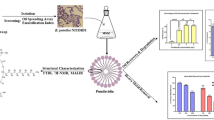
Abstract
Bacillus licheniformis TT42 produced a low-molecular weight anionic biosurfactant that reduced the surface tension of water from 72 to 27 mN/m and the interfacial tension from 12 to 0.05 mN/m against crude oil. We have earlier reported significant enhancement in oil recovery in laboratory sand pack columns and core flood studies, by biosurfactant-TT42 compared to standard strain, Bacillus mojavensis JF2. In the context of this application of the biosurfactant-TT42, its characterization was deemed important. In the preliminary studies, the biosurfactant-TT42 was found to be functionally stable at under conditions of temperature, pH, and salinity generally prevalent in oil reservoirs. Furthermore, the purified biosurfactant-TT42 was found to have a CMC of 22 mg/l. A newly developed activity staining TLC method was used for the purification of biosurfactant-TT42. Structural characterization of biosurfactant-TT42 using TLC, Fourier transform infrared spectroscopy (FTIR), GC-MS, and matrix-assisted laser desorption ionization time of flight (MALDI-TOF)/TOF suggested that it was a mixture of lipopeptide species, all having a common hydrophilic cyclic heptapeptide head with the sequence, Gln-Leu/Ileu-Leu/Ileu-Val-Asp-Leu/Ileu-Leu/Ileu linked to hydrophobic tails of different lengths of 3β-OH-fatty acids bearing 1043, 1057 and 1071 Da molecular weight, where 3β-OH-C19 fatty acid was predominant. This is the longest chain length of fatty acids reported in a lipopeptide.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Surfactants are amphipathic molecules since they possess both hydrophilic and hydrophobic moieties. They enrich at interfaces, lower surface tension (ST) and interfacial tension (IFT), and form micelle [1]. Biosurfactants are structurally diverse surfactants of microbial origin that include low-molecular weight biosurfactants (glycolipids and lipopeptides) that lower ST and high-molecular weight biosurfactants (amphipathic polysaccharides, proteins, lipopolysaccharide, lipoproteins, and complex biopolymer mixture) that stabilize emulsions. Numerous bacteria and fungi produce biosurfactants of varied chemical nature [2]. They possess emulsifying, foaming, detergency, and dispersing properties. Biosurfactants offer the possibility of replacing chemical surfactants, produced from non-renewable resources, with alternatives produced from cheap renewable feedstocks. Biosurfactants are also attractive because they are eco-friendly and robust enough for industrial use [3].
Cyclic lipopeptide classes are prominent biosurfactants, produced as secondary metabolites by Bacillus sp. having unique structure, exceptional surfactant power, antibacterial activity, and antifungal activity [4, 5]. They have advantage over the chemical surfactants due to their high activity, specificity, biodegradability, and versatility having varied applications in textile, pharmaceutical, cosmetics, crop protection, food, detergents, and oil industries, of which microbial enhanced oil recovery (MEOR) is assuming importance in recent years [6, 7]. MEOR is a cost-effective, eco-friendly, alternative tertiary recovery process which uses microorganisms or their metabolites for recovery of oil from the reservoir. Lichenysins are most potent anionic cyclic lipoheptapeptide biosurfactants produced by Bacillus licheniformis using hydrocarbonless medium with mainly glucose as carbon source. They have the capacity to lower the ST of water from 72 to ∼27 mN/m. Lichenysin is synthesized by a multienzyme complex called lichenysin synthetase. The structure of lichenysin and its operon indicated the non-ribosomal biosynthesis with the same multifunctional modular arrangement as seen in surfactin synthetase (SrfA). The lichenysin biosynthesis operon (lic) cloned and sequenced from B. licheniformis ATCC 10716 consists of three peptide synthetase genes licA, licB, and licC encoding the proteins LicA, LicB, and LicC [8]. LicA and LicB consist of three each and LicC one amino acid activating modules. All seven modules contain three domains each, adenylation (A) domain that recognizes and activates specific amino acids by adenylation at the expense of an ATP, a thioester-binding (T) domain that catalyzes the covalent binding of the activated amino acid at a specific serine site via a thioester linkage to a 4′-phosphopantetheine cofactor, and condensation (C) domain that forms a peptide bond between two activated amino acids. The DNA sequence of licA is about 98 % identical to a sequence of a lchA operon obtained from the lichenysin A producing strain B. licheniformis BNP29 [9]. Lichenysins have been characterized from varied B. licheniformis strains, namely, A [10], B [11], C [12], D [8], G [13], surfactant BL86 [14], F2.2 [15], and BC98 [16]. Reported literature indicated that the lichenysin biosynthesis operons for all lichenysin isoforms have emerged from the same origin. Alterations in the primary peptide sequence may be due to slight evolution-dependent strain-specific mutations [8, 9, 17]. In the present study, the characterization of biosurfactant from B. licheniformis TT42 was undertaken since it produced a highly surface active low-molecular weight anionic biosurfactant (i.e., biosurfactant-TT42), and it was found to possess excellent MEOR potential since it gave 34.6 ± 3.7 % additional oil recovery in laboratory-simulated sand pack columns. This is 5.2 % higher than that obtained from crude lichenysin biosurfactant produced by the standard strain, Bacillus mojavensis JF2 [18]. Purified biosurfactant-TT42 reduced the ST of water from 72 to 27 mN/m and the IFT from 12 to 0.05 mN/m against crude oil. In core flood studies, it gave 10.64 % additional oil recovery (unpublished data). In this background, its structural characterization was deemed interesting and the results of the studies undertaken thereof are presented here.
Materials and Methods
Microorganism
B. licheniformis TT42 is a Gram-positive, sporulating, rod-shaped, and motile strain, isolated from Tuva-Timba hot water spring, Gujarat, India. It was identified based on morphological, biochemical, and physiological characterization and 16S rRNA gene sequence analysis (NCBI Genbank accession number DQ922951) [19].
Production of Biosurfactant
Of sterile K. Jenny’s medium [12] in 250-ml Erlenmeyer flask, 50 ml was inoculated with 2 %, 12-h-old inoculum of B. licheniformis TT42 and incubated at 37 °C on shaker at 180 rpm for 72 h. The culture broth was centrifuged at 10,000 rpm for 30 min, and the cell-free supernatant (CFS) was used as crude biosurfactant.
ST and IFT Measurements
ST measurement is generally used as an indirect method to measure the surfactant concentration. ST of the crude biosurfactant was measured using a Du-Nuoy’s tensiometer (Win-Son & Co., Kolkata, India) based on the ring detachment technique. Its IFT was measured using a spinning drop interfacial tensiometer (Temco, USA) [18, 20].
Stability Studies of the Biosurfactant-TT42
To determine the stability of biosurfactant produced by B. licheniformis TT42 under extreme conditions, effects of temperature (65–100 °C), salinity (0–15 %), and pH (3–11) on the ST-reducing ability of the crude biosurfactant-TT42 were studied before and after incubation under the above mentioned conditions [21]. Temperature stability of the biosurfactant was studied by incubating the biosurfactant-TT42 at 65, 75, 85, and 100 °C for different time periods. For salt stability, different concentrations of NaCl were added to the biosurfactant-TT42 solution and mixed until complete dissolution, followed by 25 days incubation. To check the pH stability, pH of the biosurfactant-TT42 solution were adjusted to different pH values with NaOH or HCl, followed by incubation [21].
Extraction and Purification of the Biosurfactant-TT42
Crude biosurfactant was precipitated with 40 % (w/v) ammonium sulfate and incubated overnight at 4 °C. The precipitate, containing the biosurfactant, was then collected by centrifugation at 15,000 rpm for 30 min at 4 °C and extracted with chilled acetone. The extract was subjected to TLC (Silica gel 60F254, Merck) and resolved with a solvent system of isopropanol:water:28 % (w/v) NH4OH (90:10:6) [22]. In the present study, instead of oil spreading method used, an activity staining was used to demonstrate surface activity on the TLC plate itself. For that, the TLC plates were developed by spraying blood reagent (20 % human blood in potassium phosphate buffer, pH −7). The spot of biosurfactant shows a clear zone of hemolysis, while rest of the plate remains red.
Critical Micelle Concentration Determination of Biosurfactant-TT42
The concentration at which biosurfactant micelles begin to form is called critical micelle concentration (CMC). It is the minimum concentration of biosurfactant required to achieve the maximum surface activity [23]. Purified biosurfactant was determined by dissolving 220 mg of purified biosurfactant in 35 ml distilled water. This solution was further serially diluted (1–350-fold), and ST value of each dilution was measured. CMC was obtained from the biosurfactant concentration vs. ST plot where CMC is that ST value at which there is a sudden increase in ST.
Chemical Analysis of the Biosurfactant-TT42
Detection of Peptide Moiety
Biosurfactant-TT42 was hydrolyzed with 6 M HCl at 110 °C for 3 h. Ten microliter of the hydrolyzed and unhydrolyzed biosurfactant samples were spotted on TLC plate; solvent system used was butanol:acetic acid:water (12:3:5). Of ninhydrin (in acetone), 0.2 % was used as a developing reagent. After spraying the developing agent, the TLC plate was heated at 100 °C for 3 min. The purple-colored spot confirmed the presence of peptide moiety [24, 25].
Detection of Lipid Moiety
Biosurfactant-TT42 was spotted on TLC plate and run in the solvent system containing isopropanol:water:28 % (w/v) NH4OH (90:10:6). Lipid components were detected by subjecting the plate to iodine vapors in a chamber. A yellow-colored spot confirmed the presence of lipid moiety [26].
Fourier Transform Infrared Spectroscopy Analysis
IR spectrum of the purified biosurfactant-TT42 was recorded in potassium bromide pellets using Fourier transform infrared spectroscopy (FTIR) spectroscopy (PerkinElmer Rx 1, Department of Chemistry, M. S. University of Baroda, Vadodara, Gujarat, India).
Mass Spectrometry Analysis of the Biosurfactant-TT42
Fatty Acid Analysis
The methyl ester/trimethylsilyl (ME/TMS) derivatives of lipid moiety of biosurfactant-TT42 were formed, and their analysis was carried out by GC-MS using reported literature [22, 27].
Peptide Analysis
To determine the molecular weight of the purified biosurfactant-TT42 and the amino acid sequence of its peptide moiety, matrix-assisted laser desorption ionization time-of-flight (MALDI TOF/TOF) analysis was carried out [22]. The analysis was performed using ULTRAFLEX-TOF/TOF instrument (Bruker Daltonics, IISc, Bangalore, India).
Results
Stability Studies of the Biosurfactant-TT42
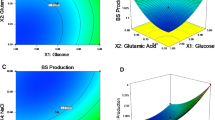
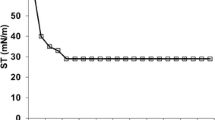
Temperature stabilities of the biosurfactant produced by B. licheniformis TT42 were checked at 65, 75, 85, and 100 °C for different time periods as depicted in Fig. 1a, b. The results implied that the biosurfactant remained stable up to 85 °C for 25 days. The stability study when extended for 90 days at 85 °C, the biosurfactant retained 75.7 % of its surface activity. The biosurfactant was also found to be stable at 100 °C for 300 min. Moreover, the ST values of crude biosurfactant were 27–30 mN/m for 25 days with no loss of activity.
Salt stability of the biosurfactant produced by B. licheniformis TT42 was checked in the presence of 5, 10, and 15 % NaCl up to 25 days of incubation. As shown in Fig. 1c, the biosurfactant was stable in presence of 5 % NaCl. In presence of 10 and 15 % NaCl concentrations, slight increase in ST values was observed. pH stability of biosurfactant produced by B. licheniformis TT42 was checked at a wide range of pH, and it was found to be stable in the pH range of 5 to 9. Slight loss of activity was observed at pH 3 and pH 11 (Fig. 1d). In the purification of biosurfactant-TT42, activity staining method was used for the detection of biosurfactant on TLC plate. It made detection and purification of biosurfactant fast and easy. Three spots, viz., A, B, and C, showed RBC lysis (clearance) against the red background of non-lysed RBC. Out of these, the spot A, showing maximum activity, was scrapped off and purified (Fig. 2).
CMC of Biosurfactant-TT42
The purified biosurfactant-TT42 was dissolved in water and used to determine its CMC value, which was found to be 22 mg/l. The amount of biosurfactant-TT42 produced by selected strain was calculated as 0.58 g/l.
Chemical Analysis of Biosurfactant-TT42
Detection of Peptide and Lipid Moieties
TLC of the hydrolyzed biosurfactant-TT42 showed a purple-colored spot when developed with ninhydrin, whereas the unhydrolyzed biosurfactant-TT42 showed no spot with ninhydrin. This indicated that the α-amino group of unhydrolyzed biosurfactant became free on hydrolysis, which reacted with ninhydrin and gave a purple-colored spot (Fig. 3a). Thus, the peptide moiety of biosurfactant-TT42 might be cyclic in nature. Presence of lipid in biosurfactant-TT42 was indicated by the yellow-colored spot on TLC when developed with iodine vapors (Fig. 3b).
FTIR analysis of the purified biosurfactant-TT42 showed band characteristics of peptide at 3400 cm−1 (−NH stretching), 1700 cm−1 (>C = O bond), and 1500 cm−1 (deformation). The bands at 2960–2940 and 1480–1450 cm−1 correspond to that of aliphatic chains (–CH3, –CH2–; Fig. 4). The results obtained indicated the presence of peptide and lipid moieties in biosurfactant-TT42.
Mass Spectrometry Analysis of Biosurfactant-TT42
ME/TMS derivatives of fatty acid moiety of biosurfactant-TT42 were analyzed by GC-MS. The individual peaks obtained by GC (Fig. 5a) were subjected to mass spectrometry analysis. The results of GC-MS analysis indicated that the fatty acid moiety of biosurfactant-TT42 contained a mixture of 3β-OH fatty acids of different chain lengths, ranging from C13 to C19. Figure 5b shows the mass spectrum of predominant peak, identified as 3β-OH-C19 fatty acid on the basis of the characteristic peaks of 3β-hydroxy fatty acid.
MALDI-TOF spectrum of the purified biosurfactant-TT42 is shown in Fig. 6. Total of three peaks with m/z of 1043, 1057, and 1071 Da were selected for further fragmentation. The fragmentation pattern (TOF/TOF) of these three peaks indicated that they were the major species of the cyclic lipopeptide biosurfactant. Amino acid sequence of the peptide moiety of the purified biosurfactant was elucidated by interpreting the TOF/TOF spectrum of the precursor ion m/z 1071.70 (Fig. 7). Subtraction of 958.63 from precursor ion m/z 1071.70 leaves a value of 113.07 (Ile/Leu), 845.54 from 958.63 leaves a value of 113.09 (Ile/Leu), 730.50 from 845.54 leaves a value of 115.04 (Asp), 631.41 from 730.50 leaves a value of 99.11 (Val), 518.31 from 631.41 leaves a value of 113.1 (Ile/Leu), 405.27 from 518.31 leaves a value of 113.04 (Ile/Leu), and 277.14 from 405.27 leaves a value of 128.13 (Gln). The most probable peptide sequence of purified biosurfactant-TT42 was deduced to be Gln-Ile/Leu-Ile/Leu-Val-Asp-Leu-Ile/Leu [10].
Discussion
B. licheniformis TT42 producing biosurfactant was isolated for application in oil reservoir to facilitate biosurfactant-mediated MEOR. The biosurfactant used in MEOR must possess good surface/interfacial activity which remained stable under extreme conditions, generally prevalent in oil reservoirs. The suitability of biosurfactant-TT42 for MEOR was determined by physiological characterization. The ST-lowering ability of biosurfactant-TT42 was as efficient (from 72 to 27 mN/m of water) as that reported till date [28, 29]. The IFT-reducing ability of biosurfactant-TT42 against crude oil was also very high, reaching a value of 0.05 from 12 mN/m. Crude biosurfactant-TT42 was observed to remain stable under extreme temperature, pH, and salinity conditions. In case of extended incubation at higher temperature, a slight loss of activity was observed. Similar results have been reported for surfactin and lichenysin [30, 31, 32]. The biosurfactant-TT42 retained its activity in the range of pH 5 to 9. A decrease in surface activity was reported at pH 3 due to biosurfactant precipitation. These results indicate that the pH range of biosurfactant-TT42 correlates with that of surfactin (pH 6 to 12) and lichenysin (pH 6.2 to 10) [31, 32].
The crude biosurfactant-TT42 gave 34.6 ± 3.7 % additional oil recovery in sand pack column studies [18]. To use biosurfactant-TT42 for MEOR from oil reservoirs, acute dermal toxicity test was performed using Wistar rats. It indicated that B. licheniformis TT42 was a non-pathogenic strain, implying that its biosurfactant was also non-toxic in nature (data not shown). This prompted the structural characterization of these metabolites. The structure and composition of the biosurfactant produced by B. licheniformis TT42 were studied by a variety of analytical techniques such as TLC, FTIR, MALDI-TOF/TOF, and GC-MS. An activity staining technique was used for the detection of biosurfactant-TT42 on TLC plate using blood reagent as a developing agent. The biosurfactant-TT42 gave a spot of clearance due to RBC lysis against the red background of non-lysed RBCs. This method developed by us makes the detection and purification of biosurfactant fast and easy as compared to the reported methods [33–35]. Lichenysins are most potent anionic cyclic lipoheptapeptide biosurfactants produced by B. licheniformis on hydrocarbonless medium with mainly glucose as carbon source. Based on species-specific variations, they are named lichenysin A, B, C, D, and G and surfactant BL86. Lichenysins have been reported to have molecular weights 993–1091 Da and CMC in 10–60-mg/l range [28, 36]. A comparison of chemical structure of the cyclic heptapeptide moiety of lichenysins reveals that they have variation at first, fifth, and seventh amino acids mostly. The fatty acid side chain is made of structure of the cyclic heptapeptide moiety of lichenysins reveals th, etc., in different lichenysins [17]. The chemical structure of the purified biosurfactant-TT42, identified by TLC and FTIR analyses, indicated that the biosurfactant-TT42 was a cyclic lipopeptide. Lipid moiety of the biosurfactant-TT42 was analyzed by GC-MS. A total of 20 peaks, obtained by GC, were further analyzed by MS. The peaks of 3β-OH-fatty acids were identified by MS of the ME/TMS derivatives giving the characteristic fragment ions. The results obtained from GC-MS indicated that the lipophilic part of biosurfactant-TT42 contained a mixture of 3c-OH fatty acids of different chain lengths, out of which the 3β-OH-C19 fatty acid, obtained at a run time of 29.009 min, was predominant. As reported, long-chain fatty acids have high surface and interfacial activity, implying that biosurfactant-TT42 was useful for MEOR [10]. Molecular weight of the biosurfactant and amino acid sequence of its peptide moiety were determined by MALDI-TOF/TOF as this analytical technique is highly efficient in characterization of the molecular structure of secondary metabolites like biosurfactants [37]. The [M + H]+ ions generated by this technique are very stable, thus leading to intense signals which are useful for rapidly determining the heterogeneity of the samples submitted to MS analysis [38]. The reported molecular weight of the major component of surfactin, lichenysin B, surfactant BL86, and lichenysin G is 1035 Da, while the molecular weights of the major components of lichenysin C and A range from 1022 to 1036 03 and 1006 to 1034 Da, respectively [10–14]. Based on GC-MS and MALDI-TOF/TOF analyses, it can be assumed that biosurfactant-TT42 was a mixture of lipopeptides with different molecular weights in the range of 1023–1071. These lipopeptides had identical heptapeptide moiety of seven amino acids. The difference in molecular weights of the lipopeptides produced by B. licheniformis TT42 was due to the difference in the chain length of lipopeptides produced by T42 C12 to C19. The sequence of heptapeptide moiety of biosurfactant-TT42, obtained by MALDI-TOF/TOF, showed similarity with lichenysins A, D, and G, a known biosurfactant produced by B. licheniformis strains BAS50, ATCC 10716, and IM1307 [8, 10, 13, 27], since the heptapeptide moiety of biosurfactant-TT42 contained glutamine (Gln) at its first position, aspartic acid (Asp) at its fifth position, and isoleucine (Ile) at its seventh position. Hence, the most probable structure of biosurfactant-TT42 is assumed to be a cyclic heptapeptide having the sequence L-Gln-L-Leu-D-Leu-L-Val-L-Asp-D-Leu-L-Ile linked to 3β-OH-fatty acids. This unique low-molecular weight chemical structure is capable of lowering the interfacial tension very low, and by virtue of this, the oil recovery is facilitated.
References
Neu, T. (1996). Significance of bacterial surface active compounds in interaction of bacteria with interfaces. Microbiological Reviews, 60, 151–166.
Desai, J., & Banat, I. (1997). Microbial production of surfactants and their commercial potential. Microbiology and Molecular Biology Reviews, 61, 47–64.
Marchant, R., & Banat, I. (2012). Microbial biosurfactants: challenges and opportunities for future exploitation. Trends in Biotechnology, 30, 558–565.
Vater, J. (1986). Lipopeptides, an attractive class of microbial surfactants. Progress Colloid and Polymer Science, 72, 2–18.
Fernandes, P., de Arruda, I., dos Santos, A., Maior, A., & Ximenes, E. (2007). Antimicrobial activity of surfactants produced by Bacillus subtilis R14 against multidrug-resistant bacteria. Brazilian Journal of Microbiology, 38, 704–709.
Makkar, R., Cameotra, S., & Banat, I. (2011). Advances in utilization of renewable substrates for biosurfactant production. AMB Express, 1, 1–19.
Monteiro, L., Mariano, R., & Souto-Maior, A. (2005). Antagonism of Bacillus spp. against Xanthomonas campestris pv. Campestris. Brazilian Archives of Biology and Technology, 48, 23–29.
Konz, D., Doekel, S., & Marahiel, M. (1999). Molecular and biochemical characterization of the protein template controlling biosynthesis of the lipopeptide lichenysin. Journal of Bacteriology, 181, 133–140.
Yakimov, M., Kroger, A., Slepak, T., Giuliano, L., Timmis, K., & Golyshin, P. (1998). A putative lichenysin a synthetase operon in Bacillus licheniformis: initial characterization. Biochimica et Biophysica Acta, 1399, 141–153.
Yakimov, M., Abraham, W., Meyer, H., Giuliano, L., & Golyshin, P. (1999). Structural characterization of lichenysin A components by fast atom bombardment tandem mass spectrometry. Biochimica Et Biophysica Acta, 1438, 273–280.
Lin, S., Minton, M., Sharma, M., & Georgiou, G. (1994). Structural and immunological characterization of a biosurfactant produced by Bacillus licheniformis JF-2. Applied and Environmental Microbiology, 60, 31–38.
Jenny, K., Kappeli, O., & Fletcher, A. (1991). Biosurfactants from Bacillus licheniformis: structural analysis and characterization. Applied Microbiology and Biotechnology, 36, 5–13.
Grangemard, I., Bonmatin, J., Bernillon, J., & Das, B. (1999). Lichemysin G, a novel family of lipopeptide biosurfactants from Bacillus licheniformis IM1307: production, isolation and structural evaluation by NMR and mass spectrometry. The Journal of Antibiotics, 52, 363–373.
Horowitz, S., & Griftin, W. (1991). Structural analysis of Bacillus licheniformis 86 surfactant. Journal of Industrial Microbiology, 7, 45–52.
Thaniyavarn, J., Roongsawang, N., Kameyama, T., & Haruki, M. (2003). Production and characterization of biosurfactants from Bacillus licheniformis F2.2. Bioscience, Biotechnology, and Biochemistry, 67, 1239–1244.
Tendulkar, S., Saikumari, Y., Patel, V., Raghotama, S., Munshi, T., Balaram, P., et al. (2007). Isolation, purification and characterization of an antifungal molecule produced by Bacillus licheniformis BC98, and its effect on phytopathogen Magnaporthe grise. Journal of Applied Microbiology, 103, 2331–2239.
Nerurkar, A. (2010). Structural and molecular characteristics of lichenysin and its relationship with surface activity. In R. Sen (Ed.), Biosurfactants (pp. 304–315). Texas: Landes Bioscience.
Suthar, H., Hingurao, K., Desai, A., & Nerurkar, A. (2008). Evaluation of bioemulsifier mediated microbial enhanced oil recovery using sand pack column. Journal of Microbiological Methods, 75, 225–230.
Joshi, S., Suthar, H., Kumar, A., Hingurao, K., & Nerurkar, A. (2013). Occurrence of biosurfactant producing Bacillus spp. in diverse habitat. ISRN Biotechnology, 2013, 1–6.
Goswami, D., Handique, P., & Deka, S. (2014). Rhamnolipid biosurfactant against Fusarium sacchari—the causal organism of pokkah boeng disease of sugarcane. Journal of Basic Microbiology, 54, 548–557.
Eddouaouda, K., Mnif, S., Badis, A., Younes, S., Cherif, S., Ferhat, S., et al. (2011). Characterization of a novel biosurfactant produced by Staphylococcus sp. strain 1E with potential application on hydrocarbon bioremediation. Journal Basic Microbiology, 52, 408–418.
Youssef, N., Duncan, K., & McInerney, M. (2005). Importance of 3-hydroxy fatty acid composition of lipopeptides for biosurfactant activity. Applied and Environmental Microbiology, 71, 7690–7695.
McInerney, M., Knapp, R., Chisholm, J., Bhupathiraju, V., & Coates, J. (1999). Proceedings of the 8th International Symposium on Microbial Ecology. In C. Bell, M. Brylinsky, & P. Johnson-Green (Eds.), Use of indigenous or injected microorganisms for enhanced oil recovery, in microbial biosystems: new frontiers (p. 377). Halifax: Atlantic Canada Society for Microbial Ecology.
Cooper, D., & Goldenberg, B. (1987). Surface-active agents from two Bacillus species. Applied and Environmental Microbiology, 53, 224–229.
Suthar, H., Hingurao, K., Desai, A., & Nerurkar, A. (2009). Selective plugging strategy based microbial enhanced oil recovery using Bacillus licheniformis TT33. Journal of Microbiology and Biotechnology, 19, 1230–1237.
Sims, R., & Larose, J. (1962). The use of iodine vapor as a general detecting agent in the thin layer chromatography of lipids. Journal of the American Oil Chemists’ Society, 39, 232.
Yakimov, M., Timmis, K., Wray, V., & Fredrickson, H. (1995). Characterization of a new lipopeptide surfactant produced by thermotolerant and halotolerant subsurface Bacillus licheniformis BAS50. Applied and Environmental Microbiology, 61, 1706–1713.
Grangemard, I., Wallach, J., Maget-Dana, R., & Peypoux, F. (2001). Lichenysin: a more efficient cation chelator than surfactin. Applied Biochemistry and Biotechnology, 90, 199–210.
Haddad, N., Wang, J., & Mu, B. (2009). Identification of a biosurfactant producing strain: Bacillus subtilis HOB2. Protein and Peptide Letters, 16, 7–13.
Arima, K., Tamura, G. and Kakinuma, A. (1972) Surfactin. US Patent, US 3687926, 8, 29
Jenneman, G., McInerney, M., Knapp, R., Clark, J., Ferro, J., Revus, D., et al. (1983). A halotolerant, biosurfactant-producing Bacillus species potentially useful for enhanced oil recovery. Developments Industrial Microbiology, 24, 485–492.
McInerney, M., Jenneman, G., Knapp, R. and Menzie, D. (1985) Biosurfactant and enhanced oil recovery. US Patent, US 4522261, 6, 11.
Afshar, S., Lotfabad, T., Roostaazad, R., Najafabadi, A., & Noghabi, K. (2008). Comparative approach for detection of biosurfactant-producing bacteria isolated from Ahvaz petroleum excavation areas in south of Iran. Annales de Microbiologie, 58, 79–83.
Satpute, S., Bhawsar, B., Dhakephalkar, P., & Chopade, B. (2008). Assessment of different screening methods for selecting biosurfactant producing marine bacteria. Indian Journal Marine Science, 37, 243–250.
Youssef, N., Duncan, K., Nagle, D., Savage, K., Knapp, R., & McInerney, M. (2004). Comparison of methods to detect biosurfactant production by diverse microorganisms. Journal of Microbiological Methods, 56, 339–347.
McInerney, M., Javaheri, M., & Nagle, D. (1990). Properties of the biosurfactant produced by Bacillus licheniformis strain JF-2. Journal of Industrial Microbiology, 5, 95–101.
Vater, J., Kablitz, B., Wilde, C., Franke, P., Mehta, N., & Cameotra, S. (2002). Matrix-assisted laser desorption ionization-time of flight mass spectrometry of lipopeptide biosurfactants in whole cells and culture filtrates of Bacillus subtilis C-1 isolated from petroleum sludge. Applied and Environmental Microbiology, 68, 6210–6219.
Kajimura, J., Fujiwara, T., Yamada, S., Suzawa, Y., Nishida, T., Oyamada, Y., et al. (2005). Identification and molecular characterization of an N-acetylmuramyl-L-alanine amidase Sle1 involved in cell separation of Staphylococcus aureus. Molecular Microbiology, 58, 1087–1101.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest Statement
The authors of the paper have seen and agreed to the submission of the paper to your journal. The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Suthar, H., Nerurkar, A. Characterization of Biosurfactant Produced by Bacillus licheniformis TT42 Having Potential for Enhanced Oil Recovery. Appl Biochem Biotechnol 180, 248–260 (2016). https://doi.org/10.1007/s12010-016-2096-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-016-2096-6