Abstract
Osteosarcoma (OS) is a high-grade intraosseous malignancy. Twenty to thirty percent of OS patients react poorly to standard therapy with a combination of surgical resection and chemotherapy. It is necessary to find molecules that play an important role in this. This study explored the role of TRIM4 in OS chemotherapy sensitivity and malignant progression. The expression of TRIM4 in OS tissues and cells was examined by RT-qPCR, immunohistochemical staining, and western blot. Specific siRNA was transfected into U2-OS and SAOS2 cells to target TRIM4. Cell biological behavior was examined by CCK-8, Transwell, and flow cytometry experiments. Cisplatin-resistant SAOS2 (SAOS2-Cis-R) cells were established, and the effect of TRIM4 expression on the cisplatin response of SAOS2 cells was tested. Knockdown of TRIM4 significantly inhibited the proliferation, migration, and invasion of U2-OS and SAOS2 cells and induced apoptosis. TRIM4 expression was significantly higher in chemotherapy-resistant OS tissues compared to chemotherapy-sensitive OS tissues. Furthermore, the expression of TRIM4 in SAOS2-Cis-R cells was significantly increased compared to parental SAOS2 cells. Moreover, overexpression of TRIM4 enhanced cisplatin resistance in parental SAOS2 cells, while the downregulation of TRIM4 expression enhanced cisplatin sensitivity of SAOS2-Cis-R cells. High TRIM4 expression might be associated with malignant progression and poor response to chemotherapy response of OS. Targeting TRIM4 may be beneficial for OS treatment or combination therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Osteosarcoma (OS) is one of the three most common primary bone tumors that also include Ewing’s sarcoma and chondrosarcoma, which is characterized by osteolytic, proliferative lesions and a propensity for lung metastasis [1]. Bone cancers are relatively uncommon cancers with 800–1000 diagnosed cases each year, and OS accounts for about 20% [1]. OS shows a bimodal distribution, the first peak appears at 15–19 years old (8 cases/million/year), the second peak appears at 75–79 years old (6 cases/million/year), and 25–59 years old is the low-medium platform period (1–2 cases/million/year) [2]. Since the 1970s, standard therapy with a combination of chemotherapy and surgical resection has raised the long-term survival rate of OS patients from less than 20 to 70% [3]. However, over the past three decades, continued progress in standard therapies to improve survival has slowed. Twenty to thirty percent of OS patients react poorly to conventional treatment. OS patients with unresectable primary tumors or metastatic tumors (mainly in the lungs) have a low 5-year survival rate, about 30% or less [4]. Therefore, there is still a need to find novel therapeutic targets, methods, and agents.
TRIM4 is an E3 ubiquitin ligase that regulates steroid hormone levels [5], oxidative stress [6, 7], mitochondrial dysfunction [6], and interferon activation after viral infection [8]. TRIM4 sensitizes cells to H2O2-induced cell death [6]. TRIM4 is expressed differently in human tissues and in most of the analyzed human cancer cell lines [6]. Furthermore, its different types of genetic mutations are found in many types of diseases. For example, in non-alcoholic steatohepatitis hepatocellular carcinoma (HCC) [9] and neural tube defects [10], the TRIM4 gene is hypomethylated and highly expressed at mRNA and protein levels. TRIM4-BRAF or TRIM4-MET fusion gene is identified in aggressive mucinous adenocarcinoma [11], glioblastoma [12, 13], gastrointestinal stromal carcinoma [14], and melanoma [15]. In addition, Bien et al. report that the risk of colorectal cancer is closely related to the single nucleotide polymorphism of TRIM4 and PYGL genes using a genome-wide analysis [16]. However, little is currently known about the role of TRIM4 in cancer.
In this study, the differential expression of TRIM4 in OS tissues with different responses to chemotherapy was investigated, and its effects on OS cell proliferation, invasion, apoptosis, and cisplatin tolerance were analyzed.
Materials and Methods
Patients and Tissue Samples
OS tissue samples were obtained from 39 patients who underwent histopathological diagnosis and surgical resection at Shandong Second Provincial General Hospital. All patients had primary OS without other cancers or major diseases, and not received chemotherapy or other therapy before surgical resection. Twenty-four patients were followed who received chemotherapy postoperatively, of which 12 were chemotherapy sensitive and 12 were resistant. The study was approved from the institutional research ethics committee and received written consent from patients.
Cell Lines and Culture
HOS, MG63, U2-OS, SAOS2, and hFOB1.19 were obtained from ATCC and cultured in DMEM (Gibco, Hyclone, UT) containing 10% FBS at 37 °C in a 5% CO2 atmosphere. Cisplatin-resistant SAOS2 (SAOS2-Cis-R) cells were established by gradually increasing the concentrations of cisplatin (0.1 to 1 μg/ml) (Sigma-Aldrich, Germany) in the culture medium. When cells grow to 60–70% confluence, cisplatin was added in steps at concentrations of 0.1, 0.2, 0.3, 0.5, and 1 μg/ml. Cells were first exposed to cisplatin for two consecutive days and then replaced with fresh medium without cisplatin until the surviving cells recover. When the cells grow again to 60–70% confluence, cisplatin was added to the medium again. Each concentration was repeated 6 times, and there were 7 passages for each concentration. The fold increase in cisplatin resistance for each variant was determined by comparing its IC50 value (the concentration of the drug that resulted in a 50% inhibition of cell growth after 96 h of in vitro treatment) with that of its corresponding parental cell line. Finally, the cells that could grow in 1 μg/ml cisplatin were SAOS2-Cis-R cells.
Plasmid, siRNA, and Transfection
The human full-length TRIM4 cDNA sequence was cloned into the pcDNA3.1 plasmid to construct the TRIM4 overexpression plasmid. The siRNA specifically targeting human TRIM4 was synthesized by Invitrogen. 1 × 106 cells were inoculated into each well of a 6-well plate and cultured in serum-free medium. Cells were required to grow to 50–65% confluence for siRNA transfection and 75–95% confluence for overexpression plasmid transfection. Two hundred fifty microliters of Opti-MEM low-serum medium diluted 100 pmol siRNA (4 μg overexpression plasmid) and Lipofectamine 2000 reagent (Invitrogen, Carlsbad, CA), respectively. Subsequently, the two dilutions were mixed and added to the cells after standing for 20 min at room temperature.
Immunohistochemistry (IHC)
Samples were formalin-fixed and embedded in paraffin. Paraffin blocks were sectioned and sequentially dewaxed and antigenically repaired. Tissue sections were blocked with 5% goat serum and incubated with TRIM4 antibody (1:500, PA5-31491, rabbit polyclonal antibody, ThermoFisher Scientific) overnight at 4 °C, followed by incubation with a secondary antibody at room temperature for 1 h. Immunoreactivity was visualized with an enhanced DAB chromogenic kit.
Immunostaining scores were given by two independent and experienced pathologists. The score was the product of the positive stained cell ratio (R) and the staining intensity (I). R: 0 (<5%), 1 (5–25%), 2 (25–50%), 3 (>51%). S: 0 (negative), 1 (weak), 2 (middle), 3 (strong). A total score less than 4 was considered as low expression, while 4–9 was considered as high expression.
RNA Isolation and RT-qPCR
Total RNA was extracted from human tissues and cultured cells using TRIzol reagent (Invitrogen, Carlsbad, CA), and reverse transcribed into cDNA using the Reverse Transcription Kit. PCR was performed using Power SYBR Green Master Mix. Relative expression levels were calculated by 2−ΔΔCq, and GAPDH was used as an internal control.
Western Blot
Cells were lysed in RIPA buffer, and cell lysates were centrifuged at 12,000 g for 10 min at 4 °C to remove impurities. Proteins were electrophoresed by SDS-PAGE, separated according to molecular weight size, and transferred to a PVDF membrane. After being blocked with 5% skim milk, the membrane was incubated with TRIM4 antibody (PA5-90823, ThermoFisher Scientific) at 4 °C overnight, followed by secondary antibody (31466, ThermoFisher Scientific) for 2 h at 37 °C. Finally, the band was visualized using an enhanced chemiluminescent HRP substrate.
CCK-8 Assay
1 × 103 cells were inoculated into each well of a 96-well plate and continued to be cultured or treated with different concentrations of cisplatin for a certain time. Ten percent CCK-8 reagent was added to each well. After incubation for 2 h at 37 °C, the absorbance was measured at 450 nm using a microplate reader.
Transwell Assay
Cells were inoculated on the top of Matrigel-coated Chambers (BD Biosciences) and incubated or treated with cisplatin for 24 h at 37 °C to analyze cell invasion. For migration assay, a chamber uncoated with Matrigel was used. Migrated or invaded cells were fixed and stained, and finally counted.
Apoptosis Detection
Cells were cultured or treated with cisplatin for 24 h. Then, cells were digested with EDTA-free trypsin and resuspended to 1 × 106 cells/ml with binding buffer. Five microliters of FITC-Annexin V and 5 μl PI were added to 1 ml of cell suspension. After gentle mixing, the assay was performed with a FACScan flow cytometry system.
Statistical Analysis
Data were analyzed using SPSS 21.0 software and performed as mean ± SD. Two-tailed Student’s t-test and ANOVA were used to evaluate statistical differences between different groups. P < 0.05 was considered statistically significant.
Results
TRIM4 Expression is Upregulated in Chemotherapy-Resistant OS Tissues
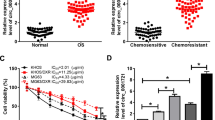
Firstly, mRNA levels of TRIM4 in 39 cases of OS tissue samples were examined using qRT-PCR (Fig. 1A and B). The TRIM4 mRNA was highly expressed in 69.23% (27/39) of OS tissue samples. Subsequently, 12 cases of samples that were sensitive to postoperative chemotherapy and 12 cases who responded poorly to chemotherapy were screened. Compared to chemotherapy-sensitive OS tissues (8/12), TRIM4 was higher expressed in chemotherapy-resistant OS tissues (11/12) (Fig. 1C and D).
TRIM4 expression is upregulated in chemotherapy-resistant OS tissues. A The mRNA levels of TRIM4 in 39 cases of OS tissue samples using qRT-PCR. The test of each tissue sample was repeated 3 times. B The sample proportion with TRIM4 mRNA low or high expression. Those lower than the average relative expression level of TRIM4 were low expression, and those higher were high expression. C The protein expression of TRIM4 in 12 cases of chemotherapy-sensitive and 12 cases of chemotherapy-resistant OS tissue samples with IHC. D The sample proportion with TRIM4 protein low or high expression
Knockdown of TRIM4 Inhibits the Malignant Progress of U2-OS and SAOS2 Cells In Vitro
Compared to hFOB1.19 cell line, the expression of TRIM4 in all HOS, MG63, U2-OS, and SAOS2 cell lines was significantly upregulated (P < 0.05, Fig. 2A and B). Subsequently, U2-OS and SAOS2 cell lines were selected for cellular experiments in vitro. The siRNA specifically targeting TRIM4 (KD) significantly reduced TRIM4 expression in U2-OS and SAOS2 cells (P < 0.05, Fig. 2C and D). Moreover, the downregulation of TRIM4 expression significantly inhibited the proliferation (Fig. 2E and F), migration (Fig. 3A), and invasion (Fig. 3B) of U2-OS and SAOS2 cells, and promoted apoptosis (Fig. 3C). The results of CCK-8 experiments showed that TRIM4-siRNA-transfected cells had much lower absorbance values at 450 nm at 48 h and 72 h after transfection than that in control cells (Fig. 2E and F). The results of Transwell assays suggested that the number of cells completing migration and invasion simulation in the KD group was much less than that in the NC group (Fig. 3A and B). In addition, the flow cytometry results displayed a significant upregulation of the proportion of apoptotic cells after TRIM4 knockdown (Fig. 3C).
The expression of TRIM4 was downregulation in U2-OS and SAOS2 cells. The mRNA (A) and protein (B) expression of TRIM4 in OS cell lines (HOS, MG63, U2-OS, and SAOS2) and normal osteoblast cell line (hFOB1.19) were examined. The specifical siRNA (KD) significantly reduced the mRNA (C) and protein (D) expression of TRIM4 in U2-OS and SAOS2 cells. The proliferation of U2-OS (E) and SAOS2 (F) by using CCK-8 assay. *P < 0.05
TRIM4 Enhances Cisplatin Resistance in SAOS2 Cells
In view of the high expression of TRIM4 in chemotherapy-resistant OS tissues, we established a cisplatin-resistant cell (SAOS2-Cis-R) to explore the role of TRIM4 in cisplatin resistance in OS cells. Figure 4A–C demonstrates the response of parental SAOS2 cells and SAOS2-CIS-R cells to cisplatin. As shown in Fig. 4A and B, the number of surviving parental SAOS2 cells decreased significantly with increasing cisplatin concentration or treatment time (Fig. 4A), while SAOS2-Cis-R cells maintained good cell viability under 1 μg/ml cisplatin treatment, and some cells survived even at 10 μg/ml cisplatin treatment for 72 h (Fig. 4B). In addition, SAOS2-Cis-R cells (Cis(−)) showed a lower apoptosis rate compared to the parental SAOS2 cells (P < 0.05, Fig. 4C). After 24 h of 5 μg/ml cisplatin treatment (Cis(+)), SAOS2-CIS-R cells still showed a lower apoptosis rate compared to parental SAOS2 cells (P < 0.05, Fig. 4C). Critically, according to the results of western blot, compared to the parental SAOS2 cells, the protein expression of TRIM4 in SAOS2-Cis-R cells was increased significantly (P < 0.05, Fig. 4D).
The cisplatin-resistant cell line (SAOS2-Cis-R) is established. The cell survival of parent SAOS2 (A) and SAOS2-Cis-R (B) cells treated with difference cisplatin concentration and treatment time by using CCK-8 assays. C The apoptosis ratio of parental SAOS2 and SAOS2-Cis-R cells treated with 5 μg/ml cisplatin for 24 h by using flow cytometry. D The protein expression of TRIM4 in parental SAOS2 and SAOS2-Cis-R cells by western blot. E The expression of TRIM4 in the parental SAOS2 cells was upregulated using the transfection of pcDNA3.1-TRIM4 plasmid (OE), and the expression of TRIM4 in SAOS2-Cis-R cells was downregulated using the transfection of siRNA (KD). NC, negative control. *P < 0.05, #P < 0.05
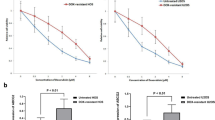
To verify the association of TRIM4 expression with cisplatin response in SAOS2 cells, TRIM4 expression in parental SAOS2 cells was upregulated using overexpression plasmid transfection (OE, Fig. 4E), and it was downregulated in SAOS2-Cis-R cells using siRNA transfection (KD, Fig. 4E). Subsequently, the effect of TRIM4 expression on cisplatin response in SAOS2 cells was examined using CCK-8 and apoptosis experiments. As shown in Fig. 5, overexpression of TRIM4 enhanced the cisplatin resistance of the parental SAOS2 cells, while the downregulation of TRIM4 enhanced the cisplatin sensitivity of SAOS2-Cis-R cells.
TRIM4 enhances the cisplatin resistance of SAOS2 cells. A The cell survival of parent SAOS2 cells upregulated the TRIM4 expression (SAOS2-TRIM4) and SAOS2-Cis-R cells downregulated the TRIM4 expression (SAOS2-Cis-R-siRNA) that treated with difference concentration of cisplatin for 24 h by using CCK-8 assays. B and C The apoptosis ratio of parent SAOS2 cells upregulated the TRIM4 expression (SAOS2-TRIM4) and SAOS2-Cis-R cells downregulated the TRIM4 expression (SAOS2-Cis-R-siRNA) that treated with 5 μg/ml cisplatin for 24 h by using flow cytometry. *P < 0.05
Discussion
In vitro cellular experiments performed in this study indicated that downregulating the expression of TRIM4 in OS cells can significantly inhibit cell proliferation, migration, and invasion and induce apoptosis. Targeting TRIM4 may be beneficial to inhibiting OS progression. Importantly, we found an association between TRIM4 expression and chemotherapy resistance in OS cells. TRIM4 appeared to be highly expressed in cisplatin-resistant OS tissues and cell lines, and altering the expression of TRIM4 in OS cell lines can affect cellular sensitivity to cisplatin.
The combination of chemotherapy and surgical removal of the primary tumor is the main reason for the cure of OS patients. Unfortunately, OS is a particularly chemotherapy-resistant tumor. These tumors respond only to high-dose chemotherapy and are able to rapidly acquire resistance, which is reflected in recurrent patients with salvage rates as low as 20%. Although the addition of cisplatin and isocyclophosphamide has improved survival rates in OS patients, there has been no improvement in OS outcomes or strategies in the ensuing decades. Chemotherapy resistance includes both primary and acquired resistance, with more than half of cancers exhibiting primary resistance and the remaining 50% acquiring resistance as a result of treatment. Targeting TRIM4 may be beneficial in improving primary and acquired resistance in OS patients. It is also of great interest to explore the response of TRIM4 protein expression to chemotherapeutics agents other than cisplatin, and continued research is needed to investigate the molecular mechanisms by which TRIM4 regulates the sensitivity and resistance of OS cells to chemotherapeutics. In addition, OS lacks clear driver mutations and therapeutic targets, and the progress of targeted therapy has been slow. The drivers and targets of TRIM4 and other family members in OS progression deserve further investigation.
The human TRIM family has over 77 members, most of which belong to the E3 ubiquitin ligase. Generally, TRIM proteins bind to ubiquitin conjugating enzyme (E2) through the N-terminal RING domain to catalyze the formation of ubiquitin chains on substrates. The expression of TRIM7, 10, 14, 46, 59, and 66 has been reported to be significantly upregulated in OS and correlated with poor prognosis [17,18,19,20,21,22]. Furthermore, TRIM7, 10, 14, and 66 not only promote the malignant progression of OS but are also responsible for the tumor chemoresistance [17,18,19]. In contrast, TRIM22 expression is reported to be downregulated in OS and associated with a better prognosis, and its overexpression could inhibit cancer progression by activating cellular autophagy [23, 24]. At present, little is known about the biological functions of TRIM4. Zhan et al. find that compared to human bronchial epithelial cell (HBE), the mRNA level of TRIM4 was upregulated in non-small cell lung cancer cell lines using RT-qRCR [25]. However, Dong et al. report the low protein expression of TRIM4 in HCC tissues and its independent correlation with prognosis [26]. Another report seems to illustrate the opposite result. Tian et al. report the DNA hypomethylation and high mRNA expression of TRIM4 in HCC tissues [9]. It is unclear whether there are post-transcriptional regulatory mechanisms in tumor tissues that make TRIM4 mRNA and protein expression trends inconsistent. In addition, there is a need to reveal the presence of TRIM4 gene mutations in cancer progression. There are some limitations in this study. Physical activity has a positive impact on patients with osteosarcoma [27]. Physical activity was also associated with trait creativity [28].
In conclusion, TRIM4 expression was associated with malignant progression and chemoresistance of OS and has the potential to be a therapeutic target for OS. However, the current studies are rather primitive and more in-depth studies are needed.
Data Availability
The datasets generated and analyzed during the current study are available from the corresponding author on reasonable request.
References
Gardner, H. L., Sivaprakasam, K., Briones, N., Zismann, V., Perdigones, N., Drenner, K., et al. (2019). Canine osteosarcoma genome sequencing identifies recurrent mutations in DMD and the histone methyltransferase gene SETD2. Communications Biology, 2, 266. https://doi.org/10.1038/s42003-019-0487-2
Savage, S. A., & Mirabello, L. (2011). Using epidemiology and genomics to understand osteosarcoma etiology. Sarcoma, 2011, 548151. https://doi.org/10.1155/2011/548151
Rickel, K., Fang, F., & Tao, J. (2017). Molecular genetics of osteosarcoma. Bone, 102, 69–79. https://doi.org/10.1016/j.bone.2016.10.017
Isakoff, M. S., Bielack, S. S., Meltzer, P., & Gorlick, R. (2015). Osteosarcoma: Current treatment and a collaborative pathway to success. Journal of Clinical Oncology, 33(27), 3029–3035. https://doi.org/10.1200/JCO.2014.59.4895
Canet, L. M., Sanchez-Maldonado, J. M., Caliz, R., Rodriguez-Ramos, A., Lupianez, C. B., Canhao, H., et al. (2019). Polymorphisms at phase I-metabolizing enzyme and hormone receptor loci influence the response to anti-TNF therapy in rheumatoid arthritis patients. The Pharmacogenomics Journal, 19(1), 83–96. https://doi.org/10.1038/s41397-018-0057-x
Tomar, D., Prajapati, P., Lavie, J., Singh, K., Lakshmi, S., Bhatelia, K., et al. (2015). TRIM4; a novel mitochondrial interacting RING E3 ligase, sensitizes the cells to hydrogen peroxide (H2O2) induced cell death. Free Radical Biology and Medicine, 89, 1036–1048. https://doi.org/10.1016/j.freeradbiomed.2015.10.425
Li, C., & He, L. (2023). The relationship between preeclampsia and arsenic concentration in the peripheral blood shows association rather than causation. Biological Trace Element Research, 201(1), 2–3. https://doi.org/10.1007/s12011-022-03157-y
Sun, X., Xian, H., Tian, S., Sun, T., Qin, Y., Zhang, S., et al. (2016). A hierarchical mechanism of RIG-I ubiquitination provides sensitivity, robustness and synergy in antiviral immune responses. Scientific Reports, 6, 29263. https://doi.org/10.1038/srep29263
Tian, Y., Arai, E., Makiuchi, S., Tsuda, N., Kuramoto, J., Ohara, K., et al. (2020). Aberrant DNA methylation results in altered gene expression in non-alcoholic steatohepatitis-related hepatocellular carcinomas. Journal of Cancer Research and Clinical Oncology, 146(10), 2461–2477. https://doi.org/10.1007/s00432-020-03298-4
Zhang, H., Guo, Y., Gu, H., Wei, X., Ma, W., Liu, D., et al. (2019). TRIM4 is associated with neural tube defects based on genome-wide DNA methylation analysis. Clinical Epigenetics, 11(1), 17. https://doi.org/10.1186/s13148-018-0603-z
Shim, H. S., Kenudson, M., Zheng, Z., Liebers, M., Cha, Y. J., Hoang Ho, Q., et al. (2015). Unique genetic and survival characteristics of invasive mucinous adenocarcinoma of the lung. Journal of Thoracic Oncology, 10(8), 1156–1162. https://doi.org/10.1097/JTO.0000000000000579
Zheng, Z., Liebers, M., Zhelyazkova, B., Cao, Y., Panditi, D., Lynch, K. D., et al. (2014). Anchored multiplex PCR for targeted next-generation sequencing. Nature Medicine, 20(12), 1479–1484. https://doi.org/10.1038/nm.3729
Shi, J., & He, L. (2023). Long-term use of anti-coronary heart disease medications may impact the serum zinc concentration. Biological Trace Element Research, 201(1), 1. https://doi.org/10.1007/s12011-022-03115-8
Vanden Bempt, I., Vander Borght, S., Sciot, R., Spans, L., Claerhout, S., Brems, H., et al. (2021). Comprehensive targeted next-generation sequencing approach in the molecular diagnosis of gastrointestinal stromal tumor. Genes, Chromosomes and Cancer, 60(4), 239–249. https://doi.org/10.1002/gcc.22923
Yeh, I., Botton, T., Talevich, E., Shain, A. H., Sparatta, A. J., de la Fouchardiere, A., et al. (2015). Activating MET kinase rearrangements in melanoma and Spitz tumours. Nature Communications, 6, 7174. https://doi.org/10.1038/ncomms8174
Bien, S. A., Su, Y. R., Conti, D. V., Harrison, T. A., Qu, C., Guo, X., et al. (2019). Genetic variant predictors of gene expression provide new insight into risk of colorectal cancer. Human Genetics, 138(4), 307–326. https://doi.org/10.1007/s00439-019-01989-8
Zhou, C., Zhang, Z., Zhu, X., Qian, G., Zhou, Y., Sun, Y., et al. (2020). N6-methyladenosine modification of the TRIM7 positively regulates tumorigenesis and chemoresistance in osteosarcoma through ubiquitination of BRMS1. EBioMedicine, 59, 102955. https://doi.org/10.1016/j.ebiom.2020.102955
Li, Y. J., Zhang, G. P., Zhao, F., Li, R. Q., Liu, S. J., Zhao, Z. R., et al. (2018). Target therapy of TRIM-14 inhibits osteosarcoma aggressiveness through the nuclear factor-kappaB signaling pathway. Experimental and Therapeutic Medicine, 15(3), 2365–2373. https://doi.org/10.3892/etm.2017.5679
Xi, X., Bao, Y., Zhou, Y., Chen, Y., Zhong, X., Liao, J., et al. (2021). Oncogenic gene TRIM10 confers resistance to cisplatin in osteosarcoma cells and activates the NF-kappaB signaling pathway. Cell Biology International, 45(1), 74–82. https://doi.org/10.1002/cbin.11468
Jiang, W., Cai, X., Xu, T., Liu, K., Yang, D., Fan, L., et al. (2020). Tripartite motif-containing 46 promotes viability and inhibits apoptosis of osteosarcoma cells by activating NF-B signaling through ubiquitination of PPAR. Oncology Research, 28(4), 409–421. https://doi.org/10.3727/096504020X15868639303417
Liang, J., Xing, D., Li, Z., Shen, J., Zhao, H., & Li, S. (2016). TRIM59 is upregulated and promotes cell proliferation and migration in human osteosarcoma. Molecular Medicine Reports, 13(6), 5200–5206. https://doi.org/10.3892/mmr.2016.5183
Chen, Y., Guo, Y., Yang, H., Shi, G., Xu, G., Shi, J., et al. (2015). TRIM66 overexpresssion contributes to osteosarcoma carcinogenesis and indicates poor survival outcome. Oncotarget, 6(27), 23708–23719. https://doi.org/10.18632/oncotarget.4291
Liu, W., Zhao, Y., Wang, G., Feng, S., Ge, X., Ye, W., et al. (2022). TRIM22 inhibits osteosarcoma progression through destabilizing NRF2 and thus activation of ROS/AMPK/mTOR/autophagy signaling. Redox Biology, 53, 102344. https://doi.org/10.1016/j.redox.2022.102344
He, L. P., Song, Y. X., Zhu, T., Gu, W., & Liu, C. W. (2022). Progress in the relationship between vitamin D deficiency and the incidence of type 1 diabetes mellitus in children. Journal of Diabetes Research, 2022, 5953562. https://doi.org/10.1155/2022/5953562
Zhan, W., Han, T., Zhang, C., Xie, C., Gan, M., Deng, K., et al. (2015). TRIM59 promotes the proliferation and migration of non-small cell lung cancer cells by upregulating cell cycle related proteins. PLoS One, 10(11), e0142596. https://doi.org/10.1371/journal.pone.0142596
Dong, Z. R., Zhou, W., Sun, D., Yan, Y. C., Yang, C. C., Yang, Y. F., et al. (2020). Role of the E3 ubiquitin ligase TRIM4 in predicting the prognosis of hepatocellular carcinoma. Journal of Cancer, 11(14), 4007–4014. https://doi.org/10.7150/jca.37164
Hao, S., Zhu, J., Zhang, X., Qiu, J., Xuan, Q., & Ye, L. (2021). Comprehensive analysis of aerobic exercise-related genes identifies CDCA4 that promotes the progression of osteosarcoma. Frontiers in Genetics, 12, 637755. https://doi.org/10.3389/fgene.2021.637755
Li, C. P., Liu, X. H., Wang, X. J., & He, L. P. (2023). Trait creativity, personality, and physical activity: A structural equation model. Annals of Palliative Medicine, 12(1), 141–149. https://doi.org/10.21037/apm-22-1310
Author information
Authors and Affiliations
Contributions
Yan Li and Jie Gao carried out the experiments and drafted the manuscript; Dong Wang contributed to the immunohistochemistry experiments; Zijin Liu was involved in the statistical analysis; Huawu Zhang managed the experimental design, reviewed the manuscript, and provided funding support. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Ethical Approval
Our study was approved by the Ethics Committee of Shandong Second Provincial General Hospital. All procedures involving people comply with the ethical standards of the institutional and/or national committee for research ethics and the 1964 Helsinki Declaration and its subsequent changes or comparable ethical standards.
Consent to Participate
The study was approved by the patient’s written consent and the institutional research ethics committee.
Consent for Publication
All other authors have read the manuscript and have agreed to submit it in its current form for consideration for publication in the Applied Biochemistry and Biotechnology.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, Y., Gao, J., Wang, D. et al. TRIM4 Expression Related to Malignant Progression and Cisplatin Resistance in Osteosarcoma. Appl Biochem Biotechnol 196, 233–244 (2024). https://doi.org/10.1007/s12010-023-04551-5
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-023-04551-5