (Table 3).
From data in Table 3, remarkably the three drug candidates exhibited stable hydrogen bonds with PHE439 with H-bond occupancy in the range 50.3–93.3%. This high H-bond occupancy indicates the state of PHE439 inside the active site of the ABCG2 transporter. Comparison of data presented in Table 3 revealed that pibrentasvir, venetoclax, and ledipasvir displayed higher stability than MZ29. Actually, pibrentasvir formed three stable hydrogen bonds with ASN436, PHE439, and GLU446 with an average H-bond distance of 2.9, 2.8, and 2.8 Å, respectively. However, venetoclax and ledipasvir exhibited only two stable hydrogen bonds with PHE439 and ASN436 with average H-bond distances of 2.9 and 3.0 and 2.8 and 2.7 Å, respectively. In comparison, MZ29 exhibited a stable hydrogen bond with ASN436 with an average of 2.8 Å and demonstrated H-bond occupancy of 90.4% of the produced MD trajectory snapshots. MZ29 also formed a moderately stable hydrogen bond with PHE439 with an average value of 2.8 Å and H-bond occupancy of 51.5%. Generally, these hydrogen bond lengths vigorously indicated the stability of the identified inhibitors complexed with ABCG2 transporter.
Center-of-mass Distance
To verify the stability of drug-ABCG2 complexes during MD simulations, center-of-mass (CoM) distances were inspected between each of pibrentasvir, venetoclax, ledipasvir, and MZ29, and PHE439 (Fig. 6). From data in Fig. 6, CoM distances demonstrated the high stability of pibrentasvir-, venetoclax-, ledipasvir-, and MZ29-ABCG2 complexes with average values of 10.2, 9.2, 5.5, and 5.5 Å, respectively. Hence, current results clearly confirmed the high stability of the identified drugs in complex with ABCG2 transporter over 100 ns MD simulations.
Root-mean-square Deviation
The overall stability of the investigated drug-ABCG2 complexes was examined by measuring the root-mean-square deviation (RMSD) of the backbone atoms relative to the initial docking structure over the 100 ns MD simulations (Fig. 7). This graph is portrayed overall stability for pibrentasvir, venetoclax and ledipasvir, and MZ29 in complex with ABCG2 transporter. As well, the average RMSD values were 0.41, 0.40, 0.34, and 0.37 nm for pibrentasvir-, venetoclax-, ledipasvir-, and MZ29-ABCG2 complexes, respectively. These findings confirmed that the proposed inhibitors are tightly bonded and do not affect the overall topology of the ABCG2 transporter.
Conclusion
The development of multidrug resistance (MDR) caused by the overexpression of ABCG2 transporter, a member of the ATP-binding cassette (ABC) family, in cancer cells continues to pose a considerable challenge to cancer chemotherapy. However, tens of drugs have been subjected to clinical-trial and investigational phases, until now, there is no therapeutic drug permitted by the U.S. Food and Drug Administration (FDA) for the modulation of ABCG2 transporter. In the present study, potentialities of 181 clinical-trial and investigation drugs were in silico assessed as ABCG2 inhibitors using combined molecular docking and molecular dynamics simulations, as well as MM-GBSA binding energy calculations. According to the MM-GBSA binding energy calculations, pibrentasvir, venetoclax, and ledipasvir demonstrated auspicious binding affinities with ΔGbinding of < −80.0 kcal/mol over 100 ns MD simulation. Compared to the binding affinity of an experimental bound inhibitor (MZ29), the binding affinities of the three identified drugs were approximately one and a half times better than MZ29. The MM-GBSA binding energies for pibrentasvir-, venetoclax-, and ledipasvir-ABCG2 complexes were estimated promptly as −97.5, −87.7, and −83.3 kcal/mol over the 100 ns MD simulations, respectively. The energetics and structural analyses displayed tight binding and stability of the three drugs towards ABCG2 transporter. Binding energies were predominated by Evdw interactions with average values of −115.8, −108.6, and −102.5 kcal/mol for pibrentasvir-, venetoclax-, and ledipasvir-ABCG2 complexes, respectively. Binding modes of all three identified drug candidates inside the active site of ABCG2 transporter over the 100 ns MD simulations revealed that the three potential drugs are capable of forming substantial hydrogen bonds with key amino acid residues involving PHE439 and ASN436. Eventually, the inhibition of ABCG2 transporter with pibrentasvir, venetoclax, and ledipasvir was unveiled as a promising option for combating multidrug resistant cancers.
References
Gottesman, M. M. (2002). Mechanisms of cancer drug resistance. Annual Review of Medicine, 53, 615–27.
Szakacs, G., Paterson, J. K., Ludwig, J. A., Booth-Genthe, C., & Gottesman, M. M. (2006). Targeting multidrug resistance in cancer. Nature Reviews Drug Discovery, 5, 219–34.
Wu, C. P., Hsieh, C. H., & Wu, Y. S. (2011). The emergence of drug transporter-mediated multidrug resistance to cancer chemotherapy. Molecular Pharmaceutics, 8, 1996–2011.
Robey, R. W., Pluchino, K. M., Hall, M. D., Fojo, A. T., Bates, S. E., & Gottesman, M. M. (2018). Revisiting the role of ABC transporters in multidrug-resistant cancer. Nature Reviews: Cancer, 18, 452–64.
Dassa, E., & Bouige, P. (2001). The ABC of ABCS: a phylogenetic and functional classification of ABC systems in living organisms. Research in Microbiology, 152, 211–29.
Vasiliou, V., Vasiliou, K., & Nebert, D. W. (2009). Human ATP-binding cassette (ABC) transporter family. Human Genomics, 3, 281–90.
Matsuo, H., Takada, T., Ichida, K., Nakamura, T., Nakayama, A., Ikebuchi, Y., Ito, K., Kusanagi, Y., Chiba, T., Tadokoro, S., Takada, Y., Oikawa, Y., Inoue, H., Suzuki, K., Okada, R., Nishiyama, J., Domoto, H., Watanabe, S., Fujita, M., Morimoto, Y., Naito, M., Nishio, K., Hishida, A., Wakai, K., Asai, Y., Niwa, K., Kamakura, K., Nonoyama, S., Sakurai, Y., Hosoya, T., Kanai, Y., Suzuki, H., Hamajima, N., & Shinomiya, N. (2009). Common defects of ABCG2, a high-capacity urate exporter, cause gout: a function-based genetic analysis in a Japanese population. Science Translational Medicine, 1, 5ra11.
Szakacs, G., Varadi, A., Ozvegy-Laczka, C., & Sarkadi, B. (2008). The role of ABC transporters in drug absorption, distribution, metabolism, excretion and toxicity (ADME-Tox). Drug Discovery Today, 13, 379–93.
Doyle, L. A., Yang, W., Abruzzo, L. V., Krogmann, T., Gao, Y., Rishi, A. K., & Ross, D. D. (1998). A multidrug resistance transporter from human MCF-7 breast cancer cells. Proceedings of the National Academy of Sciences of the United States of America, 95, 15665–70.
Litman, T., Brangi, M., Hudson, E., Fetsch, P., Abati, A., Ross, D. D., Miyake, K., Resau, J. H., & Bates, S. E. (2000). The multidrug-resistant phenotype associated with overexpression of the new ABC half-transporter, MXR (ABCG2). Journal of Cell Science, 113, 2011–21.
Perego, P., De Cesare, M., De Isabella, P., Carenini, N., Beggiolin, G., Pezzoni, G., Palumbo, M., Tartaglia, L., Pratesi, G., Pisano, C., Carminati, P., Scheffer, G. L., & Zunino, F. (2001). A novel 7-modified camptothecin analog overcomes breast cancer resistance protein-associated resistance in a mitoxantrone-selected colon carcinoma cell line. Cancer Research, 61, 6034–7.
van Hattum, A. H., Hoogsteen, I. J., Schluper, H. M., Maliepaard, M., Scheffer, G. L., Scheper, R. J., Kohlhagen, G., Pommier, Y., Pinedo, H. M., & Boven, E. (2002). Induction of breast cancer resistance protein by the camptothecin derivative DX-8951f is associated with minor reduction of antitumour activity. British Journal of Cancer, 87, 665–72.
Kage, K., Tsukahara, S., Sugiyama, T., Asada, S., Ishikawa, E., Tsuruo, T., & Sugimoto, Y. (2002). Dominant-negative inhibition of breast cancer resistance protein as drug efflux pump through the inhibition of S-S dependent homodimerization. International Journal of Cancer, 97, 626–30.
Ferreira, R. J., Bonito, C. A., Cordeiro, M., Ferreira, M. U. & Dos Santos, D. (2017). Structure-function relationships in ABCG2: insights from molecular dynamics simulations and molecular docking studies. Scientific Reports, 7, 15534–50.
Litman, T., Jensen, U., Hansen, A., Covitz, K. M., Zhan, Z. R., Fetsch, P., Abati, A., Hansen, P. R., Horn, T., Skovsgaard, T. & Bates, S. E. (2002). Use of peptide antibodies to probe for the mitoxantrone resistance-associated protein MXR/BCRP/ABCP/ABCG2. Biochimica et Biophysica Acta-Biomembranes, 1565, 6–16.
Orlando, B. J. & & Liao, M. (2020). ABCG2 transports anticancer drugs via a closed-to-open switch. Nature Communications, 11, 2264–74.
Iqbal, N. & Iqbal, N. (2014). Imatinib: a breakthrough of targeted therapy in cancer. Chemotherapy Research and Practice, 2014, 357027–36.
Homolya, L., Orban, T. I., Csanady, L. & Sarkadi, B. (2011). Mitoxantrone is expelled by the ABCG2 multidrug transporter directly from the plasma membrane. Biochimica et Biophysica Acta-Biomembranes, 1808, 154–63.
Burger, H., van Tol, H., Boersma, A. W., Brok, M., Wiemer, E. A., Stoter, G., & Nooter, K. (2004). Imatinib mesylate (STI571) is a substrate for the breast cancer resistance protein (BCRP)/ABCG2 drug pump. Blood, 104, 2940–2.
Eadie, L. N., Hughes, T. P., & White, D. L. (2014). Interaction of the efflux transporters ABCB1 and ABCG2 with imatinib, nilotinib, and dasatinib. Clinical Pharmacology and Therapeutics, 95, 294–306.
Houghton, P. J., Germain, G. S., Harwood, F. C., Schuetz, J. D., Stewart, C. F., Buchdunger, E., & Traxler, P. (2004). Imatinib mesylate is a potent inhibitor of the ABCG2 (BCRP) transporter and reverses resistance to topotecan and SN-38 in vitro. Cancer Research, 64, 2333–7.
Kannan, P., Telu, S., Shukla, S., Ambudkar, S. V., Pike, V. W., Halldin, C., Gottesman, M. M., Innis, R. B., & Hall, M. D. (2011). The “specific” P-glycoprotein inhibitor tariquidar is also a substrate and an inhibitor for breast cancer resistance protein (BCRP/ABCG2). ACS Chemical. Neuroscience, 2, 82–9.
Henrich, C. J., Robey, R. W., Bokesch, H. R., Bates, S. E., Shukla, S., Ambudkar, S. V., Dean, M., & McMahon, J. B. (2007). New inhibitors of ABCG2 identified by high-throughput screening. Molecular Cancer Therapeutics, 6, 3271–8.
Quezada, H., Martinez-Vazquez, M., Lopez-Jacome, E., Gonzalez-Pedrajo, B., Andrade, A., Fernandez-Presas, A. M., Tovar-Garcia, A., & Garcia-Contreras, R. (2020). Repurposed anti-cancer drugs: the future for anti-infective therapy? Expert Review of Anti-Infective Therapy, 18, 609–12.
Jackson, S. M., Manolaridis, I., Kowal, J., Zechner, M., Taylor, N. M. I., Bause, M., Bauer, S., Bartholomaeus, R., Bernhardt, G., Koenig, B., Buschauer, A., Stahlberg, H., Altmann, K. H., & Locher, K. P. (2018). Structural basis of small-molecule inhibition of human multidrug transporter ABCG2. Nature Structural & Molecular Biology, 25, 333–40.
Marti-Renom, M. A., Stuart, A. C., Fiser, A., Sanchez, R., Melo, F., & Sali, A. (2000). Comparative protein structure modeling of genes and genomes. Annual Review of Biophysics and Biomolecular Structure, 29, 291–325.
Gordon, J. C., Myers, J. B., Folta, T., Shoja, V., Heath, L. S., & Onufriev, A. (2005). H++: a server for estimating pKas and adding missing hydrogens to macromolecules. Nucleic Acids Research, 33, W368–71.
Wishart, D. S., Knox, C., Guo, A. C., Shrivastava, S., Hassanali, M., Stothard, P., Chang, Z., & Woolsey, J. (2006). DrugBank: a comprehensive resource for in silico drug discovery and exploration. Nucleic Acids Research, 34, D668–72.
Wishart, D. S., Feunang, Y. D., Guo, A. C., Lo, E. J., Marcu, A., Grant, J. R., Sajed, T., Johnson, D., Li, C., Sayeeda, Z., Assempour, N., Iynkkaran, I., Liu, Y., Maciejewski, A., Gale, N., Wilson, A., Chin, L., Cummings, R., Le, D., Pon, A., Knox, C., & Wilson, M. (2018). DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Research, 46, D1074–D82.
Hawkins, P. C., Skillman, A. G., Warren, G. L., Ellingson, B. A., & Stahl, M. T. (2010). Conformer generation with OMEGA: algorithm and validation using high quality structures from the Protein Databank and Cambridge Structural Database. Journal of Chemical Information and Modeling, 50, 572–84.
OMEGA 2.5.1.4. Santa Fe, NM, USA. OpenEye Scientific Software, 2013.
SZYBKI 1.9.0.3. Santa Fe, NM, USA. OpenEye Scientific Software, 2016.
Gasteiger, J., & Marsili, M. (1980). Iterative partial equalization of orbital electronegativity—a rapid access to atomic charges. Tetrahedron, 36, 3219–28.
Morris, G. M., Huey, R., Lindstrom, W., Sanner, M. F., Belew, R. K., Goodsell, D. S., & Olson, A. J. (2009). AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. Journal of Computational Chemistry, 30, 2785–91.
Forli, S., Huey, R., Pique, M. E., Sanner, M. F., Goodsell, D. S., & Olson, A. J. (2016). Computational protein-ligand docking and virtual drug screening with the AutoDock suite. Nature Protocols, 11, 905–19.
Case, D. A., Betz, R. M., Cerutti, D. S., Cheatham, T. E., Darden, T. A., Duke, R. E., Giese, T. J., Gohlke, H., Goetz, A. W., Homeyer, N., Izadi, S., Janowski, P., Kaus, J., Kovalenko, A., Lee, T. S., LeGrand, S., Li, P., Lin, C., Luchko, T., Luo, R., Madej, B., Mermelstein, D., Merz, K. M., Monard, G., Nguyen, H., Nguyen, H. T., Omelyan, I., Onufriev, A., Roe, D. R., Roitberg, A., Sagui, C., Simmerling, C. L., Botello-Smith, W. M., Swails, J., Walker, R. C., Wang, J., Wolf, R. M., Wu, X., Xiao, L., & Kollman, P. A. (2016). AMBER16. San Francisco, USA: University of California.
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A., & Case, D. A. (2004). Development and testing of a general amber force field. Journal of Computational Chemistry, 25, 1157–74.
Maier, J. A., Martinez, C., Kasavajhala, K., Wickstrom, L., Hauser, K. E., & Simmerling, C. (2015). ff14SB: improving the accuracy of protein side chain and backbone parameters from ff99SB. Journal of Chemical Theory and Computation, 11, 3696–713.
Jakalian, A., Jack, D. B., & Bayly, C. I. (2002). Fast, efficient generation of high-quality atomic charges. AM1-BCC model: II. Parameterization and validation. Journal of Computational Chemistry, 23, 1623–41.
Roux, B., & Simonson, T. (1999). Implicit solvent models. Biophysical Chemistry, 78, 1–20.
Bayly, C. I., Cieplak, P., Cornell, W. D., & Kollman, P. A. (1993). A well-behaved electrostatic potential based method using charge restraints for deriving atomic charges - the RESP model. Journal of Physical Chemistry, 97, 10269–80.
Frisch, M. J., Trucks, G. W., Schlegel, H. B., Scuseria, G. E., Robb, M. A., Cheeseman, J. R., Scalmani, G., Barone, V., Mennucci, B., Petersson, G. A., Nakatsuji, H., Caricato, M., Li, X., Hratchian, H. P., Izmaylov, A. F., Bloino, J., Zheng, G., Sonnenberg, J. L., Hada, M., Ehara, M., Toyota, K., Fukuda, R., Hasegawa, J., Ishida, M., Nakajima, T., Honda, Y., Kitao, O., Nakai, H., Vreven, T., Montgomery, J. A., Peralta, J. E., Ogliaro, F., Bearpark, M., Heyd, J. J., Brothers, E., Kudin, K. N., Staroverov, V. N., Kobayashi, R., Normand, J., Raghavachari, K., Rendell, A., Burant, J. C., Iyengar, S. S., Tomasi, J., Cossi, M., Rega, N., Millam, J. M., Klene, M., Knox, J. E., Cross, J. B., Bakken, V., Adamo, C., Jaramillo, J., Gomperts, R., Stratmann, R. E., Yazyev, O., Austin, A. J., Cammi, R., Pomelli, C., Ochterski, J. W., Martin, R. L., Morokuma, K., Zakrzewski, V. G., Voth, G. A., Salvador, P., Dannenberg, J. J., Dapprich, S., Daniels, A. D., Farkas, Ö., Foresman, J. B., Ortiz, J. V., Cioslowski, J., & Fox, D. J. (2009). Gaussian 09. Wallingford CT, USA.: Gaussian Inc.
Darden, T., York, D., & Pedersen, L. (1993). Particle mesh Ewald: AnN·log(N) method for Ewald sums in large systems. Journal of Chemical Physics, 98, 10089–92.
Berendsen, H. J. C., Postma, J. P. M., Vangunsteren, W. F., Dinola, A., & Haak, J. R. (1984). Molecular-dynamics with coupling to an external bath. Journal of Chemical Physics, 81, 3684–90.
Dassault Systèmes BIOVIA. (2019). BIOVIA Discovery Studio Visualize 2019, version 2019. San Diego, CA, USA: Dassault Systèmes BIOVIA.
Massova, I., & Kollman, P. A. (2000). Combined molecular mechanical and continuum solvent approach (MM-PBSA/GBSA) to predict ligand binding. Perspectives in Drug Discovery and Design, 18, 113–35.
Onufriev, A., Bashford, D., & Case, D. A. (2004). Exploring protein native states and large-scale conformational changes with a modified generalized born model. Proteins, 55, 383–94.
Liu, Z., Xiao, X., Wei, X., Li, J., Yang, J., Tan, H., Zhu, J., Zhang, Q., Wu, J., & Liu, L. (2020). Composition and divergence of coronavirus spike proteins and host ACE2 receptors predict potential intermediate hosts of SARS-CoV-2. Journal of Medical Virology, 92, 595–601.
Toyoda, Y., Takada, T. & Suzuki, H. (2019). Inhibitors of human ABCG2: from technical background to recent updates with clinical implications. Frontiers in Pharmacology, 10, 208–16.
De Vivo, M., Masetti, M., Bottegoni, G., & Cavalli, A. (2016). Role of molecular dynamics and related methods in drug discovery. Journal of Medicinal Chemistry, 59, 4035–61.
Kerrigan, J. E. (2013) Molecular dynamics simulations in drug design. In S. Kortagere (ed.), In Silico Models for Drug Discovery. (pp. 95–113). Totowa, NJ: Humana Press.
Acknowledgements
M.F.M. extends his appreciation to the Deanship of Scientific Research at King Khalid University for funding this work under grant no. (R.G.P.1/143/42). The computational work was completed with resources supported by the Science and Technology Development Fund, STDF, Egypt, Grants No. 5480 & 7972.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Ethical Approval
This article does not contain any studies with animals performed by any of the authors.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Ibrahim, M.A.A., Badr, E.A.A., Abdelrahman, A.H.M. et al. Prospective Drug Candidates as Human Multidrug Transporter ABCG2 Inhibitors: an In Silico Drug Discovery Study. Cell Biochem Biophys 79, 189–200 (2021). https://doi.org/10.1007/s12013-021-00985-y
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12013-021-00985-y