Abstract
Purpose of review
The cardiac conduction system, composed of pacemaker cells and conducting cardiomyocytes, orchestrates the propagation of electrical activity to synchronize heartbeats. The conduction system plays a crucial role in the development of cardiac arrhythmias. In the embryo, the cells of the conduction system derive from the same cardiac progenitors as the contractile cardiomyocytes and and the key question is how this choice is made during development.
Recent Findings
This review focuses on recent advances in developmental biology using the mouse as animal model to better understand the cellular origin and molecular regulations that control morphogenesis of the cardiac conduction system, including the latest findings in single-cell transcriptomics.
Summary
The conducting cell fate is acquired during development starting with pacemaking activity and last with the formation of a complex fast-conducting network. Cardiac conduction system morphogenesis is controlled by complex transcriptional and gene regulatory networks that differ in the components of the cardiac conduction system.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The heart, along with the digestive system, is the only organ capable of initiating its own contractions. Cardiac automaticity is made possible by the presence of a natural pacemaker within the heart, the Sinoatrial Node (SAN) [1], which spontaneously generate action potentials (AP). In mammals and birds, the AP is then transmitted to the rest of the heart by a complex network of specialized cardiomyocytes (CMs) forming the cardiac conduction system (CCS) [2]. The various components of the CCS sequentially propagate the electrical impulse throughout the heart, ensuring the coordination and stereotypy of the beats (Fig. 1).
The anatomy, conduction properties and molecular regulation of the development of the CCS. On the left, the anatomy and conduction properties of the different components of the murine or human CCS. The conduction velocity within the heart depends on the specific expression of connexins in different components of the CCS. On the right, interactions of known factors controlling the development of the VCS through the regulation of proliferation, recruitment and differentiation of conductive cells. SAN, sinus node; AVN, Atrioventriclar node; AVB, Atrioventricular bundle; RBB, Right bundle branch; LBB, Left bundle branch; PFs, Purkinje fibers
First, rapid propagation of the AP within the atria ensures synchronous contraction of both atria. The AP then reaches the unique electrical connection between the chambers: the atrioventricular node (AVN), where it is advantageously slowed, optimizing ventricular filling prior to contraction. The AP is then transmitted to the fast-conducting ventricular conduction system (VCS) or His-Purkinje. It passes through the His bundle or atrioventricular bundle (AVB), which then divides into the right and left bundle branches (RBB and LBB) on either side of the septum. The isolated RBB and LBB carry the AP along the interventricular septum to the apex and to the terminal part of the CCS: the Purkinje fiber (PF) network of each ventricle. Finally, the PF network conducts the AP along the ventricular free walls as it transmits the depolarization to the working myocardium, initiating ventricular systole from apex to base.
The electrophysiological characteristics of cardiomyocytes within the CCS result from the expression of a wide range of conduction-specific genes, including ion channels responsible for pacemaker activity and action potential characteristics, gap junctions responsible for conduction velocity, and low levels of contractile proteins [3,4,5,6]. Importantly, each compartment of the CCS is unique, with specific physiological and histological characteristics adapted to their role in cardiac conduction [7]. For example, both the SAN and the AVN rely on Hcn4, the main cation channel responsible for the Ifunny current, which underlies their automaticity [8, 9]. Furthermore, the heterogeneous conduction velocities observed in the different compartments of the CCS are made possible by the expression of a combination of gap junctions with different conductance. Specifically, the SAN and AVN, characterized by slower conduction, predominantly express connexins of low conductance, such as Cx45 and Cx30.2 (9pS) [3, 10]. In contrast, fast-conducting components like the atria, AVB, RRB LBB, LPF and RPF express Cx40, which forms high conductance gap junctions (~ 180pS) [3, 11] (Fig. 1).
Although CCS cells share characteristics with neurons, such as genetic markers, electrophysiological properties and electrical function [12], they are CMs. Indeed, retrospective clonal analysis, revealed a common cellular origin between conducting and contractile CMs [13, 14]. More precisely, the different compartments of the CCS show close lineage relationship with contractile cardiomyocytes from neighboring myocardium, suggesting that conductive and contractile properties are acquired during cardiac morphogenesis [15,16,17,18]. Thus, although all CCS compartments are integrated into a functional conducting pathway in the mature heart, their origin and developmental regulation largely differ.
Importantly, despite its small volume within the heart, the CCS plays a critical role in cardiac function and in the occurrence of cardiac arrhythmias [14, 19]. Understanding the precise regulation of CCS patterning during heart development is therefore of paramount importance. Due to the considerable influence of genetics, the mouse—which has a very similar morphology, function and molecular regulation of the CCS to that of humans [20, 21] – has become the animal model of choice for understanding the development of the CCS.
Genetic Control of SAN Development
The heart is the first functional organ in the embryo with transient calcium releases [22] and focal contractions starting as early as cardiac crescent stage [23, 24], and soon being replaced by peristaltic contractions. However, these first cardiac activities can initiate anywhere in the heart tube. A dominant pacemaker emerges at the inflow pole of the heart only after, in a region progressively enriched in Hcn4, which prefigures the emplacement of the SAN [25].
In the adult heart, SAN cells are characterized by the expression of a unique set of ion channels, i.e. Hcn4 and gap junctions, such as Cx45 and Cx30.2, which differs from the atrial myocardium that expresses Cx40, Cx43 and Nppa [26]. The SAN is divided into a large head and a tail forming a comma-shaped structure at the junction of the right atrium and superior vena cava [1]. Both parts of the SAN arise from a mesodermal pool of progenitors that diverges early from both the first and second heart fields (FHF and SHF) and that will also give rise to the sinus venosus (SV) [27,28,29,30]. The subdivision of the SAN is delineated by distinct gene programs with the expression of transcription factors Tbx18+/Nkx2-5− in the head region and Nkx2-5+/Tbx18− in the tail [30, 31]. Strengthening a bipartite model for SAN development, genetic tracing analyses using inducible Hcn4-CreERT2 mice lines indicate a progressive temporal activation of Hcn4 in the SAN, with early expression in progenitors from the tail (from E7.5) and later expression (from E8.5 onward) in the head [16].
The differentiation and maintenance of the pacemaker activity of the SAN is controlled by a gene regulatory network involving Tbx18, Tbx2, Shox2, Nkx2-5 and Isl1 [32, 33].
Illustrating the importance of Tbx18 for SAN development, inactivation of Tbx18 causes a hypoplastic SAN with absence of the head and reduced tail region [30].
At early stage of development, Short stature homeobox 2 (Shox2) gene expression is extended in the sinus venosus to cells presenting pacemaker activity and co-expressing Hcn4 and Nkx2-5 [34]. Later during development, pacemaker cells are restricted to the mature SAN under the control of a transcriptional balance between Shox2 and Nkx2-5 to regulate cell fate. Shox2 is not directly required for the pacemaker activity of the SAN but the conditional deletion of Shox2 in Nkx2-5 tail domain induces sick sinus syndrome in association with a loss of the junction between SAN and atria [34]. The T-box transcription factor Tbx3 is expressed in the developing conduction system and, similar to Shox2, acts as a transcription repressor of the working myocardium program by down regulating the expression of Gja5, Gja1 and Scn5A in pacemaker cells [35]. Conditional overexpression of Tbx3 converts atrial cells to a pacemaker phenotype characterized by a downregulation of Cx43 and upregulation of Hcn4 expression, which results in ectopic pacemakers [36]. SAN homeostasis in the adult is also tightly dependent on Tbx3 expression indicating its importance not only for pacemaker specification but also for maintenance of conduction system integrity [37]. In contrast to these transcriptional repressors, Isl1 encodes for a LIM-homeodomain transcription factor and serves as a positive transactivator for the expression of pacemaker specific genes like Tbx3, Shox2 and Hcn4 [38]. The conditional inactivation of Isl1 specifically in the SAN at different stages of development reveals that Isl1 affects the SAN size and induces bradycardia by playing a cell-autonomously role in proliferation and differentiation of pacemaker cells [26, 39]. Moreover, any disturbances of Nkx2-5 expression in the adult heart correlate with sinus diseases, showing that the regulation of this transcriptional cascade is also crucial for the maintenance of pacemaker phenotype [40]. Nkx2-5 is required to form the junction between tail and head. Indeed, inactivation of Nkx2-5 in the Shox2 domain does not hamper the development of the node but disturbs the atrial activation suggesting a precise role for this junction [41]. In summary, pacemaker cells originate from an independent mesodermal lineage that give rise to the sinus venosus and starts to synchronize heartbeats at E8.0. Defects in SAN development or dysfunction of pacemaker cells in either SAN compartment leads to bradycardia or sick sinus syndrome [42].
Genetic Control of AVN Development
The AVN is critical in slowing atrioventricular conduction, thereby introducing a necessary delay between atrial and ventricular contraction [43]. The AV delay functionally develops around E10, when the chamber-myocardium acquire a conductive phenotype [44,45,46], while the atrioventricular canal (AVC) and OFT, deprived of Cx40, remains slow [11]. As development progresses, the differentiation of the AVN is marked by the expression of Tbx3, Hcn4, Cx30.2 and Cx45, giving the AVN its pacemaker and slow conducting properties [10, 43, 47, 48]. In parallel, the reinforcement of electrical insulation through the development of fibrous tissues further isolates the AVN, leaving it as the only electrical communication between the atria and ventricles.
Genetic tracing using respectively Tbx2 or Hcn4 has shown that the AVN arises from the AVC [49, 50] and commit to a AVN fate by E10-E11 [18]. On the other hand, the annulus fibrosus arises from epithelial-mesenchymal transition (EMT) of epicardial cells at the AV junction [27].
The establishment of the AV junction relies on different signaling pathway on the right and left side. On the left side, both the transcriptional program of the AVC myocardium and establishment of the annulus fibrosus relies on Tbx2 [50]. Consequently, deletion of Tbx2 results in accessory pathways, detectable anatomically and functionally by optical mapping [51]. On the other side, the establishment of the right AV junction depends on Notch and Wnt signaling [52, 53]. Accessory pathways are detectable after Notch activation or in a Wnt loss of function (LOF) model while Wnt activation induces ectopic annulus fibrosus in ventricular myocardium [52]. Rescue experiments suggest that Notch-mediated ventricular pre-excitation results in part from downregulation of Wnt signaling [54]. Notch inhibition also regulates the AV conduction pathway by interfering with AVN maturation. Indeed, downregulation of Notch signaling leads to formation of a small AVN associated with an elevated conduction velocity between atria and ventricles [53]. However, Notch signaling does not affect the early development of the AVC or the expression of Tbx3.
The specification and development of nodal cells is very sensitive to Nkx2-5 and Tbx3 gene dosage during embryonic development [55,56,57,58]. Indeed, AVN progenitors are absent in Nkx2-5 null embryos, and a small AVN develops in Nkx2-5 haploinsufficient mice or after postnatal deletion of Nkx2-5 [56, 59]; likewise, Tbx3 hypomorphic or AVC-deleted alleles cause AV blocks and hypoplasia of the CS. Moreover, inducible deletion of Tbx3 in the adult heart causes AV blocks associated with a reduced number of AVN cells [37]. Interestingly, the severity of these AV blocks decreases after a few weeks, suggesting that the AV conduction is able to partially recover from adult depletion of Tbx3 [37]. AVN development is also specifically impaired after ventricular deletion of Gata6 by direct inhibition of its target gene Id2 [60]. Inactivation of Id2, encoding a transcriptional repressor, causes abnormal development of the AVN by perturbing the cell cycle and Hcn4 expression. As mentioned above, the maturation of the AVN is also important in the development of AV conduction pathway and this step is disturbed in MyoR mutants by interfering with Cx30.2 expression through a direct interaction with Gata4 [61].
Defective development of the AVN is associated with AV electrical insulation disturbances or AV delay that cause AV accessory pathways or AV blocks.
Genetic Control of VCS Development
Before the emergence of a specific ventricular conduction system, cardiac conduction follows a unidirectional path from the inflow tract to the LV, RV and finally outflow tract (OFT). The mature activation pattern, characterized by two parallel apex-to-base activation waves, only emerges after ventricular septation and the development of a fast-conducting compartment in the subendocardial myocardium: the Cx40 positive trabeculae [11, 45, 46, 62, 63]. Subsequently, the gradual restriction of Cx40 expression to the future LBB, RBB and PF coincides with their maturation, eventually giving rise to the well-defined VCS [11, 14].
The proximal VCS – including AVB, RBB and LBB—arises from the primary interventricular ring (PIR) [64]. Interestingly, despite the close proximity and similarities of the BB and PF networks in the adult heart, the PF network does not originate from the PIR like the BB [64]. Instead the PF network arises from the Cx40+ trabeculae [14, 65], which is consistent with the conductive function ensured by the trabeculae during development [45, 46, 62, 63]. Furthermore, the molecular signature of the trabeculae closely resembles that of mature PFs, and the progressive restriction of trabecular markers, including Cx40, Nppa, Irx3, Etv1, and Sema3a, during compaction reflects the maturation process of the PF network [11, 14, 65,66,67,68,69].
The timing of segregation also varies between the components of the VCS, occurring later in distal parts of the VCS. The AVB segregates the earliest, prior to E7.5, as shown by clonal analysis of SMA+ cells [70]. The segregation to the LBB starts prior to E7.5 [70] and extends after E10.5 [64]. Finally, the PF network grows by recruitment of new conductive cells in successive waves throughout development [65, 71]. The first committed PFs can be observed as early as the linear heart tube stage (~ E7.5) and likely define a scaffold for the formation of the mature VCS [65]. Interestingly, bipotent trabecular progenitors, which can give rise to both contractile CMs or be recruited to a PF fate, persist in the trabeculae until the time of birth. These bipotent progenitors are responsible for a second wave of PF recruitment that is key to increasing the complexity and density of the PF network [65]. After E14.5, the bipotent pool decreases dramatically and disappears by birth, marking the final segregation of contractile and conductive lineages [65]. Recently, scRNAseq at fetal stages (E16.5) identified two sub-clusters of immature PFs, one of which expresses intermediate levels of PF genes (Gja5, Etv1, Sema3a, etc.) [72]. This demonstrates heterogeneity in the level of differentiation among PFs, further confirming that conducting cells are recruited sequentially.
The early development of the LBB, like other derivatives from the PIR, is primarily regulated by Tbx3. At these early time points, Tbx3 refrains a fast conduction phenotype [57, 58], including through the inhibition of Cx40 expression. This is only around E14.5 that a gene regulatory network including Nkx2-5, Tbx5, Id2 is activated in the AVB and BB, favorizing the expression of the fast-conducting proteins including Cx40 and Nav1-5 [58, 73, 74]. Importantly, mutations of Tbx5 in these VCS components results in spontaneous ventricular tachycardia, lethal arrhythmias [73] and AV blocs [74].
The proximity of the VCS with the endocardium suggests an instructive role of this tissue in the development of the VCS. Nrg1, which is expressed by endocardial cells during development, is sufficient to convert immature CMs to a conductive phenotype without impacting proliferation [75, 76]. Nrg1 functions through the Ras MAPK pathway to activate Etv1, the most enriched TF in the developing and mature VCS [67, 76]. Consistent with the role of Nrg1 as a ventricular conduction system differentiation driver, its ectopic overactivation in embryonic CMs upregulates a subset of His-Purkinje–specific genes [77]. Additionally, other factors may predispose subendocardial CMs to be Nrg1 responsive [75], since, even under homogenous addition of Nrg1 in organ culture, conductive conversion occurs preferentially in the subendocardial myocardium. Furthermore, responsiveness to Nrg1 decreases as development proceeds accompanying the progressive segregation of conductive and contractile lineages. After conductive recruitment, Nrg1 appears to play a role in the late differentiation of PFs, as suggested by the conduction abnormalities observed following post-natal treatment with Nrg1-antagonist [76] (Fig. 1).
In parallel, Notch signaling also regulates conductive recruitment. Myocardial overactivation of Notch signaling, either in vivo throughout development, or transiently in neonatal CMs, is sufficient to induce the acquisition of a conductive phenotype by a subset of CMs without affecting their proliferation [53]. Notch induces the upregulation of both fast conducting proteins, Cx40 and Nav1-5, and nodal proteins, Cx30.2 and Hcn1, leading to the acquisition of PFs electrophysiological properties by former working CMs and thereby, to VCS hyperplasia. Interestingly, only subendocardial CMs are responsive to Notch overactivation, suggesting that endocardial derived cues, such as Nrg1, could cooperate with Notch to pattern the VCS in subendocardial regions. Conversely, CHF1/Hey2, seems to repress conductive fate in the surrounding working myocardium and consistent with the observation of VCS hyperplasia in CHF1/Hey2 KO mice [78]. Thus, Notch signaling regulates PFs patterning both by promoting a conductive fate within the VCS and by repressing conductive conversion in the neighboring working myocardium. Interestingly, conductive conversion under Notch overactivation, in the neonatal period, uncovers a conserved plasticity between conductive and contractile fates [53].
ETS Variant Transcription Factor 1 (Etv1) is directly activated by Nrg1 signaling and is necessary and sufficient to instruct a conductive phenotype. Indeed, Etv1 KO mice display a hypoplastic VCS, especially in the mid and apical PF network where the number of characteristic ellipses is reduced [76]. Conversely, overexpression of Etv1 in neonatal CMs or human induced pluripotent stem cell-derived cardiomyocytes (hiPSC-CMs) is sufficient to induce a conductive phenotype [67, 76]. Etv1 also regulates the expression of Cx40 and Scn5a in PFs, and thereby their electrophysiological properties. Thus, the Nrg1 – Etv1 axis functions as an upstream determinant of PF commitment and differentiation [67, 75, 76].
Part of the function of Etv1 appears to be mediated by the transcription factor Nkx2-5 [67, 76]. Nkx2-5 expression is up-regulated in the developing His bundle, bundle branches and PFs in a spatiotemporal correspondence with the recruitment of conductive cells [56, 67, 72, 76, 79, 80]. This upregulation is necessary for a conversion to a conductive fate as attested by hypoplasia of the AVN, His bundle, LBB and PF network in Nkx2-5 haploinsufficient mice [56, 59, 65, 81, 82]. Interestingly, the late recruitment of PFs is more sensitive to Nkx2-5 dosages than the early one. Thus, while the BB and the primitive scaffold are minorly affected in the Nkx2-5 haploinsufficient mice, the mid and apical PF network is dramatically hypoplastic, with a reduction of two to three fold in the number of ellipses [56, 59, 65, 81, 82]. Surprisingly, the differentiation of the remaining PFs in Nkx2-5 mutant doesn’t seem to be affected, as their electrophysiological and Cx40 gap junctions number appears normal. However, other studies report that Nkx2-5 directly and indirectly – in part by upregulating of the homeodomain protein HOP – regulates a number of gap junctions and ion channels, including Cx40 [83]. Consistent with this, some cells with intermediate maturation have been identified in Nkx2-5 haploinsufficient mice by their expression of EH-myomesin without Cx40 [81]. Finally, Nkx2-5 is implicated in the maintenance of a conductive phenotype, as deletion of one copy of Nkx2-5 at birth leads to a progressive loss of fast conduction proteins, including Cx40 and Hcn4 [84]. Nkx2-5 expression in conductive cells is also regulated by the co-repression of the homeobox transcription factor prospero-related homeobox protein 1 (Prox1) and HDAC3 [85].
In cooperation with Nkx2-5, Tbx5 and Irx3 orchestrate the recruitment and specification of PFs and can thus partially compensate for low levels of Nkx2-5 in some conditions (Fig. 1).
Irx3, which is expressed in the immature proximal VCS and in a subset of trabeculae during development, is restricted to the VCS of the adult heart. Similar to Nkx2-5 haploinsufficient mice, Irx3 KO mice develop hypoplastic bundle branches and PF network, defects that manifest mainly in the postnatal period [86]. Furthermore, Irx3 regulates gap junction expression in PFs by indirectly upregulating Cx40 and directly inhibiting Cx43 expression in competition with Nkx2-5 [68]. As a result, Irx3 KO mice express only half of the WT levels of Cx40 in their remaining VCS, while their LBB ectopically express Cx43, resulting in abnormal contact between the LBB and surrounding myocardium.
Similar to Nkx2-5, Tbx5 has a pleiotropic role in cardiac development causing embryonic lethality in Tbx5−/− mice and multiple cardiac defects in Tbx5 haploinsufficient mice [87]. Tbx5 expression is enriched in the mature VCS compared to the surrounding myocardium and maximal levels of Tbx5 are required for the recruitment of conductive cells in the His bundle and bundle branches [67, 74, 76]. Consequently, Tbx5 haploinsufficient mice have a hypoplastic His bundle, and immature bundle branches. In addition, Tbx5 cooperates with Nkx2-5 to synergistically activate Cx40 and Nppa expression [87]. To date, the role of Tbx5 specifically in PFs has not been investigated. The cooperative role of Nkx2-5, Tbx5 and Irx3 is illustrated by their direct physical interactions and the presence of neighboring binding sites for all three TFs in the regulatory regions of a large number of target genes including cyclin-dependent kinase inhibitors [86]. Furthermore, compensatory upregulation of Nkx2-5 and Tbx5 has been reported in Irx3−/− mice, while double or triple KO mice show increasing defects in VCS patterning [86, 88].
miR-1, a direct target of Nkx2-5, is the most abundant microRNA in the postnatal heart and plays a pleiotropic role in regulating CM electrophysiology, proliferation and VCS development [89,90,91]. During development, miR-1 represses CM proliferation through a Cdk6 /Pocket protein axis, resulting in control of myocardial and VCS growth [90]. Consequently, premature expression of miR-1 in CMs results in reduced CM proliferation and hypoplasia of the PF network. Conversely, Pocket proteins 3KO (p107−/−, p130−/−, Rbdel/del) mice have hypertrabeculated ventricles due to uncontrolled CM proliferation in both the compact layer and trabeculae resulting in thickening of the His Purkinje system [92]. These results show that increasing or decreasing the proliferation of the trabeculae, which contain the pool of PF progenitors, is sufficient to increase or decrease the production of PFs [90, 92].
Hand1 is expressed in the LV from E8.5, and, together with Hand2, regulates trabecular identity and represses trabeculae proliferation [93]. Mice lacking Hand1 expression in the LV (Hand1ΔLV/ΔLV) develop a hyperplastic VCS, which may be due to increased trabecular proliferation [94]. However, both the left and right VCS are hyperplastic in Hand1ΔLV/ΔLV mice, even if Hand1 is only expressed in the LV of developing hearts. This, together with the observation that Hand1 is expressed in the His bundle, RBB, LBB, left PFs and, potentially right PFs in mature hearts [94], suggests that Hand1 may have an additional role in PF recruitment or differentiation (Fig. 1).
PFs specifically express three proteins of the immunoglobulin family: Cntn2, Ncam-1 and Alcam [95, 96]. LOF models indicate that Ncam-1 KO mice have a hypoplastic VCS, especially at the apex, similar to of Etv1 KO, Irx3 KO or Nkx2-5 heterozygous mice, supporting a role for Ncam-1 in VCS patterning. In addition, polysialylated Ncam-1 is essential for PF differentiation by controlling the localization of Cntn2, Ncam-1, Cx40, components of the desmosomes and adherent junctions at the intercalated discs. It is interesting that an adhesion molecule can play a role in the patterning of the VCS and may reveal a feedback loop between electromechanical coupling in the PFs and PF commitment and differentiation. Furthermore, this may explain why some mutant mice develop hypoplasia of the VCS specifically in the neonatal period [81, 86], when the maturation of the intercalated discs takes place (Fig. 1).
Novel Insights Into the Differentiation Trajectories of CCS
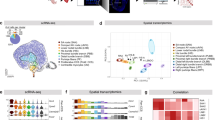
Recent advancements in single-cell transcriptomic technologies, including single-nucleus RNA sequencing (snRNA-seq) and spatial transcriptomics (STx), have significantly enhanced our understanding of the transcriptomic landscape of the CCS, improving our ability to capture cell-state heterogeneity and define developmental trajectories underlying CCS differentiation. In 2019, single-cell RNA-seq (scRNA-seq) analysis on microdissected mouse embryonic hearts, including the sinoatrial node (SAN), atrioventricular node (AVN), His bundle, and Purkinje fibers (PF), provided the first comprehensive single-cell transcriptional profiling of the developing CCS [72]. Unsupervised cell clustering along with gene enrichment analysis identified novel conductive cell markers (Igfbp5, Cpne5, Smoc2, Rgs6, Ntm) and revealed the existence of rare conduction cell subtypes in each CCS component [72]. Direct comparison of embryonic and postnatal scRNA-seq datasets with postnatal stages highlighted high transcriptional similarities between fetal and neonatal stages suggesting that CCS cell fate, particularly for the SAN, cAVN and proximal VCS, may largely occur by E16.5 [97]. Cell-state heterogeneity of the CCS, particularly of the SAN, has also been reported in the adult murine heart, including a core cell cluster functionally related to the regulation of heart rate. Among the canonical SAN markers, Vsnl1 has been identified as a novel SAN gene whose expression regulates the beating rate of hiPSC-CMs and mouse hearts [98].
The combination of single-cell analysis with unbiased, transcriptome-wide spatial gene expression information is now opening new horizons in understanding cardiac cell states in relation to their anatomic location within the different cardiac compartments. Integration of scRNA-seq and spatially resolved transcriptomic analysis in the developing murine heart has identified a novel cardiomyocyte population expressing dopamine beta-hydroxylase (Dbh + CMs), which is transcriptionally and functionally associated with the CCS in developing and mature murine hearts [99].
More recently, spatiotemporal resolved scRNA-seq analysis of the developing human heart has provided novel insights into the differentiation of the CCS [100, 101]. In particular, a detailed molecular characterization of conductive cardiomyocytes in the developing CCS compartments highlights their distinct electrophysiological properties in close relationship with spatial and functional associations with other cell types, including specialized fibroblast cell states in the nodes, endocardial cells in the ventricular conduction system, and neurons in the developing autonomic innervation [100]. The increasing generation of new multiomic datasets in both, CCS development and disease, will be instrumental for gaining novel insights into CCS development and translate this information to identify novel therapeutic approaches to treat cardiac arrhythmias.
Conclusions
The CCS is a complex tissue with its different compartments having specific functions and thus, specific electrophysiological properties in the adult. Each compartment arises from different populations and its development is regulated by distinct gene regulatory networks. The development of the CCS involves the proliferation of progenitors, the recruitment to a conductive fate, the acquisition and the maintenance of conductive properties. Though these processes are in theory independent, many factors have a pleiotropic role in the regulation of several of these steps. Dysregulation of any of these processes can result in mispatterning of one or several components of the CCS, such as hypoplasia. Importantly, all model of CCS hypoplasia—Nkx2-5+/-, Tbx5+/-, Irx3−/−, Id2−/−, Etv1−/−, Ncam-1−/− mice—present slowed conduction in the affected compartment(s) [59, 68, 74, 76, 81, 86, 96, 102,103,104]. Moreover, abnormal electrophysiological properties of conductive cells, caused by abnormal gap junction or ion channel content, can also, in combination with patterning defects or alone, result in perturbed conduction [84, 104, 105].
Data Availability
No datasets were generated or analysed during the current study.
References
Boyett MR. “And the beat goes on” the cardiac conduction system: The wiring system of the heart. Exp Physiol. 2009;94(10):1035–49. https://doi.org/10.1113/expphysiol.2009.046920.
Anderson RH, Boyett MR, Dobrzynski H, Moorman AFM. The anatomy of the conduction system: Implications for the clinical cardiologist. J Cardiovasc Transl Res. 2013;6(2):187–96. https://doi.org/10.1007/s12265-012-9433-0.
Gros DB, Jongsma HJ. Connexins in mammalian heart function. BioEssays. 1996;18(9):719–30. https://doi.org/10.1002/bies.950180907.
van Kempen MJA, Ten VI, Wessels A, et al. Differential connexin distribution accommodates cardiac function in different species. Microsc Res Tech. 1995;31(5):420–36. https://doi.org/10.1002/jemt.1070310511.
Marionneau C, Couette B, Liu J, et al. Specific pattern of ionic channel gene expression associated with pacemaker activity in the mouse heart. J Physiol. 2005;562(1):223–34. https://doi.org/10.1113/jphysiol.2004.074047.
Gaborit N, Le Bouter S, Szuts V, et al. Regional and tissue specific transcript signatures of ion channel genes in the non-diseased human heart. J Physiol. 2007;582(2):675–93. https://doi.org/10.1113/jphysiol.2006.126714.
Dobrzynski H, Anderson RH, Atkinson A, et al. Structure, function and clinical relevance of the cardiac conduction system, including the atrioventricular ring and outflow tract tissues. Pharmacol Ther. 2013;139(2):260–88. https://doi.org/10.1016/j.pharmthera.2013.04.010.
Mangoni ME, Nargeot J. Genesis and regulation of the heart automaticity. Physiol Rev. 2008;88(3):919–82. https://doi.org/10.1152/physrev.00018.2007.
Difrancesco D. The role of the funny current in pacemaker activity. Circ Res. 2010;106(3):434–46. https://doi.org/10.1161/CIRCRESAHA.109.208041.
Kreuzberg MM, Willecke K, Bukauskas FF. Connexin-Mediated Cardiac Impulse Propagation: Connexin 30.2 Slows Atrioventricular Conduction in Mouse Heart. Trends Cardiovasc Med. 2006;16(8):266–72. https://doi.org/10.1016/j.tcm.2006.05.002.
Delorme B, Dahl E, Jarry-guichard T, et al. Developmental regulation of connexin 40 gene expression in mouse heart correlates with the differentiation of the conduction system. Dev Dyn. 1995;204(4):358–71. https://doi.org/10.1002/aja.1002040403.
Gourdie RG, Harris BS, Bond J, et al. Development of the cardiac pacemaking and conduction system. Birth Defects Res Part C Embryo Today Rev. 2003;69(1):46–57. https://doi.org/10.1002/bdrc.10008.
Gourdie RG, Mima T, Thompson RP, Mikawa T. Terminal diversification of the myocyte lineage generates Purkinje fibers of the cardiac conduction system. Development. 1995;121(5):1423–31. https://doi.org/10.1242/dev.121.5.1423.
Miquerol L, Moreno-Rascon N, Beyer S, et al. Biphasic development of the mammalian ventricular conduction system. Circ Res. 2010;107(1):153–61. https://doi.org/10.1161/CIRCRESAHA.110.218156.
Miquerol L, Bellon A, Moreno N, et al. Resolving cell lineage contributions to the ventricular conduction system with a Cx40-GFP allele: A dual contribution of the first and second heart fields. Dev Dyn. 2013;242(6):665–77. https://doi.org/10.1002/dvdy.23964.
Später D, Abramczuk MK, Buac K, et al. A HCN4+ cardiomyogenic progenitor derived from the first heart field and human pluripotent stem cells. Nat Cell Biol. 2013;15(9):1098–106. https://doi.org/10.1038/ncb2824.
Liang X, Evans SM, Sun Y. Insights into cardiac conduction system formation provided by HCN4 expression. Trends Cardiovasc Med. 2015;25(1):1–9. https://doi.org/10.1016/j.tcm.2014.08.009.
Liang X, Wang G, Lin L, et al. HCN4 dynamically marks the first heart field and conduction system precursors. Circ Res. 2013;113(4):399–407. https://doi.org/10.1161/CIRCRESAHA.113.301588.
Haïssaguerre M, Shah DC, Jaïs P, et al. Role of Purkinje conducting system in triggering of idiopathic ventricular fibrillation. Lancet. 2002;359(9307):677–8. https://doi.org/10.1016/S0140-6736(02)07807-8.
Durrer D, Van Dam RT, Freud GE, Janse MJ, Meijler FL, Arzbaecher RC. Total Excitation of the Isolated Human Heart. Circulation. 1970;41(6):899–912. https://doi.org/10.1161/01.CIR.41.6.899.
van Eif VWW, Stefanovic S, Mohan RA, Christoffels VM. Gradual differentiation and confinement of the cardiac conduction system as indicated by marker gene expression. Biochim Biophys Acta - Mol Cell Res. 2020;1867(3):118509. https://doi.org/10.1016/j.bbamcr.2019.07.004.
Kobayashi T, Maeda S, Ichise N, et al. The beginning of the calcium transient in rat embryonic heart. J Physiol Sci. 2011;61(2):141–9. https://doi.org/10.1007/s12576-010-0131-x.
Nishii K, Shibata Y. Mode and determination of the initial contraction stage in the mouse embryo heart. Anat Embryol (Berl). 2006;211(2):95–100. https://doi.org/10.1007/s00429-005-0065-x.
Chen CM, Miranda AMA, Bub G, Srinivas S. Detecting cardiac contractile activity in the early mouse embryo using multiple modalities. Front Physiol. 2015;6(JAN):1–9. https://doi.org/10.3389/fphys.2014.00508.
Stieber J, Herrmann S, Feil S, et al. The hyperpolarization-activated channel HCN4 is required for the generation of pacemaker action potentials in the embryonic heart. Proc Natl Acad Sci U S A. 2003;100(25):15235–40. https://doi.org/10.1073/pnas.2434235100.
Vedantham V, Galang G, Evangelista M, Deo RC, Srivastava D. RNA Sequencing of Mouse Sinoatrial Node Reveals an Upstream Regulatory Role for Islet-1 in Cardiac Pacemaker Cells. Circ Res. 2015;116(5):797–803. https://doi.org/10.1161/CIRCRESAHA.116.305913.
Cai C-L, Liang X, Shi Y, et al. Isl1 Identifies a Cardiac Progenitor Population that Proliferates Prior to Differentiation and Contributes a Majority of Cells to the Heart. Dev Cell. 2003;5(6):877–89. https://doi.org/10.1016/S1534-5807(03)00363-0.
Mommersteeg MTM, Domínguez JN, Wiese C, et al. The sinus venosus progenitors separate and diversify from the first and second heart fields early in development. Cardiovasc Res. 2010;87(1):92–101. https://doi.org/10.1093/cvr/cvq033.
Lescroart F, Mohun T, Meilhac SM, Bennett M, Buckingham M. Lineage tree for the venous pole of the heart: Clonal analysis clarifies controversial genealogy based on genetic tracing. Circ Res. 2012;111(10):1313–22. https://doi.org/10.1161/CIRCRESAHA.112.271064.
Wiese C, Grieskamp T, Airik R, et al. Formation of the sinus node head and differentiation of sinus node myocardium are independently regulated by Tbx18 and Tbx3. Circ Res. 2009;104(3):388–97. https://doi.org/10.1161/CIRCRESAHA.108.187062.
Christoffels VM, Mommersteeg MTM, Trowe MO, et al. Formation of the venous pole of the heart from an Nkx2-5-negative precursor population requires Tbx18. Circ Res. 2006;98(12):1555–63. https://doi.org/10.1161/01.RES.0000227571.84189.65.
Ye W, Song Y, Huang Z, Zhang Y, Chen Y. Genetic regulation of sinoatrial node development and pacemaker program in the venous pole. J Cardiovasc Dev Dis. 2015;2(4):282–98. https://doi.org/10.3390/jcdd2040282.
Christoffels VM, Smits GJ, Kispert A, Moorman AFM. Development of the pacemaker tissues of the heart. Circ Res. 2010;106(2):240–54. https://doi.org/10.1161/CIRCRESAHA.109.205419.
Ye W, Wang J, Song Y, et al. A common Shox2–nkx2-5 antagonistic mechanism primes the pacemaker cell fate in the pulmonary vein myocardium and sinoatrial node. Dev. 2015;142(14):2521–32. https://doi.org/10.1242/dev.120220.
Bakker ML, Boink GJJ, Boukens BJ, et al. T-box transcription factor TBX3 reprogrammes mature cardiac myocytes into pacemaker-like cells. Cardiovasc Res. 2012;94(3):439–49. https://doi.org/10.1093/cvr/cvs120.
Hoogaars WMH, Engel A, Brons JF, et al. Tbx3 controls the sinoatrial node gene program and imposes pacemaker function on the atria. Genes Dev. 2007;21(9):1098–112. https://doi.org/10.1101/gad.416007.
Frank DU, Carter KL, Thomas KR, et al. Lethal arrhythmias in Tbx3 -deficient mice reveal extreme dosage sensitivity of cardiac conduction system function and homeostasis. Proc Natl Acad Sci. 2012;109(3). https://doi.org/10.1073/pnas.1115165109
Liang X, Zhang Q, Cattaneo P, et al. Transcription factor ISL1 is essential for pacemaker development and function. J Clin Invest. 2015;125(8):3256–68. https://doi.org/10.1172/JCI68257.
Sun Y, Liang X, Najafi N, et al. Islet 1 is expressed in distinct cardiovascular lineages, including pacemaker and coronary vascular cells. Dev Biol. 2007;304(1):286–96. https://doi.org/10.1016/j.ydbio.2006.12.048.
Nakashima Y, Yanez DA, Touma M, et al. Nkx2-5 suppresses the proliferation of atrial myocytes and conduction system. Circ Res. 2014;114(7):1103–13. https://doi.org/10.1161/CIRCRESAHA.114.303219.
Li H, Li D, Wang Y, et al. Nkx2-5 defines a subpopulation of pacemaker cells and is essential for the physiological function of the sinoatrial node in mice. Dev. 2019;146(14):1–9. https://doi.org/10.1242/dev.178145.
Vasnath V. New Approaches to Biological Pacemakers: Links to Sinoatrial Node Development. Trends Mol Med. 2015;21(12):773–9. https://doi.org/10.1016/j.molmed.2015.10.002.New.
Bakker ML, Moorman AFM, Christoffels VM. The Atrioventricular Node: Origin, Development, and Genetic Program. Trends Cardiovasc Med. 2010;20(5):164–71. https://doi.org/10.1016/j.tcm.2011.02.001.
Chen F, De Diego C, Chang MG, et al. Atrioventricular conduction and arrhythmias at the initiation of beating in embryonic mouse hearts. Dev Dyn. 2010;239(7):1941–9. https://doi.org/10.1002/dvdy.22319.
Christoffels VM, Moorman AFM. Development of the cardiac conduction system why are some regions of the heart more arrhythmogenic than others? Circ Arrhythmia Electrophysiol. 2009;2(2):195–207. https://doi.org/10.1161/CIRCEP.108.829341.
Sankova B, Benes J, Krejci E, et al. The effect of connexin40 deficiency on ventricular conduction system function during development. Cardiovasc Res. 2012;95(4):469–79. https://doi.org/10.1093/cvr/cvs210.
Horsthuis T, Buermans HPJ, Brons JF, et al. Gene expression profiling of the forming atrioventricular node using a novel Tbx3-based node-specific transgenic reporter. Circ Res. 2009;105(1):61–9. https://doi.org/10.1161/CIRCRESAHA.108.192443.
Frank M, Wirth A, Andrié RP, et al. Connexin45 provides optimal atrioventricular nodal conduction in the adult mouse heart. Circ Res. 2012;111(12):1528–38. https://doi.org/10.1161/CIRCRESAHA.112.270561.
Aanhaanen WTJ, Mommersteeg MTM, Norden J, et al. Developmental Origin, Growth, and Three-Dimensional Architecture of the Atrioventricular Conduction Axis of the Mouse Heart. Circ Res. 2010;107(6):728–36. https://doi.org/10.1161/CIRCRESAHA.110.222992.
Aanhaanen WTJ, Brons JF, Domínguez JN, et al. The Tbx2+ primary myocardium of the atrioventricular canal forms the atrioventricular node and the base of the left ventricle. Circ Res. 2009;104(11):1267–74. https://doi.org/10.1161/CIRCRESAHA.108.192450.
Aanhaanen WTJ, Boukens BJD, Sizarov A, et al. Defective Tbx2-dependent patterning of the atrioventricular canal myocardium causes accessory pathway formation in mice. J Clin Invest. 2011;121(2):534–44. https://doi.org/10.1172/JCI44350.
Gillers BS, Chiplunkar A, Aly H, et al. Canonical Wnt signaling regulates atrioventricular junction programming and electrophysiological properties. Circ Res. 2014;116(3):398–406. https://doi.org/10.1161/CIRCRESAHA.116.304731.
Rentschler S, Yen AH, Lu J, et al. Myocardial notch signaling reprograms cardiomyocytes to a conduction-like phenotype. Circulation. 2012;126(9):1058–66. https://doi.org/10.1161/CIRCULATIONAHA.112.103390.
Gomes J, Finlay M, Ahmed AK, et al. Electrophysiological abnormalities precede overt structural changes in arrhythmogenic right ventricular cardiomyopathy due to mutations in desmoplakin-A combined murine and human study. Eur Heart J. 2012;33(15):1942–53. https://doi.org/10.1093/eurheartj/ehr472.
Bakker ML, Boukens BJ, Mommersteeg MTM, et al. Transcription factor Tbx3 is required for the specification of the atrioventricular conduction system. Circ Res. 2008;102(11):1340–9. https://doi.org/10.1161/CIRCRESAHA.107.169565.
Pashmforoush M, Lu JT, Chen H, et al. Nkx2-5 Pathways and Congenital Heart Disease. Cell. 2004;117(3):373–86. https://doi.org/10.1016/S0092-8674(04)00405-2.
Hoogaars WMH, Tessari A, Moorman AFM, et al. The transcriptional repressor Tbx3 delineates the developing central conduction system of the heart. Cardiovasc Res. 2004;62(3):489–99. https://doi.org/10.1016/j.cardiores.2004.01.030.
Bhattacharyya S, Munshi NV. Development of the cardiac conduction system. Cold Spring Harb Perspect Biol. 2020;12(12):1–21. https://doi.org/10.1101/cshperspect.a037408.
Jay PY, Harris BS, Maguire CT, et al. Nkx2-5 mutation causes anatomic hypoplasia of the cardiac conduction system. J Clin Invest. 2004;113(8):1130–7. https://doi.org/10.1172/JCI19846.
Liu F, Lu MM, Patel NN, Schillinger KJ, Wang T, Patel VV. GATA-Binding Factor 6 Contributes to Atrioventricular Node Development and Function. Circ Cardiovasc Genet. 2015;8(2):284–93. https://doi.org/10.1161/CIRCGENETICS.113.000587.
Harris JP, Bhakta M, Bezprozvannaya S, et al. MyoR Modulates Cardiac Conduction by Repressing Gata4. Mol Cell Biol. 2015;35(4):649–61. https://doi.org/10.1128/mcb.00860-14.
Olejníčková V, Šaňková B, Sedmera D, Janáček J. Trabecular architecture determines impulse propagation through the early embryonic mouse heart. Front Physiol. 2019;10(JAN):1–12. https://doi.org/10.3389/fphys.2018.01876.
Olejnickova V, Kocka M, Kvasilova A, et al. Gap Junctional Communication via Connexin43 between Purkinje Fibers and Working Myocytes Explains the Epicardial Activation Pattern in the Postnatal Mouse Left Ventricle. 2021.
Mohan RA, Mommersteeg MTM, Domínguez JN, et al. Embryonic Tbx3+ cardiomyocytes form the mature cardiac conduction system by progressive fate restriction. Development. 2018;145(17). https://doi.org/10.1242/dev.167361
Choquet C, Kelly RG, Miquerol L. Nkx2-5 defines distinct scaffold and recruitment phases during formation of the murine cardiac Purkinje fiber network. Nat Commun. 2020;11(1):5300. https://doi.org/10.1038/s41467-020-19150-9. (Findings from this study suggest that PF network morphogenesis takes place in two recruitment phases).
Li Y, Tian X, Zhao H, et al. Genetic targeting of Purkinje fibres by Sema3a-CreERT2. Sci Rep. 2018;8(1):1–9. https://doi.org/10.1038/s41598-018-20829-9.
Shekhar A, Lin X, Lin B, et al. ETV1 activates a rapid conduction transcriptional program in rodent and human cardiomyocytes. Sci Rep. 2018;8(1):1–16. https://doi.org/10.1038/s41598-018-28239-7.
Zhang SS, Kim KH, Rosen A, et al. Iroquois homeobox gene 3 establishes fast conduction in the cardiac His-Purkinje network. Proc Natl Acad Sci U S A. 2011;108(33):13576–81. https://doi.org/10.1073/pnas.1106911108.
Sergeeva IA, Christoffels VM. Regulation of expression of atrial and brain natriuretic peptide, biomarkers for heart development and disease. Biochim Biophys Acta - Mol Basis Dis. 2013;1832(12):2403–13. https://doi.org/10.1016/j.bbadis.2013.07.003.
Choquet C, Marcadet L, Beyer S, Kelly R, Miquerol L. Segregation of Central Ventricular Conduction System Lineages in Early SMA+ Cardiomyocytes Occurs Prior to Heart Tube Formation. J Cardiovasc Dev Dis. 2016;3(1):2. https://doi.org/10.3390/jcdd3010002.
Sedmera D, Reckova M, DeAlmeida A, et al. Spatiotemporal pattern of commitment to slowed proliferation in the embryonic mouse heart indicates progressive differentiation of the cardiac conduction system. Anat Rec - Part A Discov Mol Cell Evol Biol. 2003;274(1):773–7. https://doi.org/10.1002/ar.a.10085.
Goodyer WR, Beyersdorf BM, Paik DT, et al. Transcriptomic profiling of the developing cardiac conduction system at single-cell resolution. Circ Res. 2019;125(4):379–97. https://doi.org/10.1161/CIRCRESAHA.118.314578.
Burnicka-Turek O, Broman MT, Steimle JD, et al. Transcriptional Patterning of the Ventricular Cardiac Conduction System. Circ Res. 2020;127(3):139–48. https://doi.org/10.1161/CIRCRESAHA.118.314460. (Findings from this study suggest that a critical balance of Tbox genes controls the slow and fast conduction properties of conducting cells).
Moskowitz IPG, Pizard A, Patel VV, et al. The T-Box transcription factor Tbx5 is required for the patterning and maturation of the murine cardiac conduction system. Development. 2004;131(16):4107–16. https://doi.org/10.1242/dev.01265.
Rentschler S, Zander J, Meyers K, et al. Neuregulin-1 promotes formation of the murine cardiac conduction system. Proc Natl Acad Sci U S A. 2002;99(16):10464–9. https://doi.org/10.1073/pnas.162301699.
Shekhar A, Lin X, Liu FY, et al. Transcription factor ETV1 is essential for rapid conduction in the heart. J Clin Invest. 2016;126(12):4444–59. https://doi.org/10.1172/JCI87968.
Grego-Bessa J, Gómez-Apiñaniz P, Prados B, Gómez MJ, Macgrogan D, De La Pompa JL. Nrg1 Regulates Cardiomyocyte Migration and Cell Cycle in Ventricular Development. Circ Res. 2023;133(11):927–43. https://doi.org/10.1161/CIRCRESAHA.123.323321.
Hartman ME, Liu Y, Zhu WZ, et al. Myocardial deletion of transcription factor CHF1/Hey2 results in altered myocyte action potential and mild conduction system expansion but does not alter conduction system function or promote spontaneous arrhythmias. FASEB J. 2014;28(7):3007–15. https://doi.org/10.1096/fj.14-251728.
Thomas PS, Kasahara H, Edmonson AM, et al. Elevated expression of Nkx-2.5 in Developing myocardial conduction cells. Anat Rec. 2001;265(3):307–13. https://doi.org/10.1002/ar.1106.
Harris BS, Spruill L, Edmonson AM, et al. Differentiation of cardiac Purkinje fibers requires precise spatiotemporal regulation of Nkx2-5 expression. Dev Dyn. 2006;235(1):38–49. https://doi.org/10.1002/dvdy.20580.
Meysen S, Marger L, Hewett KW, et al. Nkx2.5 cell-autonomous gene function is required for the postnatal formation of the peripheral ventricular conduction system. Dev Biol. 2007;303(2):740–53. https://doi.org/10.1016/j.ydbio.2006.12.044.
Choquet C, Nguyen THM, Sicard P, et al. Deletion of Nkx2–5 in Trabecular Myocardium Reveals the Developmental Origins of Pathological Heterogeneity Associated with Ventricular Non-Compaction Cardiomyopathy. Vol 14.; 2018. https://doi.org/10.1371/journal.pgen.1007502
Ismat FA, Zhang M, Kook H, et al. Homeobox protein Hop functions in the adult cardiac conduction system. Circ Res. 2005;96(8):898–903. https://doi.org/10.1161/01.RES.0000163108.47258.f3.
Choquet C, Sicard P, Vahdat J, et al. Nkx2–5 Loss of Function in the His-Purkinje System Hampers Its Maturation and Leads to Mechanical Dysfunction. 2023.
Risebro CA, Petchey LK, Smart N, et al. Epistatic rescue of Nkx2.5 adult cardiac conduction disease phenotypes by prospero-related homeobox protein 1 and HDAC3. Circ Res. 2012;111(2). https://doi.org/10.1161/CIRCRESAHA.111.260695
Kim KH, Rosen A, Hussein SMI, et al. Irx3 is required for postnatal maturation of the mouse ventricular conduction system. Sci Rep. 2014;2016(6):1–14. https://doi.org/10.1038/srep19197.
Bruneau BG, Nemer G, Schmitt JP, et al. A murine model of Holt-Oram syndrome defines roles of the T-Box transcription factor Tbx5 in cardiogenesis and disease. Cell. 2001;106(6):709–21. https://doi.org/10.1016/S0092-8674(01)00493-7.
Moskowitz IPG, Kim JB, Moore ML, et al. A Molecular Pathway Including Id2, Tbx5, and Nkx2-5 Required for Cardiac Conduction System Development. Cell. 2007;129(7):1365–76. https://doi.org/10.1016/j.cell.2007.04.036.
Qian L, Wythe JD, Liu J, et al. Tinman/Nkx2-5 acts via miR-1 and upstream of Cdc42 to regulate heart function across species. J Cell Biol. 2011;193(7):1181–96. https://doi.org/10.1083/jcb.201006114.
Samal E, Evangelista M, Galang G, Srivastava D, Zhao Y, Vedantham V. Premature microRNA-1 expression causes hypoplasia of the cardiac ventricular conduction system. Front Physiol. 2019;10(MAR):1–12. https://doi.org/10.3389/fphys.2019.00235.
Zhang Y, Sun L, Zhang Y, et al. Overexpression of microRNA-1 causes atrioventricular block in rodents. Int J Biol Sci. 2013;9(5):445–62. https://doi.org/10.7150/ijbs.4630.
Park DS, Tompkins RO, Liu F, et al. Pocket proteins critically regulate cell cycle exit of the trabecular myocardium and the ventricular conduction system. Biol Open. 2013;2(9):968–78. https://doi.org/10.1242/bio.20135785.
Vincentz JW, Toolan KP, Zhang W, Firulli AB. Hand factor ablation causes defective left ventricular chamber development and compromised adult cardiac function. PLoS Genet. 2017;13(7):1–20. https://doi.org/10.1371/journal.pgen.1006922.
Vincentz JW, Firulli BA, Toolan KP, et al. Variation in a Left Ventricle-Specific Hand1 Enhancer Impairs GATA Transcription Factor Binding and Disrupts Conduction System Development and Function. Circ Res. 2020:575–589. https://doi.org/10.1161/CIRCRESAHA.119.315313
Pallante BA, Giovannone S, Fang-Yu L, et al. Contactin-2 Expression in the Cardiac Purkinje Fiber Network. Circ Arrhythmia Electrophysiol. 2010;3(2):186–94. https://doi.org/10.1161/CIRCEP.109.928820.
Delgado C, Bu L, Zhang J, et al. Neural cell adhesion molecule is required for ventricular conduction system development. Dev. 2021;148(11). https://doi.org/10.1242/DEV.199431
van Eif VWW, Devalla HT regulation of the cardiac conduction systemD., Boink GJJ, Christoffels VM. Transcriptional Regulation of the Postnatal Cardiac Conduction System Heterogeneity. Nat Rev Cardiol. 2024;15(10):617–630. https://doi.org/10.1038/s41569-018-0031-y
Liang D, Xue J, Geng L, et al. Cellular and molecular landscape of mammalian sinoatrial node revealed by single-cell RNA sequencing. Nat Commun. 2021;12(1). https://doi.org/10.1038/s41467-020-20448-x
Sun T, Grassam-Rowe A, Pu Z, et al. Dbh + catecholaminergic cardiomyocytes contribute to the structure and function of the cardiac conduction system in murine heart. Nat Commun. 2023;14(1):1–23. https://doi.org/10.1038/s41467-023-42658-9.
Farah EN, Hu RK, Kern C, et al. Spatially Organized Cellular Communities Form the Developing Human Heart. Springer US; 2024. https://doi.org/10.1038/s41586-024-07171-z
Lázár E, Mauron R, Andrusivová Ž, et al. Spatial Dynamics of the Developing Human Heart. bioRxiv. 2024. https://doi.org/10.1101/2024.03.12.584577
Choquet C, Boulgakoff L, Kelly RG, Miquerol L. New insights into the development and morphogenesis of the cardiac purkinje fiber network: Linking architecture and function. J Cardiovasc Dev Dis. 2021;8(8). https://doi.org/10.3390/jcdd8080095
Koizumi A, Sasano T, Kimura W, et al. Genetic defects in a His-Purkinje system transcription factor, IRX3, cause lethal cardiac arrhythmias. Eur Heart J. 2016;37(18):1469–75. https://doi.org/10.1093/eurheartj/ehv449.
Van Rijen HVM, Van Veen TAB, Van Kempen MJA, et al. Impaired conduction in the bundle branches of mouse hearts lacking the gap junction protein Connexin40. Circulation. 2001;103(11):1591–8. https://doi.org/10.1161/01.CIR.103.11.1591.
Kim EE, Shekhar A, Lu J, et al. PCP4 regulates Purkinje cell excitability and cardiac rhythmicity. J Clin Invest. 2014;124(11):5027–36. https://doi.org/10.1172/JCI77495.
Funding
This work was supported by the Centre National de la Recherche Scientifique (CNRS), and by grants from the Association Française contre les Myopathies (AFM-Téléthon, #23711) and the French National Research Agency (ANR, “PurkinjeNet”).
Author information
Authors and Affiliations
Contributions
L.B. and G.D. wrote the main manuscript and L.B. prepared the figure. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Boulgakoff, L., D’Amato, G. & Miquerol, L. Molecular Regulation of Cardiac Conduction System Development. Curr Cardiol Rep 26, 943–952 (2024). https://doi.org/10.1007/s11886-024-02094-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11886-024-02094-7