Abstract
Neonatal mammalian heart has been shown to possess the capacity to regenerate substantially after an injury. This remarkable regenerative capacity is lost in a week. This transition has been marked with cardiomyocyte cell cycle arrest and induction of fibrotic response similar to what occurs after myocardial infarction in adult hearts. Recent studies outlined the function of several cardiogenic factors that play a pivotal role in neonatal cardiac regeneration. However, underlying molecular mechanisms of neonatal cardiac regeneration and other cardiogenic factors remained elusive. Here, we investigated the involvement of novel putative cardiogenic factors in neonatal cardiac regeneration and cardiomyocyte cell cycle withdrawal. We have shown that Cbl, Dnmt3a, and Itch are significantly downregulated during neonatal cardiac regeneration process after cardiac injury in vivo. Intriguingly, several of studied factors are upregulated in non-regenerative period of 7-day-old mice after cardiac injury. Knockdown of Cbl, Dnmt3a and Itch in rat neonatal cardiomyocytes lead to the induction of cardiomyocyte proliferation. Cardiomyocyte proliferation accompanies upregulation of positive regulators of cardiomyocyte division and downregulation of CDKIs. Taken together, our findings suggest that Cbl, Dnmt3a, and Itch may be involved in the regulation of cardiomyocyte cell cycle withdrawal and may represent new targets for the induction of cardiac regeneration.
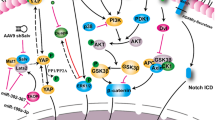
Graphic Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Studies using mammalian heart regeneration models have shown that after resection of the almost entire ventricular apex or induction of myocardial infarction (MI) by ligating left anterior descending (LAD) arteria, the neonatal mouse heart was able to regenerate completely with minimal fibrosis in the site of injury [1, 2]. Both types of injury resulted in regeneration by induction of immune cell response and switched on unique transcriptional program leading proliferation of cardiomyocytes, without fibrosis [3, 4]. Interestingly, this regenerative phenomenon overlaps with cardiomyocyte cell cycle arrest in neonatal mouse and one week after birth the mouse heart is not able to initiate regeneration after injury [1, 2]. These results have shown for the first time that a mammalian heart is able to regenerate itself completely. This model has been a powerful model to examine cardiomyocyte proliferation and neonatal heart regeneration regulators, thereby allowing the discovery of the regenerative capacity in the human heart. This mammalian cardiac regeneration model has also been used to elucidate the role of Meis1 in cardiomyocyte cell cycle arrest [5].
In our previous studies, we have discovered a new modulator (Meis1) effective in cardiomyocyte cell cycle arrest and neonatal cardiac regeneration using neonatal mammalian cardiac regeneration models [5, 6]. However, the signaling pathways and other molecular mechanisms involved in neonatal mammalian cardiac regeneration after MI has not been fully defined. Previous studies have shown that Meis1 and Hif-1α genes play an important role in the regulation of cell cycle and metabolic activities of hematopoietic stem cells (HSC) [6, 7]. Silencing Meis1 and Hif-1α genes specifically in HSCs increased their proliferative capacity. Similarly, we have shown that the expression of Meis1 gene is increased in the heart in the first week after birth resulting in cardiac cell cycle arrest [5]. In vitro knock down of Meis1 in cardiomyocytes or specific in vivo knock out have resulted in increased cardiomyocyte proliferation. Moreover, using chromatin immunoprecipitation (CHIP) experiment, Meis1 was reported to be interacting and cooperatively working with Hoxb13 to induce cardiomyocyte proliferation [8]. Our in vivo studies have shown that Meis1 is involved in cardiac regeneration following apical resection and MI in neonatal mice. In this process, we found that Meis1 transcriptionally regulates cyclin-dependent kinase inhibitors INK4a locus genes, p21, and others. To this end, to determine potent modulators of neonatal cardiac regeneration similar to Meis1, we studied and identified potent factors based on their loss-of-function in a quiescence cell type i.e., hematopoietic stem cells and their potent expression during cardiac development. Bioinformatical and literature curation studies led to identification of four potent factors; Inpp5d, Itch, Dnmt3a, and Cbl.
Inpp5d (inositol-5-phosphatase, akas SHIP1) belongs to the SH2 domain containing protein family, which demonstrates a negative regulatory role in hematopoiesis [1]. Itch, an E3 Ubiquitin ligase, acts as a transcriptional corepressor of p45/NF-E2 and is a negative regulator of hematopoietic stem cell (HSC) expansion [2]. Itch knockout mice showed an increase in HSC proliferation and repopulation [3]. Dnmt3a is involved in DNA methylation by adding methyl groups to specific CpG regions. Studies using conditional ablation of Dnmt3a in animals have shown an increase in the number of HSCs in bone marrow [4, 5]. These findings suggested that Dnmt3a is a critical regulator of the epigenetic mechanisms causing HSC proliferation. Cbl, on the other hand, encodes a RING finger E3 ubiquitin ligase thus it plays a role in ubiquitination. Cbl mutations have been shown to be associated with myeloproliferative disorders (MPD). Moreover, in a recent study, the association between the combined deletion of Cbl and Cbl-b, and expansion of the HSC pool and MPD development was reported [6]. Cell cycle analysis with HSC from Cbl knockout mice showed reduced stasis and cell cycle entry. Studies have shown that loss-of-function of these modulators play a significant role in the increase of stem cell population. Moreover, higher expression of these factors in the heart tissue is associated with microarray studies. Based on these findings and bioinformatical analysis of gene expression during cardiac development, Inpp5d, Itch, Dnmt3a, and Cbl were determined as putative cardiogenic factors and further investigated for their involvement in neonatal cardiac regeneration and cardiomyocyte cell cycle arrest.
Materials and Methods
RNA Isolation and Real-Time qPCR
Expression of putative cardiomyocyte cell cycle modulators in the cardiac tissue and other organs was analyzed with real-time qPCR method as previously described [5, 6]. Briefly, tissues were collected from mice after euthanasia and sacrifice. Collected samples were powdered in liquid nitrogen by mortar; and then, RNA was isolated by trizol method. Samples were homogenized in a solution (Trizol, Sigma, Cat. No: T9424) containing 1 mL guanidine thiocyanate with a syringe. Then, isopropanol and ethanol washes were performed. RNA concentration was determined using NanoDrop (ThermoFisher). From each tissue, total of 5 μg RNA sample was converted into cDNA using random primers and ProtoScript II First Strand cDNA Synthesis Kit (NEB, Cat. No: E656). Briefly, RNA was incubated for 5 min at 65 °C with random hexamers. After addition of enzyme and reaction mix into random hexamer and RNA mixture, incubation was applied at 25 °C for 10 min, at 50 °C for 50 min and at 85 °C for 5 min consecutively. Samples were stored at − 20 °C after dilution. Each gene specific primers (Table 1) were determined using NIH primer depot (http://mouseprimerdepot.nci.nih.gov) and ordered from Sentebiolab, Turkey. Desired gene regions were amplified from cDNAs with Bio-Rad FX96 TouchTM Real-Time qPCR Detection System (95 °C × 10 min, 95 °C × 10 s, 60 °C × 20 s, 72 °C × 30 s, 30 cycle). The expression of each amplified putative modulator gene was normalized by GAPDH content using ΔΔCt method.
Western Blot Analysis
After powdering tissue samples in liquid nitrogen by mortar, tissue samples were lysed with RIPA Lysis Buffer (Santa Cruz, CA, USA) containing PMSF and protease inhibitors and homogenized using sonicator. In addition, proteins were isolated from cardiomyocyte, fibroblast and aorta tissue of adult heart with EASYCELL-CM Langendorff Machine and 1% collagenase enzyme. After measurement of protein concentration with Pierce™ Coomassie (Bradford) Protein Assay Kit (Cat. No: 23200), protein samples were prepared as 30 μg and mixed with 4X Laemmli Sample Buffer (BIO-RAD, Cat. No: 1610747) and incubated at 95 °C for 5 min. Protein samples were loaded into 8–12% polyacrylamide gel and run at 120 V. Bands were transferred to PVDF membrane at 350 mA for 70 min. Membranes were blocked for 1 h at room temperature with either 5% milk or 5% BSA solution according to antibodies. After overnight primary antibody (Actin 1:1000 in 5% BSA, Dnmt3a 1:500 5% BSA, Cbl 1:200 in 5% milk, Ship1 1:200 5% BSA, Itch 1:200 in 5% milk) incubation at 4 °C, membranes were washed 10 min with TBST for three times. HRP-linked secondary antibody (1:1000 in 5% milk or 5% BSA solution according to primary antibody) incubation was performed for 2 h at 4 °C. After washing membrane 10 min with TBST for three times, protein expression level was detected with Cell Signaling (Signal fire) ECL and Bio-Rad ChemiDoc MP System.
Isolation and ex vivo Culture of Adult Cardiac Cells
After aorta (composed of endothelial cells and smooth muscle cells) was taken from adult mouse hearts by a micro scissor, and then samples were crushed with a mortar. Half of it was mixed with 1 mL of Trizol and stored at − 80 °C for RNA isolation, and the other half was stored in − 80° in RIPA lysis buffer for Western blot analysis. The adult cardiac tissue was treated with collagenase to obtain fibroblasts and cardiomyocytes as we have previously described [5, 6]. Fibroblasts were separated from the cardiomyocytes using PRIMARI cell plates. In addition, for collection of viable and pure cardiomyocytes from adult mouse hearts, EASYCELL-CM (Harvard apparatus) system was used. Following enzymatic perfusion of the heart, the cells were collected and left for gravity settlement. The pellet that was formed within 5 min (containing cardiomyocytes) were separated and the cells were evaluated under the microscope. Thereafter, half of the cells was mixed with 1 mL of Trizol and stored at − 80 °C for RNA isolation, and the other half was stored at -80° in RIPA lysis buffer for Western blot analysis. The solution that contains fibroblasts was seeded in two separate PRIMARI cell culture dishes (10 cm). The adhering fibroblasts were allowed to grow for 3–5 days in cell culture. One dish containing fibroblasts was used for RNA isolation with Trizol. The fibroblasts in the other dish were harvested by RIPA lysis buffer and stored at −80° for Western blot analysis. RNA isolation, Western blot analysis and RT-qPCR studies were performed as previously described. The expression of putative cardiomyocyte cell cycle modulators were analyzed in cardiomyocytes, cardiac fibroblasts, and in endothelial cells isolated from the aorta.
Immunohistochemical Staining of Cardiac Tissue
Immunohistochemistry studies were performed on paraffin sections like in previous studies [1, 5]. Adult mice hearts were incubated overnight in neutral formaldehyde at 4 °C. Similarly, 4 days after myocardial infarction, hearts were collected from 7-days-old (regenerating condition) and 11-day-old mice (non-regenerating condition). On the following day, samples were hardened by being kept in tissue embedding station before paraffin application. A slice having 5 μM thickness were cut from samples and then transferred on poly-L-lysine slides. Before incubating slides in xylene solution at 15–25 °C for 15 min twice, they were heated in the oven for 15 min at 65–75 °C. Rehydration step was performed in decreasing concentrations of ethanol; and then, slides were blocked in 1% BSA for 30 min. After 30 min boiling in sodium citrate solution (pH 6.1), slides were incubated overnight at 4 °C with cardiac troponin T (Thermoscientific, MS-295-P1, 1:200 dilution) and primary antibodies specific to putative cardiomyocyte cell cycle modulators. Next day, detection was performed using Alexa Flour 488 donkey anti-mouse and Alexa Flour 555 anti-rabbit secondary antibodies (1:500) and DNA marker Hoechst 33,342 dye. Putative modulator expression in cardiomyocytes found in TnnT + regions was observed under 40X with fluorescence microscope. On the other hand, expression of non-cardiomyocyte cells was observed in TnnT- regions.
Neonatal Myocardial Injury
Neonatal MI procedure was applied to 3- and 7-day-old mice as previously described [2, 9]. In this procedure, ligature was applied to LAD of neonatal mice with surgical polyethylene 7–0 suture in order to induce myocardial infarction. Alternatively, by apical resection, tissue damage was provided by taking 15% slice from surface of heart left ventricle. Thoracotomy was also applied without any ligature or apical resection to control groups (sham). Briefly, mice were awaited for < 5 min in ice as anesthesia and thorax was opened with thoracotomy to perform LAD ligation. With the help of 6–0 sutures, thorax was closed and described procedure was completed in less than 5 min. After surgery, mice were waited on a heat pad (37 °C) and put into cage where mother mouse was found. Furthermore, in order to prevent cannibalism, olfactory perception of mother mouse was manipulated with menthol smell.
Rat Neonatal Cardiomyocyte Isolation and Culture
Hearts of 1–2-day-old rats were removed in laminar flow hood and only ventricular parts of the hearts were put into digestion solution containing 0.1% Pancreatin. After shaking at 100–120 rpm for 20 min at 37 °C, they were centrifuged at 2000 rpm for 10 min. Fibroblasts found in pellet were removed by being seeded on 4 × 10 cm BD Falcon PRİMARİA tissue culture dish (Cat. No: 353803) and incubated for 2 h at 37 °C. Cells were collected slowly and filtered through 70–100 μm filter. 500,000 cells/mL were seeded on plate coated with gelatin with myocyte media (3:1 DMEM: M199, Pen/Strep, L-Glutamine (2 mM), 10% Horse Serum, 5% FBS) and incubated at 37 °C in incubator having 5% CO2.
siRNA Treatments and Gene Expression Analysis
12 μl lipofectamine (Invitrogen) and 50 nM siRNA for each putative cardiomyocyte cell modulators (Table 2) were incubated in 200 μL serum-free media at room temperature for 20 min. Silencer Select Pre-designed siRNAs (NegControl siRNA, Applied Biosystems, Ambion) were used as a control. siRNAs were added drop by drop and 6 h incubation was performed. 3–5 days later, cells were collected with trypsin and stored in TRIZOL until RNA isolation. RNA isolation was performed according to manufacturer’s protocol (Qiagen, RNeasy Mini Kit). 2 μg RNA was converted into cDNA with SuperScript III RT system (Invitrogen). Using primers (Table 3) which were designed for rat and SyberGreen (Applied Biosystems), real-time qPCR was performed in BioRad Mycycler. GAPDH was used as a housekeeping gene in normalization of gene expression using ΔΔCt method. Gene expression levels of CDKIs including P15INK4b, P16INK4a, P19AR, P18, P21, P27, and P57 genes in Cbl siRNA treated cardiomyocytes were determined.
Immunocytochemistry
Cardiomyocytes were grown until they form 50–70% confluency (5 × 105 cells/wells of 6-well plate). siRNA treatment was performed as previously described. After treatment, cardiomyocytes were fixed on slide with 4% PFA (5 min, at room temperature). Thereafter, cells were permeabilized in 2 min with 0.1% Triton at room temperature. After 30 min blocking with 3% goat serum and washing, they were incubated at room temperature with phospho-histone H3 (PH3, mitosis marker, Ser10, 1:100, rabbit polyclonal, Millipore, MA), Aurora B (cytokinesis marker, 1:25, rabbit polyclonal, Sigma, MO) and cardiac troponin T (TnnT, cardiomyocyte marker, Thermoscientific MS-295-P1, 1:100, mouse monoclonal). Alexa Fluor 488 donkey anti-mouse (Invitrogen, Cat. No: A-21202, 1:400) and Alexa Fluor 555 donkey antirabbit (Invitrogen, 1:400) secondary antibodies and Hoechst 33,342 (Invitrogen, CA) were used for detection. GE Cytell automated Fluorescence Microscope was used to determine Ph3 + TnnT + and AuroraB + TnnT + cardiomyocyte numbers.
Statistical Analysis
Results were analyzed with Student’s t test for any significant difference p < 0.05 was considered significant compared to control or untreated samples.
Results
Analysis of Putative Cardiomyocyte Cell Cycle Modulators in Cardiac Tissue in Comparison to Other Organs and Tissues
Bioinformatical and literature curation studies of potent cell cycle modulators related to cardiomyocytes have led us the initial identification of putative cardiomyocyte cell cycle modulators. These factors were selected based on their loss-of-function in a quiescence cell type i.e., hematopoietic stem cells and their potent expression during cardiac development. To this end, we have analyzed the expressions of four of these potent modulators including Inpp5d (SHIP), Itch, Dnmt3a, and Cbl genes in the adult heart in comparison to kidney, skeletal muscle, intestine, brain, liver, lung, bone, and spleen with real-time qPCR (Fig. 1A). Itch, c-Myc, Inpp5d, Dnmt3a, and Cbl gene expressions were validated. Furthermore, Meis-1, a known negative modulator of neonatal cardiac regeneration [5], was analyzed alongside and it was shown that it is expressed in heart and skeletal muscle as well as in other organs (Figure S1). Expression levels of each gene in heart was analyzed with real-time qPCR and normalized according to GAPDH. It was detected that Cbl is more expressed than studied genes (Fig. 1A). In addition to real-time qPCR, gene expressions were studied with Western blot in several adult mice tissues. Detection of CBL, Dnmt3a and Itch proteins in heart was validated (Fig. 1B). Expression of INPP5D in the heart was barely detected (Fig. 1B).
Analysis of potent modulators by RT-PCR and Western blot. A Analysis of expression of putative cardiomyocyte cell cycle modulators in the heart in comparison to other Organs. B Validation of expression in selected organs by Western blot. C Differential gene expression in cardiac cells by RT-PCR. D Western blot analysis in cardiac cells. n = 3
We have also characterized the expression these putative cardiomyocyte cell cycle modulators in different cardiac cell types. Expressions of Inpp5d, Itch, Dnmt3a, and Cbl genes in cardiac endothelial cells, cardiomyocytes and cardiac fibroblasts were analyzed (Fig. 1C). Inpp5d, Itch, Dnmt3a and Cbl had higher expression in cardiomyocytes in comparison to cardiac endothelial cells. Intriguingly, Itch had higher expression in cardiomyocytes in comparison to cardiac fibroblasts or endothelial cells. For the same cell types and genes, gene expression was also confirmed with Western blot (Fig. 1D). It was observed that Dnmt3a and Cbl genes were expressed at different levels in different cell types of heart. CBL protein was clearly detectable in cardiomyocytes at high level. SHIP could not be detected by Western blot in cardiomyocytes. Moreover, high expression level of Itch was observed in endothelial cells.
Expression of same genes in endothelial, cardiomyocytes and cardiac fibroblasts were verified by immunohistochemistry in the cardiac tissue sections by staining with TnnT2 as a cardiomyocyte marker (Figure S2). SHIP was detected in cardiomyocytes (TnnT + cells) (Figure S2A). Itch protein was detected in cardiomyocytes (Figure S2B). Itch protein displayed negative staining in fibroblast and endothelial cells (DAPI + TnnT- cells) (Figure S2B). Dnmt3a was found in nucleus as expected, but it was not particularly in cardiomyocytes (Figure S2C). CBL was detected in cardiomyocytes but it was not generally observed in cardiac fibroblasts (Figure S2D).
Differential Expression of Putative Cardiomyocyte Cell Cycle Modulators During Neonatal Cardiac Regeneration
Four different putative cardiomyocyte cell cycle modulators Inpp5d, Itch, Dnmt3a, and Cbl, were further investigated for their differential gene expression during neonatal cardiac regeneration. We hypothesized that, in regenerating mouse hearts, these modulators are to be reduced in expression, while in non-regenerating mouse hearts there would be an increase in expression or at least no significant change would occur. To this end, we have performed LAD ligation to induce MI during regenerative and non-regenerative windows. The surgical operations in neonatal mice were performed as we have done previously [2, 9].
MI was performed in < 3-day-old (regenerating heart, P3) and 7-day-old (non-regenerated heart, P7) neonatal mice. In neonatal mice, MI was induced by LAD ligation, which is physiologically more similar to cardiac infarction (Fig. 2A). After fours day following the operations, cardiac tissue was harvested. Gene expression levels in regenerating (P3) and non-regenerating heart (P7) tissues were determined by real-time qPCR analysis. It was determined that the expression of Inpp5d, Dnmt3a, and Cbl were decreased during cardiac regeneration as expected (Fig. 2B). While itch showed no difference in gene expression during the regenerative window in comparison to sham-operated heart, the expression of itch was significantly increased in the non-regenerative window (Fig. 2C). Similarly, in the absence of cardiac regeneration, Inpp5d, and Dnmt3a expression were increased compared to sham. These findings suggest that the decreased expression of the Inpp5d, Dnmt3a, and Cbl genes during heart regeneration and increased expression during the non-regenerative window, they could have a negative effect on cardiac regeneration. RNA expression may not always correlate with protein content. Thus, we performed Western blot analysis of putative cardiomyocyte cell cycle modulators in regenerative and non-regenerative period as well.
Neonatal cardiac injury and analysis of gene expression. A The experimental plan to study differential gene expression between regenerative and non-regenerative periods in neonatal cardiac injury mouse model. The LAD ligations were performed at postnatal day 3 (P3) and 7 (P7). Heart tissues were collected four days after operation. Quantification of selected putative cardiomyocyte cell cycle modulators during B Regenerative Period and C Non-Regenerative Period by RT-PCR. n = 3, *p < 0.05
To this end, tissue and protein samples were collected from mice in regenerative (P3 Sham vs P3 LAD groups) and non-regenerative phases (P7 Sham vs P7 LAD groups), and Western blot studies were performed for selected genes (Fig. 3A). It was confirmed that the protein content was significantly decreased during cardiac regeneration for the Itch, SHIP, Dnmt3a and CBL proteins (Fig. 3B). On the other hand, Itch and SHIP protein levels were increased in the non-regeneration phase (Fig. 3C).
Western blot analysis in cardiac regeneration. A Western blot analysis of selected putative cardiomyocyte cell cycle modulators post cardiac injuries. LAD operations were performed at P3 and P7, and samples were collected after four days. Quantification of selected putative cardiomyocyte cell cycle modulators during B Regenerative Period and C Non-Regenerative Period. n = 3, *p < 0.05, **p < 0.01
Although the amount of proteins expresses an increased in activity, changes in the localization of proteins also needs to be characterized. Therefore, immunohistochemical staining studies were performed and protein levels were investigated in samples obtained from regenerated ad non-regenerated myocardium. Inpp5d is expressed in the cytoplasm of cardiomyocytes but not in fibroblasts (Figure S3). Itch protein shows more cytoplasmic localization in the myocardium (Figure S4). No changes in localization were observed for the control and LAD operated mouse samples, both for the P3 and P7 samples. ITCH was also not detected in fibroblasts by IHC analysis. Some nuclear staining was observed in P3 samples for the Dnmt3a protein, but overall, this protein has more of cytoplasmic localization (Figure S5). Dnmt3a was not detected in fibroblast and endothelial cells. In P7 LAD samples, there is a change in expression in cardiomyocytes, and more nuclear positive cells are observed. Although we observed a decrease in the amount of protein in Western blot studies, the increased nuclear localization of Dnmt3a during the non-regenerative phase is consistent with our hypothesis. Indicating that Dnmt3a is a putative negative cardiac regeneration regulator, and this may be related to the increased nuclear localization for non-regenerative samples. The CBL protein largely shows a cytoplasmic profile in immunohistochemistry studies (Figure S6). Nuclear staining was not readily observed. CBL is highly present in cardiomyocytes, whereas a negative staining was observed in fibroblasts and endothelial cells. On the other hand, based on the overall light intensity that we observed, our results show a higher expression of CBL in P7 LAD samples compared to the other conditions.
Effect of Putative Cardiac Modulators on Cardiomyocyte Proliferation by siRNA Treatment
In order to study the effect of putative cardiac modulators on cardiomyocyte proliferation, siRNA knockdown was done. It was shown that target gene expression was decreased at least 60% by three siRNAs (Fig. 4A). Effect of siRNAs was also confirmed by Western blot (Figure S7A–D). Then, rat neonatal cardiomyocyte cells were freshly isolated and treated with corresponding siRNAs. After 3 days, cells were fixed with paraformaldehyde and proliferating cardiomyocytes were analyzed by immunostaining with Anti-Tnnt2 and Anti-pH3 antibodies (Fig. 4B). Itch and Dnmt3a siRNAs induced cardiomyocyte proliferation to some extent (Fig. 4B). Cbl siRNA showed a distinct and significant increase in proliferation (Fig. 4B). We have also analyzed molecular pathway that Cbl modulates in cardiomyocytes after Cbl siRNA treatment (Fig. 4C). It was found that the expressions of genes negatively regulated by Cbl (Stat3, Stat3a) have increased. In addition, there was an increase in the expressions of positive regulators that has effect on cardiomyocyte proliferation (Cyclin B1, Erbb4, Erbb2, Periostin, and Fgf1) and decrease in expressions of negative regulators (p15, p19arf, p21, and p27kip1). These findings outlined the identification and involvement of novel modulator of neonatal cardiac regeneration.
siRNA targeting Cbl induces cardiomyocyte division and modulates gene expression. A Real-Time PCR analysis of target gene expressions after corresponding siRNA treatments. B Quantification of proliferating cardiomyocyte number after siRNA treatment of neonatal cardiomyocytes by counting of TnnT2 & Ph3 double positive cells. C Gene expression analysis related to Cbl pathway and cardiomyocyte proliferation after Cbl sirna treatment. n = 3, **p < 0.01
Discussion
Heart failure is a severe disease with a high mortality rate that affects millions of patients worldwide. The most important factor in the pathophysiology of heart failure is that the human heart is not able to regenerate itself following an injury. Contrary to this, the functional cardiomyocytes die and are replaced by fibrotic tissue which causes the problem related to cardiac function. Although the mammalian heart has been known as a non-regenerative organ for decades, there is growing evidence that it retains the capability to renew cardiomyocytes during adulthood [10,11,12,13]. However, this is not enough to cause a significant functional recovery after injury. Regeneration of injured heart tissues requires the formation of new cardiomyocytes, and these are formed by preexisting cardiomyocyte proliferation [13]. The adult mammalian heart contains limited numbers of dividing and differentiating cells, which makes the study of mammalian cardiac regeneration modulators difficult [13]. Studies with mammalian heart regeneration models have shown complete regenerative potential in neonatal mouse heart following resection of the entire ventricular apex, myocardial infarction, or cryoinjury [1, 2, 14]. All types of injury may result in regeneration by widespread proliferation of cardiomyocytes, without hypertrophy and fibrosis. These findings have shown for the first time regeneration in a mammalian heart and opened a window to discover the potential regenerative mechanism of the stationary human heart. Moreover, after an injury, transcriptional changes in neonatal vs adult mouse heart showed epigenetic modifications including chromatin modifications around cell cycle genes are major obstacles to start regenerative program in adult ones [15]. However, there is insufficient information in literature about cardiogenic factors involved in neonatal cardiac regeneration.
In our previous studies, we have discovered a new modulator, Meis1, effective in cardiomyocyte cell cycle arrest and neonatal cardiac regeneration, using neonatal mammalian cardiac regeneration models [5, 6]. However, other signaling pathways and other molecular mechanisms involved in neonatal mammalian cardiac regeneration after MI have not been fully defined. In this direction, we aimed to identify new putative cardiomyocyte cell cycle modulators. Deletion or knockdown of Meis1 gene in cardiomyocytes induces cardiomyocyte proliferation. We studied cardiomyocyte cell cycle modulators that normally arrest cardiac muscle in vitro and in vivo and detected expression of potent putative cardiomyocyte cell cycle modulators in heart tissue, especially in cardiomyocytes. The relationship of putative cardiomyocyte cell cycle modulator expressions in regenerating < 3-day-old mice heart after MI and non-regenerating 7 days-old mice was studied.
In the first week after birth, mouse cardiomyocytes demonstrate dramatic changes in the differentiation of contractile proteins to isoforms found in adult hearts, followed by the induction of DNA synthesis without cytokinesis, and G0/G1 cell cycle arrest. These changes result in bi-nucleation of cardiomyocytes, which is a characteristic of maturation [16, 17]. While cyclin-dependent kinase inhibitors are highly prevalent in adult cardiomyocytes, the activity of positive cell cycle regulators is decreased. These factors are affecting cardiomyocyte cell cycle re-entry and have been suggested as primary limiting factors for cardiac regeneration after cardiac injury. On the other hand, some studies related to the expression of periostin, neuregulin, Fgf1, and cell cycle regulator cyclin D2 and the induction of Hippo, have demonstrated that to some extent cardiomyocytes can be induced to re-enter the cell cycle [18,19,20,21,22]. These studies show that it is possible to develop therapeutic approaches targeting cardiomyocyte cell cycle. Previous findings have reported that cardiogenic factors such as IL3, FGF10, C3orf58, Oncostatin M, TNF-induced apoptosis (TWEAK), and weak periostin inducer result in increased number of cardiomyocytes [18, 23,24,25,26]. Gene manipulation studies involving p27KIP1, mir-133a and Salvador homologue (Salv) knock out, have resulted in increased cardiomyocyte proliferation assessed with mitosis marker phospho-histone H3 (pH3) [20, 27, 28]. In addition, overexpression studies on c-myc, E1A, cyclin B1-CDC2, cyclin A2, cyclin D2, Notch signalling pathway, YAP, and ERBB2 have shown induction of cardiomyocyte proliferation [22, 29,30,31,32,33,34,35].
Cell cycle is a vital process for all cell types and is composed of a group of protein complexes. Major cell cycle regulators are cyclin-dependent kinases (CDK), CDK inhibitors (CDKI), CDK activating kinases (CAK), and retinoblastoma (Rb) family proteins. The cyclin/CDK complex has an important role as positive regulator of the cell cycle and this complex requires activation by phosphorylation of CAK. This complex is controlled by de CDKIs including the Cip/Kip family, and the Ink4 family. The members of these families are linked to Cyclin-D/E/A-dependent kinases and the CyclinD-CDK4/6 complex, respectively, and inhibit their activity.
Most of the studies have shown that CDKI levels in adult cardiomyocytes are increased, which associated with the suppression of positive cell cycle regulators. Various studies have shown that Cdk2 and c-myc are cell cycle regulators that increase the expression levels of cyclins such as CDK4 and CDKs resulting in cardiomyocyte growth. In addition, deletion of the CDKI p27Kip1 or immunodepletion of p21Cip1 in cardiomyocytes causes progression to the S phase and thus results in cardiomyocyte proliferation and an increase in heart size [21, 36]. The deletion of Meis1 induces the upregulation of CDKs and downregulation of CDKIs such as p16, p15, p19ARF, p21, and p57. Moreover, Meis1 deletion promotes upregulation of positive cell cycle regulators such as MCM3, Check1, and Ccnd2 and downregulation of the negative regulators such as APbb1, TP53 and Gpr132 [5].
Cbl proteins are highly conserved and function as ubiquitin ligase. Reported evidence showed their function as regulator of cell junctions proteins via focal adhesion protein turnover which further leads myofibril degeneration [39]. Although, c-Cbl or Cbl-b deficient mice showed no embryonic lethality, loss of both genes resulted with embryonic death [40, 41]. Interestingly, in another report, inhibition of only c-Cbl increased cardiac function after myocardial ischemia/reperfusion by suppressing EGFR and FAK pathways and lowering myocyte death in response to H2O2 [42].
Epigenetic regulations govern embryonic development as well as multiple different gene functions. Among the others, DNA methyltransferases (DNMTs) are known to be master regulators of DNA methylation DNMT1, for instance, methylate newly synthesized DNA strand and terminates transferring methylation marks during cell division [43]. On the other hand, together with DNMT3B, DNMT3A regulates DNA methylation in all different stages of cell [44]. Exceptionally, Dnmt3a and Dnmt3b deficient mice are lack of cardiac function deficiency. Using, germline knockout mice and transverse aortic constriction (TAC) injury model, there was no evidence regarding their importance for the adult mice heart function [45].
Cardiac regeneration is related with re-activation of cardiomyocyte cell cycle in zebrafish, newt and neonatal mice [1, 37, 38]. The discovery of cardiomyocyte cell cycle modulators could provide a new platform to develop new cardiovascular therapeutics targeting cardiomyocyte cell cycle. Neonatal mice surgery method was used to determine downregulation or upregulation of candidate genes. Similar to previous findings of Mahmoud et al. [9]. This method allowed us to study changes in mRNA level in regenerating (P3) and not regenerating (P7) myocardium. After surgery, downregulations of Inpp5d, Dnmt3a, and Cbl were observed during neonatal cardiac regeneration. After cardiomyocyte isolation from neonatal rats, siRNA treatment was applied in cell culture and they were analyzed on the day 4. Immunostaining was also done for TnnT2 and phospho-histone H3 (a mitosis marker) to study cardiomyocyte specific proliferation. Results showed that Cbl, Itch and Dnmt3a siRNA treatment-induced cardiomyocyte proliferation. Our study showed that Cbl along with Itch and Dnmt3a have role in mice cardiac regeneration as negative regulators and they can be targeted to induce mice cardiac regeneration. Cbl, seems to inhibit several cardiogenic factors such as Cyclin B1, Erbb4, Erbb2, Periostin, and Fgf1 that positively induce cardiomyocyte proliferation. Thus, targeting of Cbl, as we have shown here, could allow both upregulation of positive regulators of cardiomyocyte proliferation, and cardiomyocyte division as measure by TnnT2 + Ph3 + cells. This was also associated with downregulation of several CDKIs in the cardiomyocytes. Overall, the negative effects of these regulators on cardiomyocyte proliferation and cardiac regeneration are valuable targets to develop new therapies to fix cardiovascular damage.
Change history
02 February 2022
A Correction to this paper has been published: https://doi.org/10.1007/s00246-022-02833-z
References
Porrello ER, Mahmoud AI, Simpson E et al (2011) Transient regenerative potential of the neonatal mouse heart. Science 331(6020):1078–1080. https://doi.org/10.1126/science.1200708
Porrello ER, Mahmoud AI, Simpson E et al (2013) Regulation of neonatal and adult mammalian heart regeneration by the miR-15 family. Proc Natl Acad Sci USA 110(1):187–192. https://doi.org/10.1073/pnas.1208863110
Cui M, Wang Z, Chen K et al (2020) Dynamic transcriptional responses to injury of regenerative and non-regenerative cardiomyocytes revealed by single-nucleus RNA sequencing. Dev Cell 53(1):102-116.e8. https://doi.org/10.1016/j.devcel.2020.02.019
Aurora AB, Porrello ER, Tan W et al (2014) Macrophages are required for neonatal heart regeneration. J Clin Invest 124(3):1382–1392. https://doi.org/10.1172/JCI72181
Mahmoud AI, Kocabas F, Muralidhar SA et al (2013) Meis1 regulates postnatal cardiomyocyte cell cycle arrest. Nature 497(7448):249–253. https://doi.org/10.1038/nature12054
Kocabas F, Zheng J, Thet S et al (2012) Meis1 regulates the metabolic phenotype and oxidant defense of hematopoietic stem cells. Blood 120(25):4963–4972. https://doi.org/10.1182/blood-2012-05-432260
Aksoz M, Turan RD, Albayrak E, Kocabas F (2017) Emerging roles of Meis1 in cardiac regeneration, stem cells, and cancer. Curr Drug Targets 18:181–190. https://doi.org/10.2174/1389450118666170724165514
Nguyen NUN, Canseco DC, Xiao F et al (2020) A calcineurin–Hoxb13 axis regulates growth mode of mammalian cardiomyocytes. Nature 582(7811):271–276. https://doi.org/10.1038/s41586-020-2228-6
Mahmoud AI, Porrello ER, Kimura W, Olson EN, Sadek HA (2014) Surgical models for cardiac regeneration in neonatal mice. Nat Protoc 9(2):305–311. https://doi.org/10.1038/nprot.2014.021
Quaini F, Urbanek K, Beltrami AP et al (2002) Chimerism of the transplanted heart. N Engl J Med 346(1):5–15. https://doi.org/10.1056/NEJMoa012081
Canseco DC, Kimura W, Garg S et al (2015) Human ventricular unloading induces cardiomyocyte proliferation. J Am Coll Cardiol 65(9):892–900. https://doi.org/10.1016/j.jacc.2014.12.027
Bergmann O, Bhardwaj RD, Bernard S et al (2009) Evidence for cardiomyocyte renewal in humans. Science 324(5923):98–102. https://doi.org/10.1126/science.1164680
Senyo SE, Steinhauser ML, Pizzimenti CL et al (2013) Mammalian heart renewal by pre-existing cardiomyocytes. Nature 493(7432):433–436. https://doi.org/10.1038/nature11682
Darehzereshki A, Rubin N, Gamba L et al (2015) Differential regenerative capacity of neonatal mouse hearts after cryoinjury. Dev Biol 399(1):91–99. https://doi.org/10.1016/j.ydbio.2014.12.018
Quaife-Ryan GA, Sim CB, Ziemann M et al (2017) Multicellular transcriptional analysis of mammalian heart regeneration. Circulation 136(12):1123–1139. https://doi.org/10.1161/CIRCULATIONAHA.117.028252
Soonpaa MH, Kim KK, Pajak L, Franklin M, Field LJ (1996) Cardiomyocyte DNA synthesis and binucleation during murine development. Am J Physiol Heart Circ Physiol 271(540–545):2183–2189
Li F, Wang X, Capasso JM, Gerdes AM (1996) Rapid transition of cardiac myocytes from hyperplasia to hypertrophy during postnatal development. J Mol Cell Cardiol 28(8):1737–1746. https://doi.org/10.1006/jmcc.1996.0163
Kühn B, Del Monte F, Hajjar RJ et al (2007) Periostin induces proliferation of differentiated cardiomyocytes and promotes cardiac repair. Nat Med 13(8):962–969. https://doi.org/10.1038/nm1619
Gemberling M, Karra R, Dickson AL, Poss KD (2015) Nrg1 is an injury-induced cardiomyocyte mitogen for the endogenous heart regeneration program in zebrafish. Elife 4:05871. https://doi.org/10.7554/eLife.05871
Heallen T, Zhang M, Wang J et al (2011) Hippo pathway inhibits wnt signaling to restrain cardiomyocyte proliferation and heart size. Science 332(6028):458–461. https://doi.org/10.1126/science.1199010
Engel FB, Hsieh PCH, Lee RT, Keating MT (2006) FGF1/p38 MAP kinase inhibitor therapy induces cardiomyocyte mitosis, reduces scarring, and rescues function after myocardial infarction. Proc Natl Acad Sci USA 103(42):15546–15551. https://doi.org/10.1073/pnas.0607382103
Pasumarthi KBS, Nakajima H, Nakajima HO, Soonpaa MH, Field LJ (2005) Targeted expression of cyclin D2 results in cardiomyocyte DNA synthesis and infarct regression in transgenic mice. Circ Res 96(1):110–118. https://doi.org/10.1161/01.RES.0000152326.91223.4F
Kubin T, Pöling J, Kostin S et al (2011) Oncostatin M is a major mediator of cardiomyocyte dedifferentiation and remodeling. Cell Stem Cell 9(5):420–432. https://doi.org/10.1016/j.stem.2011.08.013
Novoyatleva T, Diehl F, Van Amerongen MJ et al (2010) TWEAK is a positive regulator of cardiomyocyte proliferation. Cardiovasc Res 85(4):681–690. https://doi.org/10.1093/cvr/cvp360
Beigi F, Schmeckpeper J, Pow-Anpongkul P et al (2013) C3orf58, a novel paracrine protein, stimulates cardiomyocyte cell-cycle progression through the PI3K-AKT-CDK7 pathway. Circ Res 113(4):372–380. https://doi.org/10.1161/CIRCRESAHA.113.301075
Rochais F, Sturny R, Chao CM et al (2014) FGF10 promotes regional foetal cardiomyocyte proliferation and adult cardiomyocyte cell-cycle re-entry. Cardiovasc Res 104(3):432–442. https://doi.org/10.1093/cvr/cvu232
Von Harsdorf R, Hauck L, Mehrhof F, Wegenka U, Cardoso MC, Dietz R (1999) E2F–1 overexpression in cardiomyocytes induces downregulation of p21(CIP1) and p27(KIP1) and release of active cyclin-dependent kinases in the presence of insulin-like growth factor I. Circ Res 85(2):128–136. https://doi.org/10.1161/01.RES.85.2.128
Liu N, Bezprozvannaya S, Williams AH et al (2008) microRNA-133a regulates cardiomyocyte proliferation and suppresses smooth muscle gene expression in the heart. Genes Dev 22(23):3242–3254. https://doi.org/10.1101/gad.1738708
Jackson T, Allard MF, Sreenan CM, Doss LK, Bishop SP, Swain JL (1990) The c-myc proto-oncogene regulates cardiac development in transgenic mice. Mol Cell Biol 10(7):3709–3716. https://doi.org/10.1128/mcb.10.7.3709
Agah R, Kirshenbaum LA, Abdellatif M et al (1997) Adenoviral delivery of E2F–1 directs cell cycle reentry and p53- independent apoptosis in postmitotic adult myocardium in vivo. J Clin Invest 100(11):2722–2728. https://doi.org/10.1172/JCI119817
Von Gise A, Lin Z, Schlegelmilch K et al (2012) YAP1, the nuclear target of Hippo signaling, stimulates heart growth through cardiomyocyte proliferation but not hypertrophy. Proc Natl Acad Sci U S A 109(7):2394–2399. https://doi.org/10.1073/pnas.1116136109
D’Uva G, Aharonov A, Lauriola M et al (2015) ERBB2 triggers mammalian heart regeneration by promoting cardiomyocyte dedifferentiation and proliferation. Nat Cell Biol 17(5):627–638. https://doi.org/10.1038/ncb3149
Cheng RK, Asai T, Tang H et al (2007) Cyclin A2 induces cardiac regeneration after myocardial infarction and prevents heart failure. Circ Res 100(12):1741–1748. https://doi.org/10.1161/CIRCRESAHA.107.153544
Bicknell KA, Coxon CH, Brooks G (2004) Forced expression of the cyclin B1-CDC2 complex induces proliferation in adult rat cardiomyocytes. Biochem J 382(2):411–416. https://doi.org/10.1042/BJ20031481
Campa VM, Gutiérrez-Lanza R, Cerignoli F et al (2008) Notch activates cell cycle reentry and progression in quiescent cardiomyocytes. J Cell Biol 183(1):129–141. https://doi.org/10.1083/jcb.200806104
Tseng AS, Engel FB, Keating MTT (2006) The GSK-3 inhibitor BIO promotes proliferation in mammalian cardiomyocytes. Chem Biol 13(9):957–963. https://doi.org/10.1016/j.chembiol.2006.08.004
Piatkowski T, Mühlfeld C, Borchardt T, Braun T (2013) Reconstitution of the myocardium in regenerating newt hearts is preceded by transient deposition of extracellular matrix components. Stem Cells Dev 22(13):1921–1931. https://doi.org/10.1089/scd.2012.0575
Kikuchi K, Holdway JE, Werdich AA et al (2010) Primary contribution to zebrafish heart regeneration by gata4+ cardiomyocytes. Nature 464(7288):601–605. https://doi.org/10.1038/nature08804
Parry TL, Willis MS (2016) Cardiac ubiquitin ligases: their role in cardiac metabolism, autophagy, cardioprotection and therapeutic potential. Biochim Biophys Acta Mol Basis Dis 1862(12):2259–2269. https://doi.org/10.1016/j.bbadis.2016.07.002
Naramura M, Jang IK, Kole H, Huang F, Haines D, Gu H (2002) C-Cbl and Cbl-b regulate T cell responsiveness by promoting ligand-induced TCR down-modulation. Nat Immunol 3(12):1192–1199. https://doi.org/10.1038/ni855
Murphy MA, Schnall RG, Venter DJ et al (1998) Tissue hyperplasia and enhanced T-cell signalling via ZAP-70 in c-Cbl-deficient mice. Mol Cell Biol 18(8):4872–4882. https://doi.org/10.1128/mcb.18.8.4872
Rafiq K, Kolpakov MA, Seqqat R et al (2014) C-Cbl inhibition improves cardiac function and survival in response to myocardial ischemia. Circulation 129(20):2031–2043. https://doi.org/10.1161/CIRCULATIONAHA.113.007004
Quaife-Ryan GA, Sim CB, Porrello ER, Hudson JE (2016) Resetting the epigenome for heart regeneration. Semin Cell Dev Biol 58:2–13. https://doi.org/10.1016/j.semcdb.2015.12.021
Smith ZD, Meissner A (2013) DNA methylation: roles in mammalian development. Nat Rev Genet 14(3):204–220. https://doi.org/10.1038/nrg3354
Nührenberg TG, Hammann N, Schnick T et al (2015) Cardiac myocyte de novo DNA methyltransferases 3a/3b are dispensable for cardiac function and remodeling after chronic pressure overload in mice. PLoS ONE 10(6):e0131019. https://doi.org/10.1371/journal.pone.0131019
Acknowledgements
We like to thank to the support by The Scientific and Technological Research Council of Turkey (TÜBİTAK) ARDEB 1001 [#115S185] program. FK is supported by funds provided by EU, BAGEP-2015, ICGEB, Gilead Sciences, and ERA-CVD program. DY has been supported by TÜBİTAK-BİDEB 2209A program. We like to thank Prof. Dr. Bayram Yuksel and Unal Uslu from Yeditepe University for their help in the establishment of tissue sectioning and immunostaining. We would like to thank Dr. Emrah Nikerel from Yeditepe University for their help in bioinformatical and statistical analysis.
Author information
Authors and Affiliations
Contributions
G.S.A performed neonatal cardiac injuries and performed RT-qPCR, contributed to manuscript. S.N.E performed IHC and cardiac injuries, and contributed to manuscript. S.Y and S.A. contributed to RT-qPCR and neonatal cardiomyocyte culture. A.C. contributed to IHC studies. F.P. performed and contributed to Western blot studies, and manuscript. F.K designed the all experiments and wrote the article. All authors contributed to data analysis and reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
All authors declare that they have no conflicts of interest concerning this work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original article has been revised due to addition of affiliation.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Aslan, G.S., Polat, F., Eren, S.N. et al. Identification of Novel and Potent Modulators Involved in Neonatal Cardiac Regeneration. Pediatr Cardiol 42, 1554–1566 (2021). https://doi.org/10.1007/s00246-021-02640-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00246-021-02640-y