Abstract
Purpose of the Review
The definition of arrhythmogenic cardiomyopathy (ACM) has expanded beyond desmosomal arrhythmogenic right ventricular cardiomyopathy (ARVC) to include other genetic cardiomyopathies with a significant arrhythmia burden. Emerging data on genotype–phenotype correlations has led recent consensus guidelines to urge genetic testing as a critical component of not only diagnosis but also management of ACM.
Recent Findings
Plakophilin-2 (PKP2) ARVC/ACM is most likely to meet ARVC Task Force Criteria with right sided involvement and ventricular arrhythmias, while desmoplakin (DSP) ACM may have a normal electrocardiogram (ECG) and has a subepicardial LV scar pattern. Extra-desmosomal ACM including ACM associated with transmembrane protein 43 and phospholamban variants may have characteristic ECG patterns and biventricular cardiomyopathy. Lamin A/C and SCN5A cardiomyopathy often have heart block on ECG with DCM, but are distinct from DCM in that they have significantly elevated arrhythmic risk. Newer genes, especially filamin-C (FLNC) also may have distinct imaging scar patterns, arrhythmia risk, and risk predictors.
Summary
Recognition of these key differences have implications for clinical management and reinforce the importance of genetic testing in the diagnosis and the emerging opportunities for genotype-specific management of ACM patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
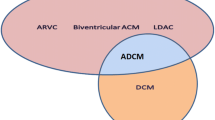
Arrhythmogenic cardiomyopathy (ACM) is an evolving diagnosis that has grown in recent years to encompass both the narrowly defined arrhythmogenic right ventricular cardiomyopathy (ARVC), and also more broadly non-ischemic cardiomyopathy (NICM) with a higher than expected arrhythmic burden [1, 2•]. ARVC is a narrow diagnosis describing an often right dominant cardiomyopathy caused by abnormalities of the desmosomal proteins meeting a set of task force criteria [1]. Expert consensus has defined the genetic basis of ARVC, highlighting its predominant association with pathogenic variants in genes encoding the cardiac desmosome [3]. The definition of ACM includes ARVC, but is broader, also including biventricular and left dominant forms of disease, and may have extra-desmosomal genetic causes. Specially, an expert consensus in the field has defined ACM as diseases that encompass structural abnormalities (both by imaging and pathology) and ventricular arrhythmia [4]. There is some debate whether this should include only genetic forms of ACM, or any form of arrhythmic cardiomyopathy, regardless of underlying cause (such as cardiac sarcoidosis) [2•]. What is clear, and documented in recent practice guidelines, is that genetic testing is a critical piece of management in these conditions. There are important genotype–phenotype correlations that inform risk stratification for malignant arrhythmias and increasingly genotype-specific therapeutic approaches indicated depending on the underlying molecular cause [5•]. Genotype informs family screening as well.
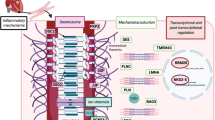
The genetic basis of ACMs began with the understanding of the genetic basis of ARVC in the early 2000s. While the first diagnostic criteria for ARVC were published in 1994 and included recognition of a familial pattern, genetic understanding lagged behind until the linkage of plakoglobin (JUP) to autosomal recessive severe ARVC in 1998 [6,7,8]. This focused etiology of ARVC to the desmosome, the protein structure adhering cellular junctions. From there, association of the more common autosomal dominant form of ARVC with the rest of the desmosomal proteins followed quickly with discovery of desmoplakin (DSP), plakophilin-2 (PKP2), desmoglein-2 (DSG2), and desmocollin-2 (DSC2) [9,10,11,12,13]. At the same time, understanding of the genetic architecture of dilated cardiomyopathy (DCM) was emerging. During this era conduction defects were linked to the lamin A/C (LMNA) gene, but it would be some years before this was formally associated with similarly arrhythmic outcomes [14, 15]. Desmin (DES) was also linked to an ACM phenotype [16]. There were additional non-desmosomal forms of ARVC identified, including a founder population in Newfoundland segregating a transmembrane protein 43 (TMEM43) variant and the Dutch founder variant in the phospholamban (PLN) gene [17, 18]. Following, more rare forms of ACM were identified, and recently, filamin-C (FLNC) has emerged as a more common cause of a very arrhythmic biventricular cardiomyopathy that often looks like ARVC [16, 19,20,21,22]. Natural history and family reports of these all these phenotypes have overlapped and also varied widely. This is not surprising, as all these genetic ACMs are associated with incomplete penetrance and highly variable expressivity [4, 23]. Therefore, large numbers of each subtype were needed to elucidate specific patterns. Now with two decades of data on the natural history of genetic ACM cohorts, clear genotype–phenotype correlations are beginning to emerge (Table 1).
PKP2
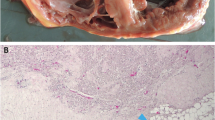
Pathogenic variants in PKP2 are the most common cause of ACM in North America [24]. PKP2-associated cardiomyopathy typically presents as the most classically described form of ACM as a right-sided, exercise-induced, very arrhythmic cardiomyopathy. PKP2-ARVC is most likely to fit the diagnostic 2010 Task Force Criteria with precordial T wave inversions on electrocardiogram (ECG), left bundle branch arrhythmias, and right ventricular dyskinesis caused by replacement with fibro-fatty scarring in the right ventricle, as shown in Fig. 1 panel A [1, 24]. Homozygous and compound heterozygote truncating variants in PKP2 are associated with a severe neonatal onset cardiomyopathy with congenital heart defects [25, 26]. Pathogenic PKP2 variants are largely loss of function variants, and there is no clear association of variant type or location with phenotype [27]. PKP2 also has the most evidence to show a clear correlation between vigorous and especially endurance exercise and disease penetrance and progression [28]. New translational research shows specifically the damage to the desmosomal reserve in Pkp2-deficient mice under the setting of regular vigorous exercise [29].
DSP
Pathogenic variants in DSP were initially identified in association with Carvajal syndrome in the recessive form with severe pediatric onset cardiomyopathy and arrhythmias with wooly hair and palmoplantar keratoses [13]. Subsequently, DSP variants were commonly found in families meeting ARVC 2010 Task Force Criteria. It was not long, however, before it became clear that DSP-associated disease could look very different from classic right sided ARVC with a tendency to present with left-sided disease and elevated rates of heart failure [24, 30]. The term arrhythmogenic left ventricular cardiomyopathy (ALVC) was often used associated with DSP. Compared to ARVC, DSP-associated ALVC more frequently shows consistent late gadolinium enhancement patterns on cardiac magnetic resonance (CMR) imaging and is less likely to have precordial T wave inversions which are often the first sign of disease in PKP2-ARVC, Fig. 1 panel B [31]. Indeed, often in DSP cardiomyopathy, the ECG is near normal [32]. As shown in Fig. 1, the most characteristic scar pattern in DSP cardiomyopathy has been described as a subepicardial ring-like scar pattern. While imaging is critical in diagnosis as referenced in Fig. 1, these individuals with often left-dominant presentation will be less likely to meet ARVC 2010 Task Force Criteria [32]. Recently, it has also has been documented that DSP cardiomyopathy will often present with chest pain, significant troponin elevation, and myocarditis-like picture [33, 34]. Scar pattern similarities between viral myocarditis and DSP cardiomyopathy can complicate diagnosis.
DSG2 and DSC2
Variants in DSG2 and DSC2 are the rarer causes of ARVC. Current evidence suggests that the phenotype is similar to PKP2 ARVC; however, there is not sufficient current case volume data to draw statistically robust comparisons. Initial evidence suggests that DSG2-associatd ARVC may have an earlier age of onset than PKP2 ARVC, and be more likely to have left-sided involvement; however, this may be complicated by selection bias in tertiary referral centers [24, 30]. There are some reports that autosomal recessive DSC2 variants may be more common in Asian populations, and there is a homozygous founder variant in the Hutterite population [35, 36].
TMEM43
TMEM43 is a membrane protein involved in nuclear envelope structure outside of the desmosome. It was identified as a candidate gene for ARVC and founder with a common haplotype initially in 15 Newfoundland ARVC families [17]. There is only one variant proven to cause ARVC in TMEM43: p.S358L [3]. Functional studies have shown that the p.S358L variant affects the expression and distribution of proteins in the intercalated disc, including JUP, resulting in an ARVC phenotype [37]. This phenotype, however, is unique in that it is particularly penetrant, and significantly arrhythmic, particularly in young males. ECG findings may be unique with notable poor R wave progression more common than precordial T wave inversion. Additionally, in TMEM43 ARVC, there is significant LV involvement as well. The most unique feature of TMEM43 ARVC is that it is appreciated to be nearly fully penetrant, whereas desmosomal ARVC penetrance rates range, but are averaged 30–50% in ARVC families [30, 38].
PLN
PLN is a transmembrane sarcoplasmic reticulum phosphoprotein that is involved with regulating calcium handling and contractility in the cardiomyocyte. It was initially described as causing familial cardiomyopathy. The Dutch founder variant p.R14del, however, was identified in causing a particularly arrhythmic cardiomyopathy and phenotype similar to desmosomal ARVC [18, 39]. It was found to be a common cause of both ARVC and familial DCM in the Netherlands, in up to 15% of cases. The phenotype involves a low-voltage ECG, and a high frequency of ventricular arrhythmias, but also a much higher risk of biventricular cardiomyopathy and end-stage heart failure than in desmosomal ARVC (Fig. 2, panel A) [30, 40]. As many PLN carriers may not meet 2010 ARVC Task Force Criteria because of the overlap with DCM, it is important to appreciate their high arrhythmic risk [2•, 39, 41].
LMNA
LMNA is a complicated nuclear protein that has been associated with over 10 different clinical syndromes, including Emery-Dreifuss muscular dystrophy and limb-girdle muscular dystrophy [42]. It was also described to cause a unique isolated cardiac phenotype with a dilated cardiomyopathy with significant conduction defects [14]. There have been several reports of LMNA carriers showing an ARVC phenotype meeting diagnostic criteria; however importantly, LMNA cardiomyopathy has shown to have signature clinical features and necessitating important risk stratification outside of the DCM phenotype [15, 43, 44]. LMNA carriers have progressive cardiac conduction defects including heart block (Fig. 2, panel B), and a higher risk of malignant arrhythmias than typical DCM patients. Risk stratification studies suggest that males with a pathogenic nonsense variant compared to a pathogenic missense variant are at increased risk of malignant arrhythmias [45,46,47]. Families with pathogenic LMNA variants also have a higher incidence of atrial arrhythmias than typically described in other ACM phenotypes [48].
SCN5A
The sodium channel (NaV1.5) is associated with wide variety of inherited arrhythmic syndromes [49]. Typically, SCN5A variants had been associated with a primarily arrhythmic phenotype in LongQT Type 3 and in Brugada syndrome. Case reports identified a specific variant that presented as a dilated cardiomyopathy conduction disorder with arrhythmias [19]. Extensive genotype–phenotype analysis in SCN5A has continued, and now, up to 18 different variants have been associated with an ACM phenotype, especially those variants located in the transmembrane voltage sensing domains of SCN5A. This phenotype is characterized by frequent ventricular ectopy, sinus node dysfunction, conduction defects including atrioventricular block, and especially atrial and ventricular arrhythmias with a subsequent identification of DCM. The arrhythmia appears to consistently precede the cardiomyopathy [49].
Recent Genes: DES, RBM20, CDH2, CTNNA3, and FLNC
Identification of rarer causes of familial ACM have implicated pathogenic changes in the desmin (DES), RNA-binding motif protein 20 (RBM20), cadherin-2 (CHD2), αT-catenin (CTNNA3), and filamin-C (FLNC) genes [16, 21, 22, 50,51,52]. Data describing these ACM phenotypes are evolving, but commonly these seem to be associated with a biventricular cardiomyopathy with a high arrhythmic risk [53]. Newer data suggests that FLNC variants may be associated with a range of specific findings including a normal ECG to more classic T wave inversions as seen in ARVC, and that subepicardial LV enhancement on MRI is common [54, 55].
Multiple Variants
Digenic, compound heterozygous and autosomal recessive inheritance, each of which are characterized by multiple pathogenic variants, have been reported to occur in an estimated 2–4% of families with ACM [30, 56]. This has important implications both for phenotyping in the family, but also for family screening. Individuals with multiple pathogenic variants consistently present at younger ages, and with more severe phenotypes. Digenic disease may also explain an inconsistency in phenotype such desmosomal variant carrier who also has an additional pathogenic cardiomyopathy variant. This information is critical for family screening, as family testing should including all pathogenic variants, and relatives carrying different combinations of variants may look quite phenotypically dissimilar depending on their unique genotype [56].
Emerging Considerations: Genome First Diagnosis
With the plummeting cost of more expansive genetic testing and the inclusion of almost all of the ACM genes as secondary findings recommended to be reported by the American College of Medical Genetics (ACMG), there is a rising number of individuals found to carry ACM risk variants in a genome first approach, and often absent of any clinical or family history [57, 58]. Studies have shown that these individuals have considerably reduced penetrance rates compared to members of ACM families in which pathogenic variants have been identified [59, 60]. While pre-clinical cases in ACM families should follow published screening guidelines, in these phenotype-absent families, caution is warranted in interpreting these variants for management and risk stratification in these individuals [61]. It has been suggested that screening and management recommendations apply differently, in a less intensive manner in this situation [62]. This finding also highlights the likely relevance of environmental and potentially genetic modifiers on outcomes of patients with a pathogenic variant [28].
Towards Genotype-specific Management
The data is now clear that identification of these specific ACM genotypes in the cardiomyopathy population is clinically useful, as arrhythmic risk is significantly higher in ACM than in typical NICM [5•]. Central themes in presentation are displayed in Fig. 3. There may be subtle clinical clues on ECG or in scar patterns to the underlying genotype, but genetic testing provides certainty. Furthermore, genotype-specific risk prediction algorithms are emerging as a way to estimate individualized malignant arrhythmia risk. Recent data suggest that there may be genotype-specific risk predictors such as scar burden and myocardial injury in DSP carriers, or ECG voltage in PLN carriers [41, 63]. Genotype-specific arrhythmia risk prediction models have been developed for PLN p.R14del ACM and LMNA cardiomyopathy and a risk calculator has been developed largely from a desmosomal ARVC cohort for prediction of incident ventricular arrhythmias (arvcrisk.com) [41, 47, 64]. Predicting progression and incidence of heart failure has remained challenging in ACM. It may be that genotype is an important predictor of trajectory [32]. Overall, many ACM families are hoping for a “cure,” and while that may not be possible, gene-specific therapies are increasing in number quickly. LMNA-related DCM was one of the first ACM clinical trials with an investigational small-molecule drug targeted towards specific genotype, but many others are now in development [65].
In families without an identifiable genetic risk factor, gene finding efforts continue. Without a clear defined genetic phenotype, however, it is becoming increasingly likely that they do not have monogenetic disease, and it may be a confluence of multiple common genetic risk alleles and environmental factors. Development of polygenic risk scores in multiple types of cardiomyopathy may aid in management in these cases [66].
Discussion and Conclusions
The promise of precision medicine is finally emerging in ACM with enough data to establish genotype–phenotype correlations. Expert consensus statements agree, and genetic counseling and genetic testing is strongly recommended as part of the diagnosis and management of ACM patients [2•, 5•, 61]. These recommendations are based upon these data that show clear differences in disease course and outcomes based on underlying genetic cause. At base, one of the most important considerations and utility of genetic testing is separating more common disease such as NICM, myocarditis, and cardiac sarcoidosis from genetic ACM. This has profound impact on patient management, especially risk stratification for malignant ventricular arrhythmias. There are clear recommendations that genetic testing is a guideline-directed part of management when diagnosing a genetic cardiomyopathy such as ARVC. These data stress the importance in identifying those with a genetic risk factor that may not have been otherwise suspected. Indeed, in many of these families due to incomplete penetrance and variable expressivity, 50% or more of individuals with a genetic ACM may not have any clear family history of disease [30]. As a result, increasingly experts are recommending genetic testing in any suspected arrhythmic cardiomyopathy presenting under the age of 50, and in many myocarditis cases, especially those with recurrent episodes and young age of presentation [34].
Identification of a genetic cause of an individual’s ACM also has critical downstream importance in family screening. Sudden cardiac death can be and is often the first symptom of disease in ACM. Therefore, identification of those at risk and regular screening is a life-saving result of genetic testing in these families. Furthermore, understanding genotype–phenotype correlations is also important in family screening, such as exercise modification in desmosomal ARVC, and screening for not only ventricular but also atrial arrhythmias in SCN5A and LMNA variant carriers [2•].
Going forward, data will continue to accumulate to refine our understanding of both ACM genotype:phenotype associations and also the interplay of genotype and environmental modifiers on ACM onset and trajectory. Individuals with genetic ACM are recommended by practice guideline to be followed in multidisciplinary expert centers who maintain an updated understanding of the latest genotype–phenotype data to incorporate into patient and family management [67]. On the horizon, however, is more complicated genotype interpretation in the non-Mendelian forms of cardiovascular disease, and the rise of polygenic risk scores for cardiac disease. The field will need to adapt once again to integration of this data into patient and family management.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance
Marcus FI, McKenna WJ, Sherrill D, Basso C, Bauce B, Bluemke DA, et al. Diagnosis of arrhythmogenic right ventricular cardiomyopathy/dysplasia: proposed modification of the task force criteria. Circulation. 2010;121:1533–41. https://doi.org/10.1161/CIRCULATIONAHA.108.840827.
• Towbin JA, McKenna WJ, Abrams DJ, Ackerman MJ, Calkins H, Darrieux FCC, et al. HRS expert consensus statement on evaluation, risk stratification, and management of arrhythmogenic cardiomyopathy. Heart Rhythm. 2019;16:e301–372. https://doi.org/10.1016/j.hrthm.2019.05.007. This critical consensus statement proposes a broad definition of arrhythmogenic cardiomyopathy and concentrates emerging data into concrete clinical recommendations for genotype-directed care.
James CA, Jongbloed JDH, Hershberger RE, Morales A, Judge DP, Syrris P, et al. International evidence based reappraisal of genes associated with arrhythmogenic right ventricular cardiomyopathy using the clinical genome resource framework. Circ Genom Precis Med 2021;14:e003273. https://doi.org/10.1161/CIRCGEN.120.003273.
Elliott PM, Anastasakis A, Asimaki A, Basso C, Bauce B, Brooke MA, et al. Definition and treatment of arrhythmogenic cardiomyopathy: an updated expert panel report. Eur J Heart Fail. 2019;21:955–64. https://doi.org/10.1002/ejhf.1534.
• Wilde AAM, Semsarian C, Marquez MF, Sepehri Shamloo A, Ackerman MJ, Ashley EA, et al. European Heart Rhythm Association (EHRA)/Heart Rhythm Society (HRS)/Asia Pacific Heart Rhythm Society (APHRS)/Latin American Heart Rhythm Society (LAHRS) Expert consensus statement on the state of genetic testing for cardiac diseases. Heart Rhythm. 2022. https://doi.org/10.1016/j.hrthm.2022.03.1225. This documentation and expert guidance detailing the impact of genetic testing in care of patients with inheirted heart disease is critical for state of the art care.
McKenna WJ, Thiene G, Nava A, Fontaliran F, Blomstrom-Lundqvist C, Fontaine G, et al. Diagnosis of arrhythmogenic right ventricular dysplasia/cardiomyopathy. Task Force of the Working Group Myocardial and Pericardial Disease of the European Society of Cardiology and of the Scientific Council on Cardiomyopathies of the International Society and Federation of Cardiology. Br Heart J 1994;71:215–218. https://doi.org/10.1136/hrt.71.3.215.
McKoy G, Protonotarios N, Crosby A, Tsatsopoulou A, Anastasakis A, Coonar A, et al. Identification of a deletion in plakoglobin in arrhythmogenic right ventricular cardiomyopathy with palmoplantar keratoderma and woolly hair (Naxos disease). Lancet. 2000;355:2119–24. https://doi.org/10.1016/S0140-6736(00)02379-5.
Coonar AS, Protonotarios N, Tsatsopoulou A, Needham EW, Houlston RS, Cliff S, et al. Gene for arrhythmogenic right ventricular cardiomyopathy with diffuse nonepidermolytic palmoplantar keratoderma and woolly hair (Naxos disease) maps to 17q21. Circulation. 1998;97:2049–58. https://doi.org/10.1161/01.cir.97.20.2049.
Rampazzo A, Nava A, Malacrida S, Beffagna G, Bauce B, Rossi V, et al. Mutation in human desmoplakin domain binding to plakoglobin causes a dominant form of arrhythmogenic right ventricular cardiomyopathy. Am J Hum Genet. 2002;71:1200–6. https://doi.org/10.1086/344208.
Gerull B, Heuser A, Wichter T, Paul M, Basson CT, McDermott DA, et al. Mutations in the desmosomal protein plakophilin-2 are common in arrhythmogenic right ventricular cardiomyopathy. Nat Genet. 2004;36:1162–4. https://doi.org/10.1038/ng1461.
Awad MM, Dalal D, Cho E, Amat-Alarcon N, James C, Tichnell C, et al. DSG2 mutations contribute to arrhythmogenic right ventricular dysplasia/cardiomyopathy. Am J Hum Genet. 2006;79:136–42. https://doi.org/10.1086/504393.
Syrris P, Ward D, Evans A, Asimaki A, Gandjbakhch E, Sen-Chowdhry S, et al. Arrhythmogenic right ventricular dysplasia/cardiomyopathy associated with mutations in the desmosomal gene desmocollin-2. Am J Hum Genet. 2006;79:978–84. https://doi.org/10.1086/509122.
Norgett EE, Hatsell SJ, Carvajal-Huerta L, Cabezas JC, Common J, Purkis PE, et al. Recessive mutation in desmoplakin disrupts desmoplakin-intermediate filament interactions and causes dilated cardiomyopathy, woolly hair and keratoderma. Hum Mol Genet. 2000;9:2761–6. https://doi.org/10.1093/hmg/9.18.2761.
Fatkin D, MacRae C, Sasaki T, Wolff MR, Porcu M, Frenneaux M, et al. Missense mutations in the rod domain of the lamin A/C gene as causes of dilated cardiomyopathy and conduction-system disease. N Engl J Med. 1999;341:1715–24. https://doi.org/10.1056/NEJM199912023412302.
Quarta G, Syrris P, Ashworth M, Jenkins S, Zuborne Alapi K, Morgan J, et al. Mutations in the Lamin A/C gene mimic arrhythmogenic right ventricular cardiomyopathy. Eur Heart J. 2012;33:1128–36. https://doi.org/10.1093/eurheartj/ehr451.
van Tintelen JP, Van Gelder IC, Asimaki A, Suurmeijer AJ, Wiesfeld AC, Jongbloed JD, et al. Severe cardiac phenotype with right ventricular predominance in a large cohort of patients with a single missense mutation in the DES gene. Heart Rhythm. 2009;6:1574–83. https://doi.org/10.1016/j.hrthm.2009.07.041.
Merner ND, Hodgkinson KA, Haywood AF, Connors S, French VM, Drenckhahn JD, et al. Arrhythmogenic right ventricular cardiomyopathy type 5 is a fully penetrant, lethal arrhythmic disorder caused by a missense mutation in the TMEM43 gene. Am J Hum Genet. 2008;82:809–21. https://doi.org/10.1016/j.ajhg.2008.01.010.
van der Zwaag PA, van Rijsingen IA, Asimaki A, Jongbloed JD, van Veldhuisen DJ, Wiesfeld AC, et al. Phospholamban R14del mutation in patients diagnosed with dilated cardiomyopathy or arrhythmogenic right ventricular cardiomyopathy: evidence supporting the concept of arrhythmogenic cardiomyopathy. Eur J Heart Fail. 2012;14:1199–207. https://doi.org/10.1093/eurjhf/hfs119.
McNair WP, Ku L, Taylor MR, Fain PR, Dao D, Wolfel E, et al. SCN5A mutation associated with dilated cardiomyopathy, conduction disorder, and arrhythmia. Circulation. 2004;110:2163–7. https://doi.org/10.1161/01.CIR.0000144458.58660.BB.
Otten E, Asimaki A, Maass A, van Langen IM, van der Wal A, de Jonge N, et al. Desmin mutations as a cause of right ventricular heart failure affect the intercalated disks. Heart Rhythm. 2010;7:1058–64. https://doi.org/10.1016/j.hrthm.2010.04.023.
Ortiz-Genga MF, Cuenca S, Dal Ferro M, Zorio E, Salgado-Aranda R, Climent V, et al. Truncating FLNC mutations are associated with high-risk dilated and arrhythmogenic cardiomyopathies. J Am Coll Cardiol. 2016;68:2440–51. https://doi.org/10.1016/j.jacc.2016.09.927.
van den Hoogenhof MMG, Beqqali A, Amin AS, van der Made I, Aufiero S, Khan MAF, et al. RBM20 mutations induce an arrhythmogenic dilated cardiomyopathy related to disturbed calcium handling. Circulation. 2018;138:1330–42. https://doi.org/10.1161/CIRCULATIONAHA.117.031947.
James CA, Syrris P, van Tintelen JP, Calkins H. The role of genetics in cardiovascular disease: arrhythmogenic cardiomyopathy. Eur Heart J. 2020;41:1393–400. https://doi.org/10.1093/eurheartj/ehaa141.
Bhonsale A, Groeneweg JA, James CA, Dooijes D, Tichnell C, Jongbloed JD, et al. Impact of genotype on clinical course in arrhythmogenic right ventricular dysplasia/cardiomyopathy-associated mutation carriers. Eur Heart J. 2015;36:847–55. https://doi.org/10.1093/eurheartj/ehu509.
Katanyuwong P, Khongkraparn A, Wattanasirichaigoon D (2021) A novel homozygous PKP2 variant in severe neonatal non-compaction and concomitant ventricular septal defect: a case report. Front Pediatr 2021;9:801491. https://doi.org/10.3389/fped.2021.801491.
Ramond F, Janin A, Di Filippo S, Chanavat V, Chalabreysse L, Roux-Buisson N, et al. Homozygous PKP2 deletion associated with neonatal left ventricle noncompaction. Clin Genet. 2017;91:126–30. https://doi.org/10.1111/cge.12780.
Dries AM, Kirillova A, Reuter CM, Garcia J, Zouk H, Hawley M, et al. The genetic architecture of Plakophilin 2 cardiomyopathy. Genet Med. 2021;23:1961–8. https://doi.org/10.1038/s41436-021-01233-7.
James CA, Bhonsale A, Tichnell C, Murray B, Russell SD, Tandri H, et al. Exercise increases age-related penetrance and arrhythmic risk in arrhythmogenic right ventricular dysplasia/cardiomyopathy-associated desmosomal mutation carriers. J Am Coll Cardiol. 2013;62:1290–7. https://doi.org/10.1016/j.jacc.2013.06.033.
Cerrone M, Marron-Linares GM, van Opbergen CJM, Costa S, Bourfiss M, Perez-Hernandez M, et al. Role of plakophilin-2 expression on exercise-related progression of arrhythmogenic right ventricular cardiomyopathy: a translational study. Eur Heart J. 2022;43:1251–64. https://doi.org/10.1093/eurheartj/ehab772.
Groeneweg JA, Bhonsale A, James CA, te Riele AS, Dooijes D, Tichnell C, et al. Clinical presentation, long-term follow-up, and outcomes of 1001 arrhythmogenic right ventricular dysplasia/cardiomyopathy patients and family members. Circ Cardiovasc Genet. 2015;8:437–46. https://doi.org/10.1161/CIRCGENETICS.114.001003.
Augusto JB, Eiros R, Nakou E, Moura-Ferreira S, Treibel TA, Captur G, et al. Dilated cardiomyopathy and arrhythmogenic left ventricular cardiomyopathy: a comprehensive genotype-imaging phenotype study. Eur Heart J Cardiovasc Imaging. 2020;21:326–36. https://doi.org/10.1093/ehjci/jez188.
Smith ED, Lakdawala NK, Papoutsidakis N, Aubert G, Mazzanti A, McCanta AC, et al. Desmoplakin cardiomyopathy, a fibrotic and inflammatory form of cardiomyopathy distinct from typical dilated or arrhythmogenic right ventricular cardiomyopathy. Circulation. 2020;141:1872–84. https://doi.org/10.1161/CIRCULATIONAHA.119.044934.
Lopez-Ayala JM, Pastor-Quirante F, Gonzalez-Carrillo J, Lopez-Cuenca D, Sanchez-Munoz JJ, Oliva-Sandoval MJ, et al. Genetics of myocarditis in arrhythmogenic right ventricular dysplasia. Heart Rhythm. 2015;12:766–73. https://doi.org/10.1016/j.hrthm.2015.01.001.
Scheel PJ III, Murray B, Tichnell C, James CA, Tandri H, Calkins H, et al. Arrhythmogenic right ventricular cardiomyopathy presenting as clinical myocarditis in women. Am J Cardiol. 2021;145:128–134. https://doi.org/10.1016/j.amjcard.2020.12.090.
Brodehl A, Weiss J, Debus JD, Stanasiuk C, Klauke B, Deutsch MA, et al. A homozygous DSC2 deletion associated with arrhythmogenic cardiomyopathy is caused by uniparental isodisomy. J Mol Cell Cardiol. 2020;141:17–29. https://doi.org/10.1016/j.yjmcc.2020.03.006.
Lorenzon A, Pilichou K, Rigato I, Vazza G, De Bortoli M, Calore M, et al. Homozygous desmocollin-2 mutations and arrhythmogenic cardiomyopathy. Am J Cardiol. 2015;116:1245–51. https://doi.org/10.1016/j.amjcard.2015.07.037.
Siragam V, Cui X, Masse S, Ackerley C, Aafaqi S, Strandberg L, et al. TMEM43 mutation p.S358L alters intercalated disc protein expression and reduces conduction velocity in arrhythmogenic right ventricular cardiomyopathy. PLoS One. 2014;9:e109128. https://doi.org/10.1371/journal.pone.0109128.
Hodgkinson KA, Connors SP, Merner N, Haywood A, Young TL, McKenna WJ, et al. The natural history of a genetic subtype of arrhythmogenic right ventricular cardiomyopathy caused by a p.S358L mutation in TMEM43. Clin Genet. 2013;83:321–331. https://doi.org/10.1111/j.1399-0004.2012.01919.x.
Haghighi K, Kolokathis F, Gramolini AO, Waggoner JR, Pater L, Lynch RA, et al. A mutation in the human phospholamban gene, deleting arginine 14, results in lethal, hereditary cardiomyopathy. Proc Natl Acad Sci USA. 2006;103:1388–1393. https://doi.org/10.1073/pnas.0510519103.
van Rijsingen IA, van der Zwaag PA, Groeneweg JA, Nannenberg EA, Jongbloed JD, Zwinderman AH, et al. Outcome in phospholamban R14del carriers: results of a large multicentre cohort study. Circ Cardiovasc Genet. 2014;7:455–65. https://doi.org/10.1161/CIRCGENETICS.113.000374.
Verstraelen TE, van Lint FHM, Bosman LP, de Brouwer R, Proost VM, Abeln BGS, et al. Prediction of ventricular arrhythmia in phospholamban p.Arg14del mutation carriers-reaching the frontiers of individual risk prediction. Eur Heart J. 2021;42:2842–2850. https://doi.org/10.1093/eurheartj/ehab294.
Di Barletta MR, Ricci E, Galluzzi G, Tonali P, Mora M, Morandi L, et al. Different mutations in the LMNA gene cause autosomal dominant and autosomal recessive Emery-Dreifuss muscular dystrophy. Am J Hum Genet. 2000;66:1407–12. https://doi.org/10.1086/302869.
Kato K, Takahashi N, Fujii Y, Umehara A, Nishiuchi S, Makiyama T, et al. LMNA cardiomyopathy detected in Japanese arrhythmogenic right ventricular cardiomyopathy cohort. J Cardiol. 2016;68:346–51. https://doi.org/10.1016/j.jjcc.2015.10.013.
Kumar S, Baldinger SH, Gandjbakhch E, Maury P, Sellal JM, Androulakis AF, et al. Long-term arrhythmic and nonarrhythmic outcomes of lamin A/C mutation carriers. J Am Coll Cardiol. 2016;68:2299–307. https://doi.org/10.1016/j.jacc.2016.08.058.
Nishiuchi S, Makiyama T, Aiba T, Nakajima K, Hirose S, Kohjitani H, et al. Gene-based risk stratification for cardiac disorders in LMNA mutation carriers. Circ Cardiovasc Genet. 2017. https://doi.org/10.1161/CIRCGENETICS.116.001603.
Sidhu K, Castrini AI, Parikh V, Reza N, Owens A, Tremblay-Gravel M, et al. The response to cardiac resynchronization therapy in LMNA cardiomyopathy. Eur J Heart Fail. 2022;24:685–93. https://doi.org/10.1002/ejhf.2463.
Wahbi K, Ben Yaou R, Gandjbakhch E, Anselme F, Gossios T, Lakdawala NK, et al. Development and validation of a new risk prediction score for life-threatening ventricular tachyarrhythmias in laminopathies. Circulation. 2019;140:293–302. https://doi.org/10.1161/CIRCULATIONAHA.118.039410.
Yoneda ZT, Anderson KC, Quintana JA, O’Neill MJ, Sims RA, Glazer AM, et al. Early-onset atrial fibrillation and the prevalence of rare variants in cardiomyopathy and arrhythmia genes. JAMA Cardiol. 2021;6:1371–9. https://doi.org/10.1001/jamacardio.2021.3370.
Peters S, Thompson BA, Perrin M, James P, Zentner D, Kalman JM, et al. Arrhythmic phenotypes are a defining feature of dilated cardiomyopathy-associated SCN5A variants: a systematic review. Circ Genom Precis Med. 2022;15:e003432. https://doi.org/10.1161/CIRCGEN.121.003432.
Hedberg C, Melberg A, Kuhl A, Jenne D, Oldfors A. Autosomal dominant myofibrillar myopathy with arrhythmogenic right ventricular cardiomyopathy 7 is caused by a DES mutation. Eur J Hum Genet. 2012;20:984–5. https://doi.org/10.1038/ejhg.2012.39.
Mayosi BM, Fish M, Shaboodien G, Mastantuono E, Kraus S, Wieland T, et al. Identification of cadherin 2 (CDH2) mutations in arrhythmogenic right ventricular cardiomyopathy. Circ Cardiovasc Genet. 2017. https://doi.org/10.1161/CIRCGENETICS.116.001605.
van Hengel J, Calore M, Bauce B, Dazzo E, Mazzotti E, De Bortoli M, et al. Mutations in the area composita protein alphaT-catenin are associated with arrhythmogenic right ventricular cardiomyopathy. Eur Heart J. 2013;34:201–10. https://doi.org/10.1093/eurheartj/ehs373.
Begay RL, Graw SL, Sinagra G, Asimaki A, Rowland TJ, Slavov DB, et al. Filamin C truncation mutations are associated with arrhythmogenic dilated cardiomyopathy and changes in the cell-cell adhesion structures. JACC Clin Electrophysiol. 2018;4:504–14. https://doi.org/10.1016/j.jacep.2017.12.003.
Hall CL, Akhtar MM, Sabater-Molina M, Futema M, Asimaki A, Protonotarios A, et al. Filamin C variants are associated with a distinctive clinical and immunohistochemical arrhythmogenic cardiomyopathy phenotype. Int J Cardiol. 2020;307:101–8. https://doi.org/10.1016/j.ijcard.2019.09.048.
Gigli M, Stolfo D, Graw SL, Merlo M, Gregorio C, Nee Chen S, et al. Phenotypic expression, natural history, and risk stratification of cardiomyopathy caused by filamin C truncating variants. Circulation. 2021;144:1600–11. https://doi.org/10.1161/CIRCULATIONAHA.121.053521.
Murray B, Tichnell C, Tandri H, Calkins H, van Tintelen JP, Judge DP, et al. Influence of panel selection on yield of clinically useful variants in arrhythmogenic right ventricular cardiomyopathy families. Circ Genom Precis Med. 2020;13:548–50. https://doi.org/10.1161/CIRCGEN.120.003020.
Miller DT, Lee K, Chung WK, Gordon AS, Herman GE, Klein TE, et al. ACMG SF v3.0 list for reporting of secondary findings in clinical exome and genome sequencing: a policy statement of the American College of Medical Genetics and Genomics (ACMG). Genet Med. 2021;23:1381–1390. https://doi.org/10.1038/s41436-021-01172-3.
Haggerty CM, James CA, Calkins H, Tichnell C, Leader JB, Hartzel DN, et al. Electronic health record phenotype in subjects with genetic variants associated with arrhythmogenic right ventricular cardiomyopathy: a study of 30,716 subjects with exome sequencing. Genet Med. 2017;19:1245–52. https://doi.org/10.1038/gim.2017.40.
Carruth ED, Beer D, Alsaid A, Schwartz MLB, McMinn M, Kelly MA, et al. Clinical findings and diagnostic yield of arrhythmogenic cardiomyopathy through genomic screening of pathogenic or likely pathogenic desmosome gene variants. Circ Genom Precis Med. 2021;14:e003302. https://doi.org/10.1161/CIRCGEN.120.003302.
Carruth ED, Young W, Beer D, James CA, Calkins H, Jing L, et al. Prevalence and electronic health record-based phenotype of loss-of-function genetic variants in arrhythmogenic right ventricular cardiomyopathy-associated genes. Circ Genom Precis Med. 2019;12:e002579. https://doi.org/10.1161/CIRCGEN.119.002579.
Hershberger RE, Givertz MM, Ho CY, Judge DP, Kantor PF, McBride KL, et al. Genetic evaluation of cardiomyopathy-a heart failure society of america practice guideline. J Card Fail. 2018;24:281–302. https://doi.org/10.1016/j.cardfail.2018.03.004.
Schwartz MLB, Buchanan AH, Hallquist MLG, Haggerty CM, Sturm AC. Genetic counseling for patients with positive genomic screening results: considerations for when the genetic test comes first. J Genet Couns. 2021;30:634–44. https://doi.org/10.1002/jgc4.1386.
Wang W, Murray B, Tichnell C, Gilotra NA, Zimmerman SL, Gasperetti A, et al. Clinical characteristics and risk stratification of desmoplakin cardiomyopathy. Europace. 2022;24:268–77. https://doi.org/10.1093/europace/euab183.
Cadrin-Tourigny J, Bosman LP, Nozza A, Wang W, Tadros R, Bhonsale A, et al. A new prediction model for ventricular arrhythmias in arrhythmogenic right ventricular cardiomyopathy. Eur Heart J. 2022. https://doi.org/10.1093/eurheartj/ehac180.
Laurini E, Martinelli V, Lanzicher T, Puzzi L, Borin D, Chen SN, et al. Biomechanical defects and rescue of cardiomyocytes expressing pathologic nuclear lamins. Cardiovasc Res. 2018;114:846–57. https://doi.org/10.1093/cvr/cvy040.
O'Sullivan JW, Raghavan S, Marquez-Luna C, Luzum JA, Damrauer SM, Ashley EA, et al. Polygenic risk scores for cardiovascular disease: a scientific statement from the American Heart Association. Circulation. 2022;101161CIR0000000000001077. https://doi.org/10.1161/CIR.0000000000001077.
Ahmad F, McNally EM, Ackerman MJ, Baty LC, Day SM, Kullo IJ, et al. Establishment of specialized clinical cardiovascular genetics programs: recognizing the need and meeting standards: a scientific statement from the American Heart Association. Circ Genom Precis Med. 2019;12:e000054. https://doi.org/10.1161/HCG.0000000000000054.
Acknowledgements
The authors wish to acknowledge that The Johns Hopkins ARVD/C Program is supported by the Leonie-Wild Foundation, the Leyla Erkan Family Fund for ARVD Research, The Hugh Calkins, Marvin H. Weiner, and Jacqueline J. Bernstein Cardiac Arrhythmia Center, the Dr. Francis P. Chiramonte Private Foundation, the Dr. Satish, Rupal, and Robin Shah ARVD Fund at Johns Hopkins, the Bogle Foundation, the Campanella family, the Patrick J. Harrison Family, the Peter French Memorial Foundation, and the Wilmerding Endowments. We are grateful to the ARVD/C patients and families who have made this work possible.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
B Murray is a consultant for MyGeneCounsel. CA James receives research support from Boston Scientific Corp and is a consultant for Pfizer, Inc (compensated), Tenaya Inc. (uncompensated), and StrideBio, Inc (uncompensated).
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Myocardial Disease
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Murray, B., James, C.A. Genotype–phenotype Correlates in Arrhythmogenic Cardiomyopathies. Curr Cardiol Rep 24, 1557–1565 (2022). https://doi.org/10.1007/s11886-022-01777-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11886-022-01777-3