Abstract
Purpose of Review
Diabetes mellitus (DM) due to toxic misfolding of proinsulin variants provides a monogenic model of endoplasmic reticulum (ER) stress. The mutant proinsulin syndrome (also designated MIDY; Mutant INS-gene-induced Diabetes of Youth or Maturity-onset diabetes of the young 10 (MODY10)) ordinarily presents as permanent neonatal-onset DM, but specific amino-acid substitutions may also present later in childhood or adolescence. This review highlights structural mechanisms of proinsulin folding as inferred from phenotype-genotype relationships.
Recent Findings
MIDY mutations most commonly add or remove a cysteine, leading to a variant polypeptide containing an odd number of thiol groups. Such variants are associated with aberrant intermolecular disulfide pairing, ER stress, and neonatal β-cell dysfunction. Non-cysteine-related (NCR) mutations (occurring in both the B and A domains of proinsulin) define distinct determinants of foldability and vary in severity. The range of ages of onset, therefore, reflects a “molecular rheostat” connecting protein biophysics to quality-control ER checkpoints. Because in most mammalian cell lines even wild-type proinsulin exhibits limited folding efficiency, molecular barriers to folding uncovered by NCR MIDY mutations may pertain to β-cell dysfunction in non-syndromic type 2 DM due to INS-gene overexpression in the face of peripheral insulin resistance.
Summary
Recent studies of MIDY mutations and related NCR variants, combining molecular and cell-based approaches, suggest that proinsulin has evolved at the edge of non-foldability. Chemical protein synthesis promises to enable comparative studies of “non-foldable” proinsulin variants to define key steps in wild-type biosynthesis. Such studies may create opportunities for novel therapeutic approaches to non-syndromic type 2 DM.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Proteotoxic diseases arise due to misfolding within or external to cells. Extracellular amyloid, for example, is a feature of diverse diseases (Table 1A) [1]. Such β-sheet-rich pathological deposits [2, 3] arise from misfolding of globular proteins, such as immunoglobulin light chains or β2-microglobulin in association with hematologic malignancies [4]; mutations in various other proteins can predispose to toxic deposition as exemplified by unstable variants of serpins, transthyretin, and lysozyme [5–7]. In neurodegenerative diseases, perineuronal plaque reflects aggregation-coupled misfolding of intrinsically disordered polypeptides [8, 9]. Principles of amyloidogenesis have recently been reviewed [10]. Of complementary importance is intracellular proteotoxicity (Table 1B). Its pathophysiologic importance has motivated studies of a series of foundational mechanisms, including nascent folding, quality control, trafficking, and degradation [11]: a dynamic regulatory network collectively designated proteostasis [12]. Distinctive protein inclusions within cells are histopathological hallmarks of neurodegenerative diseases, as exemplified by huntingtin aggregation in neuronal nuclei (Huntington’s disease [13]) and tau-related cytoplasmic neurofibrillary tangles (Alzheimer’s disease [14]).
This review highlights a monogenic form of diabetes mellitus (DM)Footnote 1—the mutant proinsulin syndrome [15••, 16••]—due to nascent protein misfolding in the endoplasmic reticulum (ER). Although monogenic DM syndromes encompass a variety of genes and molecular mechanisms, mutations in the insulin gene (INS) are of particular interest in relation to the pathway of insulin biosynthesis [17, 18]. The mutant proinsulin syndrome, also designated Mutant INS-gene-induced Diabetes of Youth (MIDY) (also known as Maturity-onset diabetes of the young 10 (MODY10) [19, 20]), is caused by toxic misfolding of variant proinsulins [15••, 16••] (for review, see [21, 22] (Table 2); the human syndrome was anticipated by the Akita mouse model [23, 24]. Clinical mutations (genetically dominant) impair secretion of both variant and wild-type insulin in trans; misfolding and aggregation activate the unfolded-protein response (UPR) and induce ER stress, leading in turn to β-cell dysfunction and death [25, 26] (for reviews, see [27•]). Here, we delineate structural mechanisms by which MIDY mutations impair the folding efficiency of proinsulin. Biophysical principles underlying ER-related proteotoxicity in this syndrome promise to provide general insight into a broad class of proteotoxic diseases [28].
Monogenic Diabetes and Proinsulin Syndrome
Monogenic DM can arise due to mutations in genes encoding key transcription factors, subunits of the β-cell potassium channel, the β-cell glucose-sensor glucokinase, or insulin itself [29, 30]. Collectively, such syndromes comprise 1–5% of DM [31]. The spectrum of phenotypes ranges from transient or permanent neonatal-onset DM (tNDM and pNDM) to MODY [32]. Whereas NDM presents within the first 6 months, MODY ordinarily has an onset between 10 and 25 years of age. Heterozygous INS mutations constitute the second most common cause of monogenic DM (after potassium channel mutations [33, 34]). Genotype–phenotype correlations in the mutant proinsulin syndrome suggest that ages of onset reflect mutational severity. Mutation-specific phenotypes are general features of other genetic diseases, such as partial or complete androgen-insensitivity syndrome and cystic fibrosis, among several others [35–40]. In addition to mutation-specific effects, clinical differences in penetrance, disease severities, or ages of onset may be influenced by modifier genes or environment as observed in other endocrine syndromes [41], including type 1 DM [42].
Biosynthesis of Insulin
The INS-gene encodes preproinsulin, a single-chain precursor polypeptide with a signal peptide-B-C-A domain N-to-C organization [18]. The signal peptide is cleaved co-translationally on ER translocation. Folding within the ER accompanies a specific pairing of three disulfide bridges. Processing of proinsulin by prohormone convertases PC1/3 and PC2 generates the two-chain hormone in glucose-regulated secretory vesicles [17]. The mature hormone’s two cystines link the A and B chains (A7-B7, B19-A20) whereas one is within the A chain (A6-A11) [43]; these are each required for stability and activity [44, 45]. Mispairing of disulfides in vitro leads to reduced stability and activity [46, 47]. The solution structure of proinsulin (as an engineered monomer) contains a native-like insulin core (51 residues) with a flexible C domain (35 residues) [48]. Whereas clinical INS mutations primarily affect nascent folding in the ER, specific mutations have been identified that selectively perturb protein trafficking, prohormone processing, and receptor binding [49, 50•].
ER Quality Control
Chemical trapping studies of insulin-related precursor polypeptides in vitro have demonstrated accumulation of one- and two-disulfide intermediates, thus providing evidence for a hierarchical disulfide pathway [51, 52]. Together, these studies suggested the initial formation of cystine B19-A20 along with hydrophobic clustering by C-terminal α-helix and central B-chain α-helix. Such a native-like structure, recapitulated in a one-disulfide peptide model [44, 45], defines a specific folding nucleus [53]. Cellular folding of proinsulin and disulfide analogs has been extensively investigated by Arvan, Kaufman, and their respective colleagues in relation to the ER oxidative-folding machinery (quality control, stress, quality control, and exit; [54] (for review, see [55]). Pairwise substitution of cysteines enabled the respective contributions of each disulfide bridge to be evaluated [54]. The results highlighted the importance of cystines [A7-B7] and [B19-A20] (but not [A6-A11]) for efficient ER export and eventual secretion. Evidence was obtained that an unpaired thiol group at CysA11 underlies the proteotoxicity of SerA6-murine proinsulin (Ins2-Munich [56]). The particularly deleterious role of a single cysteine at A11 was thus highlighted, as CysA11 can mispair with three other cysteines CysB19, CysA20, and CysB7 in the same molecule or mediate aberrant intermolecular cross-linking [54].
Molecular Rheostat of Foldability
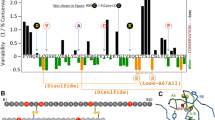
Whereas most MIDY mutations entail either loss or addition of Cys, non-Cys-related mutations highlight key determinants of foldability [50•]. Many such mutations cluster near the critical [B19-A20] disulfide bridge, particularly in the B9-B19 or A16-A19 helices. These are of biophysical interest as the variant polypeptide retains the six canonical Cys residues: impaired disulfide pairing presumably reflects general biophysical principles that underlie protein folding, structure, and stability [57, 58]. Prominent among these are (i) the efficiency of side-chain packing in a hydrophobic core [59] and (ii) the intrinsic secondary-structural propensities of the amino acids [60]. Large-to-small mutations [61], for example, can introduce destabilizing cavities in the native state [62] and by extension in a native-like specific folding nucleus [44]. Within helices, the substitution of a residue of high helical propensity by one of lower helical propensity can likewise impair stability [63, 64]. We describe in turn below clinical mutations that exemplify these principles. We chose the following subset of NCR mutations based on (a) their positioning within or near proinsulin’s specific folding nucleus [53] (Fig. 1) and (b) illustrative biophysical mechanisms of impaired foldability. A foundational structural model is provided by the crystallographic T-state monomer (PDB entry: 4INS) [43], as recapitulated in the insulin-like core of a proinsulin monomer [48].
-
(i) The side chain of LeuB6 inserts into an interchain cavity surrounded by the invariant side chains of LeuB11, LeuB15, and LeuA16 (Fig. 1). At this site, a variety of non-conservative mutations (Arg, Gln, Pro, and Val) lead to neonatal-onset DM. Each would be expected to introduce profound structural perturbations. In contrast, MODY substitution MetB6 is presumably associated with only subtle changes in packing.
-
(ii) LeuB11 contributes to segmental α-helical propensity and nascent clustering of nonpolar residues. The side chain is buried within a cavity abutting the nonpolar inner surface of the A chain. Clinical mutations are Pro or Gln, each expected to impede initial [B19-A20] disulfide pairing: ProB11 would profoundly perturb α-helical propensity, stability, and self-assembly. GlnB11 would fit within the B11-related cavity, but its carboxamide group would impose an electrostatic penalty.
-
(iii) The side chain of LeuB15 packs within a nonpolar crevice delimited by CysB19 and PheB24. Clinical mutations at B15 are Pro, His, and Val (neonatal in each case). Like ProB11 (above), ProB15 would be expected to introduce marked perturbations. Another neonatal mutation at this position (His) would insert a polar aromatic side chain into the nonpolar hydrophobic pocket, thus destabilizing the core. The β-branched side chain of ValB15 would by contrast be associated with more subtle effects due to its lower α-helical propensity and smaller volume, relative to Leu.
-
(iv) ValB18 adjoins CysB19 near the end of the central B-chain α-helix. Clinical mutations are Gly (neonatal) and Ala (MODY). Each would impair the efficiency of core packing near cystine [B19-A20] in a solvent-exposed interchain crevice. Substitution by Gly (a residue of similarly low helical propensity as Val) would create a cavity and enhance main-chain flexibility, presumably interfering with nascent [B19-A20] pairing. Interestingly the extent of these perturbations is different for Gly and Ala in terms of the severity of onset. Ala is predicted to exhibit offsetting biophysical effects: greater helical propensity but impaired packing efficiency.
-
(v) Three neonatal-onset MIDY mutations have recently been found in the A domain (ProA16, AspA19 and AsnA19) [50•, 65]. The side chain of LeuA16 is buried within the core (Fig. 1). ProA16 would perturb the segmental main-chain conformation and introduce a destabilizing cavity [66•]. TyrA19 projects from a nonpolar crevice (lined in part by cystine [B19-A20]) to expose its para-hydroxyl group; AspA19 would place a destabilizing negative charge within the core. Similarly, AsnA19 would impede the foldability by projecting the carboxamide group into the nonpolar core.
Structure and sites of clinical mutations in insulin. Ribbon model of insulin monomer showing the core residues (PDB entry 4INS [43]). Sulfur atoms in A6-A11 and B19-A20 disulfides are shown as gold spheres and A7-B7 as sticks. Other side chains are shown in dark blue (near A6-A11 cystine) or light blue (near B19-A20 cystine); residues TyrA19 and HisB5 that are at near core residue and also sites of clinical mutation are shown in magenta. All other side chains are shown in light gray (A chain) or dark gray (B chain). Right side panel shows the view rotated vertically by 90°
Position A16 has long been of interest in relation to the structure, foldability, and function of insulin [43, 67•, 68••]. Invariant within an extended vertebrate family (insulin and insulin-like growth factors [IGF-I, II]) and also among most relaxins/insulin-like peptides (ILPs) [69]), the side chain of LeuA16 is buried in the core in both free and receptor-bound states [70–72]. Packing of LeuA16 efficiently fills a potential cavity delimited by conserved nonpolar receptor-binding elements (LeuB15, IleA2, and TyrA19) and girded by cystines [A6-A11] and [B19-A20] [43, 70]. Such a “left-over space” (akin to Gould’s celebrated evolutionary metaphor of the spandrels of the San Marco cathedral in Venice [73]) rationalizes the exquisite sensitivity of insulin chain combination to A16 substitutions [67•]. LeuA16 is invariant as an “exaptation,” the only side chain able to fit in this space otherwise peripheral to the mechanism of receptor binding. Indeed, substitution of LeuA16 by Val—although rendering chain combination yield negligible and impairing the folding of proinsulin—is nonetheless compatible with native structure and function [68••]—once the folded state has been reached. Remarkably, ValA16 has recently been found in an infant in Saudi Arabia as a recessive MIDY mutation [50•] (E. De Franco, personal communication), to our knowledge the first instance of a point mutation with recessive inheritance. Additional recessive mutations may occur among MIDY patients, but lack of family history could obscure their identification (a general issue in human genetics; for review in monogenic diabetes syndromes, see [74]); ValA16 provides a prototype recessive mutation in a society notable for consanguinity [75]. It is noteworthy that detailed analysis of structure, foldability, and function of ValA16-insulin and ValA16-proinsulin [68••] preceded its clinical description [50•].
In the initial steps of proinsulin folding, the side chains of B11, B15, B18, A16, and A19 are proposed to collapse to form a specific folding nucleus guiding pairing of CysB19 and CysA20 [44, 51, 52]. Together, the above analysis supports a broad hypothesis that the variable age of onset of MIDY-related DM—and perhaps the mode of inheritance, dominant, or recession—is intrinsic to the biophysical properties of the mutations (as distinct from environmental effects or the influence of potential modifier genes as pertinent to the onset of Type 1 DM [42]). This hypothesis presumably extends to the collection of MIDY mutations as a whole and is not restricted to the above subset of substitutions.
We anticipate that one or another MIDY mutation may primarily impair pairing of any one of proinsulin’s three disulfide bridges. However, not all structural elements of a protein’s native state contribute to its folding nucleus or subsequent steps in oxidative protein folding. Like Sherlock Holmes’ famous clue: “the dog that did not bark in the night-time” [76], the absence of clinical mutations in such an element may be as informative as the presence of mutations in other elements.Footnote 2 An example is provided by proinsulin’s flexible C domain: although mutations at the dibasic cleavage sites can lead to secretion of split proinsulins [49], lack of non-Cys-related mutations in this segment implies that disulfide pairing is robust to such substitutions in accordance with both the C domain’s evolutionary variability in sequence and length and its diversification among chordate insulin-like growth factors [77].
Conclusions
The discovery of insulin in Toronto in 1921, followed the next year by its first clinical use, represents a landmark in the history of molecular medicine [78]. However transformational, the work of Banting, Best, Collip, and Macleod provided only the starting point for generations of seminal basic and translational investigations: the ensuing century of discovery is the subject of recent commemoration and review [79]. Identification of the mutant proinsulin syndrome in this century [50•] has brought together long-standing themes in diabetes research—hormone biosynthesis and structure—with foundational paradigms in human genetics, cell biology, and protein biophysics [22, 50•, 55].
NCR mutations in proinsulin associated with toxic misfolding in principle define key determinants of foldability, providing insight into how specific disulfide pairing is specified by the wild-type protein sequence [58]. Although structural studies have encountered an experimental “Catch-22” (i.e., confounded impaired folding efficiency), we anticipate that frontier synthetic methods [80, 81] may circumvent this critical barrier to provide tractable models [82, 83]. Such synthetic advances promise to enable our overarching hypothesis—that the variable age of onset among MIDY patients is due to mutation-specific biophysical mechanisms—to be rigorously tested. Furthermore, such biophysical insights may enable molecular interpretation of pathophysiologic events in the stressed ER that contribute to trans-interference with wild-type proinsulin biosynthesis and impaired glucose-stimulated insulin secretion [66•]. Key questions include how wild-type and variant folding intermediates self-associate in the ER and in turn how such aggregates block trafficking to the Golgi apparatus [55, 84, 85].
The broader significance of the mutant proinsulin syndrome pertains to non-syndromic type 2 DM. Under conditions of peripheral insulin resistance leading to INS overexpression, misfolding of even wild-type proinsulin can activate the unfolded-protein response and induce ER stress [86]. We envision that β-cell dysfunction caused by mutations in proinsulin may recapitulate, in accelerated form, the natural history of type 2 DM [27•, 83, 87]. Accordingly, studies of such variants in β-cell lines, isolated islets, and engineered mouse models promise to provide broad insights into the pathogenesis of a pandemic disease [27•]. Such models may also enable development of novel therapeutic approaches which focuses on reducing β-cell ER stress elicited by the misfolding of wild-type proinsulin [88•]. This prospect exemplifies a general paradigm in pharmacology whereby rare monogenic syndromes can open doors to innovative drug discovery [89]. It is fitting that such opportunities have arisen at the cusp of insulin’s second century [79, 90].
Note Added In Proof
Classification of clinical mutations in proinsulin based on structural mechanisms of disulfide pairing may be obtained based on equilibrium peptide models of oxidative folding intermediates [108, 109].
Notes
Abbreviations. DM, diabetes mellitus; ER, endoplasmic reticulum; MIDY, mutant INS-gene-induced Diabetes of Youth; MODY, maturity-onset diabetes of the young; NCR, non-cysteine-related; PND, permanent neonatal-onset DM; and UPR, unfolded-protein response. Residues are designated by standard three-letter code. Residue positions in insulin are shown in superscript (chain and residue number); Leu at position 15 of the B chain, for example, is denoted LeuB15. Cystine pairings are identified by brackets; the disulfide pairing between CysB19 and CysA20, for example, is [B19-A20]. Gene names are italicized.
Although insulin chain combination is in general robust to mutations in the A1-A8 α-helix [91, 92], MODY variant GluA4Lys lies on the surface of this helix. Its effect on the foldability of proinsulin may be due to disruption of a salt bridge with Arg89 (in the dibasic CA junction) in a proinsulin folding intermediate; in the solution structure of a proinsulin monomer this salt bridge appears to provide an N-cap of A1-A8 α-helix [48]. LysA4 could introduce electrostatic repulsion within this element and so attenuate nascent helix formation. A structural puzzle is posed by neonatal-onset MIDY mutation ThrA8Ser [50], also on the surface of insulin.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Dobson CM. Protein misfolding, evolution and disease. Trends Biochem Sci. 1999;24(9):329–32.
Iwata K, Fujiwara T, Matsuki Y, Akutsu H, Takahashi S, Naiki H, et al. 3D structure of amyloid protofilaments of β2-microglobulin fragment probed by solid-state NMR. Proc Natl Acad Sci USA. 2006;103(48):18119–24.
Langkilde AE, Morris KL, Serpell LC, Svergun DI, Vestergaard B. The architecture of amyloid-like peptide fibrils revealed by X-ray scattering, diffraction and electron microscopy. Acta Crystallogr Sect D Biol Crystallogr. 2015;71(4):882–95.
Eakin CM, Berman AJ, Miranker AD. A native to amyloidogenic transition regulated by a backbone trigger. Nat Struct Mol Biol. 2006;13(3):202–8.
Carrell RW, Gooptu B. Conformational changes and disease—serpins, prions and Alzheimer’s. Curr Opin Struct Biol. 1998;8(6):799–809.
Sekijima Y, Wiseman RL, Matteson J, Hammarström P, Miller SR, Sawkar AR, et al. The biological and chemical basis for tissue-selective amyloid disease. Cell. 2005;121(1):73–85.
Canet D, Last AM, Tito P, Sunde M, Spencer A, Archer DB, et al. Local cooperativity in the unfolding of an amyloidogenic variant of human lysozyme. Nat Struct Biol. 2002;9(4):308–15.
Dobson CM. Protein folding and misfolding. Nature. 2003;426(6968):884–90.
Chiti F, Dobson CM. Protein misfolding, functional amyloid, and human disease. Annu Rev Biochem. 2006;75:333–66.
Chiti F, Dobson CM. Protein misfolding, amyloid formation, and human disease: a summary of progress over the last decade. Annu Rev Biochem. 2017;86:27–68.
Volchuk A, Ron D. The endoplasmic reticulum stress response in the pancreatic β-cell. Diabetes Obes Metab. 2010;12:48–57.
Hebert DN, Molinari M. In and out of the ER: protein folding, quality control, degradation, and related human diseases. Physiol Rev. 2007;87(4):1377–408.
Roos RA. Huntington’s disease: a clinical review. Orphanet J Rare Dis. 2010;5(1):1–8.
Kumar A, Singh A. A review on Alzheimer’s disease pathophysiology and its management: an update. Pharmacol Rep. 2015;67(2):195–203.
•• Støy J, Edghill EL, Flanagan SE, Ye H, Paz VP, Pluzhnikov A, et al. Insulin gene mutations as a cause of permanent neonatal diabetes. Proc Natl Acad Sci USA. 2007;104(38):15040-4 (This study described the first ten patients with the mutant proinsulin syndrome and provided insights into their diagnosis and treatment.)
•• Colombo C, Porzio O, Liu M, Massa O, Vasta M, Salardi S, et al. Seven mutations in the human insulin gene linked to permanent neonatal/infancy-onset diabetes mellitus. J Clin Investig. 2008;118(6):2148-56. (This study (contemporaneous with ref. 15) reported seven additional heterozygous INS mutations causing permanent neonatal DM.)
Steiner D, Clark J, Nolan C, Rubenstein A, Margoliash E, Aten B, et al. Proinsulin and the biosynthesis of insulin. Recent Prog Horm Res. 1969;25:207.
Dodson G, Steiner D. The role of assembly in insulin’s biosynthesis. Curr Opin Struct Biol. 1998;8(2):189–94.
Molven A, Ringdal M, Nordbø AM, Ræder H, Støy J, Lipkind GM, et al. Mutations in the insulin gene can cause MODY and autoantibody-negative type 1 diabetes. Diabetes. 2008;57(4):1131–5.
Edghill EL, Flanagan SE, Patch A-M, Boustred C, Parrish A, Shields B, et al. Insulin mutation screening in 1,044 patients with diabetes: mutations in the INS gene are a common cause of neonatal diabetes but a rare cause of diabetes diagnosed in childhood or adulthood. Diabetes. 2008;57(4):1034–42.
Weiss MA. Diabetes mellitus due to the toxic misfolding of proinsulin variants. FEBS Lett. 2013;587(13):1942–50.
Dhayalan B, Chatterjee D, Chen Y-S, Weiss MA. Structural lessons from the mutant proinsulin syndrome. Front Endocrinol. 2021;12:754693.
Ron D. Proteotoxicity in the endoplasmic reticulum: lessons from the Akita diabetic mouse. J Clin Invest. 2002;109:443–5.
Araki E, Oyadomari S, Mori M. Impact of endoplasmic reticulum stress pathway on pancreatic β-cells and diabetes mellitus. Exp Biol Med. 2003;228(10):1213–7.
Zuber C, Fan J-Y, Guhl B, Roth J. Misfolded proinsulin accumulates in expanded pre-Golgi intermediates and endoplasmic reticulum subdomains in pancreatic beta cells of Akita mice. FASEB J. 2004;18(7):917–9.
Liu M, Hodish I, Rhodes CJ, Arvan P. Proinsulin maturation, misfolding, and proteotoxicity. Proc Natl Acad Sci USA. 2007;104(40):15841–6.
• Shrestha N, De Franco E, Arvan P, Cnop M. Pathological β-cell endoplasmic reticulum stress in type 2 diabetes: current evidence. Front Endocrinol. 2021;12:650158. (This review highlights the role of the unfolded protein response to the maintainence of ER homeostasis, β-cell function and survival.)
Wang M, Kaufman RJ. Protein misfolding in the endoplasmic reticulum as a conduit to human disease. Nature. 2016;529(7586):326–35.
Yang Y, Chan L. Monogenic diabetes: what it teaches us on the common forms of type 1 and type 2 diabetes. Endocr Rev. 2016;37(3):190–222.
Riddle MC, Philipson LH, Rich SS, Carlsson A, Franks PW, Greeley SAW, et al. Monogenic diabetes: from genetic insights to population-based precision in care. Reflections From a Diabetes Care Editors’ Expert Forum. Diabetes Care. 2020;43(12):3117–28.
Nkonge KM, Nkonge DK, Nkonge TN. The epidemiology, molecular pathogenesis, diagnosis, and treatment of maturity-onset diabetes of the young (MODY). Clin Diabetes Endocrinol. 2020;6(1):1–10.
Sousa M, Bruges-Armas J. Monogenic diabetes: genetics and relevance on diabetes mellitus personalized medicine. Curr Diabetes Rev. 2020;16(8):807–19.
Slingerland AS, Hattersley AT. Mutations in the Kir6. 2 subunit of the KATP channel and permanent neonatal diabetes: new insights and new treatment. Ann Med. 2005;37(3):186–95.
Babenko AP, Polak M, Cavé H, Busiah K, Czernichow P, Scharfmann R, et al. Activating mutations in the ABCC8 gene in neonatal diabetes mellitus. N Engl J Med. 2006;355(5):456–66.
Elhaji YA, Hui WuJ, Gottlieb B, Beitel LK, Alvarado C, Batist G, et al. An examination of how different mutations at arginine 855 of the androgen receptor result in different androgen insensitivity phenotypes. Mol Endocrinol. 2004;18(8):1876–86.
Varho TT, Alajoki LE, Posti KM, Korhonen TT, Renlund MG, Nyman SR, et al. Phenotypic spectrum of Salla disease, a free sialic acid storage disorder. Pediatr Neurol. 2002;26(4):267–73.
Nishimura G, Haga N, Kitoh H, Tanaka Y, Sonoda T, Kitamura M, et al. The phenotypic spectrum of COL2A1 mutations. Hum Mutat. 2005;26(1):36–43.
Johannesen K, Marini C, Pfeffer S, Møller RS, Dorn T, Niturad CE, et al. Phenotypic spectrum of GABRA1: from generalized epilepsies to severe epileptic encephalopathies. Neurology. 2016;87(11):1140–51.
Noone PG, Knowles MR. ‘CFTR-opathies’: disease phenotypes associated with cystic fibrosis transmembrane regulator gene mutations. Respir Res. 2001;2(6):1–5.
Remerand G, Boespflug-Tanguy O, Tonduti D, Touraine R, Rodriguez D, Curie A, et al. Expanding the phenotypic spectrum of Allan–Herndon–Dudley syndrome in patients with SLC 16A2 mutations. Dev Med Child Neurol. 2019;61(12):1439–47.
Southard-Smith EM, Angrist M, Ellison JS, Agarwala R, Baxevanis AD, Chakravarti A, et al. The Sox10Dom mouse: modeling the genetic variation of Waardenburg-Shah (WS4) syndrome. Genome Res. 1999;9(3):215–25.
Åkerblom HK, Vaarala O, Hyöty H, Ilonen J, Knip M. Environmental factors in the etiology of type 1 diabetes. Am J Med Genet. 2002;115(1):18–29.
Baker EN, Blundell TL, Cutfield JF, Dodson EJ, Dodson GG, Hodgkin DMC, et al. The structure of 2Zn pig insulin crystals at 1.5 Å resolution. Philos Trans R Soc Lond B Biol Sci. 1988;319(1195):369–456.
Hua QX, Mayer J, Jia W, Zhang J, Weiss MA. The folding nucleus of the insulin superfamily: a flexible peptide model foreshadows the native state. J Biol Chem. 2006;281:28131–42.
Narhi LO, Hua QX, Arakawa T, Fox GM, Tsai L, Rosenfeld R, et al. Role of native disulfide bonds in the structure and activity of insulin-like growth factor 1: genetic models of protein-folding intermediates. Biochemistry. 1993;32:5214–21.
Sieber P, Eisler K, Kamber B, Riniker B, Rittel W, Märki F, et al. Synthesis and biological activity of two disulphide bond isomers of human insulin:[A7-A11, A6-B7-cystine]-and [A6-A7, A11-B7-cystine] insulin (human). Biol Chem. 1978;359(1):113–24.
Hua QX, Jia W, Frank BH, Phillips NB, Weiss MA. A protein caught in a kinetic trap: structures and stabilities of insulin disulfide isomers. Biochemistry. 2002;41:14700–15.
Yang Y, Hua QX, Liu J, Shimizu EH, Choquette MH, Mackin RB, et al. Solution structure of proinsulin: connecting domain flexibility and prohormone processing. J Biol Chem. 2010;285:7847–51.
Steiner DF, Tager HS, Chan SJ, Nanjo K, Sanke T, Rubenstein AH. Lessons learned from molecular biology of insulin-gene mutations. Diabetes Care. 1990;13:600–9.
• Støy J, De Franco E, Ye H, Park S-Y, Bell GI, Hattersley AT. In celebration of a century with insulin–update of insulin gene mutations in diabetes. Mol Metab. 2021:101280. (This recent review provides an update to catalogue of INS mutations and discusses genetic causes and clinical opportunities for diagnosis and treatment.)
Miller JA, Narhi LO, Hua QX, Rosenfeld R, Arakawa T, Rohde M, et al. Oxidative refolding of insulin-like growth factor 1 yields two products of similar thermodynamic stability: a bifurcating protein-folding pathway. Biochemistry. 1993;32:5203–13.
Guo ZY, Qiao ZS, Feng YM. The in vitro oxidative folding of the insulin superfamily. Antioxid Redox Signal. 2008;10:127–40.
Finkelstein AV, Ivankov DN, Garbuzynskiy SO, Galzitskaya OV. Understanding the folding rates and folding nuclei of globular proteins. Curr Protein Peptide Sci. 2007;8(6):521–36.
Haataja L, Manickam N, Soliman A, Tsai B, Liu M, Arvan P. Disulfide mispairing during proinsulin folding in the endoplasmic reticulum. Diabetes. 2016;65(4):1050–60.
Liu M, Weiss MA, Arunagiri A, Yong J, Rege N, Sun J, et al. Biosynthesis, structure, and folding of the insulin precursor protein. Diabetes Obes Metab. 2018;20:28–50.
Herbach N, Rathkolb B, Kemter E, Pichl L, Klaften M, de Angelis MH, et al. Dominant-negative effects of a novel mutated Ins2 allele causes early-onset diabetes and severe β-cell loss in Munich Ins2C95S mutant mice. Diabetes. 2007;56(5):1268–76.
Matthews BW. Structural and genetic analysis of protein stability. Annu Rev Biochem. 1993;62(1):139–60.
Creighton TE. Protein folding: an unfolding story. Curr Biol. 1995;5(4):353–6.
Eriksson AE, Baase WA, Zhang X-J, Heinz DW, Blaber M, Baldwin EP, et al. Response of a protein structure to cavity-creating mutations and its relation to the hydrophobic effect. Science. 1992;255(5041):178–83.
Mezei M. On predicting foldability of a protein from its sequence. Proteins Struct Funct Bioinform. 2020;88(2):355–65.
Xu J, Baase WA, Baldwin E, Matthews BW. The response of T4 lysozyme to large-to-small substitutions within the core and its relation to the hydrophobic effect. Protein Sci. 1998;7:158–77.
Eriksson AE, Baase WA, Wozniak JA, Matthews BW. A cavity-containing mutant of T4 lysozyme is stabilized by buried benzene. Nature. 1992;355(6358):371–3.
O’Neil KT, DeGrado WF. A thermodynamic scale for the helix-forming tendencies of the commonly occurring amino acids. Science. 1990;250:646–51.
Padmanabhan S, Marqusee S, Ridgeway T, Laue TM, Baldwin RL. Relative helix-forming tendencies of nonpolar amino acids. Nature. 1990;344(6263):268–70.
Ortolani F, Piccinno E, Grasso V, Papadia F, Panzeca R, Cortese C, et al. Diabetes associated with dominant insulin gene mutations: outcome of 24-month, sensor-augmented insulin pump treatment. Acta Diabetol. 2016;53(3):499–501.
• Haataja L, Arunagiri A, Hassan A, Regan K, Tsai B, Dhayalan B, et al. Distinct states of proinsulin misfolding in MIDY. Cell Mol Life Sci. 2021;78:6017–31. (This study highlights the role of unpaired cysteines and proinsulin-mutant proinsulin self-association in the misfolding and ER stress using seven MIDY mutations.)
• Weiss MA, Nakagawa SH, Jia W, Xu B, Hua QX, Chu YC, et al. Protein structure and the spandrels of San Marco: insulin’s receptor-binding surface is buttressed by an invariant leucine essential for protein stability. Biochemistry. 2002;41:809-19. (This paper highlights the essential contribution of LeuA16 to chain-combination efficiency as a molecular “spandrels of San Marco. ”)
•• Liu M, Wan Z-l, Chu Y-C, Aladdin H, Klaproth B, Choquette M, et al. Crystal structure of a “nonfoldable” insulin impaired folding efficiency despite native activity. J Biol Chem. 2009;284(50):35259-72. (This paper demonstrates that substitution LeuA16→Val preserves structure and function but markedly impairs nascent disulfide pairing in proinsulin and blocks insulin chain combination in anticipation of its observation as a recessive mutation in neonatal diabetes.)
Blundell T, Humbel R. Hormone families: pancreatic hormones and homologous growth factors. Nature. 1980;287(5785):781–7.
Menting JG, Yang Y, Chan SJ, Phillips NB, Smith BJ, Whittaker J, et al. Protective hinge in insulin opens to enable its receptor engagement. Proc Natl Acad Sci U S A. 2014;111(33):E3395–404.
Scapin G, Dandey VP, Zhang Z, Prosise W, Hruza A, Kelly T, et al. Structure of the insulin receptor-insulin complex by single-particle cryo-EM analysis. Nature. 2018;556(7699):122–5.
Weis F, Menting JG, Margetts MB, Chan SJ, Xu Y, Tennagels N, et al. The signalling conformation of the insulin receptor ectodomain. Nat Commun. 2018;9(1):1–10.
Gould SJ. The exaptive excellence of spandrels as a term and prototype. Proc Natl Acad Sci USA. 1997;94(20):10750–5.
Li M, Rivière J-B, Polychronakos C. Why all MODY variants are dominantly inherited: a hypothesis. Trends Genet. 2021.
Al-Muhaizea MA, Aldeeb H, Almass R, Jaber H, Binhumaid F, Alquait L, et al. Genetics of ataxia telangiectasia in a highly consanguineous population. Ann Hum Genet. 2021;86(1):34–44.
Doyle AC, Paget S. The adventure of silver blaze: Mary McLaughlin and M. Einisman for the Scotland Yard Bookstore; 1979.
Chan SJ, Cao Q-P, Steiner DF. Evolution of the insulin superfamily: cloning of a hybrid insulin/insulin-like growth factor cDNA from amphioxus. Proc Natl Acad Sci USA. 1990;87(23):9319–23.
Bliss M. The discovery of insulin: twenty-fifth. anniversary. Chicago: University of Chicago Press; 2007. p. 310.
Flier JS, Kahn CR. Insulin: a pacesetter for the shape of modern biomedical science and the Nobel Prize. Mol Metab. 2021:101194.
Luisier S, Avital-Shmilovici M, Weiss MA, Kent SB. Total chemical synthesis of human proinsulin. Chem Commun. 2010;46(43):8177–9.
Kent SB. Novel protein science enabled by total chemical synthesis. Protein Sci. 2019;28(2):313–28.
Avital-Shmilovici M, Whittaker J, Weiss MA, Kent SB. Deciphering a molecular mechanism of neonatal diabetes mellitus by the chemical synthesis of a protein diastereomer,[D-AlaB8] human proinsulin. J Biol Chem. 2014;289(34):23683–92.
Rege NK, Liu M, Yang Y, Dhayalan B, Wickramasinghe NP, Chen Y-S, et al. Evolution of insulin at the edge of foldability and its medical implications. Proc Natl Acad Sci USA. 2020;117(47):29618–28.
Hodish I, Liu M, Rajpal G, Larkin D, Holz RW, Adams A, et al. Misfolded proinsulin affects bystander proinsulin in neonatal diabetes. J Biol Chem. 2010;285(1):685–94.
Sun J, Xiong Y, Li X, Haataja L, Chen W, Mir SA, et al. Role of proinsulin self-association in mutant INS gene–induced diabetes of youth. Diabetes. 2020;69(5):954–64.
Sun J, Cui J, He Q, Chen Z, Arvan P, Liu M. Proinsulin misfolding and endoplasmic reticulum stress during the development and progression of diabetes. Mol Aspects Med. 2015;42:105–18.
Liu M, Hodish I, Haataja L, Lara-Lemus R, Rajpal G, Wright J, et al. Proinsulin misfolding and diabetes: mutant INS gene-induced diabetes of youth. Trends Endrocrinol Metab. 2010;21(11):652–9.
• Yong J, Johnson JD, Arvan P, Han J, Kaufman RJ. Therapeutic opportunities for pancreatic β-cell ER stress in diabetes mellitus. Nat Rev Endocrinol. 2021:1–13. (This review highlights β-cell ER stress and evaluates related therapeutic opportunities to maintain functional β-cell mass in non-syndromic type 2 DM.)
Fishman MC. Power of rare diseases: found in translation. Sci Transl Med. 2013;5(201):201ps11-ps11.
Jarosinski MA, Dhayalan B, Chen Y-S, Chatterjee D, Varas N, Weiss MA. Structural principles of insulin formulation and analog design: a century of innovation. Mol Metab. 2021:101325.
Hua QX, Chu YC, Jia W, Phillips NB, Wang RY, Katsoyannis PG, et al. Mechanism of insulin chain combination. Asymmetric roles of A-chain α-helices in disulfide pairing. J Biol Chem. 2002;277:43443–53.
Weiss MA, Wan Z, Zhao M, Chu Y-C, Nakagawa SH, Burke GT, et al. Non-standard insulin design: structure-activity relationships at the periphery of the insulin receptor. J Mol Biol. 2002;315(2):103–11.
Granel B, Valleix S, Serratrice J, Cherin P, Texeira A, Disdier P, et al. Lysozyme amyloidosis: report of 4 cases and a review of the literature. Medicine (Baltimore). 2006;85(1):66–73.
Benson MD, Liepnieks J, Uemichi T, Wheeler G, Correa R. Hereditary renal amyloidosis associated with a mutant fibrinogen α–chain. Nat Genet. 1993;3(3):252–5.
Ohashi K. Pathogenesis of β2-microglobulin amyloidosis. Pathol Int. 2001;51(1):1–10.
Benson MD, Uemichi T. Transthyretin amyloidosis. Amyloid. 1996;3(1):44–56.
Merlini G, Comenzo RL, Seldin DC, Wechalekar A, Gertz MA. Immunoglobulin light chain amyloidosis. Expert Rev Hematol. 2014;7(1):143–56.
Prusiner SB. Molecular biology of prion diseases. Science. 1991;252(5012):1515–22.
Revesz T, Holton JL, Lashley T, Plant G, Frangione B, Rostagno A, et al. Genetics and molecular pathogenesis of sporadic and hereditary cerebral amyloid angiopathies. Acta Neuropathol. 2009;118(1):115–30.
Kiuru S. Gelsolin-related familial amyloidosis, Finnish type (FAF), and its variants found worldwide. Amyloid. 1998;5(1):55–66.
Leboulleux S, Baudin E, Travagli JP, Schlumberger M. Medullary thyroid carcinoma. Clin Endocrinol (Oxf). 2004;61(3):299–310.
Röcken C, Peters B, Juenemann G, Saeger W, Klein HU, Huth C, et al. Atrial amyloidosis: an arrhythmogenic substrate for persistent atrial fibrillation. Circulation. 2002;106(16):2091–7.
Sotiropoulos I, Galas M-C, Silva JM, Skoulakis E, Wegmann S, Maina MB, et al. Atypical, non-standard functions of the microtubule associated Tau protein. Acta Neuropathol Commun. 2017;5(1):1–11.
Du K, Sharma M, Lukacs GL. The ΔF508 cystic fibrosis mutation impairs domain-domain interactions and arrests post-translational folding of CFTR. Nat Struct Mol Biol. 2005;12(1):17–25.
Kereszturi É, Szmola R, Kukor Z, Simon P, Ulrich Weiss F, Lerch MM, et al. Hereditary pancreatitis caused by mutation-induced misfolding of human cationic trypsinogen: a novel disease mechanism. Hum Mutat. 2009;30(4):575–82.
Tagliavacca L, Wang Q, Kaufman RJ. ATP-dependent dissociation of non-disulfide-linked aggregates of coagulation factor VIII is a rate-limiting step for secretion. Biochemistry. 2000;39(8):1973–81.
Lang AE, Lozano AM. Parkinson’s disease. N Engl J Med. 1998;339(16):1130–43.
Dhayalan B, Glidden MD, Zaykov A, Chen Y-S, Yang Y, Phillips NB, et al. Peptide Model of the Mutant Proinsulin Syndrome. I. Design and Clinical Correlation. Front Endocrinol. 2022;13:821069
Yang Y, Glidden MD, Dhayalan B, Zaykov A, Chen Y-S, Wickramasinghe NP, et al. Peptide Model of the Mutant Proinsulin Syndrome. II. Nascent NMR Structure and Biophysical Correlation. Front Endocrinol. 2022;13:821091
Acknowledgements
The authors thank P. Arvan, T.L. Blundell, D. Chatterjee, Y.S. Chan, M. Jarosinski, S.B. Kent, F. Ismail-Beigi, M. Liu, N.B. Phillips, J. Racca, and Y. Yang for helpful discussion. We thank L. Liu, P. Arvan, G.I. Bell, L. Philipson, and E. De Franco for communication of results prior to publication. M.A.W. is grateful to the late G.G. Dodson, P.G. Katsoyannis, and D.F. Steiner for their encouragement in the early years of this research program.
Funding
This work was supported in part by a grant from the National Institutes of Health (R01 DK040949). BD is supported in part by a grant from the Diabetes Research Connection.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Pediatric Type 2 and Monogenic Diabetes
Rights and permissions
About this article
Cite this article
Dhayalan, B., Weiss, M.A. Diabetes-Associated Mutations in Proinsulin Provide a “Molecular Rheostat” of Nascent Foldability. Curr Diab Rep 22, 85–94 (2022). https://doi.org/10.1007/s11892-022-01447-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11892-022-01447-2