Abstract
Background
Polycystic ovarian syndrome (PCOS), a gynae-endocrine disorder, has a relatively high risk of differential expression of miRNA (DE-miRNA) in the disease progression.
Aims
To identify the DE-miRNA in the progression of PCOS in the ovarian cumulus cells.
Methods
The microarray dataset GSE72274 was analysed for PCOS-associated DE-miRNAs. miRNet identifies the target genes. Protein–protein interaction (PPI) network was constructed and hub genes were analysed by topology and module analysis. Transcription factors (TFs) and protein kinases (PKs) regulating the hub genes were identified using X2K tool. Biological functions were analysed using DAVID software. Finally, the DGIdb drug-gene interaction tool identifies the candidate medications.
Results
A total of 1577 DE-miRNAs linked to PCOS were identified, with 13 meeting the specified criteria. Subsequently, its 2053 target genes were retrieved through miRNet. Topology and module analysis identified the hub genes VEGFA, SOX2, KRAS, AKT1, and SMAD4 that are implicated in ovarian regulation. Notably, the study highlighted the significant role of the wnt signalling pathway, which is involved in ovarian function, specifically in follicle development, corpus luteum formation, and steroid production. Additionally, six TFs and PKs were identified as important regulators of these hub genes, and the potential medication interactions identified 11 medicines for VEGFA, KRAS, AKT1, and SMAD4 genes, while no suitable drug for SOX2 was identified.
Conclusion
Identified, hub genes are known to associate with the regulation of ovarian function such as oocyte development, and steroid synthesis via the wnt signalling pathway.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
PCOS is a gynae-endocrinological disease that affects women's health worldwide. Hyperandrogenism, menstrual irregularities, and ultrasonic polycystic ovaries are the major diagnostic manifestations of the disease [1]. This life-threatening disease is accompanied by comorbidities such as infertility, obesity, hirsutism, acne, depression, type 2 diabetes, and ovarian cancer [2]. The disease is multifactorial; however, the consequences of ovarian dysfunction are the major concern in the disease pathogenesis. Furthermore, a reduction in the development potential of oocytes is one of the significant factors observed in PCOS. In the ovary, the cumulus cells offer conduits for the passage of nutrients, regulatory chemicals, and paracrine substances during oocyte maturation which promotes nuclear and cytoplasmic maturation of the oocyte [3]. Thus, the identification of potential biomarkers in the cumulus cells of oocytes will help to predict the outcome and may create space for disease management. Furthermore, recent studies have focused more on epigenetic alterations most importantly the promising therapeutic role of microRNAs (miRNAs) in disease regulation [4].
miRNAs are endogenous single-stranded small noncoding RNA molecules that are crucial post-transcriptional controllers of cellular activities such as proliferation, apoptosis, migration, invasion, stress response, and immunological regulation [5]. Moreover, these small molecules are abundant in almost all body fluids such as serum, plasma, whole blood, follicular fluid, and ovarian cells such as granulosa cells and cumulus cells [6, 7]. Studies evidence that several external factors such as environmental disrupting chemicals, malnutrition, and mode of birth influence the miRNA expression, which can further lead to diseases in several medical conditions [8]. Furthermore, differently expressed miR-127-3p, miR-24-3p, miR-151-3p, miR-574-3p, and miRNA-361-3p have affected the pathways such as ovarian functioning, hormonal imbalance, insulin signalling, and metabolic disorder [2, 8, 9]. In the ovary, miRNA-mediated studies have much focused on the genes involved in the functioning of the granulosa cells [10]. Furthermore, miRNA regulation of genes involved in the functioning of the cumulus cells, majorly involved in the oocyte maturation and fertilization in the ovarian follicle[11], is not much explored in PCOS pathogenesis. Hence, appropriate therapeutic approaches addressing miRNA regulation in the cumulus cells of the ovary might prove potential biomarkers for PCOS treatment.
In this regard, a comprehensive, integrated network biology analysis was performed to identify differentially expressed microRNAs (DE-miRNAs) in PCOS and their potential target mRNAs, which could serve as candidate biomarkers for the condition. DE-miRNAs were initially screened from the Gene Expression Omnibus (GEO) database, and their corresponding target mRNAs were retrieved using miRNet. A network of these genes was then constructed, and hub genes were identified through topology and module analysis. Transcription factors (TFs), protein kinases (PKs), biological functions, and effective drugs that regulate these hub genes were also identified. Finally, a miRNA-mRNA regulatory interaction network was constructed using the target gene data retrieved from miRNet.
Materials and methods
Retrieval of microarray data
The clinical datasets that compared the expression of miRNA in ovarian cumulus cells with and without PCOS were extracted from the publicly accessible data repository GEO database (https://www.ncbi.nlm.nih.gov/geo) using the keywords “differentially expressed miRNA,” “PCOS,” “polycystic ovary syndrome,” and “homo sapiens.” Titles, abstracts, summary, and sample types were screened; only the dataset GSE72274 with the platform PL16543Agilent-038166 cbc_human_miR18.0 consisting of 5 PCOS and normal controls were found suitable for the integrative network analysis were considered.
Screening of DE-miRNAs and prediction of target genes
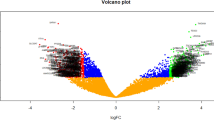
The DE-miRNAs were identified using the GEO2R analytic tool, with the following criteria: a p-value < 0.01, a log2 fold change (FC) of < − 1 for downregulated miRNAs, and > 3 for upregulated miRNAs. Subsequently, the target genes of these DE-miRNAs were identified by inputting the miRNA IDs into miRNet (http://www.mirnet.ca/) [12].
Protein–protein interaction network and identification of hub genes
The list of target genes retrieved from miRNet was subjected to the online tool (STRING) (https://www.string-db.org/) [13] to predict the interactions among proteins and to construct their PPI network. The confidence score was set to 0.4, and the PPI network file was imported into Cytoscape software (version 3.9.1) [14]. Then, the hub genes were determined through module and topology analysis of the PPI network. The network was first explored for its significant dense regions known as modules, using MCODE, a Cytoscape plug-in. Then, cytoHubba, a Cytoscape plug-in, predicted the top 10 crucial genes from five topological parameters (bottleneck, closeness, betweenness, MCC, and degree) parameter of the PPI network [14]. Finally, the genes overlapping in the modules and topological parameters were selected as the hub genes.
Construction of a regulatory network of hub genes
The regulatory network of hub genes was constructed using the X2K database (https://maayanlab.cloud/X2K/) [15]. First, the TFs of the hub genes and the intermediate proteins were identified using TF enrichment analysis, and then, the PKs responsible for controlling the hub gene expression were determined by kinase enrichment analysis.
Functional annotation and pathway enrichment analysis
The database for annotation, visualization, and integrated discovery (DAVID, http://david.abcc.ncifcrf.gov/) was used to perform gene ontology (GO) functional annotation and Kyoto encyclopedia of genes and genomes (KEGG) pathway enrichment analysis to understand the role of target genes of potential DE-miRNAs [16]. p-value < 0.05 was regarded as statistically significant.
Construction of miRNA-mRNA network and drug-gene interaction
After screening the DE-miRNAs, the miRNA-mRNA potential regulatory network was predicted using the miRNA dataset. In addition, the DGIdb drug-gene interaction tool (DGIdb, https://dgidb.org) was used to predict the potential drugs that can target the identified hub genes [17].
Results
DE-miRNA and their target genes associated with PCOS
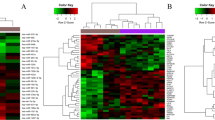
The GEO2R analysis for identifying DE-miRNAs revealed that most miRNAs were downregulated in the cumulus cells of the ovary in PCOS. The analysis identified a total of 1577 DE-miRNAs, of which 1545 were downregulated and 32 were upregulated. Among these, only 13 DE-miRNAs (7 upregulated and 6 downregulated) met the criteria set for further investigation. Subsequently, miRNet analysis revealed 1377 target genes for the 7 upregulated miRNAs and 676 target genes for the 6 downregulated DE-miRNAs (Table 1).
Identification of the hub genes
The target genes of DE-miRNAs were mapped in the STRING database to construct the PPI network, the loosely bound nodes were removed, and the network was visualized using Cytoscape (Fig. 1). MCODE analysis of the network identified a significant module of 5 genes (Table 2) and cytoHubba analysis using a novel algorithm, degree, BC, CC, MCC, bottleneck revealed the top 10 essential genes within the network. Finally, vascular endothelial growth factor A (VEGFA), sex-determining region Y box 2 (SOX2), Kirsten rat sarcoma virus (KRAS), AKT Serine/Threonine Kinase 1(AKT1), and SMAD family member 4 (SMAD4) were identified as the hub genes as these are found to be common in the module and topology analysis (Table 3).
Hub gene regulatory network
TFs, PKs, and intermediate proteins involved in the regulation of hub genes were identified using the X2K database. A total of 26 TFs were identified as regulators of the hub genes. Among these, UBTF, PPARG, RCOR1, SOX2, E2F1, and NFE2L were found to be particularly significant (Table 4). These significant TFs were shown to interact with 62 intermediate proteins (Supplementary Table S1). Additionally, KEA analysis identified 227 PKs, with CDK4, ERK2, MAPK8, CK2ALPHA, MAPK14, and MAPK3 having the highest number of connections (Supplementary Table S2). The top enriched TFs, PKs, expanded PPI, and upstream regulatory network are shown in Fig. 2.
Transcription factor enrichment analysis and their network with protein kinases and intermediate proteins. The transcription factors, regulatory network of top enriched transcription factors (TFs), protein kinase (PKs), and expanded protein–protein network (PPI) network were ranked based on their p-value
Functional annotation and pathway enrichment analysis
DEGs were functionally categorized using DAVID, and the top-five terms are indicated for GO function and enrichment analysis (Fig. 3). The results revealed that the DEGs involved in the top terms for BP enrichment analysis include, positive regulation of cell adhesion, negative regulation of fat cell differentiation and cell cycle, cellular response to amino acid stimulus and UV, response to peptide hormone and activity. Similarly, the top terms for the CC enrichment analysis include transcriptional repressor complex, euchromatin, cell surface, membrane raft, and cyclin cyclin-dependent protein kinase holoenzyme complex. Furthermore, the MF enrichment analysis of DEGs was mainly enriched in core promoter region sequence-specific DNA binding, Protein kinase C binding, SMAD binding, E-box binding, protein domain specific binding, and Hsp70 protein binding (Fig. 3a). Additionally, the pathway enrichment analysis using KEGG showed that DEGs were mainly involved in pkg signalling, Wnt signalling, adherens junction, tuberculosis, neutrophil extracellular trap formation, herpes simplex virus1 infection, IL17 signalling pathway, melanogenesis, leukocyte transendothelial migration, and salmonella infection (Fig. 3b).
Construction of potential miRNA-mRNA network in PCOS
An integrated platform linking miRNAs and their targets named miRNet was used to predict the target genes of the screened DE-miRNAs. The screened upregulated and downregulated DE-miRNAs were entered into the web platform, and the data of the potential target genes of DE-miRNAs were downloaded. Then, these data were input into the Cytoscape 3.9.1 software to construct the miRNA-mRNA network. Based on this investigation, six miRNA-mRNA networks have been identified, which include miR-183-5p/KRAS, miR-183-5p/SMAD4, miR-933/SMAD4, miR-126-3p/VEGFA, miR-126-3p/SOX2, and miR-126-3p/AKT1 (Fig. 4).
Drug-gene interactions
The DGIdb drug-gene interaction analysis identified 17 drugs targeting the four hub genes: VEGFA, KRAS, AKT1, and SMAD4. Lysine for SMAD4 and Ranibizumab for VEGFA showed the highest interaction scores 10.3 and 8.54 respectively (Table 5).
Discussion
Globally 10% of the women are suffering from PCOS, leading to exalted health and economic burden. It can be either caused by ovarian dysfunction associated with steroid imbalance, or metabolic disturbances defined by insulin resistance and obesity. More research is being undertaken to examine the influencing variables of PCOS [8]. The latest research highlights the therapeutic importance of miRNAs in several diseases. However, it has not been well explored in PCOS progression. Furthermore, the effect of oocyte maturation is also a major concern in PCOS, and the cumulus cells supply the required nutrients for oocyte maturation. Thus, the identification of potential biomarkers in the cumulus cells of oocytes may help to create space for recovery [3]. So, here we tried to explore the miRNA-mRNA network contributing to the PCOS pathogenesis in the ovarian cumulus cells. Hence, we examined differently expressed miRNAs from the cumulus cells data set of PCOS. We discovered three potential DE-miRNAs, miR-126-3p, miR-183-5p, and miR-933 contributing to the PCOS pathogenesis.
Among the identified three DE-miRNAs, relatively limited studies have been reported on miR-126-3p in PCOS. The single nucleotide polymorphisms (SNPs), haplotype combinations, and the differential expression of miR-126-3p are correlated with menstrual irregularity, increased antral follicle count, and hormonal imbalance in PCOS. Furthermore, SNPs of miR-126 are also associated with an increased risk of endometriosis. In addition, it promoted angiogenesis and attenuated GC apoptosis in the ovary in premature ovarian failure in the rat model [18,19,20]. Furthermore, the miR-183-5p and miR-933 are not been studied in PCOS so far. However, overexpressed miR-183-5p induces stress and its knockdown could protect the neurons by promoting cell proliferation and migration thereby inhibiting cell death in hepatocellular carcinoma via targeting insulin receptor substrate1 [21], while mir-933 is associated with liver cancer by enhancing pyruvate kinase isoform M2 [22]. Also, its role has been identified in the regulation of hyperglycaemic conditions and hyperinsulinism in type II diabetes mellitus [23].
The topology analysis of the PPI network revealed VEGFA, SOX2, KRAS, SMAD4, and AKT1 as the hub genes. High levels of VEGFA in various solid tumors are mainly responsible for angiogenesis through different mechanisms, such as regulating angiogenesis and vascular permeability, affecting immune cell function and modulating fibroblast function in the cancer stroma VEGFA accelerates tumorigenesis [24, 25]. In addition, VEGFA administration in animal models proved the improved outcome of increased angiogenesis and increased neuronal density in cerebral vasculature diseases and beneficial therapy for early acute kidney injury [26, 27]. SMAD4 serves as the central mediator of Transforming growth factor β (TGF-β) signalling and plays important roles in many biological processes, including cell growth, differentiation, apoptosis, migration, and cancer initiation and progression. Multiple studies have revealed that decreased SMAD4 does not initiate tumor formation alone but can promote tumor progression initiated by other genes, such as KRAS, and APC [28]. Via TGF-β signaling SMAD4 plays a role in granulosa cell function and follicular development in the ovary [29]. Even letrozole-induced PCOS mice model exhibits TGF-β/SMAAD4 pathway-mediated metabolic disturbances, sex hormone imbalance, oxidative stress, ovarian fibrosis, and reduced SMAD4 leads to arteriovenous malformations in mice models in hemorrhagic telangiectasia a genetic disorder [29, 30]. An increase in the inflammatory pathways, angiogenesis, and progression of endometriotic tissues can cause iron-mediated damage to the DNA, affecting the KRAS pathways [31, 32]. Furthermore, in the mouse model of pancreatic adenocarcinoma, KRAS inactivated by CRISPR-mediated genome editing demonstrates the ability to form tumors in mice [22]. High expression of AKT1 is associated with granulosa cell proliferation and follicular development [33]. It has even been shown to be expressed in insulin-sensitive tissues such as the liver, skeletal muscles, and adipose tissue AKT1 is crucial for initiating intracellular post-insulin receptor substrate-1 (IRS-1) phosphorylation [34]. Furthermore, it has been proved as a promising target for the treatment of angiogenesis-dependent pathologies, such as cancer and ischemic injury in mice model [35]. SOX2 as a transcriptional modulator with co-factors imposes cell-fate-determining expression patterns. Various secondary modifications have been described for SOX2 (involving phosphorylation, methylation, acetylation, ubiquitination, SUMOylation, PARPylation, and O-GlcNAcylation), but their relevance remains underexplored mainly [36]. Increased expression of SOX2 abolishes the tumorigenesis and knockdown of SOX2 has been suggested as a good treatment option for osteosarcoma in mouse-modelled studies [37]. So far, no work has been done on SOX2 in PCOS. The critical regulators of these hub genes were predicted by identifying the TFs, PKs, and intermediate proteins. 26 TFs significantly regulated these 5 hub genes, 62 intermediate proteins, and 227 protein kinases, including CDK4, ERK2, MAPK8, CK2ALPHA, MAPK14, and MAPK3, were identified. Among the 6 significant TFs, PPARG corresponded to increased androgens and LH/FSH ratio in the ovary of PCOS patients. miR-424-5p mediated inhibition of GC proliferation and cellular senescence in the ovary observed in PCOS via E2E1 pathways [38].
The biological functions of these hub genes show significantly enriched GO pathways such as cellular response to UV, fat cell differentiation, amino acid stimulus, negative regulation of cell cycle, and promoter-proximal region sequences specific DNA binding. The KEGG pathways were involved mainly in the Cushing syndrome, cGMP-PGK signalling, wnt signalling, adherens junctions, and IL-17 signalling pathway. Evidence shows the importance of wnt signalling in ovarian function related to follicle development, oocyte development, follicle maturation, corpus luteum formation, steroid production, and fertility. In addition, other studies suggest the identified DE-miRNA and the hub genes are involved majorly in ovarian functioning [39,40,41,42]. All this evidence highlights the importance of the KEGG pathway identified significant Wnt signalling pathway in PCOS.
Finally, to seek efficient therapeutic options for the treatment of PCOS, we further screened the hub gene targets of identified miRNAs for suitable drug candidates. It was found that Ranibizumab, Elmiron, Pegaptanib sodium, Bevasiranib, aflibercept, Bevasizumab, and Risuteganib were found to interact with VEGFA whereas Panitumumab and AZD-4785 were found to have interaction with KRAS. Furthermore, Ipatasertib, Chembl 480356, Chembl 1086397, Eupalinin A, Gigantol, and Perifosine were observed to interact with AKT1 whereas lysine and Alectinib were found to interact with SMAD4. The efficiency and effect of these interactions require validation through further experiments.
This in silico analysis highlights the potential role and interplay of miRNA and mRNA in the pathogenesis of PCOS. However, these findings need further validation through in-vitro and in-vivo experiments using clinical samples. Such experimental validation is crucial to confirm the functional relevance of these molecular interactions and to translate these insights into viable therapeutic strategies for PCOS.
Conclusions
Overall, the present in silico analysis highlights the crucial role of hub genes and TFs associated with miR-183-5p, miR-126-3p, and miR-933 in ovarian function. The constructed miRNA-mRNA regulatory networks, including miR-183-5p/KRAS, miR-183-5p/SMAD4, miR-933/SMAD4, miR-126-3p/VEGFA, miR-126-3p/SOX2, and miR-126-3p/AKT1, may contribute to the pathogenesis of PCOS by disrupting steroid balance and influencing oocyte development through Wnt signalling pathways. Understanding these miRNA-mRNA interactions could provide valuable insights for developing targeted clinical treatment strategies for PCOS.
Data availability
The data of the study is available upon reasonable request.
References
Huo Y, Ji S, Yang H et al (2022) Differential expression of microRNA in the serum of patients with polycystic ovary syndrome with insulin resistance. Ann Transl Med 10:762–762. https://doi.org/10.21037/ATM-22-2941
AE Butler, V Ramachandran, S Hayat, SR Dargham, TK Cunningham, M Benurwar, T Sathyapalan, SH Najafi-Shoushtari, SL Atkin (2019) Expression of microRNA in follicular fluid in women with and without PCOS. Sci Rep 9. https://doi.org/10.1038/S41598-019-52856-5.
Q Shen, M Chen, X Zhao, Y Liu, … XR-S (2020) biology in, undefined 2020, Versican expression level in cumulus cells is associated with human oocyte developmental competence, Taylor Fr. Shen, M Chen, X Zhao, Y Liu, X Ren, L ZhangSystems Biol Reprod Med 2020 Taylor Fr. 66:176–184. https://doi.org/10.1080/19396368.2020.1725685
G Rashid, NA Khan, D Elsori, RA Youness, H Hassan, D Siwan, N Seth, MA Kamal, S Rizvi, AM Babker, W Hafez (2024) miRNA expression in PCOS: unveiling a paradigm shift toward biomarker discovery. Arch Gynecol Obstet 1–17. https://doi.org/10.1007/S00404-024-07379-4/METRICS
Chen HX, Fu YF, Guo ZX, Zhou XD (2022) MicroRNA-29c-3p participates in insulin function to modulate polycystic ovary syndrome via targeting Forkhead box O 3. Bioengineered 13:4361–4371. https://doi.org/10.1080/21655979.2022.2033014
Butler AE, Ramachandran V, Hayat S et al (2019) Expression of microRNA in follicular fluid in women with and without PCOS. Sci Rep 9:1–9. https://doi.org/10.1038/s41598-019-52856-5
L Mu, X Sun, M Tu, D Zhang (2021) Non-coding RNAs in polycystic ovary syndrome: a systematic review and meta-analysis. Reprod Biol Endocrinol 19. https://doi.org/10.1186/S12958-020-00687-9
Bhandary P, Shetty PK, Manjeera L, Patil P (2022) Hormonal, genetic, epigenetic and environmental aspects of polycystic ovarian syndrome. Gene Reports 29:101698. https://doi.org/10.1016/J.GENREP.2022.101698
Y Xuan Wu, Y Shan Lin, S Chen Li, X Yao, M Cheng, L Zhu, H Ying Liu (2021) microRNA-194 is increased in polycystic ovary syndrome granulosa cell and induces KGN cells apoptosis by direct targeting heparin-binding EGF-like growth factor. Reprod Biol Endocrinol 19. https://doi.org/10.1186/S12958-021-00850-W
Alexandri C, Daniel A, Bruylants G, Demeestere I (2020) The role of microRNAs in ovarian function and the transition toward novel therapeutic strategies in fertility preservation: from bench to future clinical application. Hum Reprod Update 26:174–196. https://doi.org/10.1093/HUMUPD/DMZ039
Turathum B, Gao E-M, Chian R-C, Jessus C (2021) The function of cumulus cells in oocyte growth and maturation and in subsequent ovulation and fertilization. Mdpi Com. https://doi.org/10.3390/cells10092292
Chang L, Xia J (2023) MicroRNA regulatory network analysis using miRNet 2.0. Methods Mol Biol 2594:185–204. https://doi.org/10.1007/978-1-0716-2815-7_14
Szklarczyk D, Gable AL, Lyon D et al (2019) STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47:D607–D613. https://doi.org/10.1093/NAR/GKY1131
Zhou F, Xing Y, Cheng T et al (2022) Exploration of hub genes involved in PCOS using biological informatics methods. Med (United States) 101:E30905. https://doi.org/10.1097/MD.0000000000030905
Clarke DJB, Kuleshov MV, Schilder BM et al (2018) eXpression2Kinases (X2K) Web: linking expression signatures to upstream cell signalling networks. Nucleic Acids Res 46:W171–W179. https://doi.org/10.1093/NAR/GKY458
Sherman BT, Hao M, Qiu J et al (2022) DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res 50:W216–W221. https://doi.org/10.1093/NAR/GKAC194
Cannon M, Stevenson J, Stahl K et al (2024) DGIdb 50: rebuilding the drug-gene interaction database for precision medicine and drug discovery platforms. Nucleic Acids Res 52:D1227–D1235. https://doi.org/10.1093/NAR/GKAD1040
R Li, Y Yu, SO Jaafar, B Baghchi, M Farsimadan, I Arabipour, H Vaziri (2022) Genetic variants miR-126, miR-146a, miR-196a2, and miR-499 in polycystic ovary syndrome. Br J Biomed Sci 79. https://doi.org/10.3389/BJBS.2021.10209/FULL
Sepahi N, Kohan L, Jahromi AR et al (2017) mir-126 rs4636297 and TGFβRI rs334348 functional gene variants are associated with susceptibility to endometriosis and its severity. Gynecol Endocrinol 33:429–432. https://doi.org/10.1080/09513590.2017.1290064
X Jiang, J Li, B Zhang, J Hu, J Ma, L Cui, ZC-F and sterility, undefined 2021, Differential expression profile of plasma exosomal microRNAs in women with polycystic ovary syndrome, Elsevier. (n.d.). https://www.sciencedirect.com/science/article/pii/S0015028220307731 (accessed March 30, 2024)
Li C, Chen Y, Chen X et al (2020) MicroRNA-183-5p is stress-inducible and protects neurons against cell death in amyotrophic lateral sclerosis. J Cell Mol Med 24:8614–8622. https://doi.org/10.1111/JCMM.15490
I Ischenko, M Rao, J Li, MJ Hayman, S Powers, O Petrenko, NC Reich (n.d.) KRAS drives immune evasion in a genetic model of pancreatic cancer, Nature.ComI Ischenko, S D’Amico, M Rao, J Li, MJ Hayman, S Powers, O Petrenko, NC ReichNature Commun. 2021•nature.Com. https://doi.org/10.1038/s41467-021-21736-w
ABMMK Islam, E Mohammad, MAAK Khan (2020) Aberration of the modulatory functions of intronic microRNA hsa-miR-933 on its host gene ATF2 results in type II diabetes mellitus and neurodegenerative disease development, Hum. Genomics. 14. https://doi.org/10.1186/S40246-020-00285-1
Qin Y, Wang Y, Zhao H et al (2021) Aberrant miRNA-mRNA regulatory network in polycystic ovary syndrome is associated with markers of insulin sensitivity and inflammation. Ann Transl Med 9:1405–1405. https://doi.org/10.21037/ATM-21-1288
Y Yang, YC-S in cancer biology, undefined 2022, The impact of VEGF on cancer metastasis and systemic disease, Elsevier Yang, Y CaoSeminars Cancer Biol. 2022•Elsevier. (n.d.). https://www.sciencedirect.com/science/article/pii/S1044579X22000670 (accessed April 1, 2024)
A White (2023) GB- Biomolecules, undefined 2023, VEGFA isoforms as pro-angiogenic therapeutics for cerebrovascular diseases, Mdpi.Comal White, GJ BixBiomolecules, 2023•mdpi.Com. https://doi.org/10.3390/biom13040702
M Huang, Y Ji, J Chen, D Li, T Zhou, … PQ-AP, undefined 2023, Targeted VEGFA therapy in regulating early acute kidney injury and late fibrosis, Nature.ComM Huang, Y Ji, J Chen, D Li, T Zhou, P Qi, X Wang, X Li, Y Zhang, X Yu, L Wu, X Sun, G CaiActa Pharmacol. Sin. 2023•nature.Com. (n.d.). https://www.nature.com/articles/s41401-023-01070-1 (accessed August 10, 2024)
Wan R, Feng J, Tang L (2021) Consequences of mutations and abnormal expression of smad4 in tumours and t cells. Onco Targets Ther 14:2531–2540. https://doi.org/10.2147/OTT.S297855
Y Zhou, Y Wu, C Chong, S Zhong, ZW- Heliyon, undefined 2023, Irpex lacteus polysaccharide exhibits therapeutic potential for ovarian fibrosis in PCOS rats via the TGF-β1/smad pathway, Cell.ComYY Zhou, YQ Wu, CJ Chong, SM Zhong, ZX Wang, XH Qin, ZQ Liu, JY Liu, JL SongHeliyon, 2023•cell.Com. (n.d.). https://www.cell.com/heliyon/pdf/S2405-8440(23)05949-2.pdf (accessed April 1, 2024)
YH Kim, SW Choe, MY Chae, S Hong, SP Oh (2018) SMAD4 deficiency leads to development of arteriovenous malformations in neonatal and adult mice. J Am Heart Assoc 7. https://doi.org/10.1161/JAHA.118.009514
MA Moga, A Bălan, OG Dimienescu, V Burtea, RM Dragomir, CV Anastasiu (2019) Circulating miRNAs as biomarkers for endometriosis and endometriosis-related ovarian cancer—an overview. J Clin Med 8. https://doi.org/10.3390/JCM8050735
S Elsherif, S Faria, C Lall, … RI-J of computer, undefined 2019, Ovarian cancer genetics and implications for imaging and therapy, Journals.Lww.ComSB Elsherif, SC Faria, C Lall, R Iyer, PR BhosaleJournal Comput. Assist. Tomogr. 2019•journals.Lww.Com. (n.d.). https://journals.lww.com/jcat/fulltext/2019/11000/Ovarian_Cancer_Genetics_and_Implications_for.2.aspx (accessed April 1, 2024)
Yan MQ, Zhu BH, Liu XH et al (2023) Mitoguardin 1 and 2 promote granulosa cell proliferation by activating AKT and regulating the Hippo-YAP1 signaling pathway. Cell Death Dis 14:1–12. https://doi.org/10.1038/s41419-023-06312-y
A Alwhaibi, A Verma, M Adil, PS-P research, undefined 2019, The unconventional role of Akt1 in the advanced cancers and in diabetes-promoted carcinogenesis, ElsevierA Alwhaibi, A Verma, MS Adil, PR SomanathPharmacological Res. 2019•Elsevier. (n.d.). https://www.sciencedirect.com/science/article/pii/S1043661818318991 (accessed April 1, 2024)
Chen J, Somanath PR, Razorenova O et al (2005) Akt1 regulates pathological angiogenesis, vascular maturation and permeability in vivo. Nat Med 11:1188–1196. https://doi.org/10.1038/nm1307
T Schaefer, CL- Oncogene, undefined 2020, SOX2 protein biochemistry in stemness, reprogramming, and cancer: the PI3K/AKT/SOX2 axis and beyond, Nature.ComT Schaefer, C LengerkeOncogene, 2020•nature.Com. (n.d.). https://www.nature.com/articles/s41388-019-0997-x (accessed April 1, 2024)
G Maurizi, N Verma, A Gadi, A Mansukhani, CB- Oncogene, undefined 2018, Sox2 is required for tumor development and cancer cell proliferation in osteosarcoma, Nature.ComG Maurizi, N Verma, A Gadi, A Mansukhani, C BasilicoOncogene, 2018•nature.Com. (n.d.). https://www.nature.com/articles/s41388-018-0292-2 (accessed August 10, 2024)
D Yuan, J Luo, Y Sun, L Hao, J Zheng, ZY-C Signalling, undefined 2021, PCOS follicular fluid derived exosomal miR-424–5p induces granulosa cells senescence by targeting CDCA4 expression, ElsevierD Yuan, J Luo, Y Sun, L Hao, J Zheng, Z YangCellular Signalling, 2021•Elsevier. (n.d.). https://www.sciencedirect.com/science/article/pii/S0898656821001194 (accessed April 20, 2024)
JG- Reproduction, undefined 2015, The role of WNT signaling in adult ovarian folliculogenesis, Rep.Bioscientifica.ComJAH GiffordReproduction, 2015•rep.Bioscientifica.Com. (n.d.). https://rep.bioscientifica.com/view/journals/rep/150/4/R137.xml (accessed April 29, 2024)
Zhu M, Fan Z (2022) The role of the Wnt signalling pathway in the energy metabolism of bone remodelling. Cell Prolif 55:e13309. https://doi.org/10.1111/CPR.13309
O Habara, C Logan, … MK-A-, undefined 2021, WNT signaling in pre-granulosa cells is required for ovarian folliculogenesis and female fertility, Journals.Biologists.ComO Habar. CY Logan, M Kanai-Azuma, R Nusse, HM Tak. 2021•journals.Biologists.Com. (n.d.). https://journals.biologists.com/dev/article-abstract/148/9/dev198846/261700 (accessed September 13, 2023)
L Li, X Shi, Y Shi, Z Wang (2021) The signaling pathways involved in ovarian follicle development. Front Physiol 12. https://doi.org/10.3389/FPHYS.2021.730196/FULL
Acknowledgements
The authors are thankful to the NITTE (Deemed to be University) for providing all the support and facilities for this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bhandary, P., Alagundagi, D.B., Shetty, P.K. et al. Identification of potential miRNA-mRNA regulatory network contributing to pathogenesis of polycystic ovarian syndrome. Ir J Med Sci (2024). https://doi.org/10.1007/s11845-024-03795-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11845-024-03795-2