Abstract
Recently, DDX41 mutations have been identified both as germline and acquired somatic mutations in families with multiple cases of late-onset myelodysplastic syndrome (MDS) and/or acute myeloid leukemia. The majority of germline mutations are frameshift mutations suggesting loss of function with DDX41 acting as a tumor suppressor, and there is a common somatic missense mutation found in a majority of germline mutated tumors. Clinically, DDX41 mutations lead to development of high-risk MDS at an age similar to that observed in sporadic cohorts, presenting a unique challenge to hematologists in recognizing the familial context. Functionally, DDX41 has been shown to contribute to multiple pathways and processes including mRNA splicing, innate immunity and rRNA processing. Mutations in DDX41 result in aberrations to each of these in ways that could potentially impact on tumorigenesis—initiation, maintenance or progression. This review discusses the various molecular, clinical and biological aspects of myeloid malignancy predisposition due to DDX41 mutation and highlights how each of these suggest potential therapeutic opportunities through the use of pathway-specific inhibitors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Familial predisposition to myeloid neoplasms
Myeloid neoplasms are a heterogeneous group of diseases encompassing myeloproliferative neoplasms (MPN), myelodysplastic syndromes (MDS) and acute myeloid leukemia (AML), all of which are characterized by abnormal growth and development of cells of the myeloid blood lineage. These disease groups can be further sub-categorized by a number of different morphological and molecular approaches which may correlate with responses to specific treatments [1]. AML and MDS are generally diseases of advanced age with a median age at diagnosis of 70 years [2, 3]. Familial clustering of myeloid malignancies with autosomal dominant (AD) inheritance has been recognized phenotypically for over 100 years (reviewed in [4]), with the first molecular basis discovered through the identification of germline RUNX1 mutations associated with familial platelet disorder with predisposition to myeloid malignancy in 1999 (FPD-MM, OMIM #601399) [5]. Since that time, and recently accelerated by the advent of next-generation sequencing, a growing number of genes have been associated with AD germline predisposition to myeloid malignancies, including mutations in GATA2 [6] as described by us and others [7], as well as ANKRD26 [8], ETV6 [9], CEBPA [10], RBBP6 [11], TERT, TERC [12], DDX41 [13] and most recently mutations in SAMD9 [14] and SAMD9L [15, 16]. This growing recognition and molecular identification of the germline predisposition to a subset of myeloid malignancies has been formalized in the most recent revision to the World Health Organisation guidelines [1]. The initial identification of genetic predisposition factors for myeloid malignancies (e.g., RUNX1 [17], CEBPA [10] and GATA2 [6]) were characteristic as hematopoietic transcription factors, often associated with younger age of malignancy onset. In contrast, the recent identification of germline mutations in DDX41 in familial myeloid malignancies has caused a paradigm shift in the way we view hematological malignancy (HM) predisposition, related both to age of onset and potential function, and this is discussed in more detail below.
Mutation of DDX41 in familial hematological malignancies (FHM)
DDX41 is composed of 17 exons and is encoded on the distal end of the long arm of chromosome 5 (5q35.3). It belongs to the DEAD-box helicase family of genes that have been characterized in multiple cellular roles [18]. DDX41 was first identified as a predisposition gene for myeloid malignancy in a family with 4 family members affected by AML or MDS, where a germline frameshift mutation in DDX41 D140Gfs*2 was found to segregate in all tested affected individuals [19]. AML samples from 2 of the affected individuals had acquired an identical R525H mutation also in DDX41. In the same study cohort, further screening of both familial and sporadic myeloid malignancies identified a further 10 families or singleton cases carrying the germline D140Gfs*2 mutation, as well as identifying additional germline mutations such as I1396T, F183I, Q52fs and M155I mutations (Fig. 1) [19]. Somatic DDX41 mutations were frequently observed in myeloid tumors of germline DDX41 mutated individuals. The most common was the DDX41 R525H mutation seen in the index family and confirmed as present in trans, relative to the germline lesion in one case (i.e., biallelic). Other somatic DDX41 mutations including A225D, E247K, P321L and a splice donor site mutation were identified in tumors from other individuals, collectively identifying somatic DDX41 mutations in 50% of tumors from germline DDX41 mutated individuals (see Table 1) [19].
Germline and somatic DDX41 mutations in hematological malignancies. Two isoforms of DDX41 were identified, with the shorter p55 isoform initiating at methionine 127. Black bars conserved motifs, SA splice acceptor, SD splice donor, underlined somatic cases that are biallelic germline mutations, triangle frameshift/premature stop site mutation, square missense mutation, U ubiquitination, P phosphorylation site, # 12 biallelic cases
Following the initial description of germline DDX41 mutations [19], we reported an additional eight families with myeloid malignancies [20]. Of the eight, three harbored the germline D140Gfs*2 variant, confirming its status as the most common site of mutation in Caucasian populations (Fig. 1, 14 families) [19,20,21]. Two families carried a novel germline M1I mutation which was also seen in subsequent studies, making it the second most frequent germline DDX41 mutation in Caucasian populations (5 families) [20, 22,23,24]. Interestingly, in one family, a germline variant identical to the commonly acquired R525H mutation was identified (confirmed germline in hair). In a second family, a germline G530D mutation was identified, which was subsequently also seen as an acquired mutation in AML (COSMIC ID 5424276), as well as acquired in a familial MDS tumor with a novel germline DDX41 mutation (our unpublished data). This indicates that missense mutations in the helicase C domain may present as either germline or somatic, and care should be taken to ensure that both tumor and normal sources of DNA are used for screening to differentiate between these alternatives. Shared sites of germline and acquired mutation have precedence in other FHM, where several RUNX1 mutations (e.g., R204*, R204Q, R320*) are found both as germline mutations in families with FPD-MM and acquired in sporadic myeloid malignancy cohorts [25, 26].
In the 2 years since the initial identification of germline and somatic DDX41 mutations in myeloid malignancies, the total number of DDX41 mutant families worldwide has risen to over 70 (Fig. 1), making it one of the most frequently mutated myeloid malignancy predisposition genes, with similar numbers of reported RUNX1 mutated families [13]. Reports of germline DDX41 mutations have consistently indicated distinct ethnically associated recurrent germline mutations; currently all reports of the germline D140Gfs*2 mutations are exclusive to Caucasian populations [19, 20, 27], and the germline A500Cfs*9 has been reported in multiple families solely of Asian descent [27]. Population-based sequencing data for both D140Gfs*2 and A500Cfs*9 mutations imply the alleles are very rare (22/277,196 and 1/246,236 alleles, respectively; gnomAD database) [13, 28]. Both populations show acquisition of the DDX41 R525H mutation as the most frequent somatic event in tumors, consistent with a common germline-derived selective pressure for somatic mutations. This restriction of D140Gfs*2 and A500Cfs*9 germline mutations to different ethnic populations and their recurrence suggest that they likely derive from individual founder mutations that have spread through inheritance across generations, rather than representing different sites of de novo recurrent mutations, although this remains to be investigated.
Phenotypic, clinical and biological consequences of DDX41 mutations/loss of function in hematological malignancies
The most common phenotypes of individuals with germline DDX41 mutations are myeloid malignancy subtypes. There are some exceptions to this, including one family reported by us with lymphoma and sarcoidosis development (DDX41 R164W) [20], as well as a broader range of phenotypes associated with the M1I mutation families in addition to MDS/AML (e.g., CML, lymphoma, sarcoidosis).
Within the myeloid neoplasms, germline DDX41 mutations are associated with hypocellular bone marrow, erythroid dysplasia and higher-risk MDS (19% in higher risk vs. 6% in lower risk) and AML. Similarly, DDX41 expression level was lower in higher-risk MDS cases. Despite detection of DDX41 mutations in higher-risk MDS and associated with significantly poor survival, 70% have normal karyotype (NK). From a therapeutic point of view, response to lenalidomide, commonly used for MDS-5q− syndrome, was significantly higher in patients harboring mutations or lower expression of DDX41 [19, 20].
Biologically, shRNA-mediated knockdown of DDX41 leads to enhanced proliferation and colony-forming ability, including enhanced replating ability and hyper-responsiveness to GM-CSF [19]. Knockdown of DDX41 also impairs erythroid differentiation of K562 cells, correlating with our clinical observation that DDX41 mutant individuals often display MDS with erythroid dysplasia and progress to myeloid leukemia of an erythroblastic subtype (M6) [19, 20]. Striking findings about DDX41 in FHM, differentiating it from other known HM predisposition genes, have come through several observations. Firstly, the age of MDS onset in carriers of germline DDX41 mutations is similar to the diagnosis age of sporadic MDS, (median age of myeloid HM diagnosis of all published germline DDX41 cases to date is 65 years (range 41–88). This is unlike other predisposition syndromes which often have an early age of onset [29] (reviewed in [30]). However, we have observed that families with point mutations in the helicase C domain of DDX41 (R525H and G530D) have a significantly younger age of onset of HM than frameshift mutations, suggesting a different mechanism of action of these mutations [20]. Secondly, although the numbers are currently still small, most individuals have normal blood counts up until the point of HM development, unlike other FHM that are often characterized by extended pre-leukemic cytopenic phases. However, red cell macrocytosis, peripheral blood monocytosis and cytopenias have been observed in some carriers prior to development of malignancy [19, 20].
These above aspects are problematic for the detection of familial leukemia with germline mutation of DDX41 in several respects, as the typically advanced age of diagnosis often may not prompt a discussion by treating clinicians of family history in the way that early-onset myeloid malignancy would. Similarly, without a clear pre-malignant phenotype combined with the late age of onset, the family history is easily overlooked by individuals even when questioned about family history. Therefore the expectation is, and our unpublished data and that of Polprasert et al. [19] suggest, that many more cases of germline DDX41 mutated myeloid malignancies will be identified through retrospective screening of ‘sporadic’ cohorts, making this a significant clinical entity for hematology practitioners.
Acquired co-operating mutations and clonal evolution in germline DDX41 mutated cells
Along with the potential difficulties in the ascertainment of family history in DDX41 germline mutation carriers, due to relatively late age of onset, a predictable outcome of predisposition to high-risk MDS is the retrospective finding of asymptomatic germline DDX41 carriers being selected as stem cell transplant (SCT) donors for family members affected with malignancy. This is a treatment issue problematic for all familial leukemia syndromes; for example we and others have documented multiple cases where this has occurred for germline RUNX1 mutated families (our unpublished data and [31, 32]). Two recent studies in the literature document the outcomes of using germline DDX41 mutated donor cells for transplantation. The first described a 62-year-old asymptomatic male sibling who was used as an SCT donor for his 58-year brother diagnosed with AML, who subsequently relapsed with donor cell leukemia (DCL) 4 years post-transplant [24]. Molecular analysis identified a previously described germline M1I missense mutation in both brothers. Similarly in the second study, a 49-year-old individual who developed MDS received a bone marrow transplant from his unaffected sibling and later relapsed with DCL, where both individuals were found to be carriers of a novel DDX41 A500Cfs*9 mutation (Table 1) [33]. Of note, the donors described in both studies were not reported to have developed HM and this suggests that HSC stress (i.e., reconstitution of recipient hematopoiesis) may be a potential trigger for HM development [34]. Also importantly, in the first study, both the initial AML tumor and the relapse MDS tumor acquired independent DNMT3A mutations, suggesting that this is an important co-operating factor. Further, the T cells of the patient at the time of AML diagnosis carried the same acquired DNMT3A mutation, indicating that this individual may have had pre-existing clonal hematopoiesis. Clonal hematopoiesis is observed in individuals of advanced age without germline predisposition and is associated with increased risk of myeloid HM development [35]. Early clonal hematopoiesis may be a common feature of FHM, as it is also found at an early age in FHM individuals with germline RUNX1 mutations [29].
Studies of acquired mutations in DDX41 germline mutated tumors or individuals prior to development of malignancy are limited at this time. Aside from the observation that acquired mutation of R525H on the second DDX41 allele is the most frequent mutation observed in tumors, few recurrent co-operating mutations in other genes have been identified (Table 1). It is quite common for somatic DDX41 mutations to be seen at low variant allele frequencies (VAF) (e.g., 5–25%) in MDS and AML, which may be missed using some screening technologies. Kadono et al. showed in one patient with AML that the VAF for acquired R525H was approaching 50% in CD34+ hematopoietic stem cells (HSC), but was not seen in CD3+ T lymphocytes [36] which can constitute 40–50% peripheral leukocytes. They demonstrated growth inhibition of HSC over-expressing R525H and implied that this mutation predominantly affects myeloid differentiation.
Which co-operating gene mutations are important to trigger the clonal evolution and development of HM in DDX41 predisposed individuals is currently an open question and one of great importance. How the mutations that have been identified in DDX41 to date affect function and the mechanisms by which they could cause HM are discussed below.
DDX41 protein structure
The structure of the DDX41 protein has recently been evaluated and modeled to characterize its domains and predicted binding conformations. As denoted in Fig. 1, both germline and somatic mutations in DDX41 have been reported throughout the protein in myeloid malignancies, suggestive that DDX41 acts as a tumor suppressor gene and/or that different mutations can impact on different domains or moieties with oncogenic consequences.
DDX41 can be divided into four main structural regions; N-terminal domain, DEAD-box domain, helicase C domain and C-terminal domain (Fig. 1). Specific motifs have been distinguished within each domain. The N-terminal domain contains a putative nuclear localization sequence, while the C-terminus possesses a conserved zinc finger. The DEAD-box domain consists of seven conserved motifs (Q, I, IA, IB, IC, II (DEAD), and III) [37,38,39], while the RecA-like helicase C domain comprises four motifs (IV, IVA, V and VI), as demarcated by vertical gray lines in Fig. 1 [37]. DDX41, like other DEAD-box helicases, has been shown to interact with and bind to nucleic acids and ATP for activation [37]. Specifically, by its DNA binding activity, DDX41 acts as a sensor of dsDNA and bacterial-derived cyclic di-nucleotides (CDN) (c-di-AMP, c-di-GMP) [40]. It also binds directly to numerous proteins within the interferon response pathway in the cytoplasm, including Stimulator of interferon genes (STING) [41]. Although a member of the RNA helicase family, the majority of reported nucleic interactions of DDX41 to date involve DNA; however, it does bind to spliceosomal components in the nucleus (e.g., SF3B1 and PRPF8) implying a functional role in RNA processing (discussed further below) [19], and a recent study identified DDX41 as binding to the 3′UTR of the CDKN1A (p21) mRNA [42].
DDX41 binding interactions have been demonstrated, or proposed from structural modeling, to utilize one or a combination of motifs. DNA binding occurs via motifs Q, I and II within the DEAD-box domain [37]. 3D modeling predicts that the helicase C domain can sterically inhibit DNA or CDN binding to the DEAD-box domain. BTK phosphorylation of DDX41 per se does not alter nucleotide binding [43], but does in the cellular context [41].
Many DEAD-box helicases bind RNA via motifs Ia, Ib in the DEAD-box domain and motifs IV and V in the helicase domain [37]. However, despite conservation of these domains, DDX41 has not been shown to bind RNA directly and is dispensable for the interferon response induced by poly(I:C) in mouse dendritic cells [44]. ATP binding has been associated with activation of DDX41, where motifs Q and I are crucial for binding [43] and motifs III and VI for hydrolysis of ATP [37] in helicase activity [43]. Nucleic acids, ATP and STING binding sites as well as BTK phosphorylation sites overlap, consistent with a complex interplay between components in inducing interferon signaling via activated STING (Fig. 2b). This pathway is also regulated by ubiquitination of the N-terminus of DDX41 on K9 and K115 by TRIM21, an E3 ligase that negatively regulates the abundance of the DDX41 protein through proteosomal degradation [45].
Proposed mechanisms for DDX41 involvement in the development of myeloid malignancies. a DDX41 mutation causes aberrant mRNA splicing. U1 snRNP (U1) recognizes and binds to the splice donor region of the upstream exon, and U2 snRNP (U2) binds to the branch point within the intron adjacent to the downstream exon. This recruits the U4/U6–U5 tri-snRNP complex. U1 is displaced by U4/U6–U5 tri-snRNP complex and both U1 and U4 snRNP dissociate. DDX41 interacts with proteins associated in the spliceosomal complex, but its direct contacts and role remain to be determined. The spliceosome excises introns and ligates exons to generate mature mRNAs, and the splicing factors dissociate for recycling. DDX41 R525H promotes aberrant splicing via exon retention and exon skipping. E7107 (pladienolide B derivative) blocks U2 from binding the branch point. Sudemycins directly interact with SF3B proteins to inhibit binding of U2 to the hnRNA. b DDX41 mutation impacts STING pathway and type I interferon response against foreign nucleic acids. Entry of dsDNA or c-di-nucleotides (CDN) into the cytoplasm of the cell activates Bruton’s tyrosine kinase (BTK). Phosphorylation of DDX41 by BTK allows it to bind nucleic acids and to bind to STING. This triggers the STING pathway where STING binds TBK1, which stimulates phosphorylation of IRF3 and entry into the nucleus to initiate a type I interferon response. Ibrutinib inhibits BTK phosphorylation. STING agonists such as DMXAA (specific for mice) show nucleic acid-independent activation of the STING pathway. c DDX41 causes aberrant pre-rRNA trimming in the generation of 5.8S, 18S and 28S rRNA transcripts impacting on ribosome biogenesis. RNA Polymerase I transcribes the 47S pre-rRNA transcript with 5′ and 3′ external transcribed spacers (ETS) and internal transcribed spacers (ITS1 and ITS2). Through sequential trimming of the spacers by either Pathway A (major) or Pathway B (minor), transient intermediate pre-rRNAs 41S, 30S and 21S, and then mature rRNA transcripts 5.8S, 18S and 28S are formed. DDX41 is predominantly associated with removal of 5′ETS and ITS2. DDX41 R525H causes changes to the efficiency of this process. rRNA transcripts subsequently migrate to the cytoplasm, to be assembled into ribosomal subunits. The 18S transcript is incorporated into the smaller 40S subunit, while 5.8S, 28S and a separately generated 5S transcript (chaperoned by RPL5 and RPL11) contribute to the 60S subunit. Together, they form the completed 80S ribosome. 5S RNP (5S, RPL5, RPL11 complex) regulates the activity of MDM2, a key E3-ligase that suppresses p53 by ubiquitination, and also negatively regulates Rb. Dotted red line mechanism unknown
Numerous studies have demonstrated other post-translational modifications (e.g., sumoylation, phosphorylation, ubiquitination, methylation) on the DDX41 protein [46], but their roles in normal processes or oncogenesis remain unclear. Interestingly, missense mutations in myeloid malignancies have been identified on or near several of these (e.g., germline Y259C on BTK phosphorylation site, germline S363del adjacent to Y364 BTK phosphorylation site).
Germline mutations in DDX41 are also able to alter protein isoform usage of DDX41, as the germline mutation M1I [20] leads to a reduction in the full length 70 kDa isoform (p70) and a concomitant increased expression of a 55 kDa isoform (p55) initiating at M127 (our unpublished data and [36]) (Fig. 1). This shorter p55 isoform lacks the N-terminal nuclear localization signal (NLS) and is localized predominantly in the cytoplasm, unlike p70 which is mainly nuclear [20, 36]. Additionally, isoform p55 lacks both sites of TRIM21 ubiquitination and may be expected to have a longer half-life than the full-length protein. To date, DDX41 missense mutants tested (e.g., R525H and R164W), like WT DDX41, localize predominantly to the nucleus and, aside from M1I, express both p55 and p70 isoforms at similar levels to WT [20]. All reported premature termination mutations occur prior to the last exon. This would be predicted to reduce transcript levels via nonsense-mediated decay and hence levels of protein initiated from both p70- and p55-associated start codons derived from the mutant allele, although this is yet to be directly demonstrated.
Role of DDX41 in processes involved in the development of myeloid malignancies
DDX41 has been implicated by functional studies to be involved in at least 3 processes within the cell—mRNA splicing, innate immunity and ribosome biogenesis. These processes and how they could potentially contribute to DDX41-associated leukemogenesis are discussed below.
DDX41 in mRNA splicing
The family of DEAD-box helicases are key proteins in multiple diverse cellular pathways and processes [47]. In particular, DDX41 interacts with core splicing proteins. Binding studies in wild-type DDX41 over-expressing cells identified spliceosomal proteins as the most abundant group of interactors [19]. Of the spliceosomal proteins, numerous components of the spliceosome were found in complex with DDX41 including SF3B (SF3B1, SF3B2, SF3B3) and U2AF complexes (U2 complex), and the highly conserved PRPF8 scaffold protein and SNRNP200 (U5 complex) (Fig. 2a) [19]. The recurrent somatic DDX41 R525H mutant showed reduced binding to SF3B1 and PRPF8. Mutations in SF3B1 and PRPF8 have been seen in numerous studies of myeloid malignancies, particularly MDS [48, 49]. Somatic mutations in SF3B1 are frequently found in MDS and chronic lymphocytic leukemia (CLL) patients [50, 51]. Studies suggest that mutations in SF3B1 induce malignancy through aberrant transcription, altered pre-mRNA recognition and alternative splicing [50, 52]. Similarly, mutations in PRPF8 have been identified in MDS and AML patients where decreased PRPF8 expression is associated with increased exon skipping [53], possibly due to a splicing proofreading defect [53]. While DDX41 R525H is associated with reduced binding to SF3B1 and PRPF8, DDX41 mutations have not been reported in association with MDS with ring sideroblasts that is often characteristic of mutations in SF3B1 [50] and PRPF8 [53].
The identification of mutations in spliceosomal proteins is common in the development of myeloid malignancies [48, 54]. Analysis of mutations in 1043 sporadic myeloid malignancies identified 25 cases with DDX41 mutation—none of these had acquired mutations in commonly mutated spliceosomal proteins such as SF3B1, U2AF1, SRSF2, PRPF8 or ZRSR2, while only 2 cases harbored an LUC7L2 mutation [19]. As DDX41 mutations are largely mutually exclusive to splice factor mutations, and since splice factor mutations are relatively common in myeloid malignancies (2.4–11% for each gene in this cohort), it has been postulated that this points to an oncogenic role for mutant DDX41 via aberrant mRNA splicing [19] with the assumption that mutations in these splice factors impact on common downstream oncogenes, TSG and/or pathways.
Analysis of the influence of R525H on mRNA splicing revealed more avid exon skipping (smaller transcripts) and exon retention (larger transcripts) (Fig. 2a) within functionally relevant genes such as IKZF1, a transcription factor associated with lymphoid differentiation [55] and ZMYM2, a component of a histone deacetylase complex. Interestingly, low ZMYM2 mRNA levels are correlated with reduced expression of genes commonly mutated in myeloid malignancies such as RUNX1 and cohesins (SMC3, RAD21), as well as with low DDX41 mRNA levels [19]. Altered exon splicing may lead to decreased mRNA expression via frameshift-associated premature termination and nonsense-mediated decay or altered protein function by competition between isoforms or generation of defective proteins.
Exactly how DDX41 mutation-altered splicing drives leukemogenesis and the crucial target myeloid malignancy driver genes remains to be determined. Of interest also is whether different mutations (e.g., missense and premature termination) in different domains result in unique splicing patterns impacting on a different set of genes [54].
DDX41 in the STING–interferon pathway
Prior to its identification as a predisposition gene for HM, DDX41 was identified as a key component in the activation of the STING pathway against foreign exogenous DNA and invaders. DDX41 has been characterized as a pattern recognition receptor (PRR) against an array of dsDNA and cyclic nucleotides (CDN) produced from external pathogens or cellular sources (Fig. 2b). Some of these include bacteria [produce c-di-GMP and c-di-AMP (CDN)], and poly(dA; dT) [40], invading dsDNA viruses (e.g., adenovirus, herpes virus and papillomavirus), and nearby cells that have released dsDNA fragments by necrosis [44]. The introduction of dsDNA and CDN into the cytoplasm occurs through pathogen-specific infection mechanisms, exosomes or receptor-mediated endocytosis [56]. DDX41, in the context of binding foreign DNA (including cancer cell DNA fragments) and activation of the STING pathway and a type I interferon response, requires the binding of numerous factors via the DEAD-box domain extensively and other residues including those within the helicase C domain. Interestingly, some of the amino acids mutated in the predisposition to MDS and AML impact on several of these crucial residues or domains, implicating innate immunity in the oncogenic process.
The activation of DDX41 first requires the phosphorylation by Bruton’s tyrosine kinase (BTK). BTK phosphorylates DDX41 at the critical residue site, Y414 in the linker region between the DEAD-box domain and helicase C domain [37, 41] (Fig. 1), priming it for DNA sensing. The phosphorylated DDX41 then is able to bind to cytoplasmic foreign dsDNA via the DEAD-box domain [41]. The combination of phosphorylation by BTK and binding of dsDNA enables DDX41 to then bind to STING. Binding to STING occurs via the DEAD-box domain at Y364. DDX41 Y259 is another site of BTK phosphorylation, but its role remains unknown. This site has been identified in the Asian cohort as a residue of germline missense mutation (Y259C) in 7 families [27] (Fig. 1). Binding studies using tagged DDX41 and STING suggest that this site does not affect their affinity for each other [41]. Conversely, Y364 and Y414 are both crucial for efficient binding to STING and dsDNA.
The binding of DDX41 to STING activates and stimulates the STING pathway where host type I interferon response (IFNα and IFNβ) is initiated and drives multiple downstream pathways (Fig. 2b). These pathways include the activation of IRF3 and NF-κB, interferon regulators that trigger the inflammatory cytokine response within the cell [57]. Negative regulation of DDX41 occurs via ubiquitination by the E3-ubiquitin ligase, TRIM21, which after the STING pathway has been triggered, ubiquitinates DDX41 K9 and K115, targeting it for proteosomal degradation, terminating the innate immune response [45] (Fig. 2b).
DDX41 in post-transcriptional regulation of translation and ribosome biogenesis
A third proposed role for DDX41 lies in recent studies describing roles for post-transcriptional regulation of protein translation. Most recently, DDX41 was identified as a negative regulator of the CDKN1A (p21) protein translation via association with the 3′UTR of its mRNA. Knockdown or a helicase-dead mutation of DDX41 (G521S) led to loss of this activity and increased translation of transcripts linked to the 3′UTR. While it is not immediately clear how increased p21 protein levels in the scenario of DDX41 loss of activity fits with its proposed role as a myeloid tumor suppressor, data indicated that p53 mutation status of cells might also influence this regulation by an yet undefined mechanism, a detail that requires further investigation [42].
DDX41 has also been proposed to contribute to processing of precursor ribosomal RNA (pre-rRNA) to mature rRNAs in ribosome biogenesis [58] (Fig. 2c). Overexpression of the DDX41 R525H mutant led to decreased HSC (CD34+) growth in culture compared to WT, suggesting a possible mechanism for cytopenias seen in 3 patients with AML with this mutation [36]. Hypocellular bone marrow, consistent with perturbed HSC growth, is common in DDX41 mutant malignancies [19, 20]. Gene set enrichment analysis on DDX41 R525H-positive patient samples and R525H retroviral transduced cord blood implicated perturbation to ribosome biogenesis and pre-rRNA processing. Levels of pre-rRNAs 47S, 41S and 30S were increased and 21S decreased with increases of pre-rRNAs containing the 5′ external transcribed spacer (5′ETS) or internal transcribed spacer 2 (ITS2) compared to ITS1 containing pre-rRNAs [36] (Fig. 2c). Hence, DDX41 was implicated in the regulation of trimming of 5′ETS and ITS2, and R525H perturbed this. Trimmed pre-rRNAs are further processed into mature rRNAs 18S (small subunit) and 5.8S and 28S (large subunit) [36, 59]. Reduced assembly of ribosomal 60S subunits due to R525H resulted in increased levels of free unincorporated 5S RNP (complex of RPL5, RPL11, 5S rRNA that delivers 5S to the 60S subunit). Free 5S RNP preferentially binds to and sequesters the E3-ligase MDM2 [36], compromising its ability to ubiquitinate and negatively regulate the key tumor suppressor p53. However, the effect of DDX41 R525H on inhibition of the cell cycle was unexpectedly shown to be MDM2 and Rb dependent and not via classical p53 dependence. The mechanism by which MDM2 negatively regulates the tumor suppressor retinoblastoma protein (Rb) and its suppression of E2F, a transcription factor driver of cell cycle progression, is unclear. This complex set of interactions results in a decrease in expression of E2F, Rb and Myc target genes and cell cycling genes. DDX41 R525H acts in a dominant negative fashion where its loss of ATPase activity is crucial to the inhibitory effect on the cell cycle [36].
The overall outcome of DDX41 R525H presents as a ribosomopathy with inhibition of cell growth. Several other ribosomopathies predispose, albeit with weak penetrance, to MDS and AML including Shwachman–Diamond Syndrome [60], dyskeratosis congenita [61] and Diamond–Blackfan anemia [62]. The importance of the effect of DDX41 mutation via ribosome biogenesis on cell growth in the leukemogenic process remains to be determined.
Future handling and treatment of DDX41-associated myeloid malignancies
There are very few targeted treatment options available for AML and MDS. Induction chemotherapy, the mainstay of AML treatment, has changed very little for over 30 years, and in MDS, methyltransferase inhibitors and lenalidomide are approved in some cases [2, 3]. For both diseases, hematopoietic stem cell transplantation (HSCT) remains the treatment with the most curative potential; however, there are often contraindications that prevent implementation of this treatment option. For example, due to advanced age, many patients will not tolerate the rigorous conditioning required for HSCT, and additionally as many HSCT donors are sourced from within families, as discussed earlier, this approach presents a clear and present danger for individuals with germline predisposition and the risk of donor cell leukemia. For individuals with germline mutations in DDX41, understanding the different functional roles described in the previous section may lead to the application of targeted, better and less toxic treatments, and some current possibilities are discussed below.
Lenalidomide has become a recognized therapeutic molecule for treatment for MDS and multiple myeloma. The molecule was originally associated with its responsiveness against MDS patients with a chromosomal aberration of −5q [63], where 50% of patients showed cytogenetic remission [64]. DDX41 does not lie within the commonly deleted region of 5q [65]; however, recently it was shown that MDS patients with DDX41 mutations or low gene expression respond better to treatment with lenalidomide than MDS patients without DDX41 alteration [19]. The mechanism of this benefit is currently unknown. Several modes of action for lenalidomide have been proposed in MDS and it will be important to determine which if any of these are important for the response of DDX41 mutated malignancies [66].
Identification of a leukemogenic effect of mutant DDX41 via the STING–interferon pathway would open the way for the assessment of existing inhibitors and agonists in this pathway, as well as providing new targets for therapeutic development. There are already a number of molecules that interact with components of this pathway. For example, BTK, which phosphorylates and primes DDX41, interacts within multiple other signaling pathways including TOLL like receptor signaling, pre-B cell proliferation and signaling of chemokine receptors in B cell malignancies [67]. Accordingly, the use of a BTK inhibitor, ibrutinib, has been trialed for various lymphoid malignancies, dramatically changing management and outcome for individuals with chronic lymphocytic leukemia in particular [67, 68]. STING agonists are currently in the development phase. A STING agonist (DMXAA) potently activates the STING pathway demonstrating anti-tumor activity in mice [69]. However, due to a difference in one amino acid in the binding site between human and mouse, this molecule does not interact with human STING. The development of a human compatible DMXAA analog and other STING agonists are of great interest and in development [69, 70].
Although the exact mechanism by which DDX41 may alter mRNA splicing is still undefined, its interaction with spliceosomal proteins, including SF3B1, suggests that mutations in DDX41 may induce a deregulation of mRNA splicing. Mutations leading to deregulation of splicing are common in MDS and can lead to alterative splicing of particular genes, forming transcripts with modified domains and transcripts promoting neoplastic transformation [71]. Subsequently, potential treatment options for the deregulation of the spliceosome mechanism by DDX41 are those already associated with spliceosomal protein mutations. This includes pladienolide B, a polyketide suspected to target the binding of U2 snRNP/SF3B complex to the 3′ pre-mRNA branch point [71], preventing further formation of the spliceosome complex. Analysis of splicing in CLL cells treated with pladienolide B identified an increase in intron retention and expression of pro-apoptotic isoforms of apoptosis-related genes. Importantly, this was only observed in CLL cells and not normal blood [72]. Unfortunately, first in man, clinical trials of E7107, a derivative of pladienolide B, in solid tumors were halted due to bilateral optic neuritis and gastrointestinal toxicity [71]. A second splicing modulator of interest is H3B-8800, which selectively targets SF3B1 and leads to preferential apoptosis in SF3B1 mutant cells is currently in clinical trials for MDS (NCT02841540) [71]. Given the characterized interaction of DDX41 and SF3B1, it will be of interest to determine if DDX41 mutant cells are also sensitive to H3B-8800 activity.
Conclusions
The identification of germline DDX41 mutations in predisposition to myeloid malignancies has altered our understanding of FHM predisposition in terms of both the context in which it is found, and the potential mechanisms by which this may occur. Further research is required to determine the pathways or processes by which DDX41 mutation contributes to the oncogenic process leading to myeloid malignancies to ensure that efficacious treatments, monitoring and prevention methods can be employed for the hundreds of families affected worldwide.
References
Arber DA, Orazi A, Hasserjian R, Thiele J, Borowitz MJ, Le Beau MM, et al. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood. 2016;127(20):2391–405.
Steensma DP. Myelodysplastic Syndromes: diagnosis and treatment. Mayo Clin Proc. 2015;90(7):969–83.
Khwaja A, Bjorkholm M, Gale RE, Levine RL, Jordan CT, Ehninger G, et al. Acute myeloid leukaemia. Nat Rev Dis Primers. 2016;2:16010.
VidebæK A. Familial Leukemia1. Acta Med Scand. 1947;127(1–2):26–52.
Song WJ, Sullivan MG, Legare RD, Hutchings S, Tan X, Kufrin D, et al. Haploinsufficiency of CBFA2 causes familial thrombocytopenia with propensity to develop acute myelogenous leukaemia. Nat Genet. 1999;23(2):166–75.
Hahn CN, Chong CE, Carmichael CL, Wilkins EJ, Brautigan PJ, Li XC, et al. Heritable GATA2 mutations associated with familial myelodysplastic syndrome and acute myeloid leukemia. Nat Genet. 2011;43(10):1012–7.
Hsu AP, Sampaio EP, Khan J, Calvo KR, Lemieux JE, Patel SY, et al. Mutations in GATA2 are associated with the autosomal dominant and sporadic monocytopenia and mycobacterial infection (MonoMAC) syndrome. Blood. 2011;118(10):2653–5.
Marquez R, Hantel A, Lorenz R, Neistadt B, Wong J, Churpek JE, et al. A new family with a germline ANKRD26 mutation and predisposition to myeloid malignancies. Leuk Lymphoma. 2014;55(12):2945–6.
Topka S, Vijai J, Walsh MF, Jacobs L, Maria A, Villano D, et al. Germline ETV6 mutations confer susceptibility to acute lymphoblastic leukemia and thrombocytopenia. PLoS Genet. 2015;11(6):e1005262.
Smith ML, Cavenagh JD, Lister TA, Fitzgibbon J. Mutation of CEBPA in familial acute myeloid leukemia. N Engl J Med. 2004;351(23):2403–7.
Harutyunyan AS, Giambruno R, Krendl C, Stukalov A, Klampfl T, Berg T, et al. Germline RBBP6 mutations in familial myeloproliferative neoplasms. Blood. 2016;127(3):362–5.
Kirwan M, Vulliamy T, Marrone A, Walne AJ, Beswick R, Hillmen P, et al. Defining the pathogenic role of telomerase mutations in myelodysplastic syndrome and acute myeloid leukemia. Hum Mutat. 2009;30(11):1567–73.
Brown AL, Churpek JE, Malcovati L, Döhner H, Godley LA. Recognition of familial myeloid neoplasia in adults. Semin Hematol. 2017. (In Press). doi:10.1053/j.seminhematol.2016.11.003.
Narumi S, Amano N, Ishii T, Katsumata N, Muroya K, Adachi M, et al. SAMD9 mutations cause a novel multisystem disorder, MIRAGE syndrome, and are associated with loss of chromosome 7. Nat Genet. 2016;48(7):792–7.
Chen DH, Below JE, Shimamura A, Keel SB, Matsushita M, Wolff J, et al. Ataxia-pancytopenia syndrome is caused by missense mutations in SAMD9L. Am J Hum Genet. 2016;98(6):1146–58.
Tesi B, Davidsson J, Voss M, Rahikkala E, Holmes TD, Chiang SC, et al. Gain-of-function SAMD9L mutations cause a syndrome of cytopenia, immunodeficiency, MDS and neurological symptoms. Blood. 2017;129(16)2266–79. doi:10.1182/blood-2016-10-743302.
Michaud J, Wu F, Osato M, Cottles GM, Yanagida M, Asou N, et al. In vitro analyses of known and novel RUNX1/AML1 mutations in dominant familial platelet disorder with predisposition to acute myelogenous leukemia: implications for mechanisms of pathogenesis. Blood. 2002;99(4):1364–72.
Linder P. Dead-box proteins: a family affair—active and passive players in RNP-remodeling. Nucl Acids Res. 2006;34(15):4168–80.
Polprasert C, Schulze I, Sekeres MA, Makishima H, Przychodzen B, Hosono N, et al. Inherited and somatic defects in DDX41 in myeloid neoplasms. Cancer Cell. 2015;27(5):658–70.
Lewinsohn M, Brown AL, Weinel LM, Phung C, Rafidi G, Lee MK, et al. Novel germ line DDX41 mutations define families with a lower age of MDS/AML onset and lymphoid malignancies. Blood. 2016;127(8):1017–23.
Li R, Sobreira N, Witmer PD, Pratz KW, Braunstein EM. Two novel germline DDX41 mutations in a family with inherited myelodysplasia/acute myeloid leukemia. Haematologica. 2016;101(6):e228–31.
Cardoso SR, Ryan G, Walne AJ, Ellison A, Lowe R, Tummala H, et al. Germline heterozygous DDX41 variants in a subset of familial myelodysplasia and acute myeloid leukemia. Leukemia. 2016;30(10):2083–6.
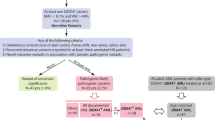
DiNardo CD, Bannon SA, Routbort M, Franklin A, Mork M, Armanios M, et al. Evaluation of patients and families with concern for predispositions to hematologic malignancies within the hereditary hematologic malignancy clinic (HHMC). Clin Lymphoma Myeloma Leuk. 2016;16(7):417–428 e412.
Berger G, van den Berg E, Sikkema-Raddatz B, Abbott KM, Sinke RJ, Bungener LB, et al. Re-emergence of acute myeloid leukemia in donor cells following allogeneic transplantation in a family with a germline DDX41 mutation. Leukemia. 2017;31(2):520–2.
Ripperger T, Steinemann D, Gohring G, Finke J, Niemeyer CM, Strahm B, et al. A novel pedigree with heterozygous germline RUNX1 mutation causing familial MDS-related AML: can these families serve as a multistep model for leukemic transformation? Leukemia. 2009;23(7):1364–6.
Greif PA, Konstandin NP, Metzeler KH, Herold T, Pasalic Z, Ksienzyk B, et al. RUNX1 mutations in cytogenetically normal acute myeloid leukemia are associated with a poor prognosis and up-regulation of lymphoid genes. Haematologica. 2012;97(12):1909–15.
Takeda J, Yoshida K, Makishima H, Yoshizato T, Shiozawa Y, Shiraishi Y, et al. Genetic predispositions to myeloid neoplasms caused by germline DDX41 mutations. Blood. 2015;126(23):2843. (Abstract 2843).
Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E, Fennell T, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016;536(7616):285–91.
Churpek JE, Pyrtel K, Kanchi KL, Shao J, Koboldt D, Miller CA, et al. Genomic analysis of germ line and somatic variants in familial myelodysplasia/acute myeloid leukemia. Blood. 2015;126(22):2484–90.
Babushok DV, Bessler M, Olson TS. Genetic predisposition to myelodysplastic syndrome and acute myeloid leukemia in children and young adults. Leuk Lymphoma. 2016;57(3):520–36.
Owen CJ, Toze CL, Koochin A, Forrest DL, Smith CA, Stevens JM, et al. Five new pedigrees with inherited RUNX1 mutations causing familial platelet disorder with propensity to myeloid malignancy. Blood. 2008;112(12):4639–45.
Buijs A, Poddighe P, van Wijk R, van Solinge W, Borst E, Verdonck L, et al. A novel CBFA2 single-nucleotide mutation in familial platelet disorder with propensity to develop myeloid malignancies. Blood. 2001;98(9):2856–8.
Kobayashi S, Kobayashi A, Osawa Y, Nagao S, Takano K, Okada Y, et al. Donor cell leukemia arising from preleukemic clones with a novel germline DDX41 mutation after allogenic hematopoietic stem cell transplantation. Leukemia. 2017;31(4):1020–2.
Link DC, Walter MJ. ‘CHIP’ping away at clonal hematopoiesis. Leukemia. 2016;30(8):1633–5.
Koeffler HP, Leong G. Preleukemia: one name, many meanings. Leukemia. 2017;31(3):534–42.
Kadono M, Kanai A, Nagamachi A, Shinriki S, Kawata J, Iwato K, et al. Biological implications of somatic DDX41 p.R525H mutation in acute myeloid leukemia. Exp Hematol. 2016;44(8):745–754 e744.
Jiang Y, Zhu Y, Qiu W, Liu YJ, Cheng G, Liu ZJ, et al. Structural and functional analyses of human DDX41 DEAD domain. Protein Cell. 2017;8(1):72–6.
Putnam AA, Jankowsky E. DEAD-box helicases as integrators of RNA, nucleotide and protein binding. Biochim Biophys Acta. 2013;1829(8):884–93.
Umate P, Tuteja N, Tuteja R. Genome-wide comprehensive analysis of human helicases. Commun Integr Biol. 2011;4(1):118–37.
Parvatiyar K, Zhang Z, Teles RM, Ouyang S, Jiang Y, Iyer SS, et al. The helicase DDX41 recognizes the bacterial secondary messengers cyclic di-GMP and cyclic di-AMP to activate a type I interferon immune response. Nat Immunol. 2012;13(12):1155–61.
Lee KG, Kim SS, Kui L, Voon DC, Mauduit M, Bist P, et al. Bruton’s tyrosine kinase phosphorylates DDX41 and activates its binding of dsDNA and STING to initiate type 1 interferon response. Cell Rep. 2015;10(7):1055–65.
Peters D, Radine C, Reese A, Budach W, Sohn D, Janicke RU. The DEAD-Box RNA helicase DDX41 is a novel repressor of p21WAF1/CIP1 mRNA translation. J Biol Chem. 2017;292(20)8331–41. doi:10.1074/jbc.M116.772327.
Omura H, Oikawa D, Nakane T, Kato M, Ishii R, Ishitani R, et al. Structural and Functional Analysis of DDX41: a bispecific immune receptor for DNA and cyclic dinucleotide. Sci Rep. 2016;6:34756.
Zhang Z, Yuan B, Bao M, Lu N, Kim T, Liu YJ. The helicase DDX41 senses intracellular DNA mediated by the adaptor STING in dendritic cells. Nat Immunol. 2011;12(10):959–65.
Zhang Z, Bao M, Lu N, Weng L, Yuan B, Liu YJ. The E3 ubiquitin ligase TRIM21 negatively regulates the innate immune response to intracellular double-stranded DNA. Nat Immunol. 2013;14(2):172–8.
PhosphoSitePlus. PhosphoSitePlus. [cited 2017 17th April]. http://www.phosphosite.org/proteinAction.action?id=6000&showAllSites=true.
Linder P, Jankowsky E. From unwinding to clamping—the DEAD box RNA helicase family. Nat Rev Mol Cell Biol. 2011;12(8):505–16.
Yoshida K, Sanada M, Shiraishi Y, Nowak D, Nagata Y, Yamamoto R, et al. Frequent pathway mutations of splicing machinery in myelodysplasia. Nature. 2011;478(7367):64–9.
Thol F, Kade S, Schlarmann C, Loffeld P, Morgan M, Krauter J, et al. Frequency and prognostic impact of mutations in SRSF2, U2AF1, and ZRSR2 in patients with myelodysplastic syndromes. Blood. 2012;119(15):3578–84.
Cazzola M, Rossi M, Malcovati L. Associazione Italiana per la Ricerca sul Cancro Gruppo Italiano Malattie M. Biologic and clinical significance of somatic mutations of SF3B1 in myeloid and lymphoid neoplasms. Blood. 2013;121(2):260–9.
Hahn CN, Venugopal P, Scott HS, Hiwase DK. Splice factor mutations and alternative splicing as drivers of hematopoietic malignancy. Immunol Rev. 2015;263(1):257–78.
Jenkins JL, Kielkopf CL. Splicing factor mutations in myelodysplasias: insights from spliceosome structures. Trends Genet. 2017;33(5):336–48.
Kurtovic-Kozaric A, Przychodzen B, Singh J, Konarska MM, Clemente MJ, Otrock ZK, et al. PRPF8 defects cause missplicing in myeloid malignancies. Leukemia. 2015;29(1):126–36.
Maciejewski JP, Padgett RA. Defects in spliceosomal machinery: a new pathway of leukaemogenesis. Br J Haematol. 2012;158(2):165–73.
de Rooij JD, Beuling E, van den Heuvel-Eibrink MM, Obulkasim A, Baruchel A, Trka J, et al. Recurrent deletions of IKZF1 in pediatric acute myeloid leukemia. Haematologica. 2015;100(9):1151–9.
Junt T, Barchet W. Translating nucleic acid-sensing pathways into therapies. Nat Rev Immunol. 2015;15(9):529–44.
Barber GN. STING: infection, inflammation and cancer. Nat Rev Immunol. 2015;15(12):760–70.
Jiang Y, Zhu Y, Liu ZJ, Ouyang S. The emerging roles of the DDX41 protein in immunity and diseases. Protein Cell. 2017;8(2):83–9.
Tafforeau L, Zorbas C, Langhendries JL, Mullineux ST, Stamatopoulou V, Mullier R, et al. The complexity of human ribosome biogenesis revealed by systematic nucleolar screening of Pre-rRNA processing factors. Mol Cell. 2013;51(4):539–51.
Donadieu J, Michel G, Merlin E, Bordigoni P, Monteux B, Beaupain B, et al. Hematopoietic stem cell transplantation for Shwachman-Diamond syndrome: experience of the French neutropenia registry. Bone Marrow Transpl. 2005;36(9):787–92.
Alter BP, Giri N, Savage SA, Rosenberg PS. Cancer in dyskeratosis congenita. Blood. 2009;113(26):6549–57.
Narla A, Ebert BL. Ribosomopathies: human disorders of ribosome dysfunction. Blood. 2010;115(16):3196–205.
List A, Dewald G, Bennett J, Giagounidis A, Raza A, Feldman E, et al. Lenalidomide in the myelodysplastic syndrome with chromosome 5q deletion. N Engl J Med. 2006;355(14):1456–65.
Fenaux P, Giagounidis A, Selleslag D, Beyne-Rauzy O, Mufti G, Mittelman M, et al. A randomized phase 3 study of lenalidomide versus placebo in RBC transfusion-dependent patients with Low-/Intermediate-1-risk myelodysplastic syndromes with del5q. Blood. 2011;118(14):3765–76.
Hosono N, Makishima H, Mahfouz R, Przychodzen B, Yoshida K, Jerez A, et al. Recurrent genetic defects on chromosome 5q in myeloid neoplasms. Oncotarget. 2017;8(4):6483–95.
Abou Zahr A, Saad Aldin E, Komrokji RS, Zeidan AM. Clinical utility of lenalidomide in the treatment of myelodysplastic syndromes. J Blood Med. 2015;6:1–16.
Hendriks RW, Yuvaraj S, Kil LP. Targeting Bruton’s tyrosine kinase in B cell malignancies. Nat Rev Cancer. 2014;14(4):219–32.
Woyach JA. How I manage ibrutinib-refractory chronic lymphocytic leukemia. Blood. 2017;129(10):1270–4.
Corrales L, Gajewski TF. Endogenous and pharmacologic targeting of the STING pathway in cancer immunotherapy. Cytokine. 2016;77:245–7.
Gao P, Zillinger T, Wang W, Ascano M, Dai P, Hartmann G, et al. Binding-pocket and lid-region substitutions render human STING sensitive to the species-specific drug DMXAA. Cell Rep. 2014;8(6):1668–76.
Brierley CK, Steensma DP. Targeting splicing in the treatment of myelodysplastic syndromes and other myeloid neoplasms. Curr Hematol Malig Rep. 2016;11(6):408–15.
Kashyap MK, Kumar D, Villa R, La Clair JJ, Benner C, Sasik R, et al. Targeting the spliceosome in chronic lymphocytic leukemia with the macrolides FD-895 and pladienolide-B. Haematologica. 2015;100(7):945–54.
Acknowledgements
The authors thank the patients and their family members for their willingness to participate in this research. This study was supported by grants APP1086302 (C.N.H., H.S.S., A.L.B), APP1024215 (C.N.H., H.S.S) and APP1023059 principal research fellowship (H.S.S) from the National Health and Medical Research Council of Australia; APP1125849 from the Cancer Council of South Australia (C.N.H., H.S.S., A.L.B); and the University of South Australia Postgraduate Research Award (J.J.C.C).
Author information
Authors and Affiliations
Corresponding authors
About this article
Cite this article
Cheah, J.J.C., Hahn, C.N., Hiwase, D.K. et al. Myeloid neoplasms with germline DDX41 mutation. Int J Hematol 106, 163–174 (2017). https://doi.org/10.1007/s12185-017-2260-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12185-017-2260-y