Abstract
Molecular dynamics simulations, binding free energy calculations, principle component analysis (PCA), and residue interaction network analysis were employed in order to investigate the molecular mechanism of M184I single mutation which played pivotal role in making the HIV-1 reverse transcriptase (RT) totally resistant to lamivudine. Results showed that single mutations at residue 184 of RT caused (1) distortion of the orientation of lamivudine in the active site due to the steric conflict between the oxathiolane ring of lamivudine and the side chain of beta-branched amino acids Ile at position 184 which, in turn, perturbs inhibitor binding, (2) decrease in the binding affinity by (~8 kcal/mol) when compared to the wild-type, (3) variation in the overall enzyme motion as evident from the PCA for both systems, and (4) distortion of the hydrogen bonding network and atomic interactions with the inhibitor. The comprehensive analysis presented in this report can provide useful information for understanding the drug resistance mechanism against lamivudine. The results can also provide some potential clues for further design of novel inhibitors that are less susceptible to drug resistance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Since acquired immunodeficiency syndrome (AIDS) was first reported in the early 1980s, this global killer has claimed the lives of an estimated 22 million individuals. Among the classified infectious diseases, the human immunodeficiency virus type 1 (HIV-1) infection, the causative agent of AIDS, remains a challenging epidemic. It has been reported that 34 million people live with HIV/AIDS globally, and in sub-Saharan Africa, approximately 22.9 million individuals contribute to the overall global estimate.
Despite extensive ongoing HIV/AIDS research, to date, no cure has been found for this disease. Currently, the most effective treatment comprises a complex “cocktail” of Food and Drug Administration (FDA)-approved and clinical trial drugs that involve multiple drug targets [1]. These drugs target and attack the virus at different stages of its lifecycle, thereby halting virus replication and reducing destruction of the immune system. They include protease inhibitors (PIs), reverse transcriptase (RT) inhibitors, integrase (IN) inhibitors, and entry inhibitors [2].
HIV-1 is a retrovirus that replicates within a host cell. The RT enzyme is an essential component for HIV-1 replication [3]. RT is responsible for converting single-stranded RNA viral genome into double-stranded DNA [3, 4]. Because of its integral role in the virus lifecycle, HIV-1 RT is a major target for drug therapy [5].
Currently, approved HIV-1 RT inhibitors used in antiretroviral therapy can be divided into two classes: (a) nucleoside analog RT inhibitors (NRTIs) [6, 7], which compete with natural nucleoside substrates for binding to the RT polymerase active site, and after their incorporation into the primer site, they act as terminators of proviral DNA synthesis; (b) non-nucleoside RT inhibitors (NNRTIs), which bind to a hydrophobic pocket close to the RT active site [4, 6]. More specifically when NRTIs are phosphorylated intracellularly, these drugs become activated and hence inhibit reverse transcription by triggering chain termination once incorporated into viral DNA [4, 7].
The initial introduction of NRTIs for treatment of HIV and later presented as the favored central core drugs of highly active antiretroviral therapy (HAART) has substantially reduced HIV-related morbidity and mortality rates [8]. Following zidovudine (AZT) as the first NRTI drug approved by the FDA for HIV therapy [8], numerous other FDA-approved NRTIs have been discovered including lamivudine (3TC), stavudine (D4T), tenofovir (TFV), and emtricitabine (FTC) among others [8, 9]. However, even though NRTIs have served as the cornerstones of successful HIV therapy, the largest problem in HIV drug therapy is that the virus mutates very quickly, leading not only to drug resistance [10], but also drug-resistant variants of the virus that have mutations in the RT target protein [11, 12]. For this reason, a molecular understanding of the impact of mutations on drug resistance will assist in the design of more potent drugs that are active against resistant strains.
An important constituent of triple-drug anti-AIDS therapy is the NRTI 2′,3′-dideoxy-3′-thiacytidine (3TC, lamivudine). It has been proven that single mutations at residue 184 of RT in HIV cause high-level resistance to 3TC and contribute to the failure of anti-AIDS combination therapy [13].
Partially due to the lack of an X-ray crystal structure of the lamivudine–RT complex, the precise mechanism by which the M184I mutant develops resistance to lamivudine is not fully understood. Based on a previously built model from “unbound” X-ray crystal structures of the wild-type and M184I mutant, it is thought that steric conflict between the oxathiolane ring of lamivudine and the side chain of beta-branched amino acids Ile at position 184 perturbs inhibitor binding, leading to a reduction in incorporation of the analog [13]. However, this assumption was based on approximated “fitted” and “static” models.
In this work, we aim to provide more comprehensive insight into the precise impact of the M184I mutation on resistance to lamivudine. We believe that, to develop potent and effective anti-HIV NRTIs against the viral variants, understanding of molecular basis of the M184I RT mutation at atomic level will be very critical.
In recent years, molecular modeling tools have become close counterparts of experiments [14]. In one of our recent papers, the interface between computation and experiment within the biological context was comprehensively highlighted [15]. Various molecular modeling approaches such as molecular docking, molecular dynamics (MD) simulation, and binding free energy calculations have proven useful in understanding the molecular basis of drug resistance to different biological systems [16–20]. This includes, but is not limited to, the drug resistance mechanisms against HIV protease [21–23], HIV reverse transcriptase (RT) [24], HIV integrase (IN) [25], influenza neuraminidase (NA) [26, 27], and HCV protease [25, 28, 29].
Standard MD analysis for large biomolecular systems can only provide limited information about the dynamic landscape of these systems. During the last few years, multiple approaches have been devised to deal with the large number of explicit degrees of freedom [30–33]. Principal component analysis (PCA) or essential dynamics analysis is one of the widely used approaches to explore the structural fluctuations of biological systems. PCA is a statistical procedure based on covariance analysis that can transform the original space of correlated variables into a reduced space of independent variables and is often used to highlight large correlated motions in macromolecules [34, 35]. It takes the trajectory of an MD simulation into account and reduces the dimensionality of the data to extract dominant nodes in the motion of a molecule [36–39]. In normal mode analysis, these pronounced motions resulting from the protein movement correspond to correlated vibrational nodes or collective group motion of atoms [40, 41]. The eigenvector for a given motion has a corresponding eigenvalue which signifies the energetic contribution of a particular component to the motion. Projection of a trajectory onto a specific eigenvector highlights the time-dependent motions that the components perform during a particular vibrational mode [37].
PCA has been used to identify the overall motion of the enzyme backbone to check the difference from one state to another with high utility in the comparison of wild-type proteins with those of mutant/resistant strains [42]. Via PCA, it is possible to identify the dominant motions observed during a simulation through visual inspection. A large portion of overall fluctuation of macromolecules can often be accounted for by a few low-frequency eigenvectors with high eigenvalues [43]. If motion between two different macromolecules is similar, then the eigenvectors coming from individual trajectories should be similar to each other. For this reason, PCA has proved an efficient tool that can be used to compare motions of different macromolecules [44].
It has also been observed that the resistant mutation or substitution in the target protein often impacts the binding ability of drugs through modification of the residue interaction network (RIN) communication of the targets [45–48]. Recently, some reports have shown that the analysis and interactive visualization of RIN of proteins can reveal important information on biological interactions in complex systems [49, 50].
In this work, using validated models of lamivudine–RT complexes, wild-type and the M184I mutant, MD simulations, post-dynamics analysis, binding free energy calculations, PCA, and RIN analysis were employed to provide an insight into the drug resistance mechanism of the RT-M184I mutant to lamivudine. To our knowledge, this report is the first account of such comprehensive computational analysis being used to reveal the precise impact of the M184I mutation on RT resistance to lamivudine.
The compilation of the computational and molecular informatics tools presented in this study could serve as powerful tools to understand protein structures and dynamics, and drug resistance and could also be incorporated in the drug discovery and development workflows.
Computational Methods
System Preparation
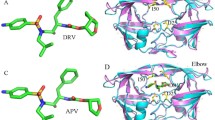
With the lack of a lamivudine–RT complex crystal structure, the construction of a reasonable initial set of 3D coordinates for the complex presented a challenge. Since molecular docking can, in many instances, produce unrealistic ligand orientations in the active site, we opted not to perform docking calculations but instead to use a “ligand fitting” strategy based on known X-ray crystal structures of similar inhibitors in complex with RT. This approach has proven successful in the past, as reported previously [51, 52]. The X-ray crystal structure of RT in complex with zidovudine, AZT, (PDB code: 1RTD) was used as the starting coordinates (Fig. 1). This choice was based on the fact that since AZT and 3TC are structurally analogous, similar binding modes would be anticipated (Fig. 1). This protein is crystalized in a polymeric form, but only one chain (chain A) was used for simulations to reduce the computational cost. Despite being distal from the active site, the X-ray crystal structure mutations, P1K, Q258C, and E478Q mutations were modified to retain the native wild-type structure. Co-crystalized solvent molecules were also deleted. Lamivudine was then superimposed against zidovudine (Fig. 2); the latter was then removed, leaving lamivudine sitting in the RT active site. Methionine (Met) at position 184 was then manually mutated into isoleucine (Ile). The Chimera software package (https://www.cgl.ucsf.edu/chimera/) was used for ligand superimposition as well as amino acid residue mutation.
Molecular Dynamics Simulation
A multiple MD-trajectory approach was adapted to understand the impact of M184I mutation on lamivudine binding with HIV-RT. A long MD run might lead to high statistical errors with denatured protein configuration during simulation period. In view of aforesaid, a multiple MD simulation acts as a high reliable alternative approach which lowers down force-field-induced artifacts, statistical bias, and computational time [53]. In this case, a multiple MD approach was adapted for both the systems from an initial starting configuration generated from a 2 ns MD run, followed by four different individual MD runs for a timescale of 5 ns with different initial velocities in each case. The multiple MD procedure adapted in this study is summarized in Fig. 3.
MD Simulations Setup and Parameters
The MD simulations for the Lamivudine–RT complexes (i.e., wild-type and mutant) were performed using the GPU version of the PMEMD engine provided with the Amber 12 package [54–57]. Two protonation states, ionized and unionized, of the triphosphate moiety attached to the ligand were considered (see “Results and Discussion” section and Supplementary Material 2). The FF99SB variant of the AMBER force field [58] was used to describe the protein. The restrained electrostatic potential (RESP) procedure [59] was used to calculate the partial atomic charges for the ligands at the HF/6-31G* level using the Gaussian 03 package [60]. The LEAP module in Amber 12 was used for addition of hydrogen atoms to the protein and the addition of counter ions for neutralization. The systems were embedded within an orthorhombic box of TIP3P [61] water molecules such that no protein atom was within 8 Å of any box edge. Periodic boundary conditions were enforced, and the long-range electrostatic interactions were treated with the particle-mesh Ewald method [62] incorporated in Amber12 with a direct space and vdW cut-off of 12 Å. The following MD procedure was applied to the initial 2 ns MD as well as the individual 5 ns MD runs (see Fig. 3). Initial energy minimization of each system, with a restraint potential of 500 kcal/mol Å2 applied to the solute, was performed for 1000 steps of steepest descent followed by 1000 steps of conjugate gradient minimization. A further 1000 steps of unrestrained conjugate gradient minimization of the entire system were then carried out. Canonical ensemble (NVT) MD simulations were then carried out for 50 ps, with gradual heating from 0 to 300 K with harmonic restraints of 5 kcal/mol Å2 applied to all solute atoms and a Langevin thermostat with a random collision frequency of 1/ps. The systems were subsequently equilibrated at 300 K in the NPT ensemble for 500 ps, during which no restrains were imposed and a Berendsen barostat was used to maintain the pressure at 1 bar. The SHAKE algorithm [63] was used to constrain the bonds of all hydrogen atoms, a time step of 2 fs, and the SPFP precision model [64] was used for all MD runs. A short production run for 2 ns was performed followed by a multiple MD production run of 4 × 5 ns from the last restart file (configuration generated from 2 ns MD run) in an isothermal isobaric (NPT) ensemble using a Berendsen barostat for each case [65] with a target pressure of 1 bar and a pressure coupling constant of 2 ps. Coordinates were saved every 1 ps, and the trajectories were analyzed every 1 ps using the PTRAJ module implemented in Amber 12.
Thermodynamic Calculations
The binding free energies of lamivudine and RTs, wild-type and mutant, were computed using the molecular mechanics/generalized born surface area (MM/GBSA) approach [66–69]. For each 5 ns trajectory, the binding free energy was averaged over 1000 snapshots at 5 ps intervals. Average values over the 4 trajectories were also computed (Table 1). The following set of equations describes the calculation of the binding free energy:
where E gas is the gas-phase energy; E int is the internal energy; and E ele and E vdW are the Coulomb and van der Waals energies, respectively. E gas is evaluated directly from the ff03 force field terms. The solvation free energy, denoted by G sol, can be decomposed into polar and nonpolar contribution states. The polar solvation contribution, G GB, is determined by solving the GB equation, whereas, G SA, the nonpolar solvation contribution is estimated from the solvent-accessible surface area (SASA) determined using a water probe radius of 1.4 Å. T and S correspond to temperature and total solute entropy, respectively.
In order to determine the individual amino acid contribution towards total binding free energy between the lamivudine inhibitor and wild-type/mutant HIV-RT types, a decomposition analysis of the interaction energy for each residue was computed by using the MM/GBSA binding free energy decomposition protocol in Amber 12.
Principle Component Analysis (PCA)
Before processing the MD trajectories for PCA, each 5 ns MD trajectories of the wild-type and mutant complexes were stripped of solvent and ions using the PTRAJ module in AMBER 12.0. The stripped trajectories were then aligned against the fully minimized structure. PCA was performed for C-α atoms on 1000 snapshots each at 5 ps intervals. Using in-house scripts, the first two principal components were calculated and the covariance matrices were generated. The first two principal components (PC1 and PC2) generated from each of the four individual 5 ns trajectories were averaged for both wild-type and M184I mutant. The first two principal components correspond to the first two Eigen vectors of the covariance matrix. The PCA scatter plots were then generated using the Xmgrace program (http://plasma-gate.weizmann.ac.il/Grace/). All structural diagrams were created using VMD [70]. The porcupine plots were generated using the ProDy interface of normal mode wizard (NMW) of VMD [71].
Residue Interaction Network (RIN) Analysis
The fully minimized structure of each system, wild and M184I mutant, was used for constructing the RIN interactively in 2D graphs. The PROBE [39] software was used to identify the contacts (i.e., non-covalent interactions) between amino acids in the proteins by evaluating their atomic packing using small-probe contact dot surfaces. PROBE uses a small virtual probe (typically 0.25 Å) that is rolled around the van der Waals surface of each atom, and an interaction (contact dot) is detected if the probe touches another non-covalently bonded atom [39].
Interactive Visual Analysis of Residue Networks
The RINs generated from the MD averaged structures were used to visualize the network using Cytoscape [72] and the plugin RINalyzer [49]. In a RIN, the nodes represent the protein amino acid residues and the edges between them represent the non-covalent interactions. The edges are labeled with an interaction type, usually including interatomic contact, hydrogen bonds, salt bridges, etc.
Results and Discussion
MD Simulations and Systems Stability
RMSD and potential energy fluctuations were monitored to ensure that the systems were well equilibrated before any further MD analysis. RMSD and potential energy plots are provided as Supplementary Material (Supplementary Materials 1).
All corresponding PDB coordinates of the simulated systems (wild and mutant for unionized and ionized phosphate moiety—4 systems in total) are provided with the supplementary material files.
Post-dynamic Analysis: Wild-Type Versus M184I Mutant
Root of Mean Square Fluctuation (RMSF)
Figure 4 shows per-residue C-α root mean square fluctuations (RMSF) of the wild-type and M184I mutant simulations. The most interesting observation is that the amino acid residues in the region 170–220 (which contains the mutation site at position 184) show higher fluctuation in the mutant than compared with the wild-type. One possible explanation is that the Ile184 residue in the mutant interacts less with the surrounding than methionine.
RMSF for the wild-type and M184I mutant lamivudine–RT complex systems: T1, T2, T3, T4, and T-Avg denote for the four individual 5 ns MD trajectories and the overall average, respectively. A zoomed view of region that contains the point mutation, 170–200, is shown in T-Avg plot. The RMSF was calculated taking in account C-α atoms of the backbone
Interestingly, the M184I mutation was also found to impact the overall dynamics of some distal amino acid regions (Fig. 4). For instances, residues 2–50 exhibit larger fluctuations, while residues 300–400 were found to show less fluctuation relative to the wild-type.
Steric Conflict Between Ile184 and Oxathiolane Ring of Lamivudine
Steric conflict between the oxathiolane ring of lamivudine and the side chain of beta-branched amino acids Ile at position 184 has traditionally been assumed to be responsible for the perturbed inhibitor binding. However, this assumption was based on static models [13]. In this study to test this assumption, we monitored the distance between the (O) atom of oxathiolane ring and the Cβ of the amino acid residue at position 184 (Met in wild-type and Ile in mutant) during the MD trajectory. Figure 5 suggests that substitution of Met with Ile results in increased steric conflict with the ligand that continues to exist during the duration of the MD simulation as evident from the increased separation between the (O) atom of oxathiolane ring and the Cβ of the amino acid residue of Ile184. These findings lend credence to the static-based assumption of Sarafianos et al. [13].
MM/PBSA Binding Free Energy Calculations
All the components of molecular mechanics and solvation energy were averaged over the a 4 × 5 ns multiple MD simulation in each case using the MM/GBSA technique, which are listed in Table 1. The calculated binding free energy (∆G bind) between lamivudine and wild-type RT is −45.6597 kcal/mol compared to −38.5251 kcal/mol in the case of M184I mutant. Such a large reduction in binding affinity (~8 kcal/mol) as a result of mutation could significantly impair ligand binding and thus the effectiveness of lamivudine against the mutant—this is in a great accordance with experimental data (the EC50 value in case of M184I mutant is ~1000 times lower than the wild-type) [73]. The calculated van der Waals contributions (∆E vdW) to the binding free energy in the lamivudine wild-type-bound RT complex (−48.2095 kcal/mol) are higher than that for the lamivudine-bound M184I mutant RT complex (−42.5814 kcal/mol). On the other hand, the calculated electrostatic contributions (∆E ele) to the binding free energy for lamivudine-bound M184I mutant RT complex (−16.9961 kcal/mol) are lower compared to that for the lamivudine wild-type-bound RT complex (−24.6420 kcal/mol). In the lamivudine wild-type-bound RT complex, the calculated solvation contributions (∆G sol. ∆G sol = ∆G SA + ∆G GB) to the binding free energy (26.4379 kcal/mol) are higher than that for the lamivudine-bound M184I mutant RT complex (21.0511 kcal/mol). The free energy components shown in Table 1 suggest that the majority of the favorable contributions observed for lamivudine binding arise from ∆E vdW and ∆E ele. The remarkable difference (~13 kcal/mol) in the ∆G gas as a result of mutation also confirmed that M184I mutation could lead to loss of efficacy of lamivudine.
For the un-protonated triphosphate moiety, interestingly, however, the same trend was observed in the overall binding affinity, and the electrostatic contribution was higher when compared to the protonated state—this might be due to higher electrostatic interactions between the active site residues with the charged triphosphate moiety (see Supplementary Material 2).
Validation of the Binding Energy Calculations
To validate the binding free energy results, another RT mutation, M184V, with available experimental data [74] was considered. Encouragingly, the calculated binding free energy was found to be in a good agreement with experimental data (see Supplementary Material). Such findings imply that the simulation protocol of this work is appropriate and reliable.
Per-residue Interaction Energy Decomposition Analysis
The binding free energy was further decomposed into contributions from each HIV-RT amino acid residue. In Table 2, the comparison of protein–ligand interaction spectra between wild-type and M184I mutant is shown. It can be observed from the energy decomposition analysis that in the lamivudine-bound wild-type HIV-RT system, the major contributions were −3.21, −2.16, and −1.14 kcal/mol from amino acid residues Arg72, Tyr115, and Met184, respectively (Table 2). On the other hand, there were some minor contributions towards the interaction energy from residues Asp110 (−0.13 kcal/mol), Ala114 (−0.79 kcal/mol), and Phe116 (−0.83 kcal/mol).
As shown in Table 2 below, the decomposed van der Waals and electrostatic energies for residues 72, 115, and 184 in the lamivudine-bound M184I mutant complex imply that the change in the van der Waals contribution is what is mostly responsible for the decrease in the interaction energy. Furthermore, when Met is mutated to Ile at position 184, the prominent van der Waals contributions towards the total binding free energy from this residue increased by twofold with a slight decrease in electrostatic contribution but it ultimately lowers the total van der Waals contributions of other prominent residues which ultimately affects the total binding free energy. This finding stands consistent with decreased van der Waals contributions from Arg72 (−3.21 to −2.07 kcal/mol) and Tyr115 (−2.16 to −2.10 kcal/mol) and an overall loss of van der Waals contribution towards binding free energy (Table 1, 2).
Interestingly, as evident from Table 2, mutation of Met to Ile at position 184 has improved the binding at the site of mutation—this might be due to better hydrophobic interactions with the Ile side chain—the overall drug binding affinity is reduced due to the negative impact of the mutation on the binding affinity of the nearby active residues. Different binding forces between the ligand and the protein, wild and mutant, are shown in Fig. 6.
Principle Component Analysis (PCA)
Figure 7 shows a PCA scatter plot generated for the wild-type and M184I mutant showing a significant difference between both systems as evident from the characteristic structures plotted along the direction of two principal components. From the scatter plot, it is clear that eigenvectors computed from the MD trajectory for both systems are quite varied which clearly indicates difference in protein motion between wild and M184I mutant.
In most cases, a few low-frequency normal modes are enough to capture major protein motions along specific directions, which are represented by eigenvectors. In this case, we generate porcupine plots with three low-frequency modes (Mode = 1, 2, and 3) to visualize the comparative motion difference between wild-type and mutant (Fig. 8). The eigenvectors show a clear difference in direction of motion, which is consistent with the PCA, scatter plot.
Residue Interaction Network (RIN)
The network analysis of the protein backbone is a new strategy to identify key residue interactions and can be used to explore the difference in RINs between different proteins including wild-type and mutants. In this work, we investigated the relationship between key residues of the wild-type and M184I mutant by generating RINs using the representative structures from the 5 ns of MD. As evident from the RIN plots (Figs. 9, 10), it is clear that the M184I mutation has distorted the overall RIN when compared to wild-type. For instance, as shown in Figs. 7 and 8, there is a hydrogen bond interaction between Tyr115 and Met184, whereas in the case of the mutant where Met184 has been mutated to Ile184, the hydrogen bond interaction with Tyr115 changes to just a close atom interaction. It is interesting to observe that in the case of the wild-type, Pro157 builds a close atom interaction with Met184, whereas in the case of the mutant, there is a close atom interaction instead between Gln161 and Ile184. Again, the M184I mutation affects the interaction network, which ultimately affects the protein backbone and consequently the drug binding landscape.
Conclusions
The precise molecular understanding of the serious impact of a single M184I on the RT resistance to lamivudine is lacking in literature. In this report, we embarked on a wide range of computational approaches in order to provide a multidimensional view on M184I resistance to lamivudine. Multiple molecular dynamics simulations, binding free energy calculations, principle component analysis (PCA), and RIN analysis led us to several findings that can explain the serious impact of M184I mutation on drug resistance. These findings verified that mutation decreased drug binding affinity by ~8 kcal/mol, distorted the ligand optimum orientation in the RT active site, affected the overall enzyme conformational landscape, and distorted the atomic interaction network with the ligand.
The findings of this report can provide potential markers for further design of novel inhibitors that are less susceptible to drug resistance.
Supplementary Materials
RMSD versus time and Potential Energy (kcal/mol) versus time data for ionized and unionized form of wild and M184I variant of HIV-RT complexed with lamivudine as well as of M184V variant is provided with Supplementary Materials 1 and Supplementary Materials 2.
References
Morah, E. U. (2007). Are people aware of their HIV-positive status responsible for driving the epidemic in subsaharan Africa? The case of Malawi. Development Policy Review, 25(2), 215–242.
Pani, A., Loi, A. G., Mura, M., Marceddu, T., La Colla, P., & Marongiu, M. E. (2002). Targeting HIV: Old and new players. Current Drug Target Infectious Disorders, 2, 17–32.
Esposito, F., Corona, A., & Tramontano, E. (2012). HIV-1 reverse transcriptase still remains a new drug target: Structure function, classical inhibitors, and new inhibitors with innovative mechanisms of actions. Molecular Biology International, 2012, 1–23.
Schinazi, R. F., Hernandez-Santiago, B. I., & Hurwitz, S. J. (2006). Pharmacology of current and promising nucleosides for the treatment of human immunodeficiency viruses (vol 71, pg 322, 2006). Antiviral Research, 72, 256.
Ilina, T., LaBarge, K., Sarafianos, S. G., Ishima, R., & Parniak, M. A. (2012). Inhibitors of HIV-1 reverse transcriptase—associated ribonuclease H activity. Biology, 1, 521–541.
Vivet-Boudou, V., Didierjean, J., Isel, C., & Marquet, R. (2006). Nucleoside and nucleotide inhibitors of HIV-1 replication. Cellular and Molecular Life Sciences, 63, 163–186.
Bauman, J. D., Das, K., Ho, W. C., Baweja, M., Himmel, D. M., Clark, A. D, Jr, et al. (2008). Crystal engineering of HIV-1 reverse transcriptase for structure-based drug design. Nucleic Acids Research, 36, 5083–5092.
Turner, D., Brenner, B., & Wainberg, M. A. (2003). Multiple effects of the M184V resistance mutation in the reverse transcriptase of human immunodeficiency virus type 1. Clinical and Diagnostic Laboratory Immunology, 10, 979–981.
Ray, A. S. (2005). Intracellular interactions between nucleos(t)ide inhibitors of HIV reverse transcriptase. AIDS Reviews, 7(2), 113–125.
Hamers, R. L., Kityo, C., Sigaloff, K. C., & de Wit, T. F. R. (2013). Pretreatment HIV-1 drug resistance in Africa. Lancet Infectious Diseases, 13, 476.
Wainberg, M. A., & Turner, D. (2004). Resistance issues with new nucleoside/nucleotide backbone options. JAIDS Journal of Acquired Immune Deficiency Syndromes, 37, S36–S43.
Gao, H. Q., Boyer, P. L., Sarafianos, S. G., Arnold, E., & Hughes, S. H. (2000). The role of steric hindrance in 3TC resistance of human immunodeficiency virus type-1 reverse transcriptase. Journal of Molecular Biology, 300, 403–418.
Sarafianos, S. G., Das, K., Clark, A. D., Ding, J. P., Boyer, P. L., Hughes, S. H., & Arnold, E. (1999). Lamivudine (3TC) resistance in HIV-1 reverse transcriptase involves steric hindrance with beta-branched amino acids. Proceedings of the National Academy of Sciences of the United States of America, 96, 10027–10032.
Purohit, R. (2014). Role of ELA region in auto-activation of mutant KIT receptor: A molecular dynamics simulation insight. Journal of Biomolecular Structure and Dynamics, 32, 1033–1046.
Bahareh, H., Govender, T., Maguire, G. E. M., Soliman, M. E. S., & Kruger, H. G. (2013). Integrated approach to structure-based enzymatic drug design: molecular modeling, spectroscopy, and experimental bioactivity. Chemical Reviews,. doi:10.1021/cr300314q.
Purohit, R., Rajendran, V., & Sethumadhavan, R. (2011). Studies on adaptability of binding residues and flap region of TMC-114 resistance HIV-1 protease mutants. Journal of Biomolecular Structure and Dynamics, 29, 137–152.
Rajendran, V., Purohit, R., & Sethumadhavan, R. (2012). In silico investigation of molecular mechanism of laminopathy caused by a point mutation (R482W) in lamin A/C protein. Amino Acids, 43, 603–615.
Purohit, R., Rajendran, V., & Sethumadhavan, R. (2011). Relationship between mutation of serine residue at 315th position in M. tuberculosis catalase-peroxidase enzyme and isoniazid susceptibility: An in silico analysis. Journal of Molecular Modeling, 17, 869–877.
Balu, K., Rajendran, V., Sethumadhavan, R., & Purohit, R. (2013). Investigation of binding phenomenon of NSP3 and p130Cas mutants and their effect on cell signalling. Cell Biochemistry and Biophysics, 67, 623–633.
Rajendran, V., & Sethumadhavan, R. (2014). Drug resistance mechanism of PncA in Mycobacterium tuberculosis. Journal of Biomolecular Structure and Dynamics, 32, 209–221.
Aruksakunwong, O., Wolschann, P., Hannongbua, S., & Sompornpisut, P. (2006). Molecular dynamic and free energy studies of primary resistance mutations in HIV-1 protease-ritonavir complexes. Journal of Chemical Information and Modeling, 46, 2085–2092.
Hou, T. J., & Yu, R. (2007). Molecular dynamics and free energy studies on the wild-type and double mutant HIV-1 protease complexed with amprenavir and two amprenavir-related inhibitors: Mechanism for binding and drug resistance. Journal of Medicinal Chemistry, 50, 1177–1188.
Stoica, I., Sadiq, S. K., & Coveney, P. V. (2008). Rapid and accurate prediction of binding free energies for saquinavir-bound HIV-1 proteases. Journal of the American Chemical Society, 130, 2639–2648.
Zhou, Z. G., Madrid, M., Evanseck, J. D., & Madura, J. D. (2005). Effect of a bound non-nucleoside RT inhibitor on the dynamics of wild-type and mutant HIV-1 reverse transcriptase. Journal of the American Chemical Society, 127, 17253–17260.
Xue, W. W., Qi, J., Yang, Y., Jin, X. J., Liu, H. X., & Yao, X. J. (2012). Understanding the effect of drug-resistant mutations of HIV-1 intasome on raltegravir action through molecular modeling study. Molecular BioSystems, 8, 2135–2144.
Chachra, R., & Rizzo, R. C. (2008). Origins of resistance conferred by the R292K neuraminidase mutation via molecular dynamics and free energy calculations. Journal of Chemical Theory and Computation, 4, 1526–1540.
Liu, H. X., Yao, X. J., Wang, C. Q., & Han, J. A. (2010). In silico identification of the potential drug resistance sites over 2009 influenza A (H1N1) virus neuraminidase. Molecular Pharmaceutics, 7, 894–904.
Guo, Z. Y., Prongay, A., Tong, X., Fischmann, T., Bogen, S., Velazquez, F., et al. (2006). Computational study of the effects of mutations A156T, D168V, and D168Q on the binding of HCV protease inhibitors. Journal of Chemical Theory and Computation, 2, 1657–1663.
Pan, D. B., Xue, W. W., Zhang, W. Q., Liu, H. X., & Yao, X. J. (2012). Understanding the drug resistance mechanism of hepatitis C virus NS3/4A to ITMN-191 due to R155K, A156V, D168A/E mutations: A computational study. Biochimica Et Biophysica Acta General Subjects, 1820, 1526–1534.
Cheng, X. L., Cui, G. L., Hornak, V., & Sinnnerling, C. (2005). Modified replica exchange simulation methods for local structure refinement. Journal of Physical Chemistry B, 109, 8220–8230.
Affentranger, R., Tavernelli, I., & Di Iorio, E. E. (2006). A novel Hamiltonian replica exchange MD protocol to enhance protein conformational space sampling. Journal of Chemical Theory and Computation, 2, 217–228.
Okur, A., Wickstrom, L., Layten, M., Geney, R., Song, K., Hornak, V., & Simmerling, C. (2006). Improved efficiency of replica exchange simulations through use of a hybrid explicit/implicit solvation model. Journal of Chemical Theory and Computation, 2, 420–433.
Liu, P., Kim, B., Friesner, R. A., & Berne, B. J. (2005). Replica exchange with solute tempering: A method for sampling biological systems in explicit water. Proceedings of the National Academy of Sciences of the United States of America, 102, 13749–13754.
Maisuradze, G. G., Liwo, A., & Scheraga, H. A. (2009). Principal component analysis for protein folding dynamics. Journal of Molecular Biology, 385, 312–329.
Thomas, J. R., Gedeon, P. C., Grant, B. J., & Madura, J. D. (2012). LeuT conformational sampling utilizing accelerated molecular dynamics and principal component analysis. Biophysical Journal, 103, L01–L03.
Amadei, A., Linssen, A. B., de Groot, B. L., van Aalten, D. M., & Berendsen, H. J. (1996). An efficient method for sampling the essential subspace of proteins. Journal of Biomolecular Structure and Dynamics, 13, 615–625.
van Aalten, D. M., Findlay, J. B., Amadei, A., & Berendsen, H. J. (1995). Essential dynamics of the cellular retinol-binding protein—Evidence for ligand-induced conformational changes. Protein Engineering Design and Selection, 8, 1129–1135.
Amadei, A., Linssen, A. B. M., & Berendsen, H. J. C. (1993). Essential dynamics of proteins. Proteins: Structure Function, and Bioinformatics, 17, 412–425.
Word, J. M., Lovell, S. C., LaBean, T. H., Taylor, H. C., Zalis, M. E., Presley, B. K., et al. (1999). Visualizing and quantifying molecular goodness-of-fit: Small-probe contact dots with explicit hydrogen atoms. Journal of Molecular Biology, 285, 1711–1733.
Case, D. A. (1994). Normal-mode analysis of protein dynamics. Current Opinion in Structural Biology, 4, 285–290.
Brooks, B., & Karplus, M. (1985). Normal-modes for specific motions of macromolecules—Application to the Hinge-Bending mode of lysozyme. Proceedings of the National Academy of Sciences of the United States of America, 82, 4995–4999.
Laine, E., de Beauchene, I. C., Perahia, D., Auclair, C., & Tchertanov, L. (2011). Mutation D816V alters the internal structure and dynamics of c-KIT receptor cytoplasmic region: Implications for dimerization and activation mechanisms. PLoS Computational Biology, 7(6), e1002068–e1002068.
Teodoro, M. L., Phillips, G. N., & Kavraki, L. E. (2003). Understanding protein flexibility through dimensionality reduction. Journal of Computational Biology, 10, 617–634.
Yang, L., Song, G., Carriquiry, A., & Jernigan, R. L. (2008). Close correspondence between the motions from principal component analysis of multiple HIV-1 protease structures and elastic network modes. Structure, 16, 321–330.
del Sol, A., Fujihashi, H., Amoros, D., & Nussinov, R. (2006). Residues crucial for maintaining short paths in network communication mediate signaling in proteins. Molecular Systems Biology, 2. doi:10.1038/msb4100063.
Welsch, C., Schweizer, S., Shimakami, T., Domingues, F. S., Kim, S., Lemon, S. M., & Antes, I. (2012). Ketoamide resistance and hepatitis C virus fitness in Val55 variants of the NS3 serine protease. Antimicrobial Agents and Chemotherapy, 56, 1907–1915.
Welsch, C., Domingues, F. S., Susser, S., Antes, I., Hartmann, C., Mayr, G., et al. (2008). Molecular basis of telaprevir resistance due to V36 and T54 mutations in the NS3-4A protease of the hepatitis C virus. Genome Biology, 9, R16.
Xue, W. W., Jin, X. J., Ning, L. L., Wang, M. X., Liu, H. X., & Yao, X. J. (2013). Exploring the molecular mechanism of cross-resistance to HIV-1 integrase strand transfer inhibitors by molecular dynamics simulation and residue interaction network analysis. Journal of Chemical Information and Modeling, 53, 210–222.
Doncheva, N. T., Klein, K., Domingues, F. S., & Albrecht, M. (2011). Analyzing and visualizing residue networks of protein structures. Trends in Biochemical Sciences, 36, 179–182.
Doncheva, N. T., Assenov, Y., Domingues, F. S., & Albrecht, M. (2012). Topological analysis and interactive visualization of biological networks and protein structures. Nature Protocols, 7, 670–685.
Ahmed, S. M., Kruger, H. G., Govender, T., Maguire, G. E., Sayed, Y., Ibrahim, M. A., et al. (2013). Comparison of the molecular dynamics and calculated binding free energies for nine FDA-approved HIV-1 PR drugs against subtype B and C-SA HIV PR. Chemical Biology and Drug Design, 81, 208–218.
Soliman, M. E. S. (2013). A hybrid structure/pharmacophore-based virtual screening approach to design potential leads: A computer-aided design of south African HIV-1 subtype C protease inhibitors. Drug Development Research, 74, 283–295.
Kanibolotsky, D. S., Novosyl’na, O. V., Abbott, C. M., Negrutskii, B. S., & El’skaya, A. V. (2008). Multiple molecular dynamics simulation of the isoforms of human translation elongation factor 1A reveals reversible fluctuations between “open” and “closed” conformations and suggests specific for eEF1A1 affinity for Ca(2+)-calmodulin. BMC Structural Biology, 8(1), 4.
Goetz, A. W., Williamson, M. J., Xu, D., Poole, D., Le Grand, S., & Walker, R. C. (2012). Routine microsecond molecular dynamics simulations with AMBER on GPUs. 1. Generalized born. Journal of Chemical Theory and Computation, 8, 1542–1555.
Salomon-Ferrer, R., Goetz, A. W., Poole, D., Le Grand, S., & Walker, R. C. (2013). Routine microsecond molecular dynamics simulations with AMBER on GPUs. 2. Explicit solvent particle mesh Ewald. Journal of Chemical Theory and Computation, 9, 3878–3888.
Salomon-Ferrer, R., Case, D. A., & Walker., R. C. (2013). An overview of the Amber biomolecular simulation package. WIREs Computational Molecular Science, 3, 198–210.
Case, D. A., Darden, T. A., Cheatham, III, T. E., Simmerling, C. L., Wang, J., Duke, R. E., et al. (2012). AMBER 12. San Francisco: University of California.
Lindorff-Larsen, K., Piana, S., Palmo, K., Maragakis, P., Klepeis, J. L., Dror, R. O., & Shaw, D. E. (2010). Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins Structure Function and Bioinformatics, 78, 1950–1958.
Cieplak, P., Cornell, W. D., Bayly, C., & Kollman, P. A. (1995). Application of the multimolecule and multiconformational RESP methodology to biopolymers: Charge derivation for DNA, and proteins. Journal of Computational Chemistry, 16, 1357–1377.
Frisch, M. J., Trucks, G. W., Schlegel, H. B., Scuseria, G. E., Robb, M. A., Cheeseman, J. R., et al. (2009). Gaussian 09, Revision D.01. Wallingford, CT: Gaussian, Inc.
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W., & Klein, M. L. (1983). Comparison of simple potential functions for simulating liquid water. Journal of Chemical Physics, 79, 926–935.
Essmann, U., Perera, L., Berkowitz, M. L., Darden, T., Lee, H., & Pedersen, L. G. (1995). A smooth particle mesh Ewald method. Journal of Chemical Physics, 103, 8577–8593.
Ryckaert, J. P., Giovanni, C., & Berendsen, H. J. C. (1977). Numerical integration of the cartesian equations of motion of a system with constraints: Molecular dynamics of n-alkanes. Journal of Computational Physics, 23, 327–341.
Le Grand, S., Götz, A. W., & Walker, R. C. (2013). SPFP: Speed without compromise—A mixed precision model for GPU accelerated molecular dynamics simulations. Computer Physics Communications, 184, 374–380.
Berendsen, H. J. C., Postma, J. P. M., van Gunsteren, W. F., DiNola, A., & Haak, J. R. (1984). Molecular dynamics with coupling to an external bath. Journal of Chemical Physics, 81, 3684–3690.
Kollman, P. A., Massova, I., Reyes, C., Kuhn, B., Huo, S. H., Chong, L., et al. (2000). Calculating structures and free energies of complex molecules: Combining molecular mechanics and continuum models. Accounts of Chemical Research, 33, 889–897.
Massova, I., & Kollman, P. A. (2000). Combined molecular mechanical and continuum solvent approach (MM-PBSA/GBSA) to predict ligand binding. Perspectives in Drug Discovery and Design, 18, 113–135.
Tsui, V., & Case, D. A. (2000). Theory and applications of the Generalized born solvation model in macromolecular simulations. Biopolymers, 56, 275–291.
Onufriev, A., Bashford, D., & Case, D. A. (2000). Modification of the generalized Born model suitable for macromolecules. Journal of Physical Chemistry B, 104, 3712–3720.
Humphrey, W., Dalke, A., & Schulten, K. (1996). VMD: Visual molecular dynamics. Journal of Molecular Graphics and Modelling, 14, 33–38.
Bakan, A., Meireles, L. M., & Bahar, I. (2011). ProDy: Protein dynamics inferred from theory and experiments. Bioinformatics, 27, 1575–1577.
Shannon, P., Markiel, A., Ozier, O., Baliga, N. S., Wang, J. T., Ramage, D., et al. (2003). Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Research, 13, 2498–2504.
Choo, H., Chong, Y., & Chu, C. K. (2003). The role of 2′,3′-unsaturation on the antiviral activity of anti-HIV nucleosides against 3TC-resistant mutant (M184V). Bioorganic and Medicinal Chemistry Letters, 13, 1993–1996.
Diallo, K., Gotte, M., & Wainberg, M. A. (2003). Molecular impact of the M184V mutation in human immunodeficiency virus type 1 reverse transcriptase. Antimicrobial Agents and Chemotherapy, 47, 3377–3383.
Acknowledgments
The authors acknowledge the School of Health Sciences, UKZN, for financial support and the Center of High Performance Computing (CHPC, www.chpc.ac.za) for computational facilities. SB acknowledges the consultancy support from Open Source Drug Design and In Silico Molecules (www.insilicomolecule.org) community. RCW acknowledges funding from the National Science Foundation (NSF) through the Scientific Software Innovations Institutes program NSF SI2-SSE (NSF114876) and a fellowship from NVIDIA Inc.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of Interest
The authors declare that they have no potential conflicts of interest.
Additional information
Suri Moonsamy and Soumendranath Bhakat have contributed equally to this work.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Moonsamy, S., Bhakat, S., Walker, R.C. et al. Single Active Site Mutation Causes Serious Resistance of HIV Reverse Transcriptase to Lamivudine: Insight from Multiple Molecular Dynamics Simulations. Cell Biochem Biophys 74, 35–48 (2016). https://doi.org/10.1007/s12013-015-0709-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12013-015-0709-2