Abstract
Purpose of Review
Deciphering the mechanisms of type 2 diabetes (T2DM) risk loci can greatly inform on disease pathology. This review discusses current knowledge of mechanisms through which genetic variants influence T2DM risk and considerations for future studies.
Recent Findings
Over 100 T2DM risk loci to date have been identified. Candidate causal variants at risk loci map predominantly to non-coding sequence. Physiological, epigenomic and gene expression data suggest that variants at many known T2DM risk loci affect pancreatic islet regulation, although variants at other loci also affect protein function and regulatory processes in adipose, pre-adipose, liver, skeletal muscle and brain. The effects of T2DM variants on regulatory activity in these tissues appear largely, but not exclusively, due to altered transcription factor binding. Putative target genes of T2DM variants have been defined at an increasing number of loci and some, such as FTO, may entail several genes and multiple tissues. Gene networks in islets and adipocytes have been implicated in T2DM risk, although the molecular pathways of risk genes remain largely undefined.
Summary
Efforts to fully define the mechanisms of T2DM risk loci are just beginning. Continued identification of risk mechanisms will benefit from combining genetic fine-mapping with detailed phenotypic association data, high-throughput epigenomics data from diabetes-relevant tissue, functional screening of candidate genes and genome editing of cellular and animal models.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Type 2 diabetes (T2DM) affects over 400 million people worldwide and is one of the major challenges in modern health [1]. Genetic association studies have identified over 100 loci that influence T2DM risk, and recent studies have argued that many additional risk loci remain to be discovered [2••]. These loci predominantly have small to moderate effects on individual predisposition to T2DM, complicating the clinical translation of genetic information directly. Each locus, however, contains molecular and cellular mechanisms through which risk variants functionally contribute to disease. Deciphering these mechanisms is paramount to a greater understanding of T2DM pathogenesis and can potentially inform novel avenues for disease prevention, therapies and treatments.
Evaluation of T2DM risk locus mechanisms involves a variety of complementary approaches that include dense T2DM genetic data, physiological trait data, genome and epigenome annotation, quantitative trait locus (QTL) mapping and cellular and animal modelling. Advances in genetics, genomics and cellular modelling technology have enabled more precise identification of risk locus mechanisms using these approaches. Here, I discuss recent advances in applying these and other approaches to identify mechanisms of T2DM risk as well as considerations for future studies.
Identification of Causal Variants at T2DM Loci
Risk loci identified in genetic studies of T2DM often contain many associated variants due to linkage disequilibrium between variants in the human genome [2••, 3, 4]. Genetic studies traditionally report the most significantly associated variant at a given locus, or ‘index’ variant [3, 5], although this variant is not necessarily causal for T2DM. The majority of associated variants at a given locus are not causal and only correlate with the true causal variant(s) because of linkage disequilibrium. Resolving which specific variants are causal at a risk locus is critical to subsequently understanding molecular functions of these loci.
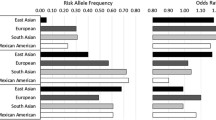
Genetic fine mapping is a common strategy for resolving causal variants, whereby a comprehensive set of genetic variants across a locus is genotyped or imputed and evaluated in large sample sets [6]. A study of genotyping data from the Metabochip microarray fine-mapped 39 known T2DM loci using Bayesian methods [7], and at the MTNR1B locus resolved a single variant, rs10830963, with >99% probability of being causal for T2DM risk [7]. Whole-genome sequencing of T2DM samples combined with imputation into GWAS data from 55 k samples fine-mapped 81 known T2DM loci and at several loci identified candidate variants with high causal probabilities that were not assayed in previous studies [2••]. One useful strategy to further improve resolution of causal variants through fine mapping is using genetic data from different ethnicities, assuming a causal variant is shared across ethnic backgrounds [8]. A study of T2DM using data of samples from European, Hispanic, East Asian and South Asian ancestry fine-mapped 10 known T2DM signals and reduced the set of variants likely causal for these signals [8].
Dense genetic data also facilitates the identification of additional risk variants at loci that have distinct effects on T2DM risk [2••, 7]. For example, in a T2DM fine-mapping study of 39 loci, conditional analyses identified 49 total distinct common variant risk signals including 5 signals at the KCNQ1 locus, 3 signals at the HNF1A locus and 2 signals at several additional loci [7]. Common risk loci also harbour lower frequency risk variants, such as at the CCND2, PPARG and IRS1 loci identified in genome sequencing of T2DM samples [2••, 9]. Moving forward, in addition to genome sequencing, studies imputing genotype data from T2DM cohorts and Biobanks into large reference panels will facilitate continued fine-mapping of causal variants [10, 11].
Physiological Association of T2DM Variants
Genetic association data of quantitative measures relevant to T2DM pathogenesis can provide key insight into the mechanisms of a diabetes risk locus.
The canonical example of physiology at T2DM signals is the FTO locus, which has a primary effect on body mass index (BMI) and obesity [12]. Studies have more broadly determined the effect of T2DM risk variants on measures of fasting glycemia such as glucose and insulin levels [13, 14]. Homeostasis model assessment (HOMA) derived from these measures provides likely effects on pancreatic beta cell function and insulin secretion (HOMA-B) and insulin resistance (HOMA-IR) [5]. Cataloguing T2DM loci based on these measures revealed a clear distinction between loci that influence insulin secretion and insulin sensitivity [5]. A larger percentage of loci from these analyses influenced insulin secretion, suggesting that this is the predominant mechanism of currently known T2DM loci. Additional glycemic measures such as HbA1C, proinsulin level, 2 h glucose response and glucose-stimulated insulin secretion (GSIS) [15,16,17,18,19], as well as phenotypes such as lipid levels [20] and anthropometric traits [21,22,23], are also relevant to T2DM pathophysiology.
Patterns of quantitative trait associations can also help identify groups of T2DM loci with shared physiology and, by extension, potential shared mechanisms. For example, many variants that influence BMI appear to affect neuronal functions and variants that influence glucose levels, HOMA-B and related phenotypes appear to affect pancreatic islet functions [22, 24]. A study used several glycemic measures to define distinct clusters of loci influencing insulin sensitivity, insulin secretion and insulin processing [25]. The majority of loci, however, could not be assigned to a cluster, suggesting that association data from larger studies of these physiological measures are needed.
Not all risk loci will necessarily affect normal physiology and instead, for example, influence progression or exacerbation of T2DM in pre-diabetic individuals. T2DM loci also influence other complex diseases such as T1D [26], other autoimmune disease [27], cancer [28] and neurodegenerative disease [29]. Finally, the increasing availability of richly phenotyped cohorts such as those from biobanks further enables ‘phenome-wide’ studies of hundreds of human phenotypes and diseases [10, 30, 31]. Moving forward, these associations can be exploited to gain additional clues into the mechanisms of T2DM loci.
Consequences of T2DM Variants on Genome Function
Determining the genomic consequences of variants causal for T2DM risk is closely linked to functional annotation of the genome.
Protein-coding variants are causal candidates for risk signals at several loci including SLC30A8, GCKR, PPARG, KCNJ11, ABCC8, HNF1A and HNF4A [3, 7]. Relatively few T2DM loci, however, are likely explained by a protein-coding variant. The majority of loci instead map to non-coding sequence, implying that they likely affect gene regulation [7]. Regulatory processes are often cell type specific and require data generated in each cell type [32, 33]. High-throughput techniques provide genome-wide maps of cell type epigenomes, including accessible chromatin (ATAC-seq, DHS-seq, FAIRE-seq), histone tail modifications and transcription factor binding (ChIP-seq) and DNA methylation (WGBS) [34,35,36]. Consortia such as ENCODE and the NIH Epigenome Roadmap have used these assays to map the epigenome of hundreds of human cell lines and primary tissues [32, 37]. Additional studies have focused on profiling the epigenome of specific diabetes-relevant tissues such as islets, skeletal muscle and adipose [24, 38,39,40].
Variants known to affect regulatory activity are listed in Table 1. A common strategy is to identify candidate variants that overlap epigenome annotations from T2DM-relevant tissues. A study of islet FAIRE-seq revealed that rs7903146 at TCF7L2 lies in islet accessible chromatin [36], which genetic studies support as the most likely causal variant for this locus [7]. Candidate T2DM variants in islet regulatory sites defined using ChIP-seq, FAIRE-seq and DHS-seq have also been reported at MTNR1B [7], JAZF1 [41], CDC123/CAMK1D [42], ZFAND3 [24], WFS1 [43], KCNK16/17 [44], KCNQ1 [45] and CENTD2 [46•, 47]. T2DM variants also overlap regulatory sites active in other tissues such as skeletal muscle at ANK1 [40], adipose at PPARG [48] and pre-adipose and brain at FTO [49••, 50, 51]. Experiments have then confirmed the allelic effects of these candidate variants on cell-type regulatory activity, for example through gene reporter assays or correlating variant genotypes to molecular outputs using allelic imbalance or QTL mapping. For example, at TCF7L2, the risk allele of rs7903146 correlated with increased accessible chromatin in islet samples and had increased reporter activity in islet cell lines [36]. At some loci, such as PPARG and FTO, multiple functional variants may exist on the same risk haplotype. These studies demonstrate that variants at specific T2DM loci affect regulatory activity in islets, as well as other disease-relevant tissues such as adipose, muscle and brain.
Characterizing patterns of genomic annotations at variants across sets of T2DM loci can reveal broad genomic consequences of T2DM risk variants, as well as help prioritize between multiple annotated variants at a locus. Several studies have identified enrichment of variants across T2DM loci in regulatory sites active in pancreatic islets [2••, 7, 24, 39]. These enrichments are particularly pronounced in regions of islet ‘stretch’-enhancer activity or in regions of highly clustered islet active enhancers [24, 39]. Studies have also identified enrichment of variants in regulatory sites for other T2DM-relevant tissues, such as liver, adipocytes and pre-adipocytes and sites bound by specific regulatory proteins within these tissues, such as FOXA2 [2••, 7, 52]. Recent methods incorporate the effects of annotations directly as priors in fine mapping to improve causal variant resolution [53,54,55]. For example, incorporation of genome and 12 cell type epigenome annotations into fine mapping of 81 T2DM loci reduced the number of candidate causal variants by over 35% [2••]. At the CENTD2 locus, incorporating epigenome priors identified a candidate causal variant in islet accessible chromatin, rs140130268, which was not as highly prioritized in genetic data alone [46•]. The use of such methods will benefit future studies to prioritize causal variants at known signals and help identify additional risk variants.
Many T2DM variants likely impact cell-type regulatory activity by altering transcription factor (TF) binding. Identification of disrupted TFs can also provide further clues towards T2DM-relevant regulatory pathways. Methods to identify specific TF binding sites within regulatory regions include in silico prediction, which can be combined with species conservation and DNA footprints, and experimental techniques such as electromobility shift assays (EMSA) and allelic ChIP [34, 56]. For example, at the CAMK1D/CDC123 locus, variant rs11257655 disrupted islet FOXA1 and FOXA2 binding in both EMSA and ChIP assays [42]. At MTNR1B, variant rs10830963 disrupted NEUROD1 binding in islets [7]. In adipose cells, rs4684847 at PPARG affects PRRX1 binding and rs1421085 at FTO affects ARID5B binding within conserved regulatory modules [48, 49••]. In brain, rs8050136 at FTO affects CUX1 binding [51]. In skeletal muscle, rs508419 at ANK1 affects TR4 binding in an accessible chromatin footprint [40]. More broadly, several T2DM loci contain candidate variants that map in islet accessible chromatin footprints for RFX [44]. Finally, a study of DHS-seq data from ENCODE identified T2DM variants mapping in footprints for MODY transcription factors [57]. Together this demonstrates that T2DM variants affect genomic binding of a variety of transcription factors acting within diabetes-relevant tissues. It is likely, however, that not all T2DM variants affect regulatory activity by disrupting transcription factor binding. Instead some may involve changes, for example, in nucleosome positioning [58], DNA methylation [59], miRNA binding [60] or gene splicing [61].
Generation of epigenome data across environmental conditions, disease states, development [62] and cellular populations [63] will continue to help identify mechanisms by which non-coding T2DM variants affect genome function. Epigenomic data from larger numbers of samples will also be useful for correlating genetic variants to chromatin, histone modifications and transcription factor binding via QTL mapping [64, 65]. Furthermore, high-throughput assays can determine allelic effects on enhancer activity or transcription factor binding across tens to hundreds of thousands of variants in a single experiment [66,67,68]. These studies have not yet been widely conducted in key diabetes-relevant tissue and will be invaluable in identifying the effects of T2DM variants on epigenome function.
Target Genes of Causal Regulatory Variants
Identifying the genes affected by T2DM variants is a critical challenge, particularly as most T2DM signals are non-coding and regulatory elements are often distal to their target genes. Genes implicated as targets of variants at T2DM signals are listed in Table 2 .
The primary approach to determine gene targets is to correlate T2DM regulatory variants to gene expression level via QTL or allele-specific expression (ASE) mapping [33]. For example, a study identified correlation of risk alleles at TCF7L2 with higher TCF7L2 expression in human islets, albeit in few samples [69]. Studies of diabetes-relevant tissue have since mapped gene expression in larger sample sizes [70, 71]. In islets, QTL mapping of RNA-seq identified potential target genes including AP3S2, STARD10, CAMK1D, HMG20A, ADCY5, DGKB, MTNR1B, NKX6–3, ZMIZ1, GPSM1, UBE2E2 and KCNK17 [44, 60, 71]. Additional putative target genes include ANK1, JAZF1, GPSM1, ABCC8, PROX1-AS, ZFAND3 in skeletal muscle [40], IRX3 in brain [50] and IRX3 and IRX5 in pre-adipocytes [49••], and PPARG, KLF14 and CCND2 in adipocytes [9, 48, 72]. A critical aspect of mapping eQTLs at disease signals is determining whether both signals share the same casual variant(s), and thus conditional association or co-localization tests are needed to confirm sharing [73, 74]. An alternate to QTL analyses is to map ASE in heterozygote coding variants linked to a T2DM variant. In islets, this has implicated additional genes such as SLC30A8, ANPEP, KCNJ11, THADA and WFS1 [71, 75]. At most loci, however, no target gene has been defined through either approach. At other loci, multiple target genes have been implicated and may involve a combination of genes. Continued efforts to define target genes will be enhanced by larger expression datasets from key tissues across a variety of environmental conditions and within specific cellular populations, as well as the use of multi-variant expression models [76, 77].
Complementary techniques such as 3C, 4C, Capture-C and Hi-C capture the spatial configuration of chromatin to identify physical links between regulatory elements and target promoters [78, 79]. For example, at the CENTD2 locus 3C analyses in islets identified interactions of T2DM risk variants with the promoter region of STARD10 [46•]. At the FTO locus, 4C-seq analyses in mouse brain regions identified long-range interactions between T2DM regions and IRX3 [50]. Data from Hi-C assays can also identify broad chromatin domains called TADs (topologically associating domains) that can be used to restrict the genomic space over which regulatory elements act [80]. For example, at the FTO locus, TAD definitions were used to restrict genes likely to be regulated by T2DM risk variants including IRX3 and IRX5 [49••]. Chromosome conformation data can also be combined with RNA-seq and allelic imbalance mapping to boost the ability to detect QTLs for T2DM variants on gene expression [81], as well as a means to identify haplotype-specific regulatory effects [82].
Rare and low frequency T2DM risk variants affecting protein function can be used to bolster support for the gene likely affected by a risk signal. For example, many T2DM loci map near genes that cause Mendelian T2D and obesity [2••] and it is likely that the majority of these genes are the targets of common variants. Rare and low frequency coding variants at T2DM loci identified through exome sequencing also influence polygenic T2D risk, such as at MTNR1B, PAX4, PPARG, PAM, RREB1 and SLC30A8 [2••, 83,84,85], supporting the likely causality of these genes. In addition, the availability of large Biobanks has enabled the recall of individuals with rare coding variants for phenotypic studies, such as for CDKN2A [86].
Finally, genome editing is increasingly an invaluable tool for determining target genes. For example, editing the risk alleles of a T2DM variant or deleting entire regulatory regions surrounding these variants followed by RNA-seq or 4C/Hi-C/Capture-C can be used to identify genes with differences in expression or promoter interactions caused by specific T2DM variants. For example, at the TCF7L2 locus, deletion of regulatory regions surrounding the T2DM risk variants revealed effects on expression and promoter interactions at ACSL5 in HCT116 cells [87]. At the FTO locus, editing the alleles of rs1421085 in pre-adipocytes confirmed effects of this variant on regulation of IRX3 and IRX5 [49••]. Future efforts to define target genes will thus benefit greatly from genome editing in cell lines and iPSC-derived models of diabetes-relevant tissue.
Cellular and Animal Models of T2DM Loci
Modelling risk variant and gene function within cellular and animal systems can provide further support for the role of a candidate target gene in T2DM pathogenesis as well as determine biological functions and pathways target genes affect.
Cellular models of candidate genes at T2DM loci include studies of primary tissue, diabetes-relevant cell lines and cells derived from induced pluripotent stem cells (iPSCs). For coding variants, one approach is to assess the activity of tagged alleles. For example, at GCKR, transfecting fluorescently labelled alleles revealed differential effects on hepatic glucose uptake [88]. As most T2DM variants are regulatory, studies of gene expression changes are more appropriate, for example using siRNA or expression vectors. Silencing CDKN2A expression in the EndoC-bH1 human islet cell line increased insulin secretion and PKA signalling [86]. At TCF7L2, adenovirus-based overexpression in human islets reduced glucose-stimulated insulin secretion [69]. Given the large number of T2DM risk loci and candidate target genes, functional screens can help expedite these evaluations. For example, a siRNA-mediated screen of genes at T2DM loci in EndoC-bH1 cells identified genes affecting insulin secretion [89]. Genome editing in both cell line and iPSC-derived models further enables characterizing the effects of specific variant alleles on cellular function. At IRX3/IRX5, editing the alleles of rs1421085 resulted in altered thermogenesis in differentiating pre-adipocytes [49••].
Animal models can help further determine the effects of altered T2DM gene activity at an organismal level. Knockouts and transgenes in rodents both globally and in tissue-specific settings have been characterized for genes at several T2DM loci, such as TCF7L2 [90], SLC30A8 [91,92,93], MC4R [94], WFS1 [95], IRS1 [96], STARD10 [46•], IRX3 [50], FTO [97] and RPGRIP1L [98]. At some loci, rodent models provide additional support for likely mechanisms of human risk variants. For example, STARD10 beta cell-specific mouse knockouts had impaired proinsulin processing and insulin secretion, and overexpression in beta cells improved glucose homeostasis, consistent with human data [46•]. In other cases, data from rodent models have complicated interpretation of risk mechanisms such as at SLC30A8, where multiple knockout models had differential effects on glucose homeostasis [84, 99]. A further challenge with rodent models is evaluating the effects of human risk variants, as most are non-coding and not often highly conserved across species. One strategy is to introduce human sequence directly in rodents, such as in bacterial artificial chromosomes (BACs) [100]. The utility of rodent models will be enhanced as more target genes are defined and expedited by the use of genome editing techniques.
Moving forward, as T2DM risk is a consequence of many variants, models of multiple T2DM signals will be informative in addition to single locus models. Furthermore, given the potential for T2DM risk variants to interact with environmental cues, these models ideally need to be evaluated in a variety of environmental contexts. Finally, in addition to rodent and human cellular models, zebrafish and drosophila are attractive models due to the relative ease of genetic manipulations, short generation time and the conservation of glucose homeostasis mechanisms across species [101, 102].
Molecular Pathways of Genes at T2DM Risk Loci
Genes affected by T2DM risk signals likely map within shared cellular pathways through which disease pathogenesis is mediated.
A study analysed genes mapping to T2DM-associated variants or involved in monogenic diabetes using protein-protein interaction (PPI) networks [3]. T2DM genes mapping within interaction networks were generally more inter-connected than expected. The most inter-connected node in the network was CREBBP, involved in chromatin remodelling of regulatory elements, suggesting the importance of this protein to T2DM-relevant pathways, even though it does not harbour known T2DM risk variants itself. Additional analyses identified an enrichment of T2DM-associated variants among genes involved in adipocytokine signalling.
An alternate approach to understand pathways of T2DM variants is to define the upstream factors broadly regulating target gene activity. For example, studies have found that T2DM-associated variants preferentially map within binding motifs for transcription factors involved in MODY [57], as well as RFX motifs [44]. A fine-mapping study of T2DM loci identified enrichment of likely causal T2DM variants in FOXA2-bound regulatory sites [7]. Candidate genes at FOXA2 loci were preferentially downregulated in knockout models of FOXA2 compared to other loci. Although binding of FOXA2 itself was not often disrupted by the variants directly, it suggests the potential importance of this protein in regulating T2DM-relevant sites.
Finally, mapping the trans effects of a T2DM variant can reveal disease-relevant gene networks. For example, variants at the KLF14 locus with maternally inherited cis effects on KLF14 expression in adipose tissue were also associated with adipose expression of an entire gene network in trans [72]. Genes regulated in trans were then further associated with insulin resistance related traits, broadly implicating the KLF14-regulated gene network in metabolic disease risk.
Thus far, however, efforts to comprehensively define pathways of T2DM genes have not been widely successful. This is due in part to the still relatively small number of T2DM loci at which the specific target gene(s) and upstream regulator(s) are known, in combination with the large number of pathways likely involved in T2DM pathophysiology. A further complication is the incomplete knowledge of biological pathways and interactions. Future studies incorporating increasing knowledge of target genes and regulators and their tissue activity will help uncover molecular pathways of T2DM risk.
Conclusions
Determining the molecular mechanisms of T2DM risk loci holds enormous promise as a means to understand the genes and pathways involved in diabetes pathophysiology. The mechanisms of the majority of T2DM risk loci, however, are currently unknown and even for loci with proposed mechanisms the stories are often incomplete. In addition to this review, Grotz et al. also discuss identifying causal mechanisms and genes at T2DM risk loci [103].
Evaluation of risk loci entails genetic fine-mapping, quantitative phenotype association, genomic and epigenomic annotation and cellular/animal models. Finding consistency across these data is critical to fully describe mechanisms as well as their potential for clinical translation. Physiological and genomic studies both suggest that many currently known T2DM loci affect pancreatic islet regulation. However, loci also affect protein function and adipose, liver, muscle and brain regulation. As the majority of T2DM risk remains to be described, continued discovery of risk loci will undoubtedly reveal a greater diversity of cellular mechanisms.
Future studies that leverage the substantial advances and opportunities in high-throughput epigenomics, functional screening, genome editing and statistical methods to integrate these data will further expedite efforts to describe mechanisms of T2DM loci.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Worth Health Organiziation. www.who.int/mediacentre/factsheets/fs312/en/. Accessed July 2017.
•• Fuchsberger C, Flannick J, Teslovich TM, Mahajan A, Agarwala V, Gaulton KJ, et al. The genetic architecture of type 2 diabetes. Nature. 2016;536(7614):41–7. doi:10.1038/nature18642. This study demonstrates that the genetic basis of type 2 diabetes is largely a consequence of many small effect variants.
Morris AP, Voight BF, Teslovich TM, Ferreira T, Segre AV, Steinthorsdottir V, et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat Genet. 2012;44(9):981–90. doi:10.1038/ng.2383.
Genomes Project C, Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, et al. A global reference for human genetic variation. Nature. 2015;526(7571):68–74. doi:10.1038/nature15393.
Voight BF, Scott LJ, Steinthorsdottir V, Morris AP, Dina C, Welch RP, et al. Twelve type 2 diabetes susceptibility loci identified through large-scale association analysis. Nat Genet. 2010;42(7):579–89. doi:10.1038/ng.609.
Wellcome Trust Case Control C, Maller JB, McVean G, Byrnes J, Vukcevic D, Palin K, et al. Bayesian refinement of association signals for 14 loci in 3 common diseases. Nat Genet. 2012;44(12):1294–301. doi:10.1038/ng.2435.
Gaulton KJ, Ferreira T, Lee Y, Raimondo A, Magi R, Reschen ME, et al. Genetic fine mapping and genomic annotation defines causal mechanisms at type 2 diabetes susceptibility loci. Nat Genet. 2015;47(12):1415–25. doi:10.1038/ng.3437.
Replication DIG, Meta-analysis C, Asian Genetic Epidemiology Network Type 2 Diabetes C, South Asian Type 2 Diabetes C, Mexican American Type 2 Diabetes C, Type 2 Diabetes Genetic Exploration by Nex-generation sequencing in muylti-Ethnic Samples C, et al. Genome-wide trans-ancestry meta-analysis provides insight into the genetic architecture of type 2 diabetes susceptibility. Nat Genet. 2014;46(3):234–44. doi:10.1038/ng.2897.
Steinthorsdottir V, Thorleifsson G, Sulem P, Helgason H, Grarup N, Sigurdsson A, et al. Identification of low-frequency and rare sequence variants associated with elevated or reduced risk of type 2 diabetes. Nat Genet. 2014;46(3):294–8. doi:10.1038/ng.2882.
Sudlow C, Gallacher J, Allen N, Beral V, Burton P, Danesh J, et al. UK biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 2015;12(3):e1001779. doi:10.1371/journal.pmed.1001779.
McCarthy S, Das S, Kretzschmar W, Delaneau O, Wood AR, Teumer A, et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat Genet. 2016;48(10):1279–83. doi:10.1038/ng.3643.
Frayling TM, Timpson NJ, Weedon MN, Zeggini E, Freathy RM, Lindgren CM, et al. A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science. 2007;316(5826):889–94. doi:10.1126/science.1141634.
Scott RA, Lagou V, Welch RP, Wheeler E, Montasser ME, Luan J, et al. Large-scale association analyses identify new loci influencing glycemic traits and provide insight into the underlying biological pathways. Nat Genet. 2012;44(9):991–1005. doi:10.1038/ng.2385.
Manning AK, Hivert MF, Scott RA, Grimsby JL, Bouatia-Naji N, Chen H, et al. A genome-wide approach accounting for body mass index identifies genetic variants influencing fasting glycemic traits and insulin resistance. Nat Genet. 2012;44(6):659–69. doi:10.1038/ng.2274.
Soranzo N, Sanna S, Wheeler E, Gieger C, Radke D, Dupuis J, et al. Common variants at 10 genomic loci influence hemoglobin A(1)(C) levels via glycemic and nonglycemic pathways. Diabetes. 2010;59(12):3229–39. doi:10.2337/db10-0502.
Saxena R, Hivert MF, Langenberg C, Tanaka T, Pankow JS, Vollenweider P, et al. Genetic variation in GIPR influences the glucose and insulin responses to an oral glucose challenge. Nat Genet. 2010;42(2):142–8. doi:10.1038/ng.521.
Dupuis J, Langenberg C, Prokopenko I, Saxena R, Soranzo N, Jackson AU, et al. New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nat Genet. 2010;42(2):105–16. doi:10.1038/ng.520.
Strawbridge RJ, Dupuis J, Prokopenko I, Barker A, Ahlqvist E, Rybin D, et al. Genome-wide association identifies nine common variants associated with fasting proinsulin levels and provides new insights into the pathophysiology of type 2 diabetes. Diabetes. 2011;60(10):2624–34. doi:10.2337/db11-0415.
Prokopenko I, Poon W, Magi R, Prasad BR, Salehi SA, Almgren P, et al. A central role for GRB10 in regulation of islet function in man. PLoS Genet. 2014;10(4):e1004235. doi:10.1371/journal.pgen.1004235.
Teslovich TM, Musunuru K, Smith AV, Edmondson AC, Stylianou IM, Koseki M, et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature. 2010;466(7307):707–13. doi:10.1038/nature09270.
Shungin D, Winkler TW, Croteau-Chonka DC, Ferreira T, Locke AE, Magi R, et al. New genetic loci link adipose and insulin biology to body fat distribution. Nature. 2015;518(7538):187–96. doi:10.1038/nature14132.
Locke AE, Kahali B, Berndt SI, Justice AE, Pers TH, Day FR, et al. Genetic studies of body mass index yield new insights for obesity biology. Nature. 2015;518(7538):197–206. doi:10.1038/nature14177.
Horikoshi M, Beaumont RN, Day FR, Warrington NM, Kooijman MN, Fernandez-Tajes J, et al. Genome-wide associations for birth weight and correlations with adult disease. Nature. 2016;538(7624):248–52. doi:10.1038/nature19806.
Pasquali L, Gaulton KJ, Rodriguez-Segui SA, Mularoni L, Miguel-Escalada I, Akerman I, et al. Pancreatic islet enhancer clusters enriched in type 2 diabetes risk-associated variants. Nat Genet. 2014;46(2):136–43. doi:10.1038/ng.2870.
Dimas AS, Lagou V, Barker A, Knowles JW, Magi R, Hivert MF, et al. Impact of type 2 diabetes susceptibility variants on quantitative glycemic traits reveals mechanistic heterogeneity. Diabetes. 2014;63(6):2158–71. doi:10.2337/db13-0949.
Onengut-Gumuscu S, Chen WM, Burren O, Cooper NJ, Quinlan AR, Mychaleckyj JC, et al. Fine mapping of type 1 diabetes susceptibility loci and evidence for colocalization of causal variants with lymphoid gene enhancers. Nat Genet. 2015;47(4):381–6. doi:10.1038/ng.3245.
Lees CW, Barrett JC, Parkes M, Satsangi J. New IBD genetics: common pathways with other diseases. Gut. 2011;60(12):1739–53. doi:10.1136/gut.2009.199679.
Frayling TM, Colhoun H, Florez JC. A genetic link between type 2 diabetes and prostate cancer. Diabetologia. 2008;51(10):1757–60. doi:10.1007/s00125-008-1114-9.
Cook JP, Morris AP. Multi-ethnic genome-wide association study identifies novel locus for type 2 diabetes susceptibility. Eur J Human Genet: EJHG. 2016;24(8):1175–80. doi:10.1038/ejhg.2016.17.
Hebbring SJ. The challenges, advantages and future of phenome-wide association studies. Immunology. 2014;141(2):157–65. doi:10.1111/imm.12195.
Stancakova A, Paananen J, Soininen P, Kangas AJ, Bonnycastle LL, Morken MA, et al. Effects of 34 risk loci for type 2 diabetes or hyperglycemia on lipoprotein subclasses and their composition in 6,580 nondiabetic Finnish men. Diabetes. 2011;60(5):1608–16. doi:10.2337/db10-1655.
Roadmap Epigenomics C, Kundaje A, Meuleman W, Ernst J, Bilenky M, Yen A, et al. Integrative analysis of 111 reference human epigenomes. Nature. 2015;518(7539):317–30. doi:10.1038/nature14248.
Consortium GT. Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science. 2015;348(6235):648–60. doi:10.1126/science.1262110.
Furey TS. ChIP-seq and beyond: new and improved methodologies to detect and characterize protein-DNA interactions. Nat Rev Genet. 2012;13(12):840–52. doi:10.1038/nrg3306.
Buenrostro JD, Giresi PG, Zaba LC, Chang HY, Greenleaf WJ. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat Methods. 2013;10(12):1213–8. doi:10.1038/nmeth.2688.
Gaulton KJ, Nammo T, Pasquali L, Simon JM, Giresi PG, Fogarty MP, et al. A map of open chromatin in human pancreatic islets. Nat Genet. 2010;42(3):255–9. doi:10.1038/ng.530.
Consortium EP. An integrated encyclopedia of DNA elements in the human genome. Nature. 2012;489(7414):57–74. doi:10.1038/nature11247.
Mikkelsen TS, Xu Z, Zhang X, Wang L, Gimble JM, Lander ES, et al. Comparative epigenomic analysis of murine and human adipogenesis. Cell. 2010;143(1):156–69. doi:10.1016/j.cell.2010.09.006.
Parker SC, Stitzel ML, Taylor DL, Orozco JM, Erdos MR, Akiyama JA, et al. Chromatin stretch enhancer states drive cell-specific gene regulation and harbor human disease risk variants. Proc Natl Acad Sci U S A. 2013;110(44):17921–6. doi:10.1073/pnas.1317023110.
Scott LJ, Erdos MR, Huyghe JR, Welch RP, Beck AT, Wolford BN, et al. The genetic regulatory signature of type 2 diabetes in human skeletal muscle. Nat Commun. 2016;7:11764. doi:10.1038/ncomms11764.
Fogarty MP, Panhuis TM, Vadlamudi S, Buchkovich ML, Mohlke KL. Allele-specific transcriptional activity at type 2 diabetes-associated single nucleotide polymorphisms in regions of pancreatic islet open chromatin at the JAZF1 locus. Diabetes. 2013;62(5):1756–62. doi:10.2337/db12-0972.
Fogarty MP, Cannon ME, Vadlamudi S, Gaulton KJ, Mohlke KL. Identification of a regulatory variant that binds FOXA1 and FOXA2 at the CDC123/CAMK1D type 2 diabetes GWAS locus. PLoS Genet. 2014;10(9):e1004633. doi:10.1371/journal.pgen.1004633.
Stitzel ML, Sethupathy P, Pearson DS, Chines PS, Song L, Erdos MR, et al. Global epigenomic analysis of primary human pancreatic islets provides insights into type 2 diabetes susceptibility loci. Cell Metab. 2010;12(5):443–55. doi:10.1016/j.cmet.2010.09.012.
Varshney A, Scott LJ, Welch RP, Erdos MR, Chines PS, Narisu N, et al. Genetic regulatory signatures underlying islet gene expression and type 2 diabetes. Proc Natl Acad Sci U S A. 2017;114(9):2301–6. doi:10.1073/pnas.1621192114.
Horikoshi M, Pasquali L, Wiltshire S, Huyghe JR, Mahajan A, Asimit JL, et al. Transancestral fine-mapping of four type 2 diabetes susceptibility loci highlights potential causal regulatory mechanisms. Hum Mol Genet. 2016;25(10):2070–81. doi:10.1093/hmg/ddw048.
• Carrat GR, Hu M, Nguyen-Tu MS, Chabosseau P, Gaulton KJ, van de Bunt M, et al. Decreased STARD10 expression is associated with defective insulin secretion in humans and mice. Am J Hum Genet. 2017;100(2):238–56. doi:10.1016/j.ajhg.2017.01.011. This study comprehensively details a mechanism for a diabetes risk locus combining genetic fine-mapping, epigenome and expression QTL mapping, and animal models.
Kulzer JR, Stitzel ML, Morken MA, Huyghe JR, Fuchsberger C, Kuusisto J, et al. A common functional regulatory variant at a type 2 diabetes locus upregulates ARAP1 expression in the pancreatic beta cell. Am J Hum Genet. 2014;94(2):186–97. doi:10.1016/j.ajhg.2013.12.011.
Claussnitzer M, Dankel SN, Klocke B, Grallert H, Glunk V, Berulava T, et al. Leveraging cross-species transcription factor binding site patterns: from diabetes risk loci to disease mechanisms. Cell. 2014;156(1–2):343–58. doi:10.1016/j.cell.2013.10.058.
•• Claussnitzer M, Hui CC, Kellis M. FTO obesity variant and adipocyte browning in humans. N Engl J Med. 2016;374(2):192–3. doi:10.1056/NEJMc1513316. This study comprehensively details a mechanism for a diabetes risk locus using contemporary epigenomic and genome editing techniques.
Smemo S, Tena JJ, Kim KH, Gamazon ER, Sakabe NJ, Gomez-Marin C, et al. Obesity-associated variants within FTO form long-range functional connections with IRX3. Nature. 2014;507(7492):371–5. doi:10.1038/nature13138.
Stratigopoulos G, LeDuc CA, Cremona ML, Chung WK, Leibel RL. Cut-like homeobox 1 (CUX1) regulates expression of the fat mass and obesity-associated and retinitis pigmentosa GTPase regulator-interacting protein-1-like (RPGRIP1L) genes and coordinates leptin receptor signaling. J Biol Chem. 2011;286(3):2155–70. doi:10.1074/jbc.M110.188482.
Trynka G, Sandor C, Han B, Xu H, Stranger BE, Liu XS, et al. Chromatin marks identify critical cell types for fine mapping complex trait variants. Nat Genet. 2013;45(2):124–30. doi:10.1038/ng.2504.
Pickrell JK. Joint analysis of functional genomic data and genome-wide association studies of 18 human traits. Am J Hum Genet. 2014;94(4):559–73. doi:10.1016/j.ajhg.2014.03.004.
Kichaev G, Yang WY, Lindstrom S, Hormozdiari F, Eskin E, Price AL, et al. Integrating functional data to prioritize causal variants in statistical fine-mapping studies. PLoS Genet. 2014;10(10):e1004722. doi:10.1371/journal.pgen.1004722.
Kichaev G, Pasaniuc B. Leveraging functional-annotation data in trans-ethnic fine-mapping studies. Am J Hum Genet. 2015;97(2):260–71. doi:10.1016/j.ajhg.2015.06.007.
Mathelier A, Shi W, Wasserman WW. Identification of altered cis-regulatory elements in human disease. Trends Genet: TIG. 2015;31(2):67–76. doi:10.1016/j.tig.2014.12.003.
Maurano MT, Humbert R, Rynes E, Thurman RE, Haugen E, Wang H, et al. Systematic localization of common disease-associated variation in regulatory DNA. Science. 2012;337(6099):1190–5. doi:10.1126/science.1222794.
Gaffney DJ, McVicker G, Pai AA, Fondufe-Mittendorf YN, Lewellen N, Michelini K, et al. Controls of nucleosome positioning in the human genome. PLoS Genet. 2012;8(11):e1003036. doi:10.1371/journal.pgen.1003036.
Elliott HR, Shihab HA, Lockett GA, Holloway JW, McRae AF, Smith GD, et al. The role of DNA methylation in type 2 diabetes aetiology—using genotype as a causal anchor. Diabetes. 2017; doi:10.2337/db16-0874.
van de Bunt M, Gaulton KJ, Parts L, Moran I, Johnson PR, Lindgren CM, et al. The miRNA profile of human pancreatic islets and beta-cells and relationship to type 2 diabetes pathogenesis. PLoS One. 2013;8(1):e55272. doi:10.1371/journal.pone.0055272.
Li YI, van de Geijn B, Raj A, Knowles DA, Petti AA, Golan D, et al. RNA splicing is a primary link between genetic variation and disease. Science. 2016;352(6285):600–4. doi:10.1126/science.aad9417.
Wang A, Yue F, Li Y, Xie R, Harper T, Patel NA, et al. Epigenetic priming of enhancers predicts developmental competence of hESC-derived endodermal lineage intermediates. Cell Stem Cell. 2015;16(4):386–99. doi:10.1016/j.stem.2015.02.013.
Ackermann AM, Wang Z, Schug J, Naji A, Kaestner KH. Integration of ATAC-seq and RNA-seq identifies human alpha cell and beta cell signature genes. Mol Metab. 2016;5(3):233–44. doi:10.1016/j.molmet.2016.01.002.
McVicker G, van de Geijn B, Degner JF, Cain CE, Banovich NE, Raj A, et al. Identification of genetic variants that affect histone modifications in human cells. Science. 2013;342(6159):747–9. doi:10.1126/science.1242429.
Kilpinen H, Waszak SM, Gschwind AR, Raghav SK, Witwicki RM, Orioli A, et al. Coordinated effects of sequence variation on DNA binding, chromatin structure, and transcription. Science. 2013;342(6159):744–7. doi:10.1126/science.1242463.
Jolma A, Yan J, Whitington T, Toivonen J, Nitta KR, Rastas P, et al. DNA-binding specificities of human transcription factors. Cell. 2013;152(1–2):327–39. doi:10.1016/j.cell.2012.12.009.
Riley TR, Slattery M, Abe N, Rastogi C, Liu D, Mann RS, et al. SELEX-seq: a method for characterizing the complete repertoire of binding site preferences for transcription factor complexes. Methods Mol Biol. 2014;1196:255–78. doi:10.1007/978-1-4939-1242-1_16.
Arnold CD, Gerlach D, Stelzer C, Boryn LM, Rath M, Stark A. Genome-wide quantitative enhancer activity maps identified by STARR-seq. Science. 2013;339(6123):1074–7. doi:10.1126/science.1232542.
Lyssenko V, Lupi R, Marchetti P, Del Guerra S, Orho-Melander M, Almgren P, et al. Mechanisms by which common variants in the TCF7L2 gene increase risk of type 2 diabetes. J Clin Invest. 2007;117(8):2155–63. doi:10.1172/JCI30706.
Lappalainen T, Sammeth M, Friedlander MR, ‘t Hoen PA, Monlong J, Rivas MA, et al. Transcriptome and genome sequencing uncovers functional variation in humans. Nature. 2013;501(7468):506–11. doi:10.1038/nature12531.
Fadista J, Vikman P, Laakso EO, Mollet IG, Esguerra JL, Taneera J, et al. Global genomic and transcriptomic analysis of human pancreatic islets reveals novel genes influencing glucose metabolism. Proc Natl Acad Sci U S A. 2014;111(38):13924–9. doi:10.1073/pnas.1402665111.
Small KS, Hedman AK, Grundberg E, Nica AC, Thorleifsson G, Kong A, et al. Identification of an imprinted master trans regulator at the KLF14 locus related to multiple metabolic phenotypes. Nat Genet. 2011;43(6):561–4. doi:10.1038/ng.833.
van de Bunt M, Manning Fox JE, Dai X, Barrett A, Grey C, Li L, et al. Transcript expression data from human islets links regulatory signals from genome-wide association studies for type 2 diabetes and glycemic traits to their downstream effectors. PLoS Genet. 2015;11(12):e1005694. doi:10.1371/journal.pgen.1005694.
Giambartolomei C, Vukcevic D, Schadt EE, Franke L, Hingorani AD, Wallace C, et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 2014;10(5):e1004383. doi:10.1371/journal.pgen.1004383.
Locke JM, Hysenaj G, Wood AR, Weedon MN, Harries LW. Targeted allelic expression profiling in human islets identifies cis-regulatory effects for multiple variants identified by type 2 diabetes genome-wide association studies. Diabetes. 2015;64(4):1484–91. doi:10.2337/db14-0957.
Gusev A, Ko A, Shi H, Bhatia G, Chung W, Penninx BW, et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat Genet. 2016;48(3):245–52. doi:10.1038/ng.3506.
Lawlor N, George J, Bolisetty M, Kursawe R, Sun L, Sivakamasundari V, et al. Single-cell transcriptomes identify human islet cell signatures and reveal cell-type-specific expression changes in type 2 diabetes. Genome Res. 2017;27(2):208–22. doi:10.1101/gr.212720.116.
Hughes JR, Roberts N, McGowan S, Hay D, Giannoulatou E, Lynch M, et al. Analysis of hundreds of cis-regulatory landscapes at high resolution in a single, high-throughput experiment. Nat Genet. 2014;46(2):205–12. doi:10.1038/ng.2871.
Jin F, Li Y, Dixon JR, Selvaraj S, Ye Z, Lee AY, et al. A high-resolution map of the three-dimensional chromatin interactome in human cells. Nature. 2013;503(7475):290–4. doi:10.1038/nature12644.
Dixon JR, Selvaraj S, Yue F, Kim A, Li Y, Shen Y, et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature. 2012;485(7398):376–80. doi:10.1038/nature11082.
Grubert F, Zaugg JB, Kasowski M, Ursu O, Spacek DV, Martin AR, et al. Genetic control of chromatin states in humans involves local and distal chromosomal interactions. Cell. 2015;162(5):1051–65. doi:10.1016/j.cell.2015.07.048.
Selvaraj S, R Dixon J, Bansal V, Ren B. Whole-genome haplotype reconstruction using proximity-ligation and shotgun sequencing. Nat Biotechnol. 2013;31(12):1111–8. doi:10.1038/nbt.2728.
Bonnefond A, Clement N, Fawcett K, Yengo L, Vaillant E, Guillaume JL, et al. Rare MTNR1B variants impairing melatonin receptor 1B function contribute to type 2 diabetes. Nat Genet. 2012;44(3):297–301. doi:10.1038/ng.1053.
Flannick J, Thorleifsson G, Beer NL, Jacobs SB, Grarup N, Burtt NP, et al. Loss-of-function mutations in SLC30A8 protect against type 2 diabetes. Nat Genet. 2014;46(4):357–63. doi:10.1038/ng.2915.
Majithia AR, Flannick J, Shahinian P, Guo M, Bray MA, Fontanillas P, et al. Rare variants in PPARG with decreased activity in adipocyte differentiation are associated with increased risk of type 2 diabetes. Proc Natl Acad Sci U S A. 2014;111(36):13127–32. doi:10.1073/pnas.1410428111.
Pal A, Potjer TP, Thomsen SK, Ng HJ, Barrett A, Scharfmann R, et al. Loss-of-function mutations in the cell-cycle control gene CDKN2A impact on glucose homeostasis in humans. Diabetes. 2016;65(2):527–33. doi:10.2337/db15-0602.
Xia Q, Chesi A, Manduchi E, Johnston BT, Lu S, Leonard ME, et al. The type 2 diabetes presumed causal variant within TCF7L2 resides in an element that controls the expression of ACSL5. Diabetologia. 2016;59(11):2360–8. doi:10.1007/s00125-016-4077-2.
Rees MG, Wincovitch S, Schultz J, Waterstradt R, Beer NL, Baltrusch S, et al. Cellular characterisation of the GCKR P446L variant associated with type 2 diabetes risk. Diabetologia. 2012;55(1):114–22. doi:10.1007/s00125-011-2348-5.
Thomsen SK, Ceroni A, van de Bunt M, Burrows C, Barrett A, Scharfmann R, et al. Systematic functional characterization of candidate causal genes for type 2 diabetes risk variants. Diabetes. 2016;65(12):3805–11. doi:10.2337/db16-0361.
da Silva XG, Mondragon A, Sun G, Chen L, McGinty JA, French PM, et al. Abnormal glucose tolerance and insulin secretion in pancreas-specific Tcf7l2-null mice. Diabetologia. 2012;55(10):2667–76. doi:10.1007/s00125-012-2600-7.
Nicolson TJ, Bellomo EA, Wijesekara N, Loder MK, Baldwin JM, Gyulkhandanyan AV, et al. Insulin storage and glucose homeostasis in mice null for the granule zinc transporter ZnT8 and studies of the type 2 diabetes-associated variants. Diabetes. 2009;58(9):2070–83. doi:10.2337/db09-0551.
Pound LD, Sarkar SA, Benninger RK, Wang Y, Suwanichkul A, Shadoan MK, et al. Deletion of the mouse Slc30a8 gene encoding zinc transporter-8 results in impaired insulin secretion. Biochem J. 2009;421(3):371–6. doi:10.1042/BJ20090530.
Wijesekara N, Dai FF, Hardy AB, Giglou PR, Bhattacharjee A, Koshkin V, et al. Beta cell-specific Znt8 deletion in mice causes marked defects in insulin processing, crystallisation and secretion. Diabetologia. 2010;53(8):1656–68. doi:10.1007/s00125-010-1733-9.
Butler AA, Cone RD. Knockout studies defining different roles for melanocortin receptors in energy homeostasis. Ann N Y Acad Sci. 2003;994:240–5.
Yamada T, Ishihara H, Tamura A, Takahashi R, Yamaguchi S, Takei D, et al. WFS1-deficiency increases endoplasmic reticulum stress, impairs cell cycle progression and triggers the apoptotic pathway specifically in pancreatic beta-cells. Hum Mol Genet. 2006;15(10):1600–9. doi:10.1093/hmg/ddl081.
Tamemoto H, Kadowaki T, Tobe K, Yagi T, Sakura H, Hayakawa T, et al. Insulin resistance and growth retardation in mice lacking insulin receptor substrate-1. Nature. 1994;372(6502):182–6. doi:10.1038/372182a0.
Church C, Lee S, Bagg EA, McTaggart JS, Deacon R, Gerken T, et al. A mouse model for the metabolic effects of the human fat mass and obesity associated FTO gene. PLoS Genet. 2009;5(8):e1000599. doi:10.1371/journal.pgen.1000599.
Stratigopoulos G, Martin Carli JF, O'Day DR, Wang L, Leduc CA, Lanzano P, et al. Hypomorphism for RPGRIP1L, a ciliary gene vicinal to the FTO locus, causes increased adiposity in mice. Cell Metab. 2014;19(5):767–79. doi:10.1016/j.cmet.2014.04.009.
da Silva XG, Bellomo EA, McGinty JA, French PM, Rutter GA. Animal models of GWAS-identified type 2 diabetes genes. J Diabetes Res. 2013;2013:906590. doi:10.1155/2013/906590.
Savic D, Ye H, Aneas I, Park SY, Bell GI, Nobrega MA. Alterations in TCF7L2 expression define its role as a key regulator of glucose metabolism. Genome Res. 2011;21(9):1417–25. doi:10.1101/gr.123745.111.
Dooley K, Zon LI. Zebrafish: a model system for the study of human disease. Curr Opin Genet Dev. 2000;10(3):252–6.
Alfa RW, Kim SK. Using Drosophila to discover mechanisms underlying type 2 diabetes. Dis Model Mech. 2016;9(4):365–76. doi:10.1242/dmm.023887.
Grotz AK, Thomsen SK. Prioritizing causal genes at type 2 diabetes risk loci. Curr Diabetes Rep. 2017;in press.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Kyle Gaulton declares that he has no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by the author.
Additional information
This article is part of the Topical Collection on Genetics
Rights and permissions
About this article
Cite this article
Gaulton, K.J. Mechanisms of Type 2 Diabetes Risk Loci. Curr Diab Rep 17, 72 (2017). https://doi.org/10.1007/s11892-017-0908-x
Published:
DOI: https://doi.org/10.1007/s11892-017-0908-x