Abstract
Purpose of Review
This review provides an overview of the molecular mechanisms underpinning the cardiac regenerative capacity during the neonatal period and the potential targets for developing novel therapies to restore myocardial loss.
Recent Findings
We present recent advances in the understanding of the molecular mechanisms of neonatal cardiac regeneration and the implications for the development of new cardiac regenerative therapies. During the early postnatal period, several cell types and pathways are involved in cardiomyocyte proliferation including immune response, nerve signaling, extracellular matrix, mitochondria substrate utilization, gene expression, miRNAs, and cell cycle progression.
Summary
The early neonatal mammalian heart has remarkable regenerative capacity, which is mediated by proliferation of endogenous cardiomyocytes, and is lost when cardiomyocytes stop dividing shortly after birth. A wide array of mechanisms that regulate this regenerative process have been proposed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
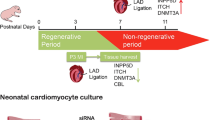
In contrast to adults, neonatal mammals have a remarkable cardiac regenerative capacity following injury. This has been demonstrated by apical resection of ~ 15% of the left ventricular myocardium [1] and by coronary ligation [2] during the first 2 days of life in mice. In these two models, the heart is able to regenerate in the ensuing 3–4 weeks after injury, with no evidence of maladaptive cardiac remodeling. However, this regenerative capacity is lost by postnatal day 7 (P7) [1, 3] which coincides with cardiomyocyte cell cycle arrest and binucleation [4]. This regenerative capacity seems to also be conserved in large animals as recently demonstrated in a neonatal pig myocardial infarction (MI) model. As in mice, the regenerative capacity of large mammals appears to be limited to the first 2 days of life [5, 6]. Fate-mapping studies have demonstrated that the regenerative capacity of the neonatal mammalian heart is mediated by proliferation of preexisting cardiomyocytes, rather than by a progenitor or stem cell population [1, 5, 7]. Following this brief window in the postnatal life, cardiac growth is achieved by hypertrophic enlargement of the cardiomyocytes, rather than cardiomyocyte hyperplasia. In the adult mammalian heart, cardiomyocyte renewal occurs at a low rate, which increases after injury, and appears to be also mediated by division of preexisting cardiomyocytes. Nevertheless, cardiomyocyte renewal in the adult heart is minimal and is insufficient for restoration of contractile function after injury [8,9,10,11,12,13]. Consequently, injuries in the adult heart fail to induce a meaningful regenerative response and often results in maladaptive cardiac remodeling and fibrosis, culminating in heart failure. Although the regenerative potential of the newborn human heart is not known, there is some evidence that the early postnatal window in humans is characterized by a measurable proliferative potential of cardiomyocytes [14]. In the adult human heart, as in mice, cardiomyocyte renewal decreases exponentially with age and occurs at around 1% per year [10]. Thus, the Achilles heel in mammalian cardiomyocyte regeneration appears to be the limited proliferative capacity of existing cardiomyocytes, which has been the focus of intense studies in recent years. The development of neonatal heart injury models has provided a much-needed set of tools for studying regulators of endogenous heart regeneration. In this review, we provide an overview of the molecular mechanisms underpinning the cardiac regenerative capacity during the neonatal period and the potential targets for development novel therapies to restore myocardial loss.
Oxidative Stress, Energy Metabolism, and Redox Regulators
There is a strong correlation between oxidation status and regenerative capacity, both in lower vertebrates and in the early postnatal mammalian development. For example, in lower vertebrates like zebrafish, which also have a notable capacity of regeneration throughout their lifetime, the circulatory system is relatively hypoxemic which is a result of arterio-venous mixing in the primitive two-chamber heart with one atrium and one ventricle. Additionally, the natural habitat aquatic environment where zebrafish lives has 1/30 the oxygen capacitance compared with the atmospheric air [15]. Similarly, the mammalian fetal heart is exposed to a relatively hypoxic environment given the shunt-dependent circulation that results in significant arterio-venous mixing [16]. However, transition from the embryonic to postnatal circulation soon after birth drastically changes the oxygenation state of cardiomyocytes [15], where the arterial pO2 increases from 30 [17,18,19] to 100 mmHg [20] within minutes. This drastic shift in oxygenation status drives a metabolic switch in energy metabolism of the postnatal heart.
During embryonic development, when cardiomyocytes rapidly proliferate, the relatively hypoxic embryonic heart utilizes anaerobic glycolysis as a main source of energy [21, 22], whereas adult cardiomyocytes utilize the oxygen-dependent mitochondrial oxidative phosphorylation as an energy source [23, 24]. The majority of the enzymes related to glycolysis are downregulated from postnatal (P) day 1 (P1) to P7, and concomitantly, the majority of the enzymes involved in mitochondrial Krebs cycle and fatty acid beta oxidation are upregulated from P1 to P7 [15]. This temporal shift from glycolytic to oxidative metabolism results an increase in mitochondrial reactive oxygen species (ROS) production. More specifically, mitochondria produce H2O2 at an elevated rate when using fatty acids relative to pyruvate as respiratory substrate [25,26,27]. The increase in mitochondrial respiration corresponds temporally with an increase in ROS in the neonatal heart, this is also associated with an increase in oxidative DNA damage and activation of DNA damage response [15]. Overall, the increased intracellular ROS induces a wide array of toxic effects by promoting damage of proteins, lipids, and DNA, such as oxidized bases as well as single- or double-strand breaks, resulting in cell cycle arrest, apoptosis, or cellular senescence [28,29,30,31]. Systemic scavenging of ROS by N-acetylcysteine (NAC) administration, or overexpression of mitochondrial catalase (mCAT) in cardiomyocytes, decreases oxidative DNA damage and prolongs the postnatal window of cardiomyocyte proliferation [15]. Moreover, pharmacological inhibition of the DNA damage response pathway extends the window of cardiomyocyte proliferation in the postnatal mouse heart [15]. These studies suggest that the oxygen-dependent mitochondrial metabolism is an important driver of cell cycle arrest in postnatal cardiomyocytes.
To extend these findings to the adult heart, our group recently demonstrated that adult mice exposed to chronic severe hypoxemia manifested metabolic and cell cycle reprogramming of adult cardiomyocytes [32•]. Severe hypoxemia was induced by gradually decreasing the fraction of inspired oxygen (FiO2) by 1% per day from 20.9 (room air oxygen) to 7% over the course of 2 weeks, followed by exposure to 7% oxygen for an additional 2 weeks. Under these conditions, several enzymes involved in mitochondrial Krebs cycle and fatty acid β oxidation were significantly decreased. In addition, adult mice exposed to severe hypoxia showed a decrease in mitochondrial ROS and decreased oxidative DNA damage in the cardiomyocytes. Overall, chronic severe hypoxia exposure induced a fetal-like metabolic reprogramming pattern characterized by upregulation of glycolytic and cell cycle genes and downregulation of fatty acid oxidation and cell cycle inhibitor genes. These genetic and metabolic alterations were associated with induction of cardiomyocyte proliferation and enhancement of re-vascularization in the infarcted zone, resulting in significant functional recovery following MI [32•].
Several genes can potentially be involved in the observed molecular and metabolic reprogramming induced by severe hypoxia. For example, exposure to 7% oxygen results in rapid stabilization of hypoxia-inducible factor 1 α (Hif1α) subunit, a master regulator of cellular hypoxic response. The Hif-1α protein contains an oxygen-dependent degradation (ODD) domain, which mediates degradation of the protein during normoxia. Upon exposure to hypoxia, the low oxygen environment prevents the prolyl hydroxylation of P402 and P564 residues within the ODD domain by prolyl hydroxylases enzymes [33]. Consequently, under hypoxia, Hif-1α is protected from degradation, resulting in transcriptional activation of hundreds of downstream target genes involved in glycolytic and mitochondrial metabolism, antioxidant enzymes, and cell cycle regulators [34, 35]. In fact, previous studies have shown that the Hif-1α protein is required for cardiac regeneration after injury in zebrafish [36] and cardiac development in the hypoxic fetal cardiomyocyte environment [37] through regulation of cell cycle, stress response pathways, and cellular metabolism [38,39,40]. On the other hand, forced activation of Hif-1α signaling by loss-of-function of prolyl hydroxylases (PHD2, PHD3) [41] or von Hippel-Lindau (VHL) [42] in cardiomyocytes led to dilated cardiomyopathy and heart failure, suggesting that hypoxia signaling should be utilized only transiently as a potential strategy for heart regeneration, and that unmitigated activation of Hif-1α signaling is likely deleterious.
Other reports support the role of antioxidant systems in myocardial regeneration. For example, activating antioxidant response after cardiac injury decrease reactive oxygen species and promote cardiac repair [43]. In cardiomyocytes, the transcripts levels of Pitx2 (paired-like homeodomain transcription factor 2) decreases postnatally [43], and the Pitx2 protein levels significantly increase after injury which is mediated by the oxidative stress response element Nrf2 (nuclear factor erythroid 2-related factor 2) [43]. Genetic ablation either Pitx2 or Nrf2 in neonatal hearts results in failure of regeneration after injury, indicating that the antioxidant stress response pathway is essential for cardiomyocyte proliferation and cardiac regeneration [43]. Importantly, Pitx2 cooperates with Hippo signaling to regulate redox-related genes. Pitx2 binds to the Hippo effector YAP (yes-associated protein) to cooperatively activate the expression of antioxidant genes [43]. Pitx2 recruits YAP to target genes in response to oxidative stress, protecting cell from ROS, promoting myocardial protection and regeneration in response to injury [43, 44].
In addition to the impact on the redox status, carbohydrate utilization in cardiomyocytes during development and early postnatal life can contribute to the cardiac regenerative potential through other mechanisms. For example, variation in cell surface and extracellular matrix glycosylation between neonatal and adult heart has been reported recently, showing a higher expression of high mannose structures and lower expression of complex N-linked glycans in the 3-day-old neonatal tissue [45]. However, a link between cell surface glycosylation and cardiomyocyte proliferation remains uncertain.
In addition, several enzymes involved in the chromatin epigenetic regulation are dependent on intermediary metabolites [46]. For example, acetyl-CoA, a key intermediary molecule of fatty acids, glucose oxidation, and amino acids metabolism, is utilized by histone acetyltransferases (HATs) as an acetyl group donor for lysine acetylation [46, 47]. The acetylation of lysine residues in histones represents a well-characterized post-translational modification associated with transcriptional regulation [47]. The histone acetyltransferases p300- or CBP-deficient mice display embryonic or neonatal lethality, with reduced left ventricular thickness, hypoplastic septum, and several cardiac malformations, which is thought to be due to the failure to activate cardiac-specific genes related to cell cycle, morphogenesis, and differentiation [48]. On the other hand, histone deacetylation is frequently associated with transcription repression. Studies in mice with global deletion of histone deacetylase 2 (Hdac2) indicate that loss of Hdac1 results in hyper proliferation of ventricular myocytes, leading to ventricle obliteration and perinatal lethality [49]. In parallel, other proteins are post-transcriptionally regulated by lysine acetylation and control cell cycle activity in cardiomyocytes. For example, acetylation regulates and stabilizes Notch1 intracellular domain (N1ICD) in neonatal rat cardiomyocyte. In one study, overexpression of the fusion protein N1ICD with the catalytic domain of p300 (N1ICD-HAT) prolonged the proliferation window of neonatal cardiomyocytes, both in vitro and in vivo. Consequently, neonatal mice overexpressing the constitutively acetylated N1ICD-HAT fusion protein enhanced myocardial regeneration after apical resection [50]. These studies suggest that the metabolic changes that occur during the postnatal development may contribute to the post-translational modification in proteins, RNA, and chromatin, leading to changes in transcription, protein/RNA stability, and cell signaling to control cardiomyocyte cell cycle arrest and growth.
In parallel to the metabolic switch from carbohydrate to fatty acid oxidation, studies using quantitative multi-omic approaches revealed significant changes in amino acid metabolism and branched chain amino acid (BCAA) catabolism during the postnatal heart development [51, 52]. The increase in amino acid metabolism correlates with the increase in protein synthesis required during the cardiomyocyte growth and maturation postnatally [51, 52]. Several studies have outlined the role of amino acids as signaling molecules regulating cardiomyocyte growth, homeostasis, and disease [53, 54]. However, the contribution of amino acid signaling to the postnatal loss of cardiac regenerative capacity remains unexplored.
Taken together, these studies suggest that the environmental and metabolic changes during the first few postnatal days contribute and mediate cardiomyocyte cell cycle arrest.
Cell Cycle and Chromatin Regulators
Cardiomyocyte cell cycle arrest during the first postnatal week is associated with downregulation of several positive cell cycle regulators (CDK2, CDK3, CDK4, CCND1, and CDK cofactors) and induction of cell cycle inhibitors (CDKI, members of the INK4 and CIP/KIP families) [55,56,57,58]. While many of these targets can be regulated by previously mentioned pathways such as DNA damage response and hypoxia signaling, the precise molecular network that regulates cardiomyocyte proliferation and maturation is not well understood. Recently, Meis homeobox 1 (MEIS1) protein, which is essential for cardiac development and homeostasis [59, 60], has been shown to regulate postnatal cardiomyocyte cell cycle arrest [3]. MEIS1 deletion in mouse cardiomyocytes was sufficient for extension of the postnatal proliferative window of cardiomyocytes and for reactivation of cardiomyocyte mitosis in the adult heart with no deleterious effect on cardiac function. In contrast, overexpression of MEIS1 in cardiomyocytes decreased neonatal myocyte proliferation and inhibited neonatal heart regeneration. Mechanistically, MEIS1 deletion resulted in downregulation of cyclin-dependent kinase inhibitors in isolated cardiomyocytes, including the members of the Ink4b–Arf–Ink4a locus (p16, p15, and p19ARF) and CIP/KIP family (p21 and p57), as well as upregulation of a number of positive regulators of the cell cycle [3]. Chromatin immunoprecipitation and luciferase reporter assay demonstrated that MEIS1 binds to a number of cyclin-dependent kinase inhibitors including p16INK4a/p19ARF/p15INK4b as well as p21CDKN1A promoters and regulates their expression [3]. Interestingly, in a separate report, MEIS1 downregulation during the postnatal development has been linked to the metabolic shift from glycolytic to oxidative metabolism [61].
The transcription factor GATA4 was also shown to promote myocardial regeneration in neonatal mice [62, 63]. GATA4 plays critical roles to promote cardiomyogenesis during heart development [64]. On the other hand, GATA4 is required to promote compensatory cardiac hypertrophy in response to pressure overload in adult heart [65, 66]. In addition, GATA4 was reported to be upregulated in proliferating cardiomyocytes, and the inhibition of GATA4 impairs heart regeneration after injury in adult zebrafish [67]. During postnatal cardiac development, GATA4 is abundant at P1 and the protein level significant decreases by P7, which coincides with cardiomyocyte cell cycle exit [62]. In one study, cardiomyocyte-specific deletion of GATA4 reduces cardiomyocyte (CM) proliferation and regenerative capacity by downregulating interleukin-13 (IL-13) [62]. Another study demonstrated that GATA4 is required for CM proliferation and neonatal cardiac regeneration after injury, by regulation of Fgf-16 [63]. As expected, these studies indicate that transcription factors play complex roles in postnatal cardiomyocyte response to injury including promoting proliferation or hypertrophy depending on the context and postnatal age.
Moreover, proteins involved in chromatin remodeling and post-translational histone modification have been demonstrated to regulate cardiomyocyte homeostasis and proliferation during postnatal heart development. For example, Ezh2 acts as a regulator of gene silencing mainly by trimethylation of histone 3 lysine 27 (H3K27Me3). In that context, Ezh2 plays a critical role regulating cell proliferation during embryonic cardiogenesis. Deletion of Ezh2 in cardiac progenitors caused postnatal myocardial pathology and dysregulated cardiac gene expression with activation of Six1-dependent skeletal muscle genes [68]. This resulted in cardiovascular defects, perinatal death, decreased cardiomyocyte proliferation, and increased apoptosis [69]. Another study demonstrated that the deletion of Ezh2 in cardiomyocytes had no effect on cardiac regeneration after apical resection or myocardial infarction in neonatal mice [70], suggesting that Ezh2 is not required for heart regeneration after injury in neonatal mice.
Hippo Pathway in Cardiac Regeneration
The Hippo signaling pathway plays an important role in tissue homeostasis and organ size control in several species. In mammals, the Hippo pathway consists of a core kinases STE20-like protein kinases 1 and 2 (Mst1 and Mst2) and the large tumor suppressor 1 and 2 (LATS1 and LATS2), the scaffolding protein Salvador homolog 1 (SAV1), and MOB domain kinase activator 1A (MOB1A) and MOB1B. Downstream the pathway includes the transcription coactivators YAP and WW domain—containing transcription regulator 1 (WWTR1 or TAZ), which when unphosphorylated, binds to transcription factors TEAD1–4 (TEA domain family members 1–4), to induce expression of genes related to cell growth and proliferation [71, 72].
Several studies have demonstrated that the Hippo signaling has a key role in regulating cell proliferation and heart growth during embryonic cardiac development. Embryonic YAP inactivation in the heart, using Nkx2.5-Cre which is expressed early in heart development, resulted in lethal myocardial hypoplasia, decreased cardiomyocyte proliferation, and embryonic lethality by embryonic day E10.5 [73]. Similar results were reported using YAP knockout mice TnnT2-cre, showing reduced ventricular chamber size, cardiac hypoplasia, and reduced cardiomyocyte proliferation with no effect in apoptosis [74].
Postnatally, the Hippo pathway controls cardiomyocyte survival and basal cardiac homeostasis. For example, conditional deletion (cKO) of YAP using cre-driver α-myosin heavy chain promoter, which is upregulated during fetal and postnatal cardiac development, resulted in cardiomyocyte apoptosis, fibrosis, progressive dilated cardiomyopathy, and premature death, with a median lifespan between 11 and 20 weeks [75, 76]. Moreover, the YAP knockout mice, subjected to the left anterior descending coronary artery ligation (LAD) at P2 and analyzed at P28, presented pathological cardiac remodeling characterized by fibrosis and cardiac dilation, showing that YAP is required for neonatal heart regeneration in response to injury [75].
On the other hand, transgenic mice overexpressing an active form of YAP under the control of the βMHC (β-myosin heavy chain) promoter, which is transcriptionally activated in the heart beginning at E9, promotes neonatal cardiomyocyte proliferation with increased heart size. Mechanistically, the substitution of Ser112 with Ala (S112A) generates a YAP protein that is constitutively active and localized to the nucleus [77]. The cardiac-specific Tg-YAPS112A mice demonstrated activation of the insulin-like growth factor (IGF) signaling pathway, resulting in inactivation of glycogen synthase kinase 3β, with increased expression of genes involved in mitosis and cytokinesis [73]. In addition, the Tg-YAPS112A showed an enhanced regenerative response, improved heart function, reduced maladaptive remodeling, and fibrosis, in response to myocardial infarction at postnatal day 7 [75]. Therefore, these results indicate that YAP gain of function controls postnatal cardiomyocyte proliferation by inducing cell cycle, DNA synthesis, and cytokinesis genes.
Finally, the protein SAV1 has been reported to play a critical role in cardiomyocyte proliferation, growth, and survival postnatally [78]. The scaffold protein SAV1 forms a complex with Mst1/Mst2 kinases that phosphorylates and activates LATS1/LATS2 kinases, which in turn phosphorylate and promote the degradation of the downstream effectors YAP/TAZ [44, 79]. Studies using SAV1 conditional knockout using an Nkx2.5-Cre show substantial cardiac overgrowth with elevated cardiomyocyte proliferation at 2 days postnatally, with no change in myocardial cell size [78]. Moreover, the SAV1-cKO hearts show reduced levels of phosphorylated inactive form of YAP but no change in total YAP [78]. In addition, SAV1 deletion extends the window of cardiomyocyte proliferation in the postnatal mouse heart and induces heart regeneration following injury either by apical resection or MI at postnatal day 8 [75, 80]. Mechanistically, Salvador deletion was associated with the activation of Wnt signaling, showing that Hippo signaling negatively regulates Wnt/β-catenin pathway to control cardiomyocyte proliferation, apoptosis, and differentiation [78]. Similar embryonic heart phenotypes were described in LATS2 and Mst1/2 mutant hearts [78]. These findings suggest that the Hippo signaling pathway is a key regulator of cardiomyocyte proliferation and heart regeneration in mammals.
miRNAs
Small non-coding RNAs are involved in a vast array of biological processes by post-transcriptional regulation of gene expression. In the heart context, miRNAs play distinct roles during heart development [81, 82] and under pathological disorders [83].
During postnatal heart development, the members of the miR-15 family of microRNAs have been reported to be upregulated and contribute to cardiomyocyte mitotic arrest [84]. More specifically, miR-195, miR-497, miR-15a, and miR-16 were all upregulated in mouse cardiac ventricles between P7 and P14 [84]. Transgenic mice overexpressing miR-195 in the embryonic heart under the control of the βMHC displayed ventricular hypoplasia and ventricular septal defects, abnormalities associated with premature cell cycle arrest [84]. Moreover, a marked reduction in the cycling cardiomyocytes undergoing mitosis and an increased proportion of multinucleated myocytes were described in the Tg-miR-195 compared with control [84]. No significant differences in cell size or apoptosis were observed. However, the neonatal Tg-miR-195 hearts fail to regenerate after LAD ligation at P1. On the other hand, inhibition of the miR-15 family in neonatal mice by using locked nucleic acid (LNA)–modified anti-miR injections at P1, P7, and P14 extends the regenerative window, increased adult CM proliferation, and improved left ventricular systolic function after adult MI [2]. Mechanistically, miR-195 regulates the expression of multiple targets including checkpoint kinase 1 (Chek1), which is a highly conserved direct target of miR-195 [84].
Similarly, the expression of miR302–367 cluster also has been related to cardiomyocyte cell cycle arrest [85]. More specifically, all five members of the miR302–367 cluster were significantly decreased after E11.5 to undetectable levels at postnatal stages. Transgenic mice overexpressing miR302–367 displayed increased levels of cardiomyocyte proliferation in embryonic and postnatal hearts, accompanied by profound cardiac enlargement at P20, resulting in premature death at P28 [85]. In addition, postnatal re-expression of miR302-367 promotes cardiomyocyte proliferation and cardiac remodeling improvement, with reduced scar formation after MI. However, prolonged postnatal expression of miR302-367 leads to cardiomyocyte dedifferentiation phenotype and compromised cardiac function [85]. Transcriptome analysis of transgenic hearts overexpressing miR302–367 displayed upregulation of cell proliferation–associated genes and downregulation of cell death, differentiation, and fatty acid metabolism genes [85]. Mechanistically, miR302–367 inhibits the Hippo pathway, by targeting the 3′ untranslated region (3′UTR) of Mst1 and LATS2 [85]. Mst1 and LATS2 are core kinase components of Hippo signaling and it has been described previously that the deletion of Mst1/2 and LATS extends the cardiomyocyte proliferation window in postnatal mice [78].
During postnatal heart growth, the upregulation of miR-128 is related to cell cycle exit [86]. In addition, cardiomyocyte-specific Dox-inducible “Tet-off” transgenic mice (α-MHC-tTA; miR-128TetRE) showed decreased cardiomyocyte proliferation, compensatory pathological cardiomyocyte hypertrophy, and cardiac contractile dysfunction at postnatal day 1 and inhibited cardiac regeneration after apical resection at P1. On the other hand, cardiac-specific deletion of miR-128 during heart development (NKX2.5Cre; miR-128fl/fl) extends the postnatal cardiomyocyte proliferation window, promotes cardiomyocyte proliferation in neonatal mice [86], and promotes cardiac regeneration in adult mice. In general, deletion of miR-128 promotes cell cycle re-entry by enhancing expression of the chromatin regulator SUZ12 (Polycomb protein SUZ12), which suppresses p27 and activates cyclin E and CDK2 [86].
Furthermore, miR-34a has been found to be a suppressor of cell proliferation and has been involved in the regulation of cardiac aging and function [87]. The miR-34a expression is increased in adult but not in neonatal mouse heart post-MI [88]. The miR-34a inhibition reduces cell death and fibrosis following acute myocardial infarction and improves the recovery of myocardial function [87, 88] . In contrast, the overexpression of miR-34a in early postnatal mice decreased cardiomyocyte proliferation and impaired cardiac regeneration after injury [88]. Mechanistically, miR-34a acts by targeting PPP1R10, which regulates telomere shortening, DNA damage response, and cardiomyocyte apoptosis, to improve functional recovery after myocardial infarction [87]. Other studies have also linked miR-34a expression to cardiomyocyte cell cycle arrest and apoptosis by targeting Bcl2, cyclin D1, and Sirt1 [88].
In contrast, high-throughput screening in neonatal rat CMs using a library of 875 synthetic miRNAs identified miRNAs that promote CM proliferation [89]. More specifically, intracardiac injection of synthetic miRNAs hsa-miR-590 and hsa-miR-199a in neonatal mice resulted in a significant increase in CM proliferation in vivo [89]. In addition, 12 days after the neonatal intraperitoneal injection of adeno-associated virus serotype 9 overexpressing the hsa-miR-590 and hsa-miR-199a miRNAs, significant enlargement of the heart with no signs of CM hypertrophy or fibrosis was demonstrated [89]. Moreover, the hsa-miR-590 and hsa-miR-199a miRNAs induced cardiac regeneration and improved cardiac function after myocardial infarction injury in adult mice [89]. Recent studies in adult pigs have shown that injections of AAV6-miR-199a in infarcted heart induces cardiomyocyte dedifferentiation and proliferation, improves heart function, and reduces scar size [90••]. However, sustained and uncontrolled long-term expression of the miR-199a in the infarcted pigs resulted in sudden cardiac death due to ventricular arrhythmias [90••].
In addition, another cluster of microRNAs was correlated with cell proliferation. The miR-17-92 cluster was originally identified as a potential human oncogene [91], by modulating E2F1 expression [92]. Global deletion of miR-17-92 in mice results in lethality shortly after birth with lung hypoplasia and a ventricular septal defect [93]. Cardiac-specific deletion of miR-17-92 in mice (NKX2.5Cre; miR-17-92fl/fl) displayed decreased CM proliferation and partial embryonic lethality [94]. On the other hand, cardiac-specific overexpression of miR-17-92 induced heart enlargement, increased heart weight, and increased CM proliferation and hyperplasia, with no signal of hypertrophy [94]. Overexpression of miR-17-92 in neonatal cardiomyocyte in vitro represses the expression and function of PTEN to promote cardiomyocyte proliferation [94]. Collectively, these results suggest that microRNAs play a central role in cardiomyocyte cell cycle regulation through a diverse set of pathways.
Immune Response Signals
The ischemic injury that occurs during myocardial infarction promotes cell death and triggers an inflammatory response which is essential for clearance of necrotic tissue and establishment of a wound healing response and fibrotic scar formation [95]. Similarly, a strong inflammatory response accompanies the early stages of cardiac regeneration after injury in neonatal mice [1, 96, 97]. However, in contrast to adult, the neonatal inflammatory response appears to contribute to promoting myocardial regeneration. There are significant differences related to the immune response and macrophage subpopulations between P1, P14, and adult heart after MI [97, 98]. After injury, the total number of macrophages and monocytes significantly increases in both neonatal and adult heart [97, 98]. However, in neonatal mice, the increase is mediated by populations of embryonic-derived resident cardiac macrophages, which display minimal inflammatory response and are required for heart regeneration. In contrast, in adults, macrophages derived from circulating monocytes are the dominant cell population in the injured myocardium, which triggers a proinflammatory response and contributes to maladaptive remodeling of the heart [98]. In one study, inhibition of monocyte recruitment after neonatal cardiac injury did not affect cardiomyocyte proliferation, angiogenesis, or alter resident macrophage accumulation [98]. On the other hand, in adult heart, inhibition of monocyte recruitment by a selective CCR2 (chemokines C–C motif receptor 2) inhibitor significantly reduced inflammation, preserved resident macrophage subsets, and enhanced myocardial repair [98]. Similarly, macrophage-depleted neonatal mice, by clodronate-liposome injection, resulted in impaired regenerative capacity after injury [97]. Therefore, it appears that resident neonatal macrophages are required for heart regeneration and angiogenesis [97].
In addition, neonatal IL-13 knockout mice showed decreased proliferative markers and upregulation of hypertrophic markers. IL-13 KO hearts also fail to regenerate after apical resection [99]. In contrast, recombinant IL-13 administration increases CM proliferation by upregulating the pro-proliferative and pro-survival Erk1/2 and Akt signaling pathways [99]. Immunosuppression, induced by dexamethasone administration in P1 mice, abolished heart repair after apical resection [96]. This study concluded that IL-13-mediated acute inflammation is required for cardiac regeneration in neonatal mice, and the STAT3/IL-6 axis is essential to promote cardiomyocyte proliferation and heart regeneration after injury in neonatal mice [96]. Taken together, the immune response is required for neonatal heart regeneration, and modulating inflammatory response may provide a key therapeutic strategy to support adult heart regeneration.
Extracellular Matrix
Myocardial injury in adult mammals invariably results in extensive remodeling of the extracellular matrix (ECM) leading to formation of fibrotic scars and consequently reduced heart function [100]. In contrast to adults, neonatal ECM components have been shown to be important modulators of cardiomyocyte proliferation and are required for myocardial regeneration [100]. For example, agrin, which is a large extracellular heparan sulfate proteoglycan which is expressed in developing axonal tracts and neuromuscular junctions, was recently shown to be a critical regulator of cardiomyocyte proliferation [100, 101]. The agrin protein level decreases postnatally [100], which coincides with the loss of cardiac regenerative capacity in mice [1]. Conditional deletion of agrin in the cardiac mesoderm showed reduced cardiomyocyte cell cycle markers at P1 and fails to completely regenerate the heart after apical resection [100]. However, agrin does not appear to be necessary for cardiac growth and homeostasis during postnatal heart development [100]. Mechanistically, agrin binds to the DGC (dystrophin–glycoprotein complex) through α-dystroglycan (Dag1), then promotes DGC disassembly, leading to sarcomere disassembly and activation of downstream signaling molecules including Yap and ERK [100].
Changes in ECM in the early postnatal heart appear to coincide with loss of neonatal regenerative response. For example, a recent study showed that apical resection performed at P2 is characterized by fibrotic scar deposition and incomplete regeneration [102], which is in sharp contrast with the regenerative capacity described in P1 mice [1, 102]. Transcriptome analysis revealed that there are major changes in the ECM composition during the first days postnatally, which has been reported to be associated with the decline in regenerative capacity and cardiomyocyte cell cycle arrest [100, 102]. Moreover, the overall stiffness of the ECM in the heart is significantly higher in P2 compared with P1 hearts, suggesting that cross-linking of ECM components plays a critical role during heart development and regeneration [102]. Pharmacological inhibition of the cross-linker enzyme LOX by using 3-aminopropionitrile (BAPN) significantly reduced ECM stiffness and prevented the robust fibrotic deposition at postnatal day 21 in response to apical resection on P3 [102]. These studies, along with others, support the notion that the postnatal environment regulates the pattern of cardiomyocyte growth and response to injury.
Nerve Signaling
The mammalian heart is innervated by parasympathetic and sympathetic fibers, which exert antagonistic effects and are responsible for regulation of an array of physiological functions including heart rate, contraction, and cardiac output. Not surprisingly, several recent reports have suggested that cardiac innervation is critical for the neonatal cardiac regenerative response [103,104,105,106]. Following ischemic myocardial infarction, several other cell types, besides cardiomyocytes, are affected by ischemia, including the cardiac ganglia which can be a major determinant of cardiac arrhythmias [107].
Recently, two independent groups reported that sympathetic and parasympathetic denervation impairs cardiomyocyte proliferation and cardiac regeneration in neonatal mice. Sympathetic ablation of sub-epicardial nerves by using 6-hydroxydopamine hydrobromide (6-OHDA) resulted in robust denervation, extensive fibrotic scar, and lack of regeneration following apical resection in neonatal mice [106]. Similarly, the parasympathetic nerve ablation by vagotomy impaired cardiomyocyte proliferation and cardiac repair in neonatal mice after apical resection and myocardial infarction [105]. In addition, the inhibition of regeneration by mechanical unilateral left vagotomy denervation was partially rescued by the administration of recombinant neuregulin1 (Ngr1) and nerve growth factor (Ngf) proteins [105]. Results from these studies demonstrate that both sympathetic and parasympathetic nerves might contribute to the regenerative capacity of the neonatal heart. Moreover, therapies that target and stimulate re-innervation might be crucial for cardiovascular disease treatment.
Conclusions
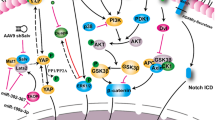
In mammals, several molecular and cellular mechanisms are involved in cardiac physiological adaptations during the postnatal period. The transition from the in utero to the postnatal environment has a significant impact on metabolism, mitochondria substrate utilization, DNA damage, transcription, cardiomyocyte proliferation, and growth. Besides myocytes, several cellular subsets have been shown to be required for neonatal heart regeneration (Fig. 1). Studies have demonstrated that targeting neonatal cardiac pathways to force cardiomyocyte proliferation is a promising approach to achieve adult heart regeneration [2, 3, 75, 89, 108]. However, the sustained activation of cell cycle activity and myocyte dedifferentiation in adult hearts can be deleterious in certain scenarios [90••]. Therefore, while our understanding of the basic biology of heart regeneration continues to grow, the neonatal mouse model is proving to be an important tool that offers a glimpse of the elegance and complexity of endogenous heart regeneration in mammals. It remains to be seen if insights learned from neonatal heart regeneration studies will 1 day translate into regenerative therapies for the human heart.
Mechanisms of neonatal cardiac regeneration. A schematic view of the key factors involved in heart regeneration by stimulating the proliferation of preexisting cardiomyocytes. Several processes have been shown to contribute to the regenerative capacity of the heart. The oxygen environment; the different cell types and the extracellular matrix contribute and create a favorable environment to induce cardiomyocyte proliferation. Intracellularly, the cardiomyocyte metabolic substrate utilization, the protein and RNA signaling, and the chromatin transcription regulation activate critical pathways that end up in the cardiomyocyte proliferation and cardiac repair. The image was produced using images modified from Servier Medical Art, licensed under the Creative Commons Attribution 3.0 Unported License
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Porrello ER, Mahmoud AI, Simpson E, Hill JA, Richardson JA, Olson EN, et al. Transient regenerative potential of the neonatal mouse heart. Science. 2011;331:1078–80.
Porrello ER, Mahmoud AI, Simpson E, Johnson BA, Grinsfelder D, Canseco D, et al. Regulation of neonatal and adult mammalian heart regeneration by the miR-15 family. Proc Natl Acad Sci U S A. 2013;110:187–92.
Mahmoud AI, Kocabas F, Muralidhar SA, Kimura W, Koura AS, Thet S, et al. Meis1 regulates postnatal cardiomyocyte cell cycle arrest. Nature. 2013;497:249–53.
Soonpaa MH, Kim KK, Pajak L, Franklin M, Field LJ. Cardiomyocyte DNA synthesis and binucleation during murine development. Am J Phys. 1996;271:H2183–9.
Ye L, D’Agostino G, Loo SJ, Wang CX, Su LP, Tan SH, et al. Early regenerative capacity in the porcine heart. Circulation. 2018;138:2798–808.
Zhu W, Zhang E, Zhao M, Chong Z, Fan C, Tang Y, et al. Regenerative potential of neonatal porcine hearts. Circulation. 2018;138:2809–16.
Elhelaly WM, Cardoso AC, Pereira AHM, Elnawasany A, Ebrahimi S, Nakada Y, et al. C-kit cells do not significantly contribute to cardiomyogenesis during neonatal heart regeneration. Circulation. 2019;139:559–61.
Bergmann O, Zdunek S, Frisen J, Bernard S, Druid H, Jovinge S. Cardiomyocyte renewal in humans. Circ Res. 2012;110:e17–8 author reply e19-21.
Laflamme MA, Myerson D, Saffitz JE, Murry CE. Evidence for cardiomyocyte repopulation by extracardiac progenitors in transplanted human hearts. Circ Res. 2002;90:634–40.
Bergmann O, Bhardwaj RD, Bernard S, Zdunek S, Barnabe-Heider F, Walsh S, et al. Evidence for cardiomyocyte renewal in humans. Science. 2009;324:98–102.
Nadal-Ginard B. Generation of new cardiomyocytes in the adult heart: prospects of myocardial regeneration as an alternative to cardiac transplantation. Revista espanola de cardiologia. 2001;54:543–50.
Quaini F, Urbanek K, Beltrami AP, Finato N, Beltrami CA, Nadal-Ginard B, et al. Chimerism of the transplanted heart. N Engl J Med. 2002;346:5–15.
Hsieh PC, Segers VF, Davis ME, MacGillivray C, Gannon J, Molkentin JD, et al. Evidence from a genetic fate-mapping study that stem cells refresh adult mammalian cardiomyocytes after injury. Nat Med. 2007;13:970–4.
Mollova M, Bersell K, Walsh S, Savla J, Das LT, Park SY, et al. Cardiomyocyte proliferation contributes to heart growth in young humans. Proc Natl Acad Sci U S A. 2013;110:1446–51.
Puente BN, Kimura W, Muralidhar SA, Moon J, Amatruda JF, Phelps KL, et al. The oxygen-rich postnatal environment induces cardiomyocyte cell-cycle arrest through DNA damage response. Cell. 2014;157:565–79.
Dawes GS, Mott JC, Widdicombe JG. The foetal circulation in the lamb. J Physiol. 1954;126:563–87.
Lawrence J, Xiao D, Xue Q, Rejali M, Yang S, Zhang L. Prenatal nicotine exposure increases heart susceptibility to ischemia/reperfusion injury in adult offspring. J Pharmacol Exp Ther. 2008;324:331–41.
Mitchell JA, Van Kainen BR. Effects of alcohol on intrauterine oxygen tension in the rat. Alcohol Clin Exp Res. 1992;16:308–10.
Reynolds JD, Penning DH, Dexter F, Atkins B, Hrdy J, Poduska D, et al. Ethanol increases uterine blood flow and fetal arterial blood oxygen tension in the near-term pregnant ewe. Alcohol. 1996;13:251–6.
Webster WS, Abela D. The effect of hypoxia in development. Birth Defects Res C Embryo Today. 2007;81:215–28.
Fisher DJ, Heymann MA, Rudolph AM. Myocardial oxygen and carbohydrate consumption in fetal lambs in utero and in adult sheep. Am J Phys. 1980;238:H399–405.
Lopaschuk GD, Collins-Nakai RL, Itoi T. Developmental changes in energy substrate use by the heart. Cardiovasc Res. 1992;26:1172–80.
Wisneski JA, Gertz EW, Neese RA, Gruenke LD, Morris DL, Craig JC. Metabolic fate of extracted glucose in normal human myocardium. J Clin Invest. 1985;76:1819–27.
Gertz EW, Wisneski JA, Stanley WC, Neese RA. Myocardial substrate utilization during exercise in humans. Dual carbon-labeled carbohydrate isotope experiments. J Clin Investig. 1988;82:2017–25.
Anderson EJ, Yamazaki H, Neufer PD. Induction of endogenous uncoupling protein 3 suppresses mitochondrial oxidant emission during fatty acid-supported respiration. J Biol Chem. 2007;282:31257–66.
Rindler PM, Plafker SM, Szweda LI, Kinter M. High dietary fat selectively increases catalase expression within cardiac mitochondria. J Biol Chem. 2013;288:1979–90.
Seifert EL, Estey C, Xuan JY, Harper ME. Electron transport chain-dependent and -independent mechanisms of mitochondrial H2O2 emission during long-chain fatty acid oxidation. J Biol Chem. 2010;285:5748–58.
Moos PJ, Edes K, Fitzpatrick FA. Inactivation of wild-type p53 tumor suppressor by electrophilic prostaglandins. Proc Natl Acad Sci U S A. 2000;97:9215–20.
Marnett LJ, Riggins JN, West JD. Endogenous generation of reactive oxidants and electrophiles and their reactions with DNA and protein. J Clin Invest. 2003;111:583–93.
Hoeijmakers JH. DNA damage, aging, and cancer. N Engl J Med. 2009;361:1475–85.
Elhelaly WM, Lam NT, Hamza M, Xia S, Sadek HA. Redox regulation of heart regeneration: an evolutionary tradeoff. Frontiers In Cell And Developmental Biology. 2016;4:137.
• Nakada Y, Canseco DC, Thet S, Abdisalaam S, Asaithamby A, Santos CX, et al. Hypoxia induces heart regeneration in adult mice. Nature. 2017;541:222–7 Findings from this study suggest that oxygen metabolism is a critical regulator of cardiomyocyte cell cycle.
Ivan M, Kondo K, Yang H, Kim W, Valiando J, Ohh M, et al. HIFalpha targeted for VHL-mediated destruction by proline hydroxylation: implications for O2 sensing. Science. 2001;292:464–8.
Prabhakar NR, Semenza GL. Oxygen sensing and homeostasis. Physiology. 2015;30:340–8.
Kaelin WG Jr. The von Hippel-Lindau protein, HIF hydroxylation, and oxygen sensing. Biochem Biophys Res Commun. 2005;338:627–38.
Jopling C, Sleep E, Raya M, Marti M, Raya A, Izpisua Belmonte JC. Zebrafish heart regeneration occurs by cardiomyocyte dedifferentiation and proliferation. Nature. 2010;464:606–9.
Dunwoodie SL. The role of hypoxia in development of the mammalian embryo. Dev Cell. 2009;17:755–73.
Breckenridge RA, Piotrowska I, Ng KE, Ragan TJ, West JA, Kotecha S, et al. Hypoxic regulation of hand1 controls the fetal-neonatal switch in cardiac metabolism. PLoS Biol. 2013;11:e1001666.
Guimaraes-Camboa N, Stowe J, Aneas I, Sakabe N, Cattaneo P, Henderson L, et al. HIF1alpha represses cell stress pathways to allow proliferation of hypoxic fetal cardiomyocytes. Dev Cell. 2015;33:507–21.
Menendez-Montes I, Escobar B, Palacios B, Gomez MJ, Izquierdo-Garcia JL, Flores L, et al. Myocardial VHL-HIF signaling controls an embryonic metabolic switch essential for cardiac maturation. Dev Cell. 2016;39:724–39.
Moslehi J, Minamishima YA, Shi J, Neuberg D, Charytan DM, Padera RF, et al. Loss of hypoxia-inducible factor prolyl hydroxylase activity in cardiomyocytes phenocopies ischemic cardiomyopathy. Circulation. 2010;122:1004–16.
Lei L, Mason S, Liu D, Huang Y, Marks C, Hickey R, et al. Hypoxia-inducible factor-dependent degeneration, failure, and malignant transformation of the heart in the absence of the von Hippel-Lindau protein. Mol Cell Biol. 2008;28:3790–803.
Tao G, Kahr PC, Morikawa Y, Zhang M, Rahmani M, Heallen TR, et al. Pitx2 promotes heart repair by activating the antioxidant response after cardiac injury. Nature. 2016;534:119–23.
Liu S, Martin JF. The regulation and function of the Hippo pathway in heart regeneration. Wiley Interdiscip Rev Dev Biol. 2019;8:e335.
Contessotto P, Ellis BW, Jin C, Karlsson NG, Zorlutuna P, Kilcoyne M, et al. Distinct glycosylation in membrane proteins within neonatal versus adult myocardial tissue. Matrix Biol. 2019.
Etchegaray JP, Mostoslavsky R. Interplay between metabolism and epigenetics: a nuclear adaptation to environmental changes. Mol Cell. 2016;62:695–711.
Jenuwein T, Allis CD. Translating the histone code. Science. 2001;293:1074–80.
Shikama N, Lutz W, Kretzschmar R, Sauter N, Roth JF, Marino S, et al. Essential function of p300 acetyltransferase activity in heart, lung and small intestine formation. EMBO J. 2003;22:5175–85.
Montgomery RL, Davis CA, Potthoff MJ, Haberland M, Fielitz J, Qi X, et al. Histone deacetylases 1 and 2 redundantly regulate cardiac morphogenesis, growth, and contractility. Genes Dev. 2007;21:1790–802.
Collesi C, Felician G, Secco I, Gutierrez MI, Martelletti E, Ali H, et al. Reversible Notch1 acetylation tunes proliferative signalling in cardiomyocytes. Cardiovasc Res. 2018;114:103–22.
Talman V, Teppo J, Poho P, Movahedi P, Vaikkinen A, Karhu ST, et al. Molecular atlas of postnatal mouse heart development. J Am Heart Assoc. 2018;7:e010378.
Lalowski MM, Bjork S, Finckenberg P, Soliymani R, Tarkia M, Calza G, et al. Characterizing the key metabolic pathways of the neonatal mouse heart using a quantitative combinatorial omics approach. Front Physiol. 2018;9:365.
Broer S, Broer A. Amino acid homeostasis and signalling in mammalian cells and organisms. Biochem J. 2017;474:1935–63.
Neinast M, Murashige D, Arany Z. Branched chain amino acids. Annu Rev Physiol. 2019;81:139–64.
Walsh S, Ponten A, Fleischmann BK, Jovinge S. Cardiomyocyte cell cycle control and growth estimation in vivo--an analysis based on cardiomyocyte nuclei. Cardiovasc Res. 2010;86:365–73.
Pasumarthi KB, Nakajima H, Nakajima HO, Soonpaa MH, Field LJ. Targeted expression of cyclin D2 results in cardiomyocyte DNA synthesis and infarct regression in transgenic mice. Circ Res. 2005;96:110–8.
Gude N, Muraski J, Rubio M, Kajstura J, Schaefer E, Anversa P, et al. Akt promotes increased cardiomyocyte cycling and expansion of the cardiac progenitor cell population. Circ Res. 2006;99:381–8.
Sdek P, Zhao P, Wang Y, Huang CJ, Ko CY, Butler PC, et al. Rb and p130 control cell cycle gene silencing to maintain the postmitotic phenotype in cardiac myocytes. J Cell Biol. 2011;194:407–23.
Azcoitia V, Aracil M, Martinez AC, Torres M. The homeodomain protein Meis1 is essential for definitive hematopoiesis and vascular patterning in the mouse embryo. Dev Biol. 2005;280:307–20.
Stankunas K, Shang C, Twu KY, Kao SC, Jenkins NA, Copeland NG, et al. Pbx/Meis deficiencies demonstrate multigenetic origins of congenital heart disease. Circ Res. 2008;103:702–9.
Lindgren IM, Drake RR, Chattergoon NN, Thornburg KL. Down-regulation of MEIS1 promotes the maturation of oxidative phosphorylation in perinatal cardiomyocytes. FASEB J. 2019;33:7417–26.
Malek Mohammadi M, Kattih B, Grund A, Froese N, Korf-Klingebiel M, Gigina A, et al. The transcription factor GATA4 promotes myocardial regeneration in neonatal mice. EMBO Mol Med. 2017;9:265–79.
Yu W, Huang X, Tian X, Zhang H, He L, Wang Y, et al. GATA4 regulates Fgf16 to promote heart repair after injury. Development. 2016;143:936–49.
Zeisberg EM, Ma Q, Juraszek AL, Moses K, Schwartz RJ, Izumo S, et al. Morphogenesis of the right ventricle requires myocardial expression of Gata4. J Clin Invest. 2005;115:1522–31.
Bisping E, Ikeda S, Kong SW, Tarnavski O, Bodyak N, McMullen JR, et al. Gata4 is required for maintenance of postnatal cardiac function and protection from pressure overload-induced heart failure. Proc Natl Acad Sci U S A. 2006;103:14471–6.
Oka T, Maillet M, Watt AJ, Schwartz RJ, Aronow BJ, Duncan SA, et al. Cardiac-specific deletion of Gata4 reveals its requirement for hypertrophy, compensation, and myocyte viability. Circ Res. 2006;98:837–45.
Kikuchi K, Holdway JE, Werdich AA, Anderson RM, Fang Y, Egnaczyk GF, et al. Primary contribution to zebrafish heart regeneration by gata4(+) cardiomyocytes. Nature. 2010;464:601–5.
Delgado-Olguin P, Huang Y, Li X, Christodoulou D, Seidman CE, Seidman JG, et al. Epigenetic repression of cardiac progenitor gene expression by Ezh2 is required for postnatal cardiac homeostasis. Nat Genet. 2012;44:343–7.
Chen L, Ma Y, Kim EY, Yu W, Schwartz RJ, Qian L, et al. Conditional ablation of Ezh2 in murine hearts reveals its essential roles in endocardial cushion formation, cardiomyocyte proliferation and survival. PLoS One. 2012;7:e31005.
Ahmed A, Wang T, Delgado-Olguin P. Ezh2 is not required for cardiac regeneration in neonatal mice. PLos One. 2018;13:e0192238.
Porrello ER, Olson EN. A neonatal blueprint for cardiac regeneration. Stem Cell Res. 2014;13:556–70.
Piccolo S, Dupont S, Cordenonsi M. The biology of YAP/TAZ: hippo signaling and beyond. Physiol Rev. 2014;94:1287–312.
Xin M, Kim Y, Sutherland LB, Qi X, McAnally J, Schwartz RJ, et al. Regulation of insulin-like growth factor signaling by Yap governs cardiomyocyte proliferation and embryonic heart size. Sci Signal. 2011;4:ra70.
von Gise A, Lin Z, Schlegelmilch K, Honor LB, Pan GM, Buck JN, et al. YAP1, the nuclear target of Hippo signaling, stimulates heart growth through cardiomyocyte proliferation but not hypertrophy. Proc Natl Acad Sci U S A. 2012;109:2394–9.
Xin M, Kim Y, Sutherland LB, Murakami M, Qi X, McAnally J, et al. Hippo pathway effector Yap promotes cardiac regeneration. Proc Natl Acad Sci U S A. 2013;110:13839–44.
Del Re DP, Yang Y, Nakano N, Cho J, Zhai P, Yamamoto T, et al. Yes-associated protein isoform 1 (Yap1) promotes cardiomyocyte survival and growth to protect against myocardial ischemic injury. J Biol Chem. 2013;288:3977–88.
Nishioka N, Inoue K, Adachi K, Kiyonari H, Ota M, Ralston A, et al. The Hippo signaling pathway components Lats and Yap pattern Tead4 activity to distinguish mouse trophectoderm from inner cell mass. Dev Cell. 2009;16:398–410.
Heallen T, Zhang M, Wang J, Bonilla-Claudio M, Klysik E, Johnson RL, et al. Hippo pathway inhibits Wnt signaling to restrain cardiomyocyte proliferation and heart size. Science. 2011;332:458–61.
Wang J, Liu S, Heallen T, Martin JF. The hippo pathway in the heart: pivotal roles in development, disease, and regeneration. Nat Rev Cardiol. 2018;15:672–84.
Heallen T, Morikawa Y, Leach J, Tao G, Willerson JT, Johnson RL, et al. Hippo signaling impedes adult heart regeneration. Development. 2013;140:4683–90.
Chen JF, Murchison EP, Tang R, Callis TE, Tatsuguchi M, Deng Z, et al. Targeted deletion of Dicer in the heart leads to dilated cardiomyopathy and heart failure. Proc Natl Acad Sci U S A. 2008;105:2111–6.
Zhao Y, Ransom JF, Li A, Vedantham V, von Drehle M, Muth AN, et al. Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1-2. Cell. 2007;129:303–17.
Tatsuguchi M, Seok HY, Callis TE, Thomson JM, Chen JF, Newman M, et al. Expression of microRNAs is dynamically regulated during cardiomyocyte hypertrophy. J Mol Cell Cardiol. 2007;42:1137–41.
Porrello ER, Johnson BA, Aurora AB, Simpson E, Nam YJ, Matkovich SJ, et al. MiR-15 family regulates postnatal mitotic arrest of cardiomyocytes. Circ Res. 2011;109:670–9.
Tian Y, Liu Y, Wang T, Zhou N, Kong J, Chen L, et al. A microRNA-Hippo pathway that promotes cardiomyocyte proliferation and cardiac regeneration in mice. Sci Transl Med. 2015;7:279ra238.
Huang W, Feng Y, Liang J, Yu H, Wang C, Wang B, et al. Loss of microRNA-128 promotes cardiomyocyte proliferation and heart regeneration. Nat Commun. 2018;9:700.
Boon RA, Iekushi K, Lechner S, Seeger T, Fischer A, Heydt S, et al. MicroRNA-34a regulates cardiac ageing and function. Nature. 2013;495:107–10.
Yang Y, Cheng HW, Qiu Y, Dupee D, Noonan M, Lin YD, et al. MicroRNA-34a plays a key role in cardiac repair and regeneration following myocardial infarction. Circ Res. 2015;117:450–9.
Eulalio A, Mano M, Dal Ferro M, Zentilin L, Sinagra G, Zacchigna S, et al. Functional screening identifies miRNAs inducing cardiac regeneration. Nature. 2012;492:376–81.
•• Gabisonia K, Prosdocimo G, Aquaro GD, Carlucci L, Zentilin L, Secco I, et al. MicroRNA therapy stimulates uncontrolled cardiac repair after myocardial infarction in pigs. Nature. 2019;569:418–22 Findings from this study suggests that heart regeneration through cardiomyocyte proliferation is achievable in large animals, but caution is warranted since widespread cardiomyocyte proliferation can have serious side effects.
He L, Thomson JM, Hemann MT, Hernando-Monge E, Mu D, Goodson S, et al. A microRNA polycistron as a potential human oncogene. Nature. 2005;435:828–33.
O’Donnell KA, Wentzel EA, Zeller KI, Dang CV, Mendell JT. c-Myc-regulated microRNAs modulate E2F1 expression. Nature. 2005;435:839–43.
Ventura A, Young AG, Winslow MM, Lintault L, Meissner A, Erkeland SJ, et al. Targeted deletion reveals essential and overlapping functions of the miR-17 through 92 family of miRNA clusters. Cell. 2008;132:875–86.
Chen J, Huang ZP, Seok HY, Ding J, Kataoka M, Zhang Z, et al. mir-17-92 cluster is required for and sufficient to induce cardiomyocyte proliferation in postnatal and adult hearts. Circ Res. 2013;112:1557–66.
Frangogiannis NG. Inflammation in cardiac injury, repair and regeneration. Curr Opin Cardiol. 2015;30:240–5.
Han C, Nie Y, Lian H, Liu R, He F, Huang H, et al. Acute inflammation stimulates a regenerative response in the neonatal mouse heart. Cell Res. 2015;25:1137–51.
Aurora AB, Porrello ER, Tan W, Mahmoud AI, Hill JA, Bassel-Duby R, et al. Macrophages are required for neonatal heart regeneration. J Clin Invest. 2014;124:1382–92.
Lavine KJ, Epelman S, Uchida K, Weber KJ, Nichols CG, Schilling JD, et al. Distinct macrophage lineages contribute to disparate patterns of cardiac recovery and remodeling in the neonatal and adult heart. Proc Natl Acad Sci U S A. 2014;111:16029–34.
Wodsedalek DJ, Paddock SJ, Wan TC, Auchampach JA, Kenarsary A, Tsaih SW, et al. IL-13 promotes in vivo neonatal cardiomyocyte cell cycle activity and heart regeneration. Am J Physiol Heart Circ Physiol. 2019;316:H24–34.
Bassat E, Mutlak YE, Genzelinakh A, Shadrin IY, Baruch Umansky K, Yifa O, et al. The extracellular matrix protein agrin promotes heart regeneration in mice. Nature. 2017;547:179–84.
Tsen G, Halfter W, Kroger S, Cole GJ. Agrin is a heparan sulfate proteoglycan. J Biol Chem. 1995;270:3392–9.
Notari M, Ventura-Rubio A, Bedford-Guaus SJ, Jorba I, Mulero L, Navajas D, et al. The local microenvironment limits the regenerative potential of the mouse neonatal heart. Sci Adv. 2018;4:eaao5553.
Kumar A, Brockes JP. Nerve dependence in tissue, organ, and appendage regeneration. Trends Neurosci. 2012;35:691–9.
Garikipati VN, Verma SK, Kishore R. The nervous heart: role of sympathetic reinnervation in cardiac regeneration. Circ Res. 2015;117:980–1.
Mahmoud AI, O’Meara CC, Gemberling M, Zhao L, Bryant DM, Zheng R, et al. Nerves regulate cardiomyocyte proliferation and heart regeneration. Dev Cell. 2015;34:387–99.
White IA, Gordon J, Balkan W, Hare JM. Sympathetic reinnervation is required for mammalian cardiac regeneration. Circ Res. 2015;117:990–4.
Vaseghi M, Lux RL, Mahajan A, Shivkumar K. Sympathetic stimulation increases dispersion of repolarization in humans with myocardial infarction. Am J Physiol Heart Circ Physiol. 2012;302:H1838–46.
Mohamed TMA, Ang YS, Radzinsky E, Zhou P, Huang Y, Elfenbein A, et al. Regulation of cell cycle to stimulate adult cardiomyocyte proliferation and cardiac regeneration. Cell. 2018;173(104–116):e112.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Alisson C. Cardoso, Ana Helena M. Pereira, and Hesham A. Sadek declare that they have no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Regenerative Medicine
Rights and permissions
About this article
Cite this article
Cardoso, A.C., Pereira, A.H.M. & Sadek, H.A. Mechanisms of Neonatal Heart Regeneration. Curr Cardiol Rep 22, 33 (2020). https://doi.org/10.1007/s11886-020-01282-5
Published:
DOI: https://doi.org/10.1007/s11886-020-01282-5